Abstract
The rabbit has long been a model for studies of the immune system. Work using rabbits contributed both to the battle against infectious diseases such as rabies and syphilis, and to our knowledge, of antibodies' structure, function, and regulated expression. With the description of rabbit Ig allotypes, the discovery of different gene segments encoding immunoglobulins became possible. This challenged the “one gene–one protein” dogma. The observation that rabbit allotypic specificities of the variable regions were present on IgM and IgG molecules also led to the hypothesis of Ig class switching. Rabbit allotypes contributed to the documentation of phenomena such as allelic exclusion and imbalance in production of allelic gene products. During the last 30 years, the rabbit Ig allotypes revealed a number of unique features, setting them apart from mice, humans, and other mammals. Here, we review the most relevant findings concerning the rabbit IGHV. Among these are the preferential usage of one VH gene in VDJ rearrangements, the existence of trans-species polymorphism in the IGHV locus revealed by serology and confirmed by sequencing IGHV genes in Lepus, the unusually large genetic distances between allelic lineages and the fact that the antibody repertoire is diversified in this species only after birth. The whole genome sequence of a rabbit, plus re-sequencing of additional strains and related genera, will allow further evolutionary investigations of antibody variation. Continued research will help define the roles that genetic, allelic, and population diversity at antibody loci may play in host-parasite interactions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Historic perspective
Studies of the rabbit immune system have greatly contributed to our knowledge of the structure, function, and regulation of antibodies. During the late nineteenth and early twentieth centuries, the use of rabbit in immunological research was crucial to the development of the rabies vaccine by Louis Pasteur and to the battle against syphilis (reviewed in Dubiski 1987). In the first half of the twentieth century, the foundations of molecular immunology were laid with almost exclusive use of the rabbit. The antigenic polymorphism of serum components was described in this species as early as 1902, although the antigens involved were characterized only 50 years later (see Kelus and Gell 1967). In 1956, Oudin (1960) demonstrated and defined allotypy of immunoglobulins in the rabbit. Allotypes were proven to be hereditary traits in rabbit by Dubiski et al. (1959) prior to the establishment of the two major linkage groups “a” and “b”, corresponding to the H and light chain loci, respectively, and even before the heterodimeric structure of the antibody molecule was clearly established.
The existence of genetic markers of the different gene fragments encoding the antibody H chain was unique to this species and the rabbit Ig allotypes challenged the “one gene–one protein” dogma. Indeed, Todd (1963) and others found that rabbit allotypic specificities of the variable regions were present on both IgM and IgG molecules, suggesting that the same VH gene segment can be translocated to different constant region genes. This observation opened the road to confirmation of the concept of multiple “germline” VH gene segments that can be joined to a limited number of genes encoding the different heavy chain constant regions (Dreyer and Bennett 1965), and led to the hypothesis of Ig class switching (Kearney et al. 1976; van der Loo et al. 1979; reviewed in Severinson et al. 1982). Hamers and co-workers (1966) reported cis-expression of the VH and CH genes by using allelic markers on both protein domains. The markers in the CH2 domain were re-discovered by Dubiski (1969) and renamed the e14 and e15 allotypes. Mage and co-workers (1971) reported the first of a number of crossing-over events observed by laboratories during breeding. Findings of genetic recombination between the genes controlling the rabbit VH and CH (estimated recombination frequency of 0.1%; (Mage 1979; Kelus and Steinberg 1991)) confirmed the model of Dreyer and Bennett (1965).
Documentation of the current V-D-J-C model therefore originated through studies of the rabbit allotypes. In addition to genetic linkage of VH and CH genes, and the expression of apparently identical VH regions on different classes of Ig, it was through studies of rabbit immunoglobulin markers that phenomena such as allelic exclusion and imbalance in production of allelic gene products were described (e.g., Davie et al. 1971; Loor and Kelus 1978; Schmale et al. 1969; Wolf et al. 1971). During the last 30 years, the rabbit Ig allotypes revealed a number of unique features, setting them apart from mice, humans, and other mammals. Among these are the preferential usage of one VH gene in VDJ rearrangements, the evidence for the existence of trans-species polymorphism in the IGHV, IGHG, and IGKC1 loci, the unusually large genetic distances between the allelic lineages and the fact that the antibody repertoire is diversified in this species only after birth.
Lagomorph taxonomy
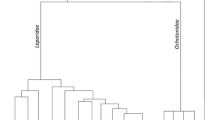
The order Lagomorpha comprises two families: Ochotonidae and Leporidae. The family Ochotonidae includes 30 species restricted to the genus Ochotona (Pikas). The family Leporidae can be divided into two groups: hares and rabbits. According to Chapman and Flux (1990), the hare group encompasses a single genus, Lepus, whereas the rabbit group includes ten genera (Brachylagus, Bunolagus, Caprolagus, Nesolagus, Oryctolagus, Pentalagus, Poelagus, Pronolagus, Romerolagus, and Sylvilagus) and 25 species (Alves and Hackländer 2008). There is not a consensus estimate of the divergence time between rabbits and hares. Analyses of several nuclear and mitochondrial markers suggest that the genera Oryctolagus and Lepus diverged between 6 and 20 Mya (Biju-Duval et al. 1991; Halanych and Robinson 1999; Matthee et al. 2004) (Fig. 1), although fossil data suggests an earlier divergence (between 2.5 and 3.5 Mya)) (Lopez-Martinez 2008).
Evolutionary topology reflecting the relationships within the Lagomorpha group based on a molecular super matrix (adapted from Matthee et al. 2004)
The genus Oryctolagus is monospecific (Oryctolagus cuniculus). In the Iberian Peninsula, where the fossil record suggests the European rabbit originated (Pages 1980; Lopez-Martinez 1989, 2008; Corbet 1994), two morphologically differentiated subspecies of European rabbit have been distinguished: O. cuniculus algirus and O. cuniculus cuniculus (Cabrera 1914). These two subspecies diverged ~1.8 Mya (reviewed in Carneiro et al. 2009). O. c. algirus inhabits the southwestern Iberian Peninsula, while O. c. cuniculus is present in the northeastern Iberian Peninsula. O. c. cuniculus later expanded its range north towards France, likely after the last glacial peak (Queney et al. 2001), where it still remains present. The Pleistocene Ice ages forced many temperate species to retreat into southern refugia, leading to high levels of diversity and endemism in these areas (Hewitt 1996). The Iberian Peninsula was one of these refugia in Europe (Taberlet et al. 1998). This is reflected in the significantly lower genetic diversity of the wild French O. c. cuniculus populations compared to the Iberian populations (van der Loo et al. 1991, 1999; Queney et al. 2001; Esteves et al. 2004; Ferrand and Branco 2007; Surridge et al. 2008; Carneiro et al. 2011). The European rabbit gene pool has been manipulated through a recent single domestication event of French origin, and therefore, all domestic rabbits belong to subspecies O. c. cuniculus (reviewed in Ferrand and Branco 2007; Carneiro et al. 2011). Today, through mediated anthropic dispersal, the European rabbit is present in Continental Europe, England, Australia, New Zealand, North and South America, and North Africa.
