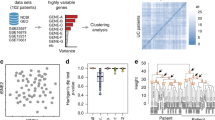
Abstract
An approach based on analysis of variance was applied to raw expression data on 44,760 transcripts in order to identify those with significant differential expression across ileum and colon in Crohn's disease (CD) and ulcerative colitis (UC). The design treated tissue as a block effect, thereby removing this effect statistically and increasing the power to test for effects of disease states (control, CD, and UC). A significant F-statistic for the disease effect was not correlated with the ratios CD/control or UC/control, evidently because many transcripts with high-expression ratios to the control showed inconsistent patterns across tissues. Of 1,053 transcripts showing a significant effect of disease state at the 1% level by the bootstrap test, 508 showed significant difference at the 1% level in a post hoc test for difference between the mean scores for CD and control. These included a number of genes relevant to the mechanism of pathogenesis of CD and a number of genes mapping to genomic regions that have previously shown linkage to CD in association studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Inflammatory bowel disease (IBD), including Crohn's disease (CD) and ulcerative colitis (UC), is characterized by chronic inflammation of the intestine in the absence of an obvious pathogenic cause, and the underlying disease mechanisms remain poorly understood. There is evidence that both genetic and environmental factors play a role in the etiology of IBD (Watts and Satsangi 2002; Bouma and Strober 2003), and that IBD may be a complex of diseases with different etiologies (Gasche et al. 2003). A significant development in recent years has been the discovery of an association between certain polymorphisms at the CARD15/NOD2 locus on chromosome 16 and increased susceptibility to CD (Hugot et al. 2001). This locus, which maps to chromosome 16q12, enodes a protein (CARD15) that uses leucine-rich repeats (LRR) to bind bacterial peptidoglycan and subsequently is involved in the activation of NF-κB Russell et al. 2004). There is evidence of at least six other susceptibility loci for IBD, including one on chromosome 12 (mapped to 12p13.2–q24.1), one on chromosome 19 (mapped to 19p13), one on chromosome 1 (1p36), one on chromosome 5 (5q31), and one on chromosome 14 (mapped to 14q11–q12), as well as the HLA region on chromosome 6 (Cho et al. 2000; Watts and Satsangi 2002; Girardin et al. 2003; van Heel et al. 2005; Negoro et al. 2005).
The analysis of gene expression by techniques such as microarray holds promise for increasing our understanding of both the causes and the pathology of complex diseases such as IBD (Devauchelle et Chiocchia 2004; Dieckgraefe et al. 2000; Heller et al. 1997; Kok et al. 2004; Langmann et al. 2004; Mannick et al. 2004). However, gene expression data pose difficult problems of interpretation and analysis. First of all, gene expression itself is a complex phenomenon, with potential variation arising not only from differences among tissue types and disease states but also from individual genetic differences and environmental effects. In addition, because of the cost of gene expression experiments, a typical microarray data set contains information on the expression levels of numerous transcripts, but usually, the number of replicates is small. Moreover, certain highly expressed transcripts show the most marked expression level differences between disease and normal tissues. Yet expression levels of these highly expressed transcripts may be subject to substantial stochastic error, and thus, the observed differences may not be biologically significant.
One approach to overcoming these problems in microarray data interpretation is to make use of comparisons among different tissues as well as among different states of disease. Using analysis of variance, it is possible to test for differences among disease states controlling statistically for the difference among tissues. Such an approach can be used to detect transcripts which are consistently increased or decreased in a given disease state across tissues. The identification of transcripts showing a consistent pattern across tissues serves to minimize the effects of stochastic variations in the expression of highly expressed transcripts in a given experiment.
Here I apply this approach to analyze data on gene expression in IBD from a published study that focused on dysregulation of pregnane X receptor target genes (Langmann et al. 2004). The data are raw expression scores for both ileum and colon in controls, CD patients, and UC patients. Note that, because UC is a disease of the colon, it was not expected that there would be many transcripts with significant differential expression across both ileum and colon in UC. Nonetheless, the inclusion of data from UC has the desirable property of increasing the power of the statistical analysis, by providing what amounts to an additional control and by increasing the error degrees of freedom for the analysis of variance.
