Abstract
The Major Histocompatibility Complex (Mhc) class II DRB locus of vertebrates is highly polymorphic and some alleles may be shared between closely related species as a result of balancing selection in association with resistance to parasites. In this study, we developed a new set of PCR primers to amplify, clone, and sequence overlapping portions of the Mhc class II DRB-like gene from the 5′UTR end to intron 3, including exons 1, 2, and 3 and introns 1 and 2 in four species (20 Humboldt, six African, five Magellanic, and three Galapagos penguins) of penguin from the genus Spheniscus (Sphe). Analysis of gene sequence variation by the neighbor-joining method of 21 Sphe sequences and 20 previously published sequences from four other penguin species revealed overlapping clades within the Sphe species, but species-specific clades for the other penguin species. The overlap of the DRB-like gene sequence variants between the four Sphe species suggests that, despite their allopatric distribution, the Sphe species are closely related and that some shared DRB1 alleles may have undergone a trans-species inheritance because of balancing selection and/or recent rapid speciation. The new primers and PCR assays that we have developed for the identification of the DRB1 DNA and protein sequence variations appear to be useful for the characterization of the molecular evolution of the gene in closely related Penguin species and might be helpful for the assessment of the genetic health and the management of the conservation and captivity of these endangered species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Penguins (order Sphenisciformes, family Spheniscidae) are aquatic, flightless birds with their native habitats located mostly in the Southern Hemisphere between latitudes 45° and 60° S. Phylogenetically, they are closely related to the Procellariiformes order of seabirds such as albatrosses, shearwaters, and petrels (Ho et al. 1976; Sparks and Soper 1987; Hackett et al. 2008). The penguins, based on molecular and morphological phylogenetic evidence and fossil taxa, are believed to have originated in Gondwanaland from an ancestor of the flying seabirds about 71 mya with the emergence of the extant basal lineage Aptenodytes about 40–49 mya (Davis and Darby 1990; Sibley et al. 1988; Baker et al. 2006). The geographical expansion and radiation of most of the present-day penguin species probably did not occur until between 35 and 4 mya (Baker et al. 2006). It was proposed recently on the basis of multiple gene analysis that the modern penguins expanded from Antarctica by way of the circumpolar current to oceanic islands and eventually to the southern continents after Antarctica became ice-encrusted 35–25 mya (Baker et al. 2006), possibly involving two separate dispersals to the equatorial regions (Clarke et al. 2007).
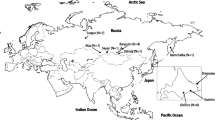
The extant penguins are classified into six genera and 16–18 species (del Hoyo et al. 1992; Jouventin 1982; Baker et al. 2006; Clarke et al. 2007), but inconsistencies and questions still remain about their classification and phylogenetic relationships particularly with respect to the closely related species, such as the Humboldt penguin (Sphe humboldt), the Magellanic penguin (Sphe magellanicus), the African penguin (Sphe demersus), and the Galapagos penguin (Sphe mendiculus). A geographic map of wild penguin location shows that the Sphe species are distributed widely in the subantarctic and their locations range from the southern coastal regions of South Africa to South America, but also as far north as the equatorial Galapagos Islands of the Pacific Ocean (Fig. 1). Although these four species have distinctly different geographic habitats, they still share a number of common traits, including morphological similarities, living in temperate climates and nesting in burrows. They are collectively known as “the banded penguins” because of their similar coloration and external forms. Furthermore, the Sphe appears to be a penguin genus that is capable of interspecies breeding (Thumser and Karron 1994; Williams 1995; McCarthy 2006).
A global distribution map of wild penguins in their natural habitats. This map was modified from a figure by Mackintosh (1960) and Williams (1995). Breeding places of Spheniscus (Humboldt penguin, Magellanic penguin, Galapagos penguin, and African penguin), Pygoscelis (Adelie penguin, chinstrap penguin, and Gentoo penguin), and the little penguin are indicated
The Mhc is a high-variable genomic region within and between different vertebrate species due to duplicated gene clusters involved in the gene control of the immune response and the influence of positive (balancing) selection (Hughes and Nei 1989a, b; Ellis and Ballingall 1999; Ellis 2004). The Mhc DRB-like gene is known to be a useful marker of trans-species allele distribution between closely related species of ungulates, canivores, birds, primates, and various other mammals (Bollmer et al. 2007; Cutera and Lacey 2007). Most research on the vertebrate Mhc class II DRB-like genes has been done on mammals (Trowsdale 1995), and studies on the avian Mhc DRB-like genes have been restricted primarily to gallinaceous birds (Zoorob et al. 1993; Kaufman et al. 1999; Wittzell et al. 1999; Hosomichi et al. 2006; Chaves et al. 2007; Strand et al. 2007), several passerines (Edwards et al. 1995a, b, 1998, 2000; Gasper et al. 2001; Hess et al. 2000; Aguilar et al. 2006), raptors (Alcaide et al. 2008; Burri et al. 2008), a shore bird (Ekblom et al. 2007), and some penguins (Tsuda et al. 2001; Kikkawa et al. 2005; Bollmer et al. 2007).