For other leporids, the study of immunoglobulin genes is mainly restricted to the genera Lepus and Sylvilagus (Cazenave et al. 1977; Teherani and Mandy 1976a, 1976b; Teherani et al. 1979; Teherani et al. 1982; Bouton and van der Loo 1997; Esteves et al. 2002a, 2005, 2006). Lepus is a polytypic, cosmopolitan genus, comprising 24 to 30 currently recognized species (Corbet and Hill 1980; Flux and Angermann 1990; Alves and Hackländer 2008), that, like Sylvilagus, most probably originated in North America (Lopez-Martinez 2008; Matthee et al. 2004). Lepus species that have been used in immunogenetic studies include Lepus americanus, Lepus timidus, Lepus granatensis, and Lepus europaeus. L. americanus is distributed throughout North America and occupies a basal position in the Lepus group (Matthee et al. 2004). L. granatensis is endemic to the Iberian Peninsula, covering the whole peninsula except the Northeast where it is replaced by L. europaeus, which spans a wide range throughout Europe, with several introduced populations in different regions, like South America, Australia, New Zealand, and several islands including Barbados, Reunion, and Falklands (reviewed in Alves and Hackländer 2008). L. timidus is a Palearctic relict species with an extensive distribution area in Europe and Asia, from Ireland and Scandinavia in the west, across Siberia, Mongolia, and China to the northern islands of Japan in the east. Sylvilagus is restricted to the American continent and comprises 17 species (reviewed in Alves and Hackländer 2008). Within this genus, the most studied species is Sylvilagus floridanus, a species that occupies Southern and Western North America.
Rabbit preferentially expresses IGHV1
The rabbit IGHV allotypes are highly divergent and behave as Mendelian alleles (Oudin 1956, 1960; Dubiski et al. 1959; Dray et al. 1963; Kim and Dray 1972; Mage et al. 1984). Serological surveys of domestic rabbits defined three allotypic lineages, the so-called VHa allotypes a1, a2 and a3. VH allotypic markers have not been found in other species, and the Mendelian behavior of rabbit IGHV allotypes was puzzling for many years, as rabbit serological data revealed a complex situation where homozygous VHa rabbits were found to express distinct VH genes. Some of them were devoid of VHa allotype-specific determinants (the so-called VHa negative or VHn, alias VHx, VHy, and VHz; Horng et al. 1976). The VHa-positive molecules (i.e., displaying motifs of VHa-characteristic determinants) also showed variation which revealed the existence of multiple VHa-related gene fragments (Brezin and Cazenave 1975; Mage et al. 1976; van der Loo et al. 1977). This was later confirmed by genomic mapping of the rabbit VH region which revealed the existence of over 200 VH elements in the germline (Currier et al. 1988; Gallarda et al. 1985; Ros et al. 2004). This may, in part, explain why for many years, it was so difficult to understand the presence of allelic allotypic markers on most serum immunoglobulins of all classes (and probably the skepticism of researchers less acquainted with serological methods).
An important contribution to understanding the mechanisms underlying preferential VDJ rearrangement in the rabbit was the study of a rabbit strain, called “Alicia”, detected as a mutant during breeding at the Basel Institute, Switzerland (Kelus and Weiss 1986). The mutant rabbits derived from a VH1a2 parental rabbit in which a 10 Kb segment of genomic DNA containing the VH1 gene was deleted (Knight and Becker 1990; Allegrucci et al. 1990; Ros et al. 2004). In contrast to normal individuals of the a2 lineage, the young homozygous ali/ali mutant Alicia rabbits produced only trace amounts of a2 molecules, and their serum contained mostly Ig resulting from rearranged genes not encoding VHa allotype-associated epitopes (VHa negative or VHn genes) (Kelus and Weiss 1986; DiPietro et al. 1990; Chen et al. 1993). As the genes encoding the VHn allotypes map at least 100 Kb upstream of VH1 (Mage et al. 2006), it is intriguing that VDJ rearrangements of these appeared to be more prevalent than VDJ rearrangements of the D-proximal functional VHa positive VH4 gene in young Alicia rabbits. Further analysis showed that the VH4 gene is in fact the predominantly rearranged VH gene found in the bone marrow of young Alicia rabbits, but for some reason most VH4-utilizing B-cells are eliminated. Thus, the immunoglobulins of the young rabbits are produced by B-cells that utilize VHn. As the young “Alicia” rabbit ages, the proportions of B cells expressing serological a2 specificities increase (Kelus and Weiss 1986; DiPietro et al. 1990; Allegrucci et al. 1990; Chen et al. 1993; Pospisil et al. 1995). Analysis of nucleotide sequences of the promoter region showed that more than 80% of the VDJ rearrangements in older Alicia rabbits utilize either the functional VH4 or VH7 genes localized upstream of VH1. The VH4 and the VH7 genes have 7 (out of 11) specific nucleotides associated with the allotype a2, while the other nucleotides that characterize a2 are gained through somatic gene conversion using VH9 or a VH9-like germline gene as donor (Sehgal et al. 1998; Zhu et al. 1999).
The study of this mutant rabbit strain showed that the Mendelian inheritance of the VHa allotypes in normal rabbit is explained by the preferential usage of only one VH gene in VDJ rearrangements, i.e., the D-proximal VH gene segment VH1 (see Fig. 2), which is deleted from the “ali” genome (Knight and Becker 1990; Knight 1992; Allegrucci et al. 1990; Ros et al. 2004). Despite having more than 200 VH genes (Currier et al. 1988; Gallarda et al. 1985; Ros et al. 2004), over 50% are “non-functional” and, apart from VH1, an even smaller fraction seems to encode a-positive proteins (Ros et al. 2004). About 80% to 90% of circulating Ig molecules are derived from the VH1 gene and express the VHa allotypic markers (e.g., Kindt 1975; Margolies et al. 1977). The VH regions of the remaining 10–20% of Ig molecules are encoded by the VHn genes (Kim and Dray 1973; Roux 1981), which are localized at least 100 Kb upstream of VH1 (Mage et al. 2006). Thus far, the basis for the preferential usage of the VH1 gene in VDJ rearrangements remains unanswered. Enhanced chromatin accessibility of this region of the DNA may be responsible (Mage et al. 2006). Furthermore, we can speculate that the retention and occasional usage of the VHn genes in 10–20% of VDJ rearrangements may represent an evolutionary relic of Lagomorphs ancestral immunologic response to pathogens.