Methods
Raw expression data from microarray experiments were downloaded from the Gene Expression Omnibus (GEO) database (Barrett et al. 2005). A given data set in the GEO database (a GDS record) represents a collection of biologically and statistically comparable samples. Two data sets were used: GDS559, derived from Affymetrix (Santa Clara, CA) GeneChip Human Genome U133 Array Set HG-U133Al and GDS560, derived from Affymetrix GeneChip Human Genome U133 Array Set HG-U133B. These chips provide a broad coverage of transcripts from the human genome. Each set contained measurements for two tissues, terminal ileum and colon transversum, from unaffected controls, from patients with CD, and from patients with UC. For each of the six combinations of tissue and disease state, tissue was obtained by pooling tissue from four donors. GDS559 provided data for 22,283 transcripts, and GDS560 provided data for 22,645 transcripts. Only 168 transcripts were in common between the two data sets. By examination of functional annotations, these 168 transcripts did not appear to be atypical of the data set as a whole. In the case of these 168 transcripts, I averaged the scores for these two data sets. Thus, the final data matrix contained measurements for 44,760 transcripts, providing extensive coverage of well-substantiated human genes.
The 168 transcripts shared between the data sets provided a test for the comparability of the results in the two data sets. For the six combinations of tissue and disease state, the correlations between the raw scores for these 168 transcripts in the two data sets ranged between 0.953 and 0.987 (P<0.001 in all cases). This result supports the hypothesis that experimental conditions in the two data sets were comparable.
For each transcript, analysis of variance was conducted in a block design. The tissue (ileum or colon) constituted the block effect (Supplementary Table S1). I tested for differences between disease states (control, CD, and UC) after removing the effect of difference among tissues. A randomization procedure was used to provide probability levels for F-statistics. Data vectors were generated for 1,000,000 simulated transcripts by sampling (with replacement) from each column of the original data matrix. The F-statistic was then calculated for each simulated transcript, and the distribution of the F-statistics for the simulated transcripts was used as a reference to assess significance of F-statistics computed from the real data. Each F-statistic computed from the real transcripts was considered significant at the α level if 100α% or fewer of the simulated transcripts showed F-statistics greater than that value. For transcripts showing a significant F-statistic, post hoc comparisons among individual disease state means were conducted by Tukey's honestly significant difference (HSD) method (Sokal and Rohlf 1981). Significance for HSD was also assessed by comparison with those calculated for the simulated transcripts. In order to correct for multiple testing, I applied the step-up false-discovery rate (FDR) method of Benjamini and Hochberg (1995) to both F tests and HSD.
Results
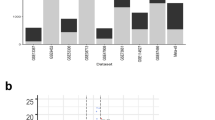
When analysis of variance was applied to expression data for 44,760 transcripts in the two tissues (ileum and colon) and three disease states (control, CD, and UC), there were striking differences among transcripts with respect to the proportion of the total sum of squares accounted for by differences among tissues and the proportion of the total sum of squares accounted for by disease state (Fig. 1). Almost every possible combination of values was seen (Fig. 1). There were transcripts for which disease state accounted for a very high proportion (nearly 100%) of the total sum of squares and tissue accounted for very little of the total sum of squares (Fig. 1). Conversely, there were transcripts for which disease state accounted for very little of the total sum of squares, whereas tissue accounted for a high proportion (Fig. 1). A group of 5,046 transcripts (11.3% of total) showed significant effects of disease state at the 5% level by the F test and a FDR of less than 5%. A group of 1,053 transcripts (2.4%) showed significant effects of disease state at the 1% level and FDR of less than 1%.
