Abstract
Understanding the rules that govern the successions of gut microbiota is prerequisite for testing general ecological theories and sustaining a desirable microbiota. However, the ignorance of microeukaryotes raises the question of whether gut microeukaryotes are assembled according to the same rules as bacteria. We tracked the shrimp gut bacterial and microeukaryotic communities by a longitudinal dense sampling. The successions of both domains were significantly correlated with host age, with relatively stable microeukaryotic communities in adult shrimp. Gut microeukaryotes exhibited significantly higher turnover rate, but fewer transient species, lower proportion of temporal generalists, and narrower habitat niche breadth than bacteria. The γ-diversity partitioning analysis revealed that the successions of gut microbiotas were primarily ascribed to the high dissimilarity as shrimp aged (\( \overline{\beta} \)IntraTimes), whereas the relative importance of \( \overline{\beta} \)IntraTimes was significantly higher for microeukaryotes than that for bacteria. Compared with contrasting ecological processes in governing free-living bacteria and microeukaryotes, the ecological patterns were comparable between host-associated gut counterparts. However, the gut microeukaryotes were governed more strongly by deterministic selection relative to nestedness compared with the gut bacteria, which supports the “size-plasticity” hypothesis. Our results highlight the importance of independently interpreting free-living and host-associated meta-communities for a comprehensive understanding of the processes that govern microbial successions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A fundamental concern in microbial ecology is what processes govern the temporal dynamics of species [1], since knowledge of these processes enables prediction and manipulation of forthcoming states of a community. This is a particular significance for the gut microbial communities, as they contribute indispensable roles in regulating host health, fitness, development, and lifespan [2,3,4,5,6,7]. It has been found that the microbiotas are highly temporal dynamic, regardless of host phylogeny [8,9,10,11,12]. However, due to the variety of gut commensals and the complicated role of host factors [8, 13], an open question remains on how gut commensals colonize and are maintained as host aged.
Gut commensals colonize either from external species in the environment or from internal sources within the same host individual. A few studies have evaluated to what extent rearing water and sediment bacterial communities affects the gut microbiota of aquatic animals, illustrating that hosts source little of their gut commensals from surrounding environments [8, 14]. Building on this, we infer that gut microbes could be derived from its younger host (that is, nestedness). Nestedness is species gain (or loss) from younger to older [15], which often links with ordered extinction–colonization dynamics, such as over host development [16]. Generally, α-diversity in the gut microbiota is significantly lower than that in surrounding environments [17, 18], indicating that the host filters its gut microbial members. In a given community, some members exhibit broad environmental adaptations (generalists), while others show specific and narrow environmental tolerances (specialists) over spatial and temporal heterogeneities [19, 20]. By this logic, specialists tend to be sorted, while generalists are persistent over spatiotemporal successions. However, current estimations of the microbial succession mainly compare communities in a pairwise way and consider all species as equivalent to community composition, while ignore the different categories and traits among species.
To quantify the multiple facets of microbial biodiversity, a quantitative framework has been proposed to partition the dynamics of a meta-community into: local (α), regional (γ), and the overall difference between local communities (β) over space and time [21]. The novelty of this approach lies in the fact that it expresses the three components (α, β, and γ) in the same unit, thus enables the calculation of the relative contributions of α- and β-diversity to the γ-diversity. By comparing these components, it has been exemplified that bacterial communities in fish gut, rearing water, and sediment are functionally redundant, given that the functional diversity of the meta-community primarily arises from local communities [22]. Clearly, β-diversity partitioning moves beyond simple descriptions of community structure towards the inferring of mechanisms that govern community diversity. However, such quantitative partitioning of regional γ-diversity is scarce, especially for host-associated communities over timescales.