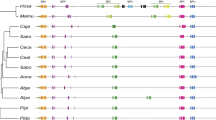
Rabbit IGHV (regions from VH haplotype (a2 F-I)) for which nucleotide sequence is available from sequenced, assembled BAC clones. Depicted are V genes, D and J regions and also Cμ. VH functional genes are represented in white rectangles, VH possible pseudogenes in gray and VH pseudogenes in black, identified according to criteria described in Ros et al. (2004). The D-proximal VH gene, VH1, used in the majority of VDJ rearrangements is indicated. Identified above are the non-overlapping BAC clones 38A2 and 225P18 (GenBank accession numbers AY386694 and AY386697), and 219D23 (GenBank accession number AY386695). The 3′ end of BAC 225P18 overlaps the 5′end of BAC 219D23 (adapted from Ros et al. 2004)
Genetic diversity at the heavy chain variable region IGHV locus (a locus)
In the domestic rabbit, the three serologically defined allotypic lineages, VHa allotypes a1, a2, and a3, are highly divergent (±20% amino acid sequence differences). The allelic specificities of a1 and a2 are correlated with several amino acid differences in framework regions 1 (positions 5, 8, 11, 13, 14, 17, and 18) and 3 (positions 74, 76, 79, 80, 83, 84, 96, and 97) (Fig. 3) (Tonnelle et al. 1983; Mage et al. 1984; Knight and Becker 1990). Serologically distinct VHa allotypes were discovered in wild rabbits and named VHa100-VHa109 by Cazenave and coworkers (reviewed in Cazenave et al. 1987). Partial protein sequences of purified antibodies of the VHa100 type (Tonnelle et al. 1983), suggest these are related to the more recently described and fully sequenced VHa allele(s) corresponding to the a4 lineage (Esteves et al. 2004). Studies by double immunodiffusion of sera from domestic breeds and wild rabbits of Continental Europe (North of Pyrenees Mountains), Great Britain and overseas showed the presence of allotypes a1, a2 and a3, as well as a rare allotype with partial reaction against a3-specific antiserum, found in the French populations. In all studied populations, the gene frequencies of a1 and a3 were similar (±40%) and higher than a2 (±20%) (van der Loo 1993). Interestingly, these allelic frequencies reflect the “pecking order” of their relative expression in heterozygous animals (Mage et al. 1967; Lummus et al. 1967). Quite surprisingly, the study of wild rabbit populations from the Iberian Peninsula belonging to the subspecies O. c. algirus and O. c. cuniculus revealed a different picture with a much higher prevalence of the “a-blank” allele. Also, a gradient in the gene frequencies of this allotype was noted, decreasing from southwestern to northeastern populations (Fig. 4) (Esteves et al. 2004). Sequencing of rearranged VH genes expressed in O. c. algirus rabbits that were typed as “a-blank” showed that these rabbits preferentially use VH genes that, although clearly related to the known VHa genes, defined a new major allotypic lineage, designated a4. The a4 sequences display the rabbit VHa hallmark residues together with a number of unprecedented amino acid changes in FR2 and FR3, which may explain the lack of reactivity with known alloantisera (Fig. 3). The net protein distances between the VH-a4 and the VH-a1, a2, and a3 lineages, was 20%, 29%, and 21%, respectively, similar to the net protein distances between VH-a1, a2, and a3 lineages. As such, a new VHa allele was found in Iberian O. c. algirus populations, constituting the fourth of the distantly related lineages of the rabbit VHa-locus, one of which, the a4 lineage, seems to be endemic in the Iberian range (Esteves et al. 2004). The data for the VHa lineages also suggested that some lineages, namely VHa a1, a2 and a4, showed higher evolutionary rates than the VHa a3 and VHn lineages. This increased evolutionary rate may account for the accumulation of a considerable fraction of the amino acid differences characterizing the a4 lineage after the separation of the two O. cuniculus subspecies, explaining its association with subspecies O. c. algirus (Esteves et al. 2004).
Amino acid sequences of European rabbit (O. cuniculus) proteins (encoded by VH1a1-a3, VHa4.1–4.2, VHx, VHy, VHz) and hare (VHa2L, VHsLe, VHsLg) VH genes. Germline elements are italicized (Accession numbers: VH1a1-M93171; VH1a2-M93172; VH1a3-M93173; VHx-L03846; VHy-L03890; VHz-AF264469.). VHa4.1–4.2, VHa2L, VHsLe and VHsLg represent consensus protein sequences obtained by Esteves et al. 2004, 2005. Framework regions (FR1, FR2, FR3) and each CDR (CDR1 and CDR2) are defined according to the IMGT numbering system (also shown). CDRs are shaded. The amino acids that are characteristic of a1, a2 and a4 lineages are marked with black, gray, and white arrows, respectively. Hallmark rabbit VHa residues are highlighted in bold and underlined. Note that the Lepus VHa2L shares these residues, as well as five of the VH1a2 lineage characteristic amino acids. Dots indicate identity with VH1a1, dashes represent indels and question marks represent ambiguous positions in consensus sequences
Geographic distribution of VHa locus allotype frequencies, serologically determined for populations of European wild rabbit (O. cuniculus) from the Iberian Peninsula and France. The subspecies O. c. algirus occupies the southwestern part of Iberia, which is indicated by A. Rabbits of the Northwestern areas of Iberia and of the rest of Europe (indicated by B) belong to the subspecies O. c. cuniculus. The contact zone between the two subspecies is indicated in gray. The coloring of the disks reflects the relative allele frequencies per locality, analyzed as a two-allele locus with a-blank (white) and a-positive (black) alleles. Locality abbreviations are as follows: prt Portimão, san Santarém, alc Alcochete, ida Idanha, cab Cabreira, Ll Las Lomas, hue Huelva, don Doñana, cre Ciudad Real, inf Infantado, vv Vila Viçosa, am Amoladeras, tol Toledo, bra Bragança, tar Tarragona, alt Alicante, nav Navarra, lle Lerida, per Perpigan, cam Camargue, ver Versailles, arj Arjuzanx, gra Grax, hel Helene
Studies on the IGHV locus diversity for other leporids are limited to a few data obtained for some Lepus species, namely L. americanus, L. europaeus, and L. granatensis. For L. americanus, only serological analyses have been conducted; these showed cross-reaction with rabbit anti-a1, anti-a2, and anti-a3 antisera and the absence of individuals with no cross-reaction to rabbit antisera (De Poorter 1984). As for L. europaeus and L. granatensis, a serological analysis of several populations using allo-antisera against rabbit a1, a2, and a3 allotypes revealed only two phenotypes: partial reaction to anti-a2 antisera and no reaction to any rabbit antiserum (Table 1) (Esteves 2003). Sequences of expressed VH genes were obtained for only five Lepus specimens of both species, three of which expressed a2-cross-reacting serum proteins, and these could be grouped into two different lineages (see below “The trans-specific polymorphism of leporids IGHV”). Extending the study of IGHV diversity to more Lepus individuals and species could unravel more VH lineages, as shown by the rabbit VH1-a4 lineage found only in O. c. algirus.
The trans-specific polymorphism of leporid IGHV
Such large interallelic distances as those observed at the rabbit VHa locus can be the outcome of unusually long allele persistence times and/or increased mutant recruitment rates. The first indications that the different VH1 polymorphisms could be trans-specific came from studies showing serological cross-reactivity of Lepus sera with rabbit VHa-allotype-specific allo-antisera (van der Loo et al. 1977; Horng et al. 1980; van der Loo 1987). Indeed, for L. americanus large-scale serological analysis, comprising several hundreds of specimens, showed cross-reaction with rabbit anti-a1, anti-a2, and anti-a3 antisera, revealing at least three alleles with hierarchical frequency distributions similar to those observed for feral rabbits (De Poorter 1984; van der Loo 1987). Su and Nei (1999) compared the extent of sequence divergence between the rabbit a1, a2, and a3 allotypes with that between human and mouse VH gene sequences and concluded that assuming a “normal” mutation rate, the rabbit VH1 polymorphism has persisted for about 50 My. Since the Oryctolagus and Lepus genera diverged 12 Mya (Matthee et al. 2004), the allelic lineages present in one leporid species should be more related to some of the alleles expressed in the other species than to their conspecific allelic counterparts. Serological cross-reactivity can depend upon a small number of amino acid replacements, and allo-antisera may reveal plesiomorphic character states (i.e., which became antigenic in the recipient strain when they were altered), and so the hypothesis that the VH1 polymorphism preceded the rabbit–hare split remained controversial until sequence data was obtained.