In the analysis of variance conducted here, it was not possible to test for interactions of tissue and disease state because of the lack of replication. In order to assess the possible impact of replication on these data, an analysis of variance testing for main effects (tissue and disease state) plus their interaction was applied to the 168 transcripts which were replicated in the GDS559 and GDS560 data sets (see Methods). None showed a significant tissue-by-disease-state interaction at the 5% level. Yet when the values for the two data sets were averaged to provide overall scores for these 168 transcripts, 15 of 168 (9.8%) showed a significant effect of disease state at the 5% level, and 3 (1.8%) showed a significant effect of disease state at the 1% level. These percentages are similar to those for the complete data set of 44,760 transcripts.
Transcripts showing a high value of the F-statistic for disease state did not necessarily show unusually high ratios of the scores for CD or UC to the control. When the F-statistic for disease state was plotted against the natural logarithm of the mean ratio CD/control, there was no correlation (r=0.006; ns; Fig. 2a). Rather, the transcripts with both the highest and lowest log ratios showed very low F-statistics, whereas the transcripts with high F-statistics tended to have log ratios close to zero (Fig. 2a). In the case of UC, there was a small but significant negative correlation between the F-statistic for disease state and the natural logarithm of the mean ratio UC/control (r=−0.016′; P=0.001; Fig. 2b). Here also, transcripts with both very high and very low log ratios tended to have low F-statistics, while those with high F-statistics tended to have log ratios close to zero (Fig. 2b). On the other hand, the natural logarithm of the mean ratio CD/control was highly, positively correlated with the natural logarithm of the ratio UC/control (r=0.559; P<0.001; Fig. 3).
A similar pattern was seen when the logarithm of maximum ratio for the two tissues of the score for CD to that of controls was correlated with the F-statistic; there was no significant linear relationship (r=−0.007; ns). In the case of UC, there was a small but significant negative correlation (r=−0.031; P<0.001) between the logarithm of maximum ratio for the two tissues and the F-statistic. These results show that neither mean nor maximum of the ratio of the scores for either CD or UC to those for controls was a good predictor of the magnitude of the effect of disease state in the analysis of variance.
Within the group of 5,046 transcripts showing a significant effect of disease state at the 5% level and FDR less than 5%, there was a group of 1,647 transcripts showing a significant HSD at the 5% level between means for CD and control and FRD less than 5%. By contrast, there were only 63 transcripts showing a significant HSD at the 5% level between means for UC and control and FDR of less than 5%. Within the group of 1,053 transcripts showing a significant effect of disease state at the 1% level and FDR of less than 1%, 508 showed significant HSD at the 1% level between means for CD and control and FRD of less than 1%. None showed an HSD between UC and control that was significant at the 1% level. Table 1 lists all transcripts with annotated protein function and map location from the RefSeq database (Pruitt et al. 2005) that showed a significant effect of disease state at the 1% level and a significant HSD at the 1% level, with FDR of 1% or less in each case. These included 22 known or putative transcription factors and 12 genes mapping to genomic regions that have shown evidence of association with IBD (Table 1).
Discussion
An approach based on analysis of variance was applied to microarray data from a publicly available database in order to identify transcripts with significant differential expression across ileum and colon in inflammatory bowel disease (IBD). Statistically significant differences in expression levels between Crohn's disease (CD) and control were observed for numerous transcripts. Such differences were more rarely seen in the case of ulcerative colitis (UC), as is expected, since the latter is not expected to affect the ileum. By combining data from two disease states and two tissues, this approach achieved the statistical power to detect transcripts with consistently altered expression across ileum and colon in CD. Furthermore, the analysis of variance design used tissue as a block effect, thereby removing this effect statistically and increasing the power to test for effects of disease states (control, CD, and UC). This approach made it possible to extract information on gene expression changes in CD from a data set lacking independent replicates from CD-affected patients.
The magnitude of the detectable difference among disease states, as measured by the F-statistic for the effect of disease state, was not strongly correlated with the ratio of raw expression scores between CD and control or between UC and control. Rather, the transcripts with the highest F-statistics often had low ratios of disease scores to control scores, and vice versa. This surprising result evidently occurred because many of the transcripts with high ratios of disease to control were transcripts lacking a consistentpattern of expression change in disease state across ileum and colon.