Our recent work has shown that the successional pattern of host-associated gut microbiota is significantly different from that of free-living bacterioplankton. Specifically, the gut bacterial communities are governed by species replacement (analogues of temporal turnover that consists in the substitution of species in one time by different species in the subsequent time [15]), while bacterioplanktons are equally shaped by replacement and nestedness [8]. Thus, host-associated and free-living microorganisms are likely governed by differed ecological processes. At this stage, prior meta-community researches have focused exclusively on host-associated bacteria [23, 24]. Consequently, our understanding on gut microeukaryotes is just beginning and lags far behind that on the bacteria. This ignorance is unwarranted because microeukaryotes account for most microbial communities due to their larger size and biomass, thereby could contribute disproportionate roles in gut microbiome [25,26,27]. Indeed, prokaryotes and eukaryotes are markedly different in biological and ecological characters, such as sexual reproduction, low metabolic flexibility, and predatory life strategies of eukaryotes [28]. Furthermore, host elements and microorganisms’ traits could complicate the ecological assembly [13, 29, 30]. Thus, open questions remain on how the host-associated microeukaryotic and bacterial communities are structured or whether the two communities are governed by contrasting rules.
Available gut microbiota studies primarily focus on humans or other vertebrates [9, 10]. Nevertheless, the lack of adaptive immunity in invertebrates doubts the extension of vertebrate gut microbiota to invertebrates [6, 31]. Additionally, the collections of fecal sample and/or partial intestine certainly underestimate gut microbial diversity. By contrast, the small size of shrimp (Litopenaeus vannamei) gut enables us to explore the intact gut microbiotas. To avoid biases from temporally limited sampling, and given the high value of shrimp in global aquaculture, we characterized the shrimp gut bacterial and microeukaryotic communities over an entire culture cycle. In contrast to the unidirectional dispersal of free-living microorganisms over spatial scale, gut commensals are orderly selected as host aged [8, 30]. Additionally, it seems that ecological determinism increases with organism size [32]. For these reasons, we hypothesized that similar ecological processes governed the successions of gut bacteria and microeukaryotes, whereas larger microeukaryotes were more strongly structured by host filtering than smaller bacteria.
Materials and Methods
Experimental Design and Sample Collection
Over the period of 8 April to 15 July, greenhouse-reared shrimp were collected from six identical greenhouse ponds at Ningbo, Eastern China (29° 32′ N, 121° 31′ E). A longitudinal dense sampling (Bi/weekly sampling along shrimp development, 7, 14, 28, 42, 56, 70, 84, 91, and 98 days post larval shrimp inoculation (DPI)) was carried out to explore the temporal dynamics of gut microbiota. Shrimp individuals were aerated in tanks with their rearing water during sampling, and then were immediately transported to the lab. Given that insufficient amount of DNA was extracted from a single intestine of larval shrimp in the initial trial runs, every three shrimp intestines from each pond were aseptically dissected and pooled to compose one biological sample. In total, 54 gut samples were collected, which were stored at – 80 °C prior to DNA extraction.
Water temperature, salinity, pH, and dissolved oxygen (DO) were monitored in situ by using corresponding probes (Oxi 340i; WTW, Weilheim, Germany). The rearing water chemical oxygen demand (COD), total nitrogen (TN), total phosphorus (TP), NO3−, NO2−, NH4+, PO43+ and SiO43+ levels were measured following seawater analysis standard of China [33].
DNA Extraction, Bacterial, and Eukaryotic rRNA Gene Sequencing
Genomic DNA was extracted using the FAST DNA Spin kit (MoBio Laboratories, Carlsbad, CA, USA) following the manufacturer’s protocols. The concentration and purity of DNA extracts were evaluated by using a NanoDrop ND-2000 spectrophotometer (NanoDrop Technologies, Wilmington, USA). The V3–V4 regions of bacterial 16S rDNA [34] and the V4 region of eukaryotic 18S rDNA [35] genes were amplified separately using the same DNA extract for each sample. This design enabled a paired comparison between bacteria and eukaryotes, thereby controlling variations among biological samples. Polymerase chain reaction (PCR) conditions were employed as described elsewhere [17, 25]. To avoid biases introduced during PCR amplification, each sample was amplified in triplicate. Amplicons for each sample were pooled and purified using a PCR fragment purification kit.