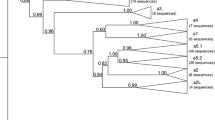
Focusing on this question, Esteves and co-workers (2005) sequenced rearranged VH gene products from five Lepus specimens of both L. granatensis and L. europaeus species, three of which expressed a2-cross-reacting serum proteins, and compared them to known rabbit VH sequences of different allotypes. Within the Lepus VH genes, two lineages were observed, one of which (a2L) showed some of the most outstanding characteristics of the rabbit VHa genes, furthermore possessing 5 out of 11 amino acid residues that characterize the allotype VHa2 (Mage et al. 1984). This lineage was only obtained from specimens showing rabbit a2 antisera cross-reaction. A second lineage displayed more similarity to the rabbit VHn gene fragments (sL). Phylogenetic inference methods clearly placed the hare a2L sequences into the same monophyletic group with the rabbit a2 sequences (Fig. 5), thus showing that these sequences were more closely related to the rabbit VH1-a2 allele than VH1-a2 was to its allelic counterparts, VH1-a1 and VH1-a3. Also, the genetic distance measured between different rabbit genomic VHa sequences was approximately 1.5 times larger than between the Lepus a2L consensus and genes of the rabbit a2 allotype. Together, these findings strongly suggest that the allotype split predates the Oryctolagus–Lepus species split, supporting previous evidence of the trans-species nature of the VH1 polymorphism. The alternative interpretation that the evolutionary rate of the rabbit a2 lineage has become significantly slower than at the other lineages, was not supported by the data, as the a2 sequences were rather more derived compared to the other allotypes, in particular, the a3 lineage.
Neighbor-joining tree for Oryctolagus cuniculus, Lepus europaeus (Le), and Lepus granatensis (Lg) VH genes, showing the transpecies polymorphism of VHa2. VH genes from human class 3 (VH3) determine the root of the tree. Numbers represent the bootstrap values (BP, in bold), and the confidence probability values (CP, in italics). Expressed VH genes (cDNA) are labeled by black (Le), gray (Lg), and white (O. cuniculus) circles. Germline VHa gene sequences (VH1-VH4) of rabbit allotypes a1, a2, and a3 are marked by black triangles. White squares indicate VHn germline genes. Black squares indicate human VH3 germline genes. “With kind permission from Springer Science + Business Media: Immunogenetics, The evolution of the immunoglobulin heavy chain variable region (IgVH) in leporids: an unusual case of trans-species polymorphism., 57(11), 2005, 874, Esteves PJ, Lanning D, Ferrand N, Knight KL, Zhai SK, van der Loo W, Fig. 1”
The large inter-allelic differences at the VH1 locus
Regarding the two different hypotheses invoked to explain the very large inter-allelic distances observed at the IGHV1 locus, the results obtained seem to be contradictory. On the one hand, the association of specific alleles with subspecific markers suggests that the large differences between a4 and domestic alleles (a1, a2, and a3) may have accumulated after the separation of the subspecies, with different evolutionary rates among lineages. On the other hand, the confirmation of the trans-specific nature of the VH1 polymorphism between Oryctolagus and Lepus supports the hypothesis that very long lineage persistence times have contributed to allelic divergence. The evidence obtained points to two major conclusions, (1) the allelic lineages can be maintained in the genome for a long time, and (2) mutation rates can differ between allelic lineages. The origin of the selection forces that regulate the VH1 polymorphism is unknown. However, the requirement for exogenous factors, such as the intestinal microbiota, for diversification of the primary antibody repertoire in rabbits suggests that this mechanism might impose constraints on the evolution of the VH1 alleles.
Diversification of the primary Ab repertoire in the rabbit
Among vertebrate species, two different strategies to generate the primary Ab repertoire have been adopted. Human and mouse preferentially use combinatorial rearrangements of a large number of V, D, and J gene segments. In contrast, several species possess or use only a limited number of germline V segments. Birds, together with several mammalian species (rabbit, sheep, pig, and bovine), use strategies of primary repertoire development that overcome this limitation. In these species, the primary antibody repertoire is diversified post-rearrangement by one or both of two mechanisms: somatic gene conversion and somatic hypermutation.
In chicken, somatic gene conversion is the major mechanism of diversification of immunoglobulin heavy and light chain variable regions. A set of pseudogenes is used as donors and the unique rearranged V(D)J gene acts as acceptor (Reynaud et al. 1985; 1989). In sheep, B-cell diversification occurs in early development in the ileal Peyer's patches and results from somatic hypermutation rather than somatic gene conversion (Reynaud et al. 1991; 1995). Both processes are used to diversify the antibody repertoire in cattle (Parng et al. 1996; Lucier et al. 1998).
Becker and Knight (1990) showed that the rearrangement process in the rabbit involves a single preferentially rearranged VH gene that is diversified mainly through somatic gene conversion, using other upstream VH genes as donors (Becker and Knight 1990; reviewed in Knight and Crane 1994). Somatic hypermutation also takes place, distributing point mutations throughout the entire VDJ gene (Short et al. 1991; Weinstein et al. 1994; Sehgal et al. 2000). In rabbits, B cells generated in the fetal liver and bone marrow subsequently migrate to the appendix and other gut-associated lymphoid tissues (GALT), where they proliferate and are intensively diversified. The VH genes upstream of VH1 contribute to the expressed VH1 diversification by somatic gene conversion. They evolved to some extent in concert, which resulted in the presence of the allotypic motifs on other VH gene segments of the Ig heavy chain locus (haplotype polymorphism).
In contrast to species such as chickens (e.g., Reynaud et al. 1989), sheep (Reynaud et al. 1995), and cattle (e.g., Lucier et al. 1998), the diversification of the primary antibody repertoire in rabbits is not developmentally programmed. It has been shown that the peripheral Ab repertoire is diversified between 4 weeks and 2 months after birth (Cooper et al. 1968; Weinstein et al. 1994). Surgical removal of the appendix and sacculus rotundus on the day of birth and the Peyer's patches at 3 weeks of age (Cooper et al. 1968) more recently showed that GALT is essential for the generation of the primary antibody repertoire (Vajdy et al. 1998). It has also been shown that exogenous factors, such as the intestinal microbiota, are required for Ab repertoire diversification in the rabbit (Lanning et al. 2000a, 2000b; Sehgal et al. 2002). The rabbit is a species with altricial young (naked and blind at birth). Other lagomorph species, such as hares (genus Lepus) have precocial young. The unique situation of post-natal diversification in rabbit could be related to the shortening of gestation time (4 weeks in rabbit and 6 weeks in hares). Considering the differences in reproductive biology between the genera Oryctolagus and Lepus, it would be interesting to investigate whether in Lepus, the Ab repertoire is diversified neonatally or during fetal life.