While some of the latter possibly represented genes with a pattern of tissue-specific differential expression in one or both diseases, the available data did not make it possible to test statistically for a tissue-specific expression difference in most cases. On the other hand, in the case of 168 transcripts for which replicated data were available, there were no significant results in tests for tissue-by-disease-state interaction. Yet these 168 transcripts showed significant effects of disease state at rates comparable to the other transcripts, suggesting that they were not atypical of the data set as a whole. The absence of detectable tissue-by-disease-state interactions suggests that inconsistent patterns of expression between the two tissues may often have been due simply to stochastic fluctuations without biological importance.
The analysis of variance identified numerous transcripts with differential expression in CD. These included transcripts from a number of genes with known roles in regulating gene expression in signal transduction and in immune recognition, all processes likely to be involved in CD. The transcripts with significant results at the 1% level and annotated function from the RefSeq database included a number of potential interest for both the mechanism of causation and the pathology of CD (Table 1). These included 22 known or putative transcription factors, among them five zinc finger proteins (Table 1). Eleven of the 22 transcription factors showed significantly higher expression levels in CD than in the control, while 11 showed significantly lower expression levels in CD than in the control (Table 1). Two α-polypeptides from cholinergic receptors showed significantly lower expression in CD than in controls (nos. 75 and 106, Table 1).
Among the most interesting genes in Table 1 were 12 genes mapping to genomic regions that have shown evidence of linkage to IBD (Table 1). Of the seven regions with the strongest association to CD (16q12, 12p13.2–q24.1, 19p13, 1p36, 5q31, 14q11–12, and the HLA region on chromosome 6), all but 16q12 and 14q11–12 are represented by one or more transcripts in Table 1. DPβ1, which was showed significantly increased expression in CD (no. 26, Table 1), maps to the HLA region. 19p13 included three genes with annotated function and significant evidence of differential expression in CD: the zinc finger protein 442 (no. 109, Table 1) and EDG5 (no. 9, Table 1), with increased expression in CD, and CLEC4M (no. 30, Table 1), with decreased expression in CD. EDG5 is a G-protein-coupled receptor involved in cell proliferation (An et al. 2000). CLEC4M forms part of an evolutionarily conserved cluster of type II membrane-associated C-type lectins, belonging to the CD209 family and expressed on dendritic cells (Geijtenbeck et al. 2000; Bashirova et al. 2003).
Table 1 included six genes mapping to the broad region of chromosome 12 (12p13.2–q24.1) that shows association with IBD. Among these was BTG1 (no. 17, Table 1), which has an anti-proliferative function (Iwai et al. 2004) and showed increased expression in CD. Likewise, showing increased expression in CD was HSPC128 (no. 55, Table 1), a transcript identified from hematopoietic stem/progenitor cells (Zhang et al. 2000). TAC3 (no. 89, Table 1) encodes a protein known as tachykinin 3 or neurokinin-B that encode molecules modulating physiological processes via G-protein-coupled receptors (Pal et al. 2004), and this gene also showed evidence of increased expression in CD. Also of interest with regard to chromosome 12 linkage was CLEC4E, which maps to 12p13.31, just outside the region associated with CD. CLEC4E is a close relative of CLEC4M on chromosome 19, but these two C-type lectin genes showed contrasting patterns in CD (Table 1). Whereas CLEC4M showed significantly decreased expression in CD, CLEC4E showed significantly increased expression in CD (Table 1).
Another functionally interesting gene mapping to a region linked with CD (1p36) is TNFRSF4 (no. 18, Table 1), which showed significantly increased expression in CD (Table 1). The protein product, also known as CD134 and OX40, is important for T cell proliferation and is upregulated in multiple sclerosis (Kashiwakura et al. 2004; Carboni et al. 2003). Procadherin beta 13 (no. 87, Table 1), with significantly increased expression in CD, forms part of a cluster in the 5q31 region encoding members of the protocadherin family, which are involved in cell adhesion (Wu et al. 2001).