In this study, we have further examined the Mhc class II DRB-like gene as a potential evolutionary and adaptive molecular marker in four closely related penguin Sphe species. We sequenced 21 new penguin alleles of the Mhc class II DRB-like gene sequence from seven Humboldt, six African, five Magellanic, and three Galapagos penguins and examined their allele distribution and variation. Allele sharing of the DRB-like gene sequences in the genus Sphe emphasizes that they are closely related species with an allopatric distribution and that some common alleles have undergone trans-species inheritance probably because of balancing selection or recent rapid speciation.
Materials and methods
Collection of penguin samples, DNA extraction
Our field collaborators obtained blood samples from 24 captive penguins at aquariums and zoological garden in Japan and Korea and from ten wild penguins in Chile, Argentina, and the Galapagos Islands of Ecuador (Table 1). The 20 Humboldt penguin samples listed in Table 1 are the same as previously described by Kikkawa et al. (2005). A liver sample for RT-PCR was obtained from a captive Humboldt penguin at Oji Zoo in Japan.
DNA was extracted from peripheral blood cells of seven Humboldt penguins (six captive and one wild), six captive African penguins, five Magellanic penguins (three captive and two wild), and three wild Galapagos penguins by the standard phenol–chloroform extraction method after proteinase K treatment (Inoko et al. 1986).
Genomic and RT-PCR
Total RNA was isolated from the liver sample of a captive Humboldt penguin using the TRIZOL reagent method. The cDNA was synthesized using a GeneAmp RNA PCR kit (Applied Biosystems). The primers RT1-ex2F and RT3-ex3R were newly designed for the RT-PCR assay (Fig. 2). These primers were also used to amplify a 568 bp genomic segment of exon 2 to exon 3, including intron 2 of the Humboldt penguin DRB-like gene. The PCR amplification was performed in 20 μl volumes containing 10× Ex Taq buffer, 0.2 mM of each dNTP, 0.5 μM of each primer, 0.5 units Ex Taq (TaKaRa, Shiga, Japan), and 50–100 ng of cDNA or genomic DNA. Amplification included an initial incubation at 96°C for 3 min followed by 35 cycles with each cycle temperature profile consisting of 96°C for 30 s, 60°C for 30 s, 72°C for 1 min, and then a final cycle at 72°C for 5 min. The same DNA and cDNA sample was amplified using b-antin-F and b-actin-R primers (Shimizu et al. 2004) as a control. The PCR products were loaded on a 1.5% agarose gel with TBE buffer and run at 100 V for 45 min at room temperature.
Information and gene map of the PCR oligonucleotide primer locations used to amplify portions of the penguin Mhc class II DRB1-like gene. The previously published five primers (LP1, Lpen.hum1F, Lpen.hum2F, Lpen.hum2R, and Lpen.hum3R; Kikkawa et al. 2005) used for PCR and sequencing are included in the figure. The six newly designed primers for PCR and sequencing (RT1, RT2, Lpen.5′UTR-F, Lpen.hum1F2, Lpen.hum 2F2, and Lpen.hum 3R2) are boxed. The LP1 primer was designed on the basis of exon 1 sequences of birds, but also included a portion of the 5′UTR sequence
Amplification and sequencing
A total of nine PCR primers were used to amplify selected portions of the Sphe DRB1-like gene by PCR for direct sequencing or sequencing after cloning the PCR products. Figure 2 shows the PCR primer sequences, the gene sequence location of the primers relative to 5′UTR and intron 3 of the DRB1 gene, and primer orientation (sense and antisense). The previously designed five primers (Kikkawa et al. 2005) and four newly designed primers (Lpen.5′UTR-F, Lpen.hum.1F2, Lpen.hum.2F2, and Lpen.hum.3R2) were used for PCR amplification.
The DNA sequences from the 5′UTR to intron 2 of the DBR1-like gene were amplified from 34 penguin genomic DNA samples (Table 1) using the previously published LP1 and Lpen.hum 2R primer pairs (Kikkawa et al. 2005). The PCR amplification was performed in 25 μl volumes containing 50–100 ng of genomic DNA, 10× NH4 buffer, 2 mM MgCl2, 0.2 mM of each dNTP, 0.12 μM of each primer, and 0.02 units of BIO Taq (BIO LINE). The PCR product of the expected target length (568 bp) was not obtained successfully with other Taq commercial products, Ampli Taq and Gold Taq (Applied Biosystems). Amplification included 96°C incubation for 3 min followed by 35 cycles with each cycle at 96°C for 30 s, 61°C for 30 s, 72°C for 2 min, and then a final cycle at 72°C for 5 min. Similarly, the 34 penguin genomic DNA samples were used to amplify the DRB1-like sequences of intron 1 to intron 3 using two other primer sets, the previously published Lpen.hum1F and Lpen.hum.3R (Kikkawa et al. 2005) and the new set Lpen.hum.1F2 and Lpen.hum.3R2 (Fig. 2). The PCR condition was the same as described above for the LP1 and Lpen.hum 2R primer pairs, except that only the annealing temperature was changed to 67°C for the Lpen.hum1F2 and Lpen.hum.3R2 primer set.