Evolution of VH gene families
The study of VH genes from several vertebrate species showed that VH genes from the same species could belong to different groups. In tetrapods, the IGHV sequences have been classified into three clans, clans I, II, and III (Schroeder et al. 1990; Kirkham et al. 1992; Das et al. 2008). The human and mouse possess VH genes that are very diversified and may cluster with either clans I, II, or III (polyphyletic), whereas VH genes of chicken, rabbit, horse, and artiodactyls (cattle, sheep, and swine) are monophyletic. Chicken, rabbit, and swine VH genes each form distinct clusters within group III, whereas those of horse, cattle, and sheep each form distinct clusters within group II. Since VH genes of different artiodactyl species cluster within different groups (II and III), their common ancestor must have possessed VH genes from both groups (Sitnikova and Su 1998). Thus, the presence of IGHV genes of only one clan in several tetrapod species reflects an evolutionary loss of IGHV genes.
Monofunctional multigene families are generally believed to undergo processes of genetic exchange, like gene conversion and unequal crossover, which homogenize the DNA sequences (Smith et al. 1971; Smith 1974; Zimmer et al. 1980). This has been called “concerted” evolution. The concerted evolution model has been invoked to explain the evolution of VH genes (Hood et al. 1975; Ohta 1983). However, the studies of VH gene families in human and mouse (Gojobori and Nei 1984; Tutter and Riblet 1989), and studies on much longer evolutionary time scales, showed that the pattern of VH gene evolution could be better explained by the so-called “birth-and-death” model of evolution (Ota and Nei 1994; Nei et al. 1997; Sitnikova and Su 1998; Das et al. 2008), in which the number of genes in a family (or “library”) are allowed to expand and contract. This model is similar to the “accordion model” of MHC evolution proposed by Klein et al. (1993) and postulates that depending upon the need to protect the host from ever-changing groups of parasites, some VH gene libraries are duplicated and can diverge functionally, while others become pseudogenes and/or are deleted from the genome. The end result of this process is a mixture of divergent and highly homologous groups of genes, and the maintenance of a substantial number of pseudogenes (Ota and Nei 1994; Nei et al. 1997). According to this model, the tetrapod ancestor possessed three VH gene groups (I, II, and III) and library contraction events occurred in several lineages independently. In species that have inherited only one group of VH genes, showing low levels of IGHV gene sequence variation, antibody diversification is mainly due to somatic hypermutation and/or somatic gene conversion. It appears that VH gene library contraction is associated with the development of a specific organ for extensive somatic diversification of the Ab repertoire (bursa of Fabricius in chicken; ileal Peyer's patches in sheep, cattle and probably swine and horse; appendix in rabbit).
Genome sequencing and future prospects
Whole genome sequencing (WGS) at ~7× coverage and assembly of a high-quality draft rabbit genome sequence was recently completed at the Broad Institute, Boston (OryCun2.0). The ENCODE Project, with ~1% of rabbit genomic sequence from a different, normal NZW animal, includes some genes of immunological interest but none of the regions with immunoglobulin heavy or light chain loci. The Thorbecke rabbit strain chosen by Broad for WGS was partially inbred. Although the rabbits accepted skin grafts, they were still segregating for the VHa allotypes a1 and a2. The rabbit chosen for sequencing was heterozygous at the immunoglobulin heavy chain locus. The IGH locus is not assigned to a chromosome in the Broad assembly. The IGHV allele encoding a1 (VH1a1) is located in unplaced scaffold chrUn0742 and the IGHV allele encoding a2 (VH1a2) is located in unplaced scaffold chrUn0439. Unfortunately, there are missing traces or traces of low quality between the telomere and the IGH. Until new sequences are obtained, the current available traces from Broad OryCun2 may not allow further assembly. There are ~75 additional unplaced scaffolds that appear to contain VH genes.
Little is known about the evolution of the IGHV genes in the Lagomorph group. Thus far, only the rabbit O. cuniculus has been extensively studied and some sequences of IGHV genes have been obtained for two Lepus species, L. europaeus and L. granatensis. The study of these species has yielded some interesting results and led to new insights on the evolution of IGHV genes in the Lagomorph group. Also, the few data obtained so far for the Lagomorph IGHG hinge region and CH2 domain has shown some particular patterns of genetic diversity possibly linked to resistance against pathogens (Esteves et al. 2002a; Esteves et al. 2002b; Esteves et al. 2006). Extending our knowledge about both the IGHV genes and IGH constant regions to other Lagomorph species would certainly contribute further to our understanding of the evolution of IGHV genes and the roles that genetic diversity of immunoglobulins may play in host resistance to pathogens.
References
Allegrucci M, Newman BA, Young-Cooper GO, Alexander CB, Meier D, Kelus AS, Mage RG (1990) Altered phenotypic expression of immunoglobulin heavy-chain variable-region (VH) genes in Alicia rabbits probably reflects a small deletion in the VH genes closest to the joining region. Proc Natl Acad Sci U S A 87(14):5444–5448
Alves PC, Hackländer K (2008) Lagomorph species: geographical distribution and conservation status. In: Alves PC, Ferrand N, Hackländer K (eds) Lagomorph biology. Springer, Berlin Heidelberg, pp 395–405
Becker RS, Knight KL (1990) Somatic diversification of immunoglobulin heavy chain VDJ genes: evidence for somatic gene conversion in rabbits. Cell 63(5):987–997
Biju-Duval C, Ennafaa H, Dennebouy N, Monnerot M, Mignotte F, Soriguer RC, Gaaïed AE, Hili AE, Mounolou J-C (1991) Mitochondrial DNA evolution in lagomorphs: origin of systematic heteroplasmy and organization of diversity in European rabbits. J Mol Evol 33(1):92–102
Bouton C, van der Loo W (1997) The trans-species nature of rabbit b locus polymorphism is supported by studies on the snow-shoe hare. Immunogenetics 45(6):444–446
Brezin C, Cazenave PA (1975) Cross reaction between allotype Aa1 of rabbit immunoglobulins and antibodies derived against allotype Aa3: participation of variants of Aa1 specificity in this cross reaction. Immunochemistry 12(3):241–247