Current models of CD implicate sensing of peptidoglycan and/or other bacterial cell wall components as a key event in the causation of disease, a line of investigation encouraged by the discovery that CARD15/NOD2 is associated with CD (Girardin et al. 2003). Since CD is believed to be a complex genetic disease with at least seven susceptibility loci (Girardin et al. 2003), genes with differential expression in CD that map to regions previously showing association with CD would seem plausible candidates for further association studies. This would seem to be especially true in the case of genes that play a role in bacterial sensing or the transduction of signals from such sensing. The present analysis revealed a number of genes having these characteristics, and investigation of polymorphism at these loci may yield further insights into the mechanism of causation of CD.
References
An S, Zheng Y, Bleu T (2000) Sphingosine 1-phosphate-induced cell proliferation, survival, and related signaling events mediated by G protein-coupled receptors Edg3 and Edg5. J Biol Chem 275:288–296
Barrett T, Suzek TO, Troup DB, Wilhite SE, Ngau W-C, Ledoux P, Rudnev D, Lash AE, Fijibuchi W, Edgar R (2005) NCBI GEO: mining millions of expression profiles—database and tools. Nucleic Acids Res 33:D562–D566
Bashirova AA, Wu L, Cheng J, Martin TD, Martin MP, Benveniste RE, Lifson JD, KewalRamani VN, Hughes A, Carrington M (2003) Novel member of the CD209 (DC-SIGN) gene family in primates. J Virol 77:217–227
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B 57:289–300
Bouma G, Strober W (2003) The immunological and genetic basis of inflammatory bowel disease. Nat Rev Immunol 3:521–533
Carboni S, Aboul-Enein F, Waltzinger C, Killenn N, Lassmann H, Peña-Rossi C (2003) CD134 plays a crucial role in the pathogenesis of EAE and is upregulated in the CNS of patients with multiple sclerosis. J Neuroimmunol 145:1–11
Cho JH, Nicolae DL, Ramos R, Fields CT, Rabenau K, Corradino S, Brant SR, Espinosa R, LeBeau M, Hanauer SB, Bodzin J, Bonen DK (2000) Linkage and linkage disequilibrium in chromosome band 1p36 in American Chaldeans with inflammatory bowel disease. Hum Mol Genet 9:1425–1432
Devauchelle V, Chiocchia G (2004) Quelle place pour les puces à AND dans les maladies inflammatoires? Rev Med Interne 25:732–739
Dieckgraefe BK, Stenson WF, Korzenik JR, Swanson PE, Harrington CA (2000) Analysis of mucosal gene expression in inflammatory bowel disease by parallel nucleotide arrays. Physiol Genomics 4:1–11
Gasche C, Alizadeh BZ, Peña AS (2003) Genotype–phenotype correlations: how many disorders constitute inflammatory bowel disease? Eur J Gastroenterol Hepatol 15:599–608
Geijtenbeck TB, Terensma R, van Vliet SJ, van Duijnhoven GC, Adema GJ, van Kooyk Y, Figdor CG (2000) identification of DC-SIGN, a novel dendritic cell-specific ICAM-3 receptor that supports primary immune responses. Cell 100:575–585
Girardin SE, Hugot J-P, Sansonetti PJ (2003) Lessons from Nod2 studies: towards a link between Crohn's disease and bacterial sensing. Trends Immunol 24:652–658
Heller RA, Schena M, Chai A, Shalon D, Bedilion T, Gilmore J, Woolley DE, Davis RW (1997) Discovery and analysis of inflammatory disease-related genes using cDNA microarrays. Proc Natl Acad Sci U S A 94:2150–2155