Equimolar amounts of the purified amplicons (bacteria or eukaryotes) from each sample were sequenced on a single run using an Illumina MiSeq 2 × 300 bp platform (Illumina, San Diego, CA, USA), generating paired-end reads. Paired-end sequences for bacteria and microeukaryotes were merged respectively with a maximum of 10% mismatches in the overlap region using FLASH [36]. For bacteria communities, the merged sequences were processed following the QIIME (v1.9.0) pipeline [37]. Briefly, the sequences with ambiguous bases or truncated at any site of more than three consecutive bases receiving a Phred quality score (Q) < 25 were excluded. Chimeras were deleted using VSEARCH [38]. Sequences with a distance-based similarity of 97% or greater were grouped into operational taxonomic units (OTUs) using UCLUST [39]. The most abundant sequence from each OTU was selected as its representative, and then was taxonomically assigned an open reference (Silva database, release 132) [40]. Furthermore, OTUs that were assigned to Archaea, chloroplasts, and those unaffiliated with the Bacteria domain, as well as singletons, were discarded from the dataset. Similar procedures were applied for microeukaryotic communities, with the exceptions that OTUs affiliated with Metazoa, unassigned to Eukaryota, and the contamination of shrimp were excluded. To correct for uneven sequencing efforts, the OTU table for bacteria and microeukaryotes was 10 × randomly rarefied subset of 14,000 and 4080 clean reads per sample in subsequent analyses, respectively. The bacterial 16S rRNA and microeukaryotic 18S rRNA sequence data have been deposited in the Sequence Read Archive at DDBJ under accession numbers DRA005256 and DRA005998.
Statistical Analysis
To compare the successional patterns of gut bacteria and microeukaryotes as shrimp aged, ecological approaches were employed to explore the multiple facets of microbial biodiversity (Fig. S1) in R 3.4.1 project, unless otherwise stated [41]. Specifically, a non-metric multidimensional scaling (NMDS) analysis was conducted to visualize the weighted UniFrac distances of microbial communities among samples. Then, a Procrustes analysis was used to detect the correlation between each pair of bacterial and eukaryotic communities [42]. The significance among groups was tested using an analysis of similarity (ANOSIM) [43]. The time-decay of community similarity relationship was plotted as logarithmic similarity against logarithmic age and a linear regression performed to obtain the temporal turnover rate [44]. This form of time lag regression could test whether communities are undergoing directional change. In addition, we treated ponds (the origin of shrimp samples, pond as a conditional factor) as replicates, thus enabled us to test significance (using paired t test to improve statistical power) in turnover rate between bacterial and eukaryotic communities.
Habitat Niche Breadth
Niche breadth is a crucial trait that influences the relative importance of species sorting and temporal nestedness in affecting communities [19]. A wider niche breadth (i.e., generalist) is expected to be more metabolically flexible, which is persistent at the regional community level over time. In contrast, the habitat specialists are more restricted to specific times [45]. Here, the generalists and specialists of the gut bacterial and microeukaryotic communities as shrimp aged were identified respectively based on the Levins’ niche breadth index (B):
Specifically, Bj indicates the niche breadth of OUTj in a community; N is the total number of species across the community; Pij is the relative abundance of OTUj in community i [19]. For a given taxon, it was categorized into specialist or generalist corresponding to B > 3 or B < 1.5. We further calculated the average B values across all taxa in bacterial or microeukaryotic community (Bcom) as an index of its habitat niche breadth [19].
Variation Partitioning Analysis
A forward selection procedure was employed to identify the most important variables in shaping the microbial communities in a distance-based multivariate linear model (DistLM), in which sequentially added one variable that improves the selection criterion (R2) the most at each step, until no improvement was reached in R2 [46, 47]. The key variables that governed both bacterial and eukaryotic communities were selected. Since host age can be used as a proxy for the unmeasured temporal host variables, i.e., immune maturity as host aged [8, 30], a partial distance-based redundancy analysis (partial db-RDA) was used to further evaluate the relative contribution of environmental and temporal host factors on the variation in gut microbiotas [48]. Variation partitioning was carried out to partition the community variation into environmental effects and temporal effects. The pure environmental variation without a temporal component (E|T) represents the strength of species sorting; the pure temporal variation without an environmental component (T|E) is interpreted as the effect of nestedness [8, 19]. To equitably evaluate the relative importance of species sorting versus replacement at different microbial groups, we compared the ratio of (E|T)/(T|E) (that is, species sorting/nestedness effect ratio), instead of their absolute values. The ratio enabled the comparison of the species sorting and nestedness dominances of bacterial and eukaryotic communities in a relative sense [19, 49].