Cabrera (1914) Fauna Ibérica. Museo Nacional de Ciências Naturales de Madrid, Madrid
Carneiro M, Ferrand N, Nachman MW (2009) Recombination and speciation: loci near centromeres are more differentiated than loci near telomeres between subspecies of the European rabbit (Oryctolagus cuniculus). Genetics 181(2):593–606
Carneiro M, Afonso S, Geraldes A, Garreau H, Bolet G, Boucher S, Tircazes A, Queney G, Nachman MW, Ferrand N (2011) The genetic structure of domestic rabbits. Mol Biol Evol. doi:10.1093/molbev/msr003, In press
Cazenave PA, Brézin C, Puget A, Mandy W (1977) Phylogeny of the e allotypic system of rabbit immunoglobulins: study of determinants recognized on ochotona IgG by anti-e15 antisera. Ann Immunol 128(1–2):323–327
Cazenave PA, Bennamar A, Sogn JA, Kindt TJ (1987) Immunoglobulin genes in feral population. In: Dubiski S (ed) The rabbit in contemporary immunological research. Longman Scientific & Technical, p 148
Chapman J, Flux J (1990) Introduction and overview of the lagomorphs. In: Chapman J, Flux J (eds) Rabbits, hares and pikas, status, survey and conservation action plan. UICN/SSC Lagomorph Specialist Group, Gland, pp 1–6
Chen HT, Alexander CB, Young-Cooper GO, Mage RG (1993) VH gene expression and regulation in the mutant Alicia rabbit. Rescue of VHa2 allotype expression. J Immunol 150(7):2783–2793
Cooper MD, Perey DY, Gabrielsen AE, Sutherland DE, McKneally MF, Good RA (1968) Production of an antibody deficiency syndrome in rabbits by neonatal removal of organized intestinal lymphoid tissues. Int Arch Allergy Appl Immunol 33(1):65–88
Corbet GB (1994) Taxonomy and origins. In: Tompson HV, King CM (eds) The European rabbit. Oxford Science, Oxford, pp 1–6
Corbet GB, Hill JE (1980) A world list of mammalian species. Oxford University Press, Oxford
Currier SJ, Gallarda JL, Knight KL (1988) Partial molecular genetic map of the rabbit VH chromosomal region. J Immunol 140(5):1651–1659
Das S, Nozawa M, Klein J, Nei M (2008) Evolutionary dynamics of the immunoglobulin heavy chain variable region genes in vertebrates. Immunogenetics 60(1):47–55
Davie JM, Paul WE, Mage RG, Goldman MB (1971) Membrane-associated immunoglobulin of rabbit peripheral blood lymphocytes: allelic exclusion at the b locus. Proc Natl Acad Sci U S A 68(2):430–434
De Poorter M (1984) An experimental test of predictions from different hypothesis of self regulation in the snowshoe hare (Lepus americanus Erxleben, 1777). Vrije University, Brussels
DiPietro LA, Short JA, Zhai SK, Kelus AS, Meier D, Knight KL (1990) Limited number of immunoglobulin VH regions expressed in the mutant rabbit “Alicia”. Eur J Immunol 20(6):1401–1404
Dray S, Young GO, Nisonoff A (1963) Distribution of allotypic specificities among rabbit gamma-globulin molecules genetically defined at two loci. Nature 199:52–55
Dreyer WJ, Bennett JC (1965) The molecular basis of antibody formation: a paradox. Proc Natl Acad Sci U S A 54(3):864–869
Dubiski S (1969) Immunochemistry and genetics of a “new” allotypic specificity Ae14 of rabbit gamma-G immunoglobulins: recombination in somatic cells. J Immunol 103(1):120–128
Dubiski S (1987) Foreword. In: Dubiski S (ed.) The rabbit in contemporary immunological research. Longman Scientific & Technical, Wiley pp. ix–xii
Dubiski S, Dudziak Z, Skalba D (1959) Serum groups in rabbits. Immunology 2(1):84–92
Esteves PJ (2003) Molecular and population genetic analysis of polymorphism at the antibody loci IgGCH2 and IgVH in lagomorphs Faculdade de Ciências Universidade do Porto, Porto
Esteves PJ, Alves PC, Ferrand N, van der Loo W (2002a) Hotspot variation at the CH2-CH3 interface of leporid IgG antibodies (Oryctolagus, Sylvilagus and Lepus). Eur J Immunogenet 29(6):529–535
Esteves PJ, Alves PC, Ferrand N, van der Loo W (2002b) Restriction fragment alleles of the rabbit IGHG genes with reference to the rabbit IGHGCH2 or e locus polymorphism. Anim Genet 33(4):309–311
Esteves PJ, Lanning D, Ferrand N, Knight KL, Zhai SK, van der Loo W (2004) Allelic variation at the VHa locus in natural populations of rabbit (Oryctolagus cuniculus, L.). J Immunol 172(2):1044–1053
Esteves PJ, Lanning D, Ferrand N, Knight KL, Zhai SK, van der Loo W (2005) The evolution of the immunoglobulin heavy chain variable region (IgVH) in leporids: an unusual case of transspecies polymorphism. Immunogenetics 57(11):874–882
Esteves PJ, Carmo C, Godinho R, van der Loo W (2006) Genetic diversity at the hinge region of the unique immunoglobulin heavy gamma (IGHG) gene in leporids (Oryctolagus, Sylvilagus and Lepus). Int J Immunogenet 33(3):171–177
Ferrand N, Branco M (2007) The evolutionary history of the european rabbit (Oryctolagus cuniculus): major patterns of population differentiation and geographic expansion inferred from protein polymorphism. In: Weiss S, Ferrand N (eds) Phylogeography of Southern Europe Refugia. Springer, pp 207–235
Flux JEC, Angermann R (1990) The hares and jackrabbits. In: Chapman JA, Flux JEC (eds) Rabbits, hares and pikas: status conservation action plan. International Union for Conservation of Nature and Natural Resources, Gland, pp 61–94
Gallarda JL, Gleason KS, Knight KL (1985) Organization of rabbit immunoglobulin genes. I. Structure and multiplicity of germ-line VH genes. J Immunol 135(6):4222–4228
Gojobori T, Nei M (1984) Concerted evolution of the immunoglobulin VH gene family. Mol Biol Evol 1(2):195–212
Halanych KM, Robinson TJ (1999) Multiple substitutions affect the phylogenetic utility of cytochrome b and 12S rDNA data: examining a rapid radiation in leporid (lagomorpha) evolution. J Mol Evol 48(3):369–379
Hamers R, Hamers-Casterman C, Lagnaux S (1966) A new allotype in the rabbit linked with Asl which may characterize a new class of IgG. Immunology 10:399–408
Hewitt GM (1996) Some genetic consequences of ice ages, and their role, in divergence and speciation. Biol J Linn Soc 58(3):247–276
Hood L, Campbell JH, Elgin SC (1975) The organization, expression, and evolution of antibody genes and other multigene families. Annu Rev Genet 9:305–353
Horng WJ, Knight KL, Dray S (1976) Heavy chain variable region allotypic sub-specificities of rabbit immunoglobulins. I. Identification of three subpopulations of a1 IgG molecules. J Immunol 116(1):117–125
Horng WJ, Papagiannes E, Dray S, Rodkey LS (1980) Expression of cross-reacting determinants of the immunoglobulin heavy chain variable region a3 allotype in Oryctolagus and Lepus. Mol Immunol 17(1):111–117
Kearney JF, Cooper MD, Lawton AR (1976) B cell differentiation induced by lipopolysaccharide. IV. Development of immunoglobulin class restriction in precursors of IgG-synthesizing cells. J Immunol 117(5 Pt 1):1567–1572