The nucleotide sequences of the PCR products were determined directly by using the Big Dye Terminator Cycle Sequencing Ready Reaction kit (Applied Biosystems) with the LP1, Lpen primers, and the ABI PRISM 3130 automated sequencer (Applied Biosystems). The PCR products were also subcloned into the plasmid pT7Blue T vector (Novagen, CA, USA) for sequencing using the Ligation convenience kit (NIPPON GENE, Tokyo, Japan) and introduced by transformation into the competent cell DH5a (TOYOBO, Osaka, Japan). At least ten subclones per PCR product were used to determine their nucleotide sequences with all ten primers (Fig. 2).
Sequence data analysis
All sequences were assembled using a SEQUENCHER and analyzed using the GENETYX MAC software package for Apple computers. The gene trees were constructed using the neighbor-joining method with the Kimura 2 parameter settings provided by the MEGA 2 software. Sequences that were homologous to our sequences were found in the GenBank database (www.ncbi.nlm.nih.gov) by using online BLAST searches. We submitted our sequences to DDBJ and obtained accession numbers as part of the international nucleotide sequence database collaboration with EBI/EMBL and NCBI/GenBank (www.ddbj.nig.ac.jp/Welcome-j.html).
Southern hybridization
Genomic DNA (15 μg) was digested with 60 U of the restriction enzyme PstI (NEB, England) and electrophoresed on 0.8% TAE agarose gel. The DNA was transferred from the agarose gel to Hybond-N+ membrane (Amersham, England) for hybridization with the Lpen.hum1F/2R PCR product and RT1/RT2 PCR product used as probes. The probes were labeled using the PCR DIG Probe synthesis kit (Roche, Germany). The hybridizations were detected by using AlkPhos Direct Labelling and Detection System with CDP-Star (Amersham, England).
Results and discussion
Genomic- and RT-PCR and Southern hybridization
The newly designed primers RT1-ex2F and RT3-ex3R (Fig. 2) were used successfully in a separate RT-PCR and genomic PCR assay to amplify a 313 bp cDNA and a 568 bp genomic segment from the expressed liver cDNA and genomic fragment of the Humboldt penguin DRB-like gene, respectively (Fig. 3). Sequencing data of the PCR products (not shown) revealed that the 313-bp cDNA band expressed by the liver represented the nucleotide sequence of exon 2 and exon 3 of the DRB-like gene, whereas the 568 bp single band represented exon 2 to exon 3, including intron 2 that was amplified from the Humboldt penguin DRB-like gene of the genomic DNA extracted from the liver. The homogeneity of the direct sequencing results (data not shown) showed that these and other primers (Kikkawa et al. 2005) amplified sequences from a single encoding DRB1-like gene and not multiple heterologous DRB-1-like gene sequences that may or may not be present in penguins because of DRB1 gene duplications as in chickens and other birds (Shiina et al. 2004). Indeed, single band was obtained as a result of Southern hybridization (Fig. 4).
Gene expression analysis by RT-PCR. The DNA and cDNA isolated from the liver of a Humboldt penguin were amplified using the RT1 and RT3 primer pair and b-actin primers as control. The RT1 and RT3 primer pair amplified a PCR product of 568 bp using genomic DNA in a standard PCR and 313 bp using cDNA in a RT-PCR. The b-actin primer pair amplified a product of 887 bp using genomic DNA in a standard PCR and 353 bp using cDNA in a RT-PCR
Nucleotide and amino acid sequence diversity and allele frequency variation
About 1.5 kb of nucleotide sequence from the 5′UTR of exon 1 to intron 3 of the DRB1-like gene was determined for 34 genomic DNA samples from the four Sphe species (Table 2). A comparison of nucleotide sequence differences detected within intron 1, exon 2, intron 2, exon 3, and intron 3 of the 22 haplotypes of the DRB1-like gene of the Galapagos penguin, Humboldt penguin, African penguin, and Magellanic penguin is shown in Fig. 5. All four species had the identical nucleotide sequences at exon 1 (70 bp). The three alleles Sphu 001, 002, and 0011 of the Humboldt penguins had deletions at nucleotide position 94 in intron 1 and a “C” stretch of >11 bp (351 to 361 bp or 352 to 362 bp) where the precise copy number of cytosines was unknown. Thus, the complete lengths of intron 1 for the Humboldt penguins were estimated to be about 514 or 515 bp. There were eight or nine deletion sites within intron 1 of the Magellanic penguins, but no evidence of frameshift mutations or premature stop codons in any of the samples that we sequenced. The number of different alleles that we identified was eight for the Humboldt penguin, four for the African penguins, eight for the Magellanic penguins, and two for the Galapagos penguins. However, the sequences of the Sphu004 allele (AB301950) in a Humboldt penguin and the Spma001 allele (AB302843) in a Magellanic penguin were identical and varied by only three nucleotides from the Spma007 allele (AB303945) of a Magellanic penguin. A 198-bp portion of the exon 2 sequences for alleles Spma001, Spma004, Spme001, and Spme002 have been previously published as accession numbers (alleles) EF212010 (Spma1), EF212011 (Spma2), EF212008 (Spme2), and EF212007 (Spme1) by Bollmer et al. (2007). Overall, the nucleotide differences between the four penguin species varied from 1 (0.07%) to 27 (1.83%).