The Rao quadratic entropy (a measure of diversity that integrates species-relative abundances and pairwise interspecies differences) [21] was applied to decompose the (ɤEcosystem) into local diversity (α) and the difference among local communities (β). The advantage of this framework is the calculation of the relative contributions of α- and β-diversity to the γ-diversity [50]. Specifically, regional diversity of microbial community was expressed as the sum of inter-time difference, the mean intra-time difference, and the mean local community diversity with: ɤEcosystem = βInterTimes +\( \overline{\beta} \)IntraTimes + \( \overline{\upalpha} \)LocalCommunities [21]. Furthermore, we quantified the relative importances of the ecological processes: selection, drift, and dispersal using the methodology as proposed before [14]. Though the original definition of these ecological processes arises from “spatial context,” similar concepts have been extrapolated in “temporal” meta-communities as host aged [10, 29]. The framework consists of two main steps: the first step applies phylogenetic turnover between communities to evaluate the contribution of selection, and the second step uses OTU turnover to assess the role of dispersal and drift [14].
Results
Successions of Gut Bacterial and Microeukaryotic Communities as Shrimp Aged
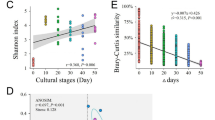
The α-diversity of gut bacterial communities was relatively stable as shrimp aged, whereas microeukaryotic communities exhibited decreasing diversity over the same timeframe (Fig. S2a–d). There was a clear separation of gut bacterial communities according to shrimp age (days post inoculation) (Fig. 1a). This also held true for gut microeukaryotes, with the exception that microeukaryotic communities tended to be stable at the adult stage (i.e., DPI70 and thereafter) (Fig. 1b). This difference was confirmed by the ANOSIM, illustrating that the gut bacterial communities were significantly distinct between every pair of shrimp ages (Table S1), while there were no significant differences in microeukaryotic communities among adults, i.e., DPI70 (70 days post larval shrimp inoculation) vs. DPI 98, and DPI91 vs. DPI 98 (Table S2). The Procrustes analysis revealed that the β-diversity patterns of bacteria and microeukaryotes were moderately correlated (M 2 = 0.598, P = 0.101) (Fig. 1c). Consistently, there was a marginally significant association (Mantel test ρ = 0.316, P = 0.081) between the temporal dynamics of the two domains (Fig. S3). Together, these results depicted comparable β-diversity patterns between gut bacterial and microeukaryotic communities as shrimp aged.
Non-metric multidimensional scaling (NMDS) ordination of the dissimilarity in the gut bacterial (a) and microeukaryotic (b) communities as shrimp aged. Samples are coded by shrimp age (days post inoculation). (c) Procrustes superimposition plot of bacterial (open circles) and microeukaryotic communities (end point of solid lines). Solid lines represent Procrustes residuals from both domains
The host age–decay rate of gut microbiotas was calculated as the slope of a linear regression on the relationship between shrimp age and gut community similarity for bacteria and microeukaryotes, respectively. Both gut bacterial and microeukaryotic communities exhibited significant shrimp age–decay for similarity relationship (Fig. S4). When the age-decay rates were calculated between the first sampling and onwards ones, a paired t tests (N = 6) revealed that the slope of microeukaryotic communities was significantly (P < 0.001) steeper than that of bacterial counterparts (Fig. S4).
Defining the Microbial Community by Frequency
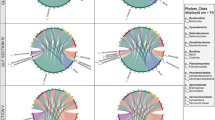
To identify the persistent and transient species, we partitioned ecological communities into their abundance and prevalence. For the prevalence, we arbitrarily divided the microbial community into the following three ecological categories: persistent (≥ 80% analyzed samples over shrimp development), intermittent (ranged 20–80%) and transient (≤ 20%) OTUs (Fig. 2). Positive relations between the mean abundance and prevalence were detected, which were best fitted using the exponential equation: y = 0.0002e0.106x (R2 = 0.722, P < 0.001) for bacteria, and y = 0.0008e0.136x (R2 = 0.628, P < 0.001) for microeukaryotes (Fig. 2a). In this regard, persistent OTUs were more abundant than intermittent and transient OTUs for both domains, although the former ecological categories harbored a much smaller number of OTUs (Fig. 2b). The distribution and frequency were similar between bacterial and microeukaryotic OTUs. That is, persistent bacterial OTUs possessed 0.59% of the total OTUs but accounted for 55.3% bacterial reads. Similarly, only 0.36% of microeukaryotic OTUs occupied 48.5% microeukaryotic sequences (Fig. 2b). Conversely, transient OTUs harbored 93.5% bacterial OTUs but only occupied 13.8% of the total sequences, while 93.4% microeukaryotic OTUs merely accounted for 10.8% (Fig. 2b). Thus, there were highly diverse but minority gut bacterial and microeukaryotic taxa as shrimp aged.