Kelus AS, Gell PG (1967) Imunoglobulin allotypes of experimental animals. Prog Allergy 11:141–184
Kelus AS, Steinberg CM (1991) Is there a high rate of mitotic recombination between the loci encoding immunoglobulin VH and CH regions in gonial cells? Immunogenetics 33(4):255–259
Kelus AS, Weiss S (1986) Mutation affecting the expression of immunoglobulin variable regions in the rabbit. Proc Natl Acad Sci U S A 83(13):4883–4886
Kim BS, Dray S (1972) Identification and genetic control of allotypic specificities on two variable region subgroups of rabbit immunoglobulin heavy chains. Eur J Immunol 2(6):509–514
Kim BS, Dray S (1973) Expression of the a, x, and y variable region genes of heavy chains among IgG, IgM, and IgA molecules of normal and a locus allotype-suppressed rabbits. J Immunol 111(3):750–760
Kindt TJ (1975) Rabbit immunoglobulin allotypes: structure, immunology, and genetics. Adv Immunol 21:35–86
Kirkham PM, Mortari F, Newton JA, Schroeder HW Jr (1992) Immunoglobulin VH clan and family identity predicts variable domain structure and may influence antigen binding. EMBO J 11(2):603–609
Klein J, Ono H, Klein D, O'hUigin C (1993) The accordion model of MHC evolution. Prog immunology 8:137–143
Knight KL (1992) Restricted VH gene usage and generation of antibody diversity in rabbit. Annu Rev Immunol 10:593–616
Knight KL, Becker RS (1990) Molecular basis of the allelic inheritance of rabbit immunoglobulin VH allotypes: implications for the generation of antibody diversity. Cell 60(6):963–970
Knight KL, Crane MA (1994) Generating the antibody repertoire in rabbit. Adv Immunol 56:179–218
Lanning D, Sethupathi P, Rhee KJ, Zhai SK, Knight KL (2000a) Intestinal microflora and diversification of the rabbit antibody repertoire. J Immunol 165(4):2012–2019
Lanning D, Zhu X, Zhai SK, Knight KL (2000b) Development of the antibody repertoire in rabbit: gut-associated lymphoid tissue, microbes, and selection. Immunol Rev 175:214–228
Loor F, Kelus AS (1978) Allelic exclusion in the B lineage cells of the rabbit. Eur J Immunol 8(5):315–324
Lopez-Martinez N (1989) Revisión sistemática y biostratigráfica de los Lagomorpha (Mammalia) del Terciario y Cuatemario de España vol 3. Memorias del Museo Paleontologico de la Universidad de Zaragoza. Diputación General de Aragón
Lopez-Martinez N (2008) The lagomorph fossil record and the origin of the European rabbit. In: Alves PC, Ferrand N, Hackländer K (eds) Lagomorph biology. Springer, Berlin Heidelberg, pp 27–46
Lucier MR, Thompson RE, Waire J, Lin AW, Osborne BA, Goldsby RA (1998) Multiple sites of V lambda diversification in cattle. J Immunol 161(10):5438–5444
Lummus Z, Cebra JJ, Mage R (1967) Correspondence of the relative cellular distribution and serum concentration of allelic allotypic markers in normal and “allotype-suppressed” heterozygous rabbits. J Immunol 99(4):737–743
Mage RG (1979) A new look at the biological and genetic significance of rabbit heavy chain allotypes. Ann Immunol (Paris) 130(2):105–114
Mage R, Young GO, Dray S (1967) An effect upon the regulation of gene expression: allotype suppression at the a locus in heterozygous offspring of immunized rabbits. J Immunol 98(3):502–509
Mage RG, Young-Cooper GO, Alexander C (1971) Genetic control of variable and constant regions of immunoglobulin heavy chains. Nat New Biol 230(10):63–64
Mage RG, Young-Cooper GO, Rejnek J, Ansari AA, Alexander CB, Appella E, Carta-Sorcini M, Landucci-Tosi S, Tosi RM (1976) Rabbit immunoglobulin allotypes: complexities of their genetics, expression, structural basis and evolution. Cold Spring Harbor Symposia on Quantitative Biology 41:677–686
Mage RG, Bernstein KE, McCartney-Francis N, Alexander CB, Young-Cooper GO, Padlan EA, Cohen GH (1984) The structural and genetic basis for expression of normal and latent VHa allotypes of the rabbit. Mol Immunol 21(11):1067–1081
Mage RG, Lanning D, Knight KL (2006) B cell and antibody repertoire development in rabbits: the requirement of gut-associated lymphoid tissues. Dev Comp Immunol 30(1–2):137–153
Margolies MN, Cannon LE, Kindt TJ, Fraser B (1977) The structural basis of rabbit VH allotypes: serologic studies on a1 H chains with defined amino acid sequence. J Immunol 119(1):287–294
Matthee CA, van Vuuren BJ, Bell D, Robinson TJ (2004) A molecular supermatrix of the rabbits and hares (Leporidae) allows for the identification of five intercontinental exchanges during the Miocene. Syst Biol 53(3):433–447
Nei M, Gu X, Sitnikova T (1997) Evolution by the birth-and-death process in multigene families of the vertebrate immune system. Proc Natl Acad Sci U S A 94(15):7799–7806
Ohta T (1983) On the evolution of multigene families. Theor Popul Biol 23(2):216–240
Ota T, Nei M (1994) Divergent evolution and evolution by the birth-and-death process in the immunoglobulin VH gene family. Mol Biol Evol 11(3):469–482
Oudin J (1956) The allotype of certain blood protein antigens. C R Hebd Seances Acad Sci 242(21):2606–2608
Oudin J (1960) Allotypy of rabbit serum proteins. I. Immuno-chemical analysis leading to the individualization of seven main allotypes. J Exp Med 112:107–124
Pages MV (1980) Essai de reconstitution de l'histoire du lapin de garenne en Europe. Bull Mens Off Natl Chasse Sp. Scien. Techn:13–21
Parng CL, Hansal S, Goldsby RA, Osborne BA (1996) Gene conversion contributes to Ig light chain diversity in cattle. J Immunol 157(12):5478–5486
Pospisil R, Young-Cooper GO, Mage RG (1995) Preferential expansion and survival of B lymphocytes based on VH framework 1 and framework 3 expression: “positive” selection in appendix of normal and VH-mutant rabbits. Proc Natl Acad Sci U S A 92(15):6961–6965
Queney G, Ferrand N, Weiss S, Mougel F, Monnerot M (2001) Stationary distributions of microsatellite loci between divergent population groups of the European rabbit (Oryctolagus cuniculus). Mol Biol Evol 18(12):2169–2178
Reynaud CA, Anquez V, Dahan A, Weill JC (1985) A single rearrangement event generates most of the chicken immunoglobulin light chain diversity. Cell 40(2):283–291
Reynaud CA, Dahan A, Anquez V, Weill JC (1989) Somatic hyperconversion diversifies the single Vh gene of the chicken with a high incidence in the D region. Cell 59(1):171–183
Reynaud CA, Mackay CR, Muller RG, Weill JC (1991) Somatic generation of diversity in a mammalian primary lymphoid organ: the sheep ileal Peyer's patches. Cell 64(5):995–1005
Reynaud CA, Garcia C, Hein WR, Weill JC (1995) Hypermutation generating the sheep immunoglobulin repertoire is an antigen-independent process. Cell 80(1):115–125