Comparison of nucleotide sequences obtained for exon 1, intron 1, exon 2, intron 2, and exon 3 of 22 haplotypes of the DRB1-like gene of the Spheniscus species (Humboldt penguin, African penguin, and Magellanic penguin, Galapagos penguin). This figure showed only variation sites, sequence sites number (bp) above the Spme001 sequence, and determined sequence length of each gene allele on the right side. There was no variation site on exon 1 sequence. Asterisks of position 94 on intron 1 indicate the deletion sites. The number and percentage difference of nucleotides for each gene allele relative to the Sphu001 sequence is listed in the column on the right
Figure 6 shows a comparison of the amino acid sequences translated from the nucleotide sequences of exon 2 for 22 allele variants of the DRB-like gene of Sphe and the DRB1*0101 allele sequence of the human that was included here as an outgroup reference. Three hypervariable regions (HV1 to HV3) were identified in the penguin sequences that corresponded to the well-defined hypervariable regions within the human DRB1*0101 sequence (Stern et al. 1994; Brown et al. 1993). We obtained identical amino acid sequences in exon 2 of the Magellanic penguin alleles Spma001 and Spma007 and the Humboldt penguin allele Sphu004.
Comparison of amino acid sequences obtained for exon 2 of 22 alleles of the DRB-like gene of Spheniscus (Humboldt penguin, African penguin, Magellanic penguin, and Galapagos penguin). The bottom sequence is the DRB1*0101 allele sequence of human. The vertical boxed columns indicate the hypervariable regions (hypervariable region I (HV1), hypervariable region II (HV2), and hypervariable region III (HV3)) for human. The number and percentage difference of amino acid sites for each allele relative to the Sphu001 amino acid sequence is listed in the column on the right. Sphu004 and Spma001 have the same nucleotide sequence
The number and percentage difference of the amino acids within the different haplotype sequences of Sphe varied from 4 (4.5%) to 15 (16.9%). In addition, the base difference between alleles of exon 2 (270 bp) within each of the Sphe species and the human DRB1 alleles was calculated. The average value of the base difference between Japanese high frequency DRB1 alleles (Hashimoto et al. 1994) was 21.2 bp (maximum 37 bp and minimum 1 bp), whereas it was 15.1 bp (21 and 5 bp) for the Magellanic penguin and 13.6 bp (19 and 1 bp) and 20.3 bp (28 and 8 bp) for the African penguin.
The allele percentage frequencies of the DRB-like gene of ten wild and 24 captive Sphe ranged between 2.5% and 75% and are shown in Table 2. The greatest number of alleles (eight each) was for the Humboldt and Magellanic penguins that both reside naturally on the southern coasts of South America (Fig. 1). Four of the five Magellanic penguins and 50% of the 20 Humboldt penguins were heterozygous when applying our PCR conditions. The homozygosity for our set of primers was high at 75% and 83% for the six African penguins and three Galapagos penguins, respectively, although there is a possibility that our PCR assay did not amplify all allelic forms and that some of these homozygotes may in fact be heterozygotes.
The sample numbers in our study are too low to be representative of actual native population frequencies, which still need to be determined for all penguin species. Therefore, we cannot conclude with certainty whether there is an unnatural increase in the homozygosity of the DRB1 gene in any of the Sphe species and whether there are fewer sequence variations within these species that are due in part to population or environmental factors or an increase in inbreeding activity. Because of the absence and/or small sample sizes for these species, we may have underestimated the actual number of DRB1 alleles and the genetic diversity among the study taxa. However, our results are consistent with the findings of Bollmer et al. (2007) who recently reported a low genetic variation for exon 2 at the DRB1 locus of 30 Galapagos penguins. Taken together, these findings confirm that the polymorphic DRB1 gene is an important marker for assessing the genetic health of these endangered species and its genetic variations may have an application for conservation management of wild animals and the efficient breeding management of captive penguins.
Nonsynonymous and synonymous nucleotide substitution rates at exons 2 and 3
The detection of nonsynonymous and synonymous substitution ratios within the Mhc class II amino acid sequences with values of greater than one provides strong evidence for positive or balancing selection acting on the gene sequence. The nonsynonymous (d N) and synonymous (d S) substitution rates and ratios for exon 2 (90 AA) and 3 (94 AA) of DRB-like gene for each species of Sphe are shown in Table 3. The d N/d S ranged in value from 11 to possibly greater than 16 for the region of exon 2 for all four species of Spheniscus, and the ratio was one and three for exon 3 of the Humboldt penguin and the Magellanic penguin, respectively, confirming the affects of balancing selection on the penguin DRB-like gene as previously indicated for exon 2 by Bollmer et al. (2007). The large nonsynonymous substitution rates at exon 2 of the Spheniscus DRB1 gene is consistent with strong positive selection forces acting on the genetic region of antigen presentation in order to cope with various infections and parasites.