Defining the gut microbial communities as shrimp aged. The left and right panels represent bacterial and microeukaryotic communities, respectively. (a) The mean abundances of OTUs regress against the occurrence frequencies. The prevalence is calculated by dividing the number of samples in which an OTU is detected by the enrolled 52 samples. (b) The number and the relative abundance of OTUs of different occurrence frequencies (persistent, intermittent, and transient) are shown
Partitioning the Microbial Community into Temporal Generalists and Specialists
There were comparable proportions of temporal generalists between bacterial and microeukaryotic communities, 55.7% vs. 47.1%. In contrast, microeukaryotic specialists occupied 23.7% of the total diversity, which was much higher than for bacteria, where they merely accounted for 10.0% (Fig. 3a and b). Consistently, the averaged community niche breadths (Bcom) for microeukaryotic communities were significantly lower than that of bacterial counterparts (Fig. 3c). These results revealed microeukaryotes were more sensitive to host age, as compared with bacteria.
The mean abundances of OTUs of Bacteria (a) or microeukaryotes (b) versus habitat niche breadth. (c) Boxplots illustrating mean habitat niche breadth (Bcom) from all taxa in each sample of bacterial versus microeukaryotic communities. The mean Bcom value for bacteria is significantly higher than that of microeukaryotes
Overall, we found that both the host age and the rearing water variables governed the gut bacterial and microeukaryotic communities (Table S3). For the two total communities, the variations were better explained by the pure environmental component (> 27% variation) than by the pure age effect (< 2% variation). A similar pattern emerged when we focused on the temporal generalists (Table 1). In contrast, a divergent trend was detected when we restricted the analysis to temporal specialists. That is, the pure age effect was a more influential contributor than the environmental component in governing the age specialists (Table 1). Additionally, the species sorting/nestedness effect ratios in microeukaryotes were consistently higher than bacteria for the three sub-communities (Fig. S5).
Partitioning Regional Diversity and Underlying Ecological Processes of Microbial Communities
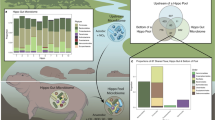
To quantitatively compare the relative contributions of α- and β-diversity to the γ-diversity, we partitioned the γEcosystem of gut microbiotas over temporal scales. The local \( \overline{\upalpha} \)LocalCommunities contributed 35.5% variation in the bacterial communities (Fig. 5b), which was significantly greater (paired t test, P = 0.048) than that in the microeukaryotic communities, \( \overline{\upalpha} \)LocalCommunities = 25.6%. Similarly, the contribution of βInterTimes on bacterial communities (21.6%) was higher than microeukaryotic counterparts, though the difference was insignificant (Fig. 4c). In contrast, the contributions of \( \overline{\beta} \)IntraTimes exhibited the opposing trend (P = 0.039), with \( \overline{\beta} \)IntraTimes = 42.9% in bacterial communities, and 57.6% in microeukaryotic communities (Fig. 5). However, for both domains, the contributions of \( \overline{\beta} \)IntraTimes outweighed (P < 0.001 in both cases) βInterTimes (Fig. 4), suggesting the fundamental role of host age in governing the succession of gut microbiotas.