Hugot JP, Chamaillard M, Zouali H, Lesage S, Cezard JP, Belaiche J et al (2001) Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn's disease. Nature 411:599–603
Iwai K, Hirata K, Ishida T, Takeuchi S, Hirase T, Rikitake Y, Kojima Y, Inoue N, Kawashima S, Yokoyama M (2004) An anti-proliferative gene BTG1 regulates angiogenesis in vitro. Biochem Biophys Res Comm 316:628–635
Kashiwakura J, Yokoi H, Saito H, Okayama Y (2004) T cell proliferation by direct crosstalk between OX40 ligand on human mast cells and OX40 on human T cells: comparison of gene expression profiles beetween human tonsillar and lung cultured mast cells. J Immunol 173:5247–5257
Kok K, Stokkers P, Reitsma PH (2004) Genomics and proteomics: implications for inflammatory bowel diseases. Inflamm Bowel Dis 10(Suppl 1):S1–S6
Langmann T, Moehle C, Mauerer R, Scharl M, Liebisch G, Zahn A, Stremmel W, Schmitz G (2004) Loss of detoxification in inflammatory bowel disease: dysregulation of pregnane X receptor target genes. Gastroenterology 127:26–40
Mannick EE, Bonomolo JC, Horswell R, Lentz JJ, Serano M-S, Zapata-Velandia A, Gastanaduy M, Himel JL, Rose SL, Udall JN Jr, Hornick CA, Liu Z (2004) Gene expression in mononuclear cells from patients with inflammatory bowel disease. Clin Immunol 112:247–257
Negoro K, McGovern DPB, Kinouchi Y, Takahashi S, Lench NJ, Shimosegawa T, Carey A, Cardon LR, Jewell DP, van Heel DA (2005) Analysis of the IBD5 locus and potential gene–gene interactions in Crohn's disease. Gut 52:541–546
Pal S, Nemeth MJ, Bodine D, Miller JL, Svaren J, Thein SL, Lowry PJ, Bresnick EH (2004) Neurokinin-B transcription in erythroid cells. J Biol Chem 279:31348–31356
Pruitt KD, Tausova T, Maglott DR (2005) NCBI reference sequence (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res 33:D501–D504
Russell RK, Nimmo ER, Satsangi J (2004) Molecular genetics of Crohn's disease. Curr Opin Genet Dev 14:264–270
Sokal RR, Rohlf FJ (1981) Biometry, 2nd edn. Freeman, San Francisco, CA
Van Heel DA, Fisher SA, Kirby A, Daly MJ, Rioux JD, Lewis CM, Genome Scan Meta-Analysis Group of the IBD International Genetics Consortium (2005) Inflammatory bowel disease susceptibility loci defined by genome scan meta-analysis of 1952 affected relative pairs. Hum Mol Genet 13:763–770
Watts DA, Satsangi J (2002) The genetic jigsaw of inflammatory bowel disease. Gut 50:31–36
Wu Q, Zhang T, Cheng J-F, Kim Y, Grimwood J, Schmutz J, Dickson M, Noonan JP, Zhang MQ, Myers RM, Maniatis T (2001) Comparative DNA sequence analysis of mouse and human protocadherin gene clusters. Genome Res 11:389–404
Zhang Q-H, Ye M, Wu X-Y, Ren S-X, Zhao M, Zhao C-J, Fu G, Shen Y, Fan H-Y, Lu G, Zhong M, Xu X-R, Han Z-G, Zhang J-W, Tao J, Huang Q-H, Zhao J, Hu G-X, Gu J, Chen S-J, Chen Z (2000) Cloning and functional analysis of cDNAs with open reading frames for 300 previously undefined genes expressed in CD34+ hematopoietic stem/progenitor cells. Genome Res 10:1546–1560
Acknowledgements
This research was supported by grant GM43940 from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Hughes, A.L. Consistent across-tissue signatures of differential gene expression in Crohn's disease. Immunogenetics 57, 709–716 (2005). https://doi.org/10.1007/s00251-005-0044-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-005-0044-7