Trans-species allele sharing of the penguin DRB1 gene
The reconstruction of a gene tree by the neighbor-joining method of exons 1, 2, and 3 nucleotide sequences (622 bp) of 20 DRB-like gene sequences from the four species of Spheniscus and one chicken Mhc class II beta chain sequence (chosen as the outgroup) is shown in Fig. 7. The gene tree branching has revealed allele sharing between the Humboldt penguin and the Magellanic and Galapagos penguins and some allelic overlaps between the African penguin and the Galapagos and Magellanic penguin sequences. The Humboldt penguin sequence Sphu004 that is identical to the Magellanic penguin sequence Spma001 clustered with other Magellanic penguin sequences Spma006 and Spma007. The low bootstrap percentages (<70%) seen on the tree branches highlight the uncertainty of the branching nodal origins.
Rooted gene tree of complete exons 1, 2, and 3 sequences (622 bp) for the DRB-like gene for four species of Spheniscus. Gene tree analysis was performed by the neighbor-joining method. The chicken Mhc class II beta chain sequence was used as an outgroup (Xu et al. 1989) to root the tree. The numbers on the branches show the bootstrap values as a percentage calculated after 1,000 replications. Allele names of Spheniscus were indicated as Sphu for the Humboldt penguin, Spma for the Magellanic penguin, Spde for the African penguin, and Spme for the Galapagos penguin
In order to view the extent of the trans-species polymorphisms of the DRB-like gene alleles in penguins, we used the neighbor-joining method to reconstruct an unrooted gene tree of nucleotide sequences (198 bp) from exon 2 for 21 representative DRB-like gene sequences from the four species of Spheniscus and 20 previously reported orthologous sequences from four other penguin species, the little penguin (Eudyptula minor), the Adelie penguin (Pygoscelis adeliae), the chinstrap penguin (Pygoscelis antartica), and the Gentoo penguin (Pygoscelis papua). The unrooted gene tree is shown in Fig. 8, and it can be seen that the gene sequences formed into distinct clades for all the species, except for the Spheniscus species. The gene alleles obtained from the genus Spheniscus formed a separate clade to Pygoscelis and Eudyptula, but the gene alleles obtained from the four species within the genus Spheniscus grouped into intermingling clusters. The gene tree also shows that the DRB1 alleles of the little penguin genus Eudyptula is intermediate between the sequences obtained from the genera Spheniscus and Pygoscelis.
Unrooted gene tree of the DRB-like gene partial exon 2 sequences (198 bp) for the four species of Spheniscus and the orthologous sequences for four other penguin species taken from Tsuda et al. (2001). Gene tree analysis was performed by the neighbor-joining method. Allele names of Spheniscus are indicated as Sphu for the Humboldt penguin, Spma for the Magellanic penguin, Spde for the African penguin, and Spme for the Galapagos penguin. The abbreviations Gentoo_N and Gentoo_S refer to the Gentoo penguins of the northern and southern Antarctic regions
The sharing of identical Mhc class II alleles has been documented in primates (Kenter et al. 1992; Doxiadis et al. 2006; Otting et al. 2002), the European bison (Radwan et al. 2007), and in the Humboldt and Magallenic penguins (Bollmer et al. 2007 and present study). We found that the allele sequences, named Sphu004 for the Humboldt penguin and Spma001 for the Magellanic penguin, were identical from the 5′UTR end to intron 3 end of the DRB1 gene. These identical allelic sequences were homozygotes in a captive Humboldt and a Magellanic penguin, but their frequency in the wild penguin populations is not known. According to Bollmer et al. (2007), the identical alleles may have arisen because of an ancestral hybridization event as the distribution of these two species overlap in the wild.
Although the sharing of DRB1 gene variants between species may imply interspecific hybridization between members of different species despite their allopatric isolation, recent rapid speciation events, gene conversion, or maintenance of trans-species alleles because of balancing selection provide alternative explanations (Meyer-Lucht et al. 2008). Allele or haplotype sharing between species suggests an inherent closeness and recent speciation or divergence events. Speciation within Spheniscus based on the reconstructed phylogeny of nuclear and mitochondrial DNA sequences is considered to have occurred mainly within the last 4 myrs with the African and Magellanic penguin sister groups and the Galapagos and Humboldt penguin sister groups emerging on the islands and coastal land masses of the Atlantic Ocean and Pacific Ocean, respectively (Fig. 1), well after the Spheniscus and Eudyptula split about 25 mya (Baker et al. 2006). Therefore, recent speciation is a reasonable explanation for the trans-species allele sharing of the DRB1 genes within Spheniscus, particularly if speciation was completed before fixation of the segregating alleles. With recent, rapid speciation events, closely related species can be expected to share alleles or sequences more frequently than more distantly related species as supported by the results of our reconstructed gene tree for the DRB1 sequences of the four closely related Spheniscus species and the more distantly related four Eudyptula and Pygoscelis species.