Ros F, Puels J, Reichenberger N, van Schooten W, Buelow R, Platzer J (2004) Sequence analysis of 0.5 Mb of the rabbit germline immunoglobulin heavy chain locus. Gene 330:49–59
Roux KH (1981) A fourth heavy chain variable region subgroup, w, with 2 variants defined by an induced auto-antiserum in the rabbit. J Immunol 127(2):626–632
Schmale J, Costea N, Dray S, Heller P, Yakulis V (1969) Allelic exclusion of light chain allotypes in rabbit IgM cold agglutinins. Exp Biol Med 130(1):48–50
Schroeder HW Jr, Hillson JL, Perlmutter RM (1990) Structure and evolution of mammalian VH families. Int Immunol 2(1):41–50
Sehgal D, Mage RG, Schiaffella E (1998) VH mutant rabbits lacking the VH1a2 gene develop a2+ B cells in the appendix by gene conversion-like alteration of a rearranged VH4 gene. J Immunol 160(3):1246–1255
Sehgal D, Schiaffella E, Anderson AO, Mage RG (2000) Generation of heterogeneous rabbit anti-DNP antibodies by gene conversion and hypermutation of rearranged VL and VH genes during clonal expansion of B cells in splenic germinal centers. Eur J Immunol 30(12):3634–3644
Sehgal D, Obiakor H, Mage RG (2002) Distinct clonal Ig diversification patterns in young appendix compared to antigen-specific splenic clones. J Immunol 168(11):5424–5433
Severinson E, Bergstedt-Lindqvist S, van der Loo W, Fernandez C (1982) Characterization of the IgG response induced by polyclonal B cell activators. Immunol Rev 67:73–85
Short JA, Sethupathi P, Zhai SK, Knight KL (1991) VDJ genes in VHa2 allotype-suppressed rabbits. Limited germline VH gene usage and accumulation of somatic mutations in D regions. J Immunol 147(11):4014–4018
Sitnikova T, Su C (1998) Coevolution of immunoglobulin heavy- and light-chain variable-region gene families. Mol Biol Evol 15(6):617–625
Smith GP (1974) Unequal crossover and the evolution of multigene families. Cold Spring Harbor Symposia on Quantitative Biology 38:507–513
Smith GP, Hood L, Fitch WM (1971) Antibody diversity. Annu Rev Biochem 40:969–1012
Su C, Nei M (1999) Fifty-million-year-old polymorphism at an immunoglobulin variable region gene locus in the rabbit evolutionary lineage. Proc Natl Acad Sci U S A 96(17):9710–9715
Surridge AK, van der Loo W, Abrantes J, Carneiro M, Hewitt GM, Esteves PJ (2008) Diversity and evolutionary history of the MHC DQA gene in leporids. Immunogenetics 60(9):515–525
Taberlet P, Fumagalli L, Wust-Saucy A-G, Cosson J-F (1998) Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol 7(4):453–464
Teherani J, Mandy WJ (1976a) Constant region IgG allotypes in cottontail rabbits: group E allelic polymorphism. Immunochemistry 13(3):221–227
Teherani J, Mandy WJ (1976b) Constant region IgG allotypes in hares: group e allelic polymorphism. Immunochemistry 13(5):401–406
Teherani J, Capra JD, Aggarwal S, Mandy WJ (1979) Amino acid sequence analysis of group e allotype-related peptides derived from lagomorph IgG. Eur J Immunol 9(9):690–695
Teherani J, Capra JD, Mandy WJ (1982) Amino acid sequence of the CH2 domain from various lagomorph IgGs. Mol Immunol 19(7):841–846
Todd CW (1963) Allotypy in rabbit 19S protein. Biochem Biophys Res Commun 11:170–175
Tonnelle C, Cazenave PA, Brezin C, Moinier D, Fougereau M (1983) Structural correlates to the rabbit immunoglobulin heavy chain a100 allotype. Mol Immunol 20(7):753–761
Tutter A, Riblet R (1989) Evolution of the immunoglobulin heavy chain variable region (Igh-V) locus in the genus Mus. Immunogenetics 30(5):315–329
Vajdy M, Sethupathi P, Knight KL (1998) Dependence of antibody somatic diversification on gut-associated lymphoid tissue in rabbits. J Immunol 160(6):2725–2729
van der Loo W (1987) “Population genetical studies on the adaptive significance of the immunoglobulin allotypes in wild rabbit” In: Dubiski S (Ed) “The rabbit in contemporary immunological research”, Longman Scientific & Technical, John Wiley, pp. 101–126
van der Loo W (1993) Variance analysis of immunoglobulin alleles in natural populations of rabbit (Oryctolagus cuniculus): the extensive interallelic divergence at the b locus could be the outcome of overdominance-type selection. Genetics 135(1):171–187
van der Loo W, De Baetselier P, Hamers-Casterman C, Hamers R (1977) Evidence for quasi-silent germline genes coding for phylogenetically ancient determinants of the rabbit a locus allotypes. Eur J Immunol 7(1):15–22
van der Loo W, Gronowicz ES, Strober S, Herzenberg LA (1979) Cell differentiation in the presence of cytochalasin B: studies on the “switch” to IgG secretion after polyclonal B cell activation. J Immunol 122(4):1203–1208
van der Loo W, Ferrand N, Soriguer RC (1991) Estimation of gene diversity at the b locus of the constant region of the immunoglobulin light chain in natural populations of European rabbit (Oryctolagus cuniculus) in Portugal, Andalusia and on the Azorean Islands. Genetics 127(4):789–799
van der Loo W, Mougel F, Sanchez MS, Bouton C, Castien E, Fonseca A, Ferrand N, Soriguer R, Monnerot M (1999) Cytonuclear disequilibria in wild populations of rabbit (Oryctolagus cuniculus L.) suggest unequal allele turnover rates at the b locus (IGKC1). Immunogenetics 49(7–8):629–643
Weinstein PD, Anderson AO, Mage RG (1994) Rabbit IgH sequences in appendix germinal centers: VH diversification by gene conversion-like and hypermutation mechanisms. Immunity 1(8):647–659
Wolf B, Janeway CA Jr, Coombs RR, Catty D, Gell PG, Kelus AS (1971) Immunoglobulin determinants on the lymphocytes of normal rabbits. 3. As4 and As6 determinants on individual lymphocytes and the concept of allelic exclusion. Immunology 20(6):931–944
Zhu X, Boonthum A, Zhai SK, Knight KL (1999) B lymphocyte selection and age-related changes in VH gene usage in mutant Alicia rabbits. J Immunol 163(6):3313–3320
Zimmer EA, Martin SL, Beverley SM, Kan YW, Wilson AC (1980) Rapid duplication and loss of genes coding for the alpha chains of hemoglobin. Proc Natl Acad Sci U S A 77(4):2158–2162
Acknowledgments
This work was supported by grants of Foundation for Science and Technology—Portugal to A.P. and P.J.E. (SFRH/BD/71252/2010 and SFRH/BPD/27021/2006), respectively, and the intramural program of the NIAID, NIH (R.G.M.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pinheiro, A., Lanning, D., Alves, P.C. et al. Molecular bases of genetic diversity and evolution of the immunoglobulin heavy chain variable region (IGHV) gene locus in leporids. Immunogenetics 63, 397–408 (2011). https://doi.org/10.1007/s00251-011-0533-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-011-0533-9