Numerous previous studies have implicated balancing selection in the retention of Mhc alleles between closely related species for much longer periods than expected under neutral evolution due to a population's interaction with infectious agents (Klein 1987; Takahata 1993; Cutrera and Lacey 2008). However, it is often difficult to distinguish between the effects of balancing selection, gene conversion, and rapid speciation on allele retention between species, which in reality probably depends on one or all of these processes to some degree (Meyer-Lucht et al. 2008).
Conclusion
We have developed a PCR amplification method to sequence RT-PCR products of the expressed DRB1 gene and overlapping portions of the DRB-like gene from the end of the 5′UTR to intron 3, including exons 1, 2, and 3 and introns 1 and 2 in four species of Spheniscus. These are the longest Mhc class II B gene sequences (946 to 1,586 bp) obtained so far in a study of Spheniscus. In the sequence analysis, we confirmed that there was little nucleotide sequence variation within the exon 1 and exon 3 regions of the gene, but that there were many polymorphic or variable sites and a high rate of nonsynonymous substitutions within exon 2 that corresponds to the hypervariable regions of the human DRB1 gene. It is known that the hypervariable regions of the human DRB1 gene are binding sites that interact with a variety of short antigenic peptide sequences involved in the adaptive immune regulatory system (Klein 1986). In addition, balancing selection is believed to contribute to the allele sequence variation of the DRB1 genes at Mhc loci where the rates of nonsynonymous base pair substitutions are usually greater than expected for neutral loci (Hughes and Nei 1988, 1989a, b). On the basis of these analyses and the comparison of the Spheniscus sequences to the human DRB1 gene, we conclude that the penguin sequences using our PCR amplification method represent a classical single copy class II gene of the Mhc. The specificity of our PCR method however does not exclude the possibility that penguins have duplicated copies of the DRB1-like sequence that have yet to be identified using other detection methods. The reconstruction of the penguin DRB1 gene tree could be greatly improved in future studies by incorporating greater sample numbers and providing a more representative analysis of all 16–18 extant penguin species. The addition of the DRB1-like sequence to other gene sequences with neutral evolutionary histories for a multigenic approach to molecular phylogeny also will be important in further analysis and assessments of penguin evolutionary history and Mhc molecular dating, but possibly not as a reliable marker for the phylogenetic classification of penguins.
References
Aguilar A, Edwards SV, Smith TB, Wayne RK (2006) Patterns of variation in MHC class II beta loci of the little greenbul (Andropadus virens) with comments on MHC evolution in birds. J Hered 97:133–142. doi:10.1093/jhered/esj013
Alcaide M, Edwards SV, Negro JJ, Serrano D, Tella JL (2008) Extensive polymorphism and geographical variation at a positively selected MHC class II B gene of the lesser kestrel (Falco naumanni). Mol Ecol 17:2652–2665. doi:10.1111/j.1365-294X.2008.03791.x
Baker AJ, Pereira SL, Haddrath OP, Edge KA (2006) Multiple gene evidence for expansion of extant penguins out of Antarctica due to global cooling. Proc Biol Sci. Jan 7273:11–17
Bollmer JL, Vargas FH, Parker PG (2007) Low MHC variation in the endangered Galapagos penguin (Spheniscus mendiculus). Immunogenetics 59:593–602. doi:10.1007/s00251-007-0221-y
Brown JH, Jardetzky TS, Gorga JC, Stern LJ, Urban RG, Strominger JL, Wiley DC (1993) Three-dimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature 364:33–39
Burri R, Hirzel HN, Salamin N, Roulin A, Fumagalli L (2008) Evolutionary patterns of MHC class II B in owls and their implications for the understanding of avian MHC evolution. Mol Biol Evol 25:1180–1191. doi:10.1093/molbev/msn065
Chaves LD, Krueth SB, Reed KM (2007) Characterization of the turkey MHC chromosome through genetic and physical mapping. Cytogenet Genome Res 117:213–220. doi:10.1159/000103182
Clarke JA, Ksepka DT, Stucchi M, Urbina M, Giannini N, Bertelli S, Narváez Y, Boyd CA (2007) Paleogene equatorial penguins challenge the proposed relationship between biogeography, diversity, and Cenozoic climate change. Proc Natl Acad Sci USA 104:11545–11550
Cutrera AP, Lacey EA (2007) Trans-species polymorphism and evidence of selection on class II MHC loci in tuco-tucos (Rodentia: Ctenomyidae). Immunogenetics 59:937–948
Cutrera AP, Lacey EA (2008) Trans-species polymorphism and evidence of selection on class II MHC loci in tuco-tucos (Rodentia: Ctenomyidae). Immunogenetics 59:937–948
Davis LS, Darby JT (eds) (1990) Penguin biology. Academic, San Diego
del Hoyo J, Elliott A, Sargatal J (eds) (1992) Handbook of the birds of the world, vol 1, Ostrich to Ducks. Lynx Editions, Barcelona
Doxiadis GG, Rouweler AJ, de Groot NG, Louwerse A, Otting N, Verschoor EJ, Bontrop RE (2006) Extensive sharing of MHC class II alleles between rhesus and cynomolgus macaques. Immunogenetics 58:259–268
Edwards SV, Grahn M, Potts WK (1995a) Dynamics of MHC evolution in birds and crocodilians: amplification of class II genes with degenerate primers. Mol Ecol 4:719–729. doi:10.1111/j.1365-294X.1995.tb00272.x
Edwards SV, Wakeland EK, Potts WK (1995b) Contrasting histories of avian and mammalian MHC genes revealed by class II sequences from songbirds. Proc Natl Acad Sci USA 92:12200–12204. doi:10.1073/pnas.92.26.12200
Edwards SV, Gasper J, March M (1998) Genomics and polymorphism of Agph-DAB1, an MHC class II gene in red-winged blackbirds (Agelaius phoecineus). Mol Biol Evol 15:236–250
Edwards SV, Gasper J, Garrigan D, Martindale D, Koop BF (2000) A 39-kb sequence around a blackbird MHC class II gene: ghost of selection past and songbird genome architecture. Mol Biol Evol 17:1384–1395
Ekblom R, Saether SA, Jacobsson P, Fiske P, Sahlman T, Grahn M, Kalas JA, Hoglund J (2007) Spatial pattern of MHC class II variation in the great snipe (Gallinago media). Mol Ecol Apr 16:1439–1451
Ellis S (2004) The cattle major histocompatibility complex: is it unique? Vet Immunol Immunopathol 102:1–8
Ellis SA, Ballingall KT (1999) Cattle MHC: evolution in action? Immunol Rev 167:159–168
Gasper JS, Shiina T, Inoko H, Edwards SV (2001) Songbird genomics: analysis of 45 kb upstream of a polymorphic MHC class II gene in redwinged blackbirds (Agelaius phoeniceus). Genomics 75:26–34. doi:10.1006/geno.2001.6596
Hackett SJ, Kimball RT, Reddy S, Bowie RC, Braun EL, Braun MJ, Chojnowski JL, Cox WA, Han KL, Harshman J, Huddleston CJ, Marks BD, Miglia KJ, Moore WS, Sheldon FH, Steadman DW, Witt CC, Yuri T (2008) A phylogenomic study of birds reveals their evolutionary history. Science 320:1763–1768
Hashimoto M, Kinoshita T, Yamasaki M, Tanaka H, Imanishi T, Ihara H, Ichikawa Y, Fukunishi T (1994) Gene frequencies and haplotypic associations within the HLA region in 916 unrelated Japanese individuals. Tissue Antigens 44:166–173
Hess CM, Gasper J, Hoekstra HE, Hill CE, Edwards SV (2000) MHC class II pseudogene and genomic signature of a 32-kb cosmid in the house finch (Carpodacus mexicanus). Genome Res 10:613–623. doi:10.1101/gr.10.5.613
Ho CY, Prager EM, Wilson AC, Osuga DT, Feeney RE (1976) Penguin evolution: protein comparisons demonstrate phylogenetic relationship to flying aquatic birds. J Mol Evol 8:271–282
Hosomichi K, Shiina T, Suzuki S, Tanaka M, Shimizu S, Iwamoto S, Hara H, Yoshida Y, Kulski JK, Inoko H, Hanzawa K (2006) The major histocompatibility complex (Mhc) class IIB region has greater genomic structural flexibility and diversity in the quail than the chicken. BMC Genomics 7:322
Hughes AL, Nei M (1988) Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 352:167–170. doi:10.1038/335167a0
Hughes AL, Nei M (1989a) Nucleotide substitutions at major histocompatability complex class II loci: evidence for overdominant selection. Proc Natl Acad Sci USA 86:958–962. doi:10.1073/pnas.86.3.958
Hughes AL, Nei M (1989b) Evolution of the major histocompatibility complex: independent origin of nonclassical class I genes in different groups of mammals. Mol Biol Evol 6:559–579, Review
Inoko H, Ando A, Ito M, Tsuji K (1986) Southern hybridization analysis of DNA polymorphisms in the HLA-D region. Hum Immunol 16:304–312. doi:10.1016/0198-8859(86) 90058-3
Jouventin P (ed) (1982) Visual and vocal signals in penguins, their evolution and adaptive characters. Parey, Berlin
Kaufman J, Milne S, Gobel TWF, Walker BA, Jacob JP, Auffray C, Zoorob R, Beck S (1999) The chicken B locus in a minimal-essential major histocompatibility complex. Nature 401:923–925. doi:10.1038/44856
Kenter M, Otting N, Anholts J, Leunissen J, Jonker M, Bontrop RE (1992) Evolutionary relationships among the primate Mhc-DQA1 and DQA2 alleles. Immunogenetics 36:71–78. doi:10.1007/BF00215282
Kikkawa EF, Tsuda TT, Naruse TK, Sumiyama D, Fukuda M, Kurita M, Murata K, Wilson RP, LeMaho Y, Tsuda M, Kulski JK, Inoko H (2005) Analysis of the sequence variations in the Mhc DRB1-like gene of the endangered Humboldt penguin (Spheniscus humboldti). Immunogenetics 57:99–107. doi:10.1007/s00251-005-0774-6
Klein J (1986) Natural history of the histocompatibility complex. Wiley, New York
Klein J (1987) Origin of the major histocompatibility complex polymorphism: the trans-species hypothesis. Hum Immunol 19:155–162. doi:10.1016/0198-8859(87) 90066-8
Mackintosh NA (1960) The pattern of distribution of Antarctic fauna. Proc R Soc Lond B Biol Sci 152:624–631
McCarthy EM (2006) Handbook of avian hybrids of the world. Oxford University US Press, Oxford, pp 194–195
Meyer-Lucht Y, Otten C, Püttker T, Sommer S (2008) Selection, diversity and evolutionary patterns of the MHC class II DAB in free-ranging Neotropical marsupials. BMC Genet 9:39
Otting N, de Groot NG, Doxiadis GG, Bontrop RE (2002) Extensive Mhc-DQB variation in humans and non-human primate species. Immunogenetics 54:230–239
Radwan J, Kawałko A, Wójcik JM, Babik W (2007) MHC-DRB3 variation in a free-living population of the European bison, Bison bonasus. Mol Ecol 16:531–540
Shiina T, Shimizu S, Hosomichi K, Kohara S, Watanabe S, Hanzawa K, Beck S, Kulski JK, Inoko H (2004) Comparative genomic analysis of two avian (quail and chicken) MHC regions. J Immunol 172:6751–6763
Shimizu S, Shiina T, Hosomichi K, Takahashi S, Koyama T, Onodera T, Kulski JK, Inoko H (2004) MHC class IIB gene sequences and expression in quails (Coturnix japonica) selected for high and low antibody responses. Immunogenetics 56:280–291
Sibley CG, Ahlquist JE, Monroe BL Jr (1988) A classification of the living birds of the world based on DNA-DNA hybridization studies. Auk 105:409–423
Sparks J, Soper T (1987) Penguins. Facts On File, New York
Stern LJ, Brown JH, Jardetzky TS, Gorga JC, Urban RG, Strominger JL, Wiley DC (1994) Crystal structure of the human class II MHC protein HLA-DR1 complexed with an influenza virus peptide. Nature 368:215–221
Strand T, Westerdahl H, Höglund J, Alatalo RV, Siitari H (2007) The Mhc class II of the Black grouse (Tetrao tetrix) consists of low numbers of B and Y genes with variable diversity and expression. Immunogenetics 59:725–734. doi:10.1007/s00251-007-0234-6
Takahata N (1993) Relaxed natural selection in human populations during the Pleistocene. pn. J Genet 68:539–547
Thumser NM, Karron JD (1994) Patterns of genetic polymorphism in five species of penguin. Auk 111:1018–1022
Trowsdale J (1995) “Both man & bird & beast”: comparative organization of MHC genes. Immunogenetics 41:1–17. doi:10.1007/BF00188427
Tsuda TT, Tsuda M, Naruse T, Kawata H, Ando A, Shiina T, Fukuda M, Kurita M, LeMaho I, Kulski JK, Inoko H (2001) Phylogenetic analysis of penguin (Spheniscidae) species based on sequence variation in Mhc class II gene. Immunogenetics 53:712–716. doi:10.1007/s002510100369
Williams TD (1995) The penguins Spheniscidae. Oxford University Press, Oxford
Wittzell H, Bernot A, Auffray C, Zoorob R (1999) Concerted evolution of two MHC class II b loci in pheasants and domestic chickens. Mol Biol Evol 16:479–490
Xu YX, Pitcovski J, Peterson L, Auffray C, Bourlet Y, Gerndt BM, Nordskog AW, Lamont SJ, Warner CM (1989) Isolation and characterization of three class II MHC genomic clones from the chicken. J Immunol 142:2122–2132
Zoorob R, Bernot A, Renoir DM, Choukri F, Auffrey C (1993) Chicken major histocompatability complex class II genes: analysis of interallelic and interlocus sequence variance. Eur J Immunol 23:1139–1145. doi:10.1002/eji.1830230524
Acknowledgments
We are grateful to the JAZGA Penguin TAG (Japan Association of Zoological Gardens and Aquariums, Penguin Taxon Advisory Group) for providing us with the precious penguin samples, especially captive individual shown in Table 1. This study was supported by the High-Tech Research Center Project of the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
The nucleotide sequence and amino acid sequence data reported in this paper have been submitted to the DDBJ database and have been assigned the accession numbers AB301478, AB301944–AB301950, AB302087–AB302090, AB302190–AB302192, AB302843, AB302844, and AB303942–AB303945.
Rights and permissions
About this article
Cite this article
Kikkawa, E.F., Tsuda, T.T., Sumiyama, D. et al. Trans-species polymorphism of the Mhc class II DRB-like gene in banded penguins (genus Spheniscus). Immunogenetics 61, 341–352 (2009). https://doi.org/10.1007/s00251-009-0363-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-009-0363-1