Partitioning microbial γ-diversity into local α-diversity and the difference among local communities, β-diversity. (a) General outline. The n local communities correspond to the same community sampled along different times. (b) Bacterial communities, (c) Microeukaryotic communities. The total compositional diversity within the shrimp gut was expressed as the sum of inter-time compositional differences (βInterTimes), the mean intra-time compositional difference (\( \overline{\beta} \)IntraTimes), and the mean local compositional diversity (\( \overline{\upalpha} \)LocalCommunities) with ɤEcosystem = βInterTimes +\( \overline{\beta} \)IntraTimes + \( \overline{\upalpha} \)LocalCommunities. * indicates significant of a given biodiversity between bacteria and microeukaryotes by using paired t test at P < 0.05 level
Quantification of the relative roles of selection (homogeneous and heterogeneous), dispersal limitation, homogenizing dispersal and drift in governing the gut bacterial and microeukaryotic communities as shrimp aged. Values indicate percentage of community turnover structured by each ecological process
We further quantified the relative roles of homogeneous or heterogeneous selection, dispersal limitation, homogenizing dispersal, and drift in governing the bacterial and microeukaryotic communities, respectively (Fig. 5). Homogeneous selection was the most important process in governing both bacterial (48.7% of the overall community turnover) and microeukaryotic (62.3%) communities. In contrast, the relative importance of drift exhibited an opposing trend, with 32.7% of the overall community turnover for bacteria, and 16.5% for microeukaryotes (Fig. 5). In both bacteria and microeukaryotes, homogenizing dispersal contributed little to community structures. Additionally, variable selection exerted comparable effects on the two microbial domains (Fig. 5).
Discussions
Contrasting ecological processes in governing the spatial distribution of free-living bacterial and microeukaryotic communities have been widely observed in diverse ecosystems [49, 51, 52]. By contrast, very little is known about the rules that govern gut microbiota as host aged, while this knowledge could guide the design of a desirable gut microbiota. Here, we compared the gut bacterial and microeukaryotic communities during shrimp development. The findings provide new insights into the successions of host-associated communities imposed by biotic and abiotic gradients, i.e., host maturity and rearing water variables, thus allowing community differentiation.
Both the gut bacterial and microeukaryotic communities markedly changed over shrimp development, though the microeukaryotic communities tended to be stable at the adult stage (Fig. 1). Indeed, ample evidence has shown that gut bacterial communities are primarily affected by host age in diverse aquatic animals [10, 18, 29, 30]. By contrast, we know little about the role of host-associated microeukaryotes outside of the view of parasitology [26, 53]. The stable gut microeukaryotic communities could be attributed to the filtering of sensitive members as adult maturity. Consistent with this assertion, gut microeukaryotic diversity linearly decreased as shrimp aged (Fig. S3). Alternatively, the diversification of gut microeukaryotes is delayed as compared with bacteria, a pattern that has been observed in human gut microbiotas [54]. As a consequence, there was an insignificant association between the β-diversity of the two domains (Fig. 1c), which in turn suggests different β-diversity patterns.
To explain this pattern, we compared the temporal turnover for bacteria versus microeukaryotes as shrimp age. We found that microeukaryotes had significantly steeper turnover (more rapid deviation from original to new state) than bacteria (Fig. S4), again indicating that gut microeukaryotes were more sensitive to host development. Why was a higher turnover rate observed for microeukaryotes? A few studies have proposed that body size determines the distance decay of community similarity, while the mechanisms behind them are divergent [32, 55, 56]. One explanation is that the turnover rate is positively associated with body size, a pattern that has been observed over spatial scale [56]. It has been proposed that an increasing temporal turnover compensates a decreasing nestedness fraction [12]. By this logic, bacterial communities are more affected by nestedness process in relation to microeukaryotes. This is in agreement with our analyses of prevalence, of which revealed a lower slope and a more even distribution for bacteria (Fig. 2a). Alternatively, larger organisms (microeukaryotes) are metabolically less flexible to persist in a reversible state [28, 32], thereby limiting their competence in adapting environmental heterogeneity, i.e., increased maturity as host aged [9, 11, 17, 30]. As a result, there was a smaller proportion of transient species in microeukaryotic assemblages than that in bacterial counterparts (Fig. 2b).
Given the unequal tolerance of microbial members to changing environments [19, 20], we further categorized the species into generalists and specialists. Bacteria harbored a higher proportion of generalists and fewer specialists than microeukaryotes (Fig. 3a and b). Consistently, bacteria exhibited significantly wider niche breadths than microeukaryotes (Fig. 3c), which confirms the results from other habitats [20, 49]. In addition, the species sorting/nestedness effect ratios were consistently higher in microeukaryotes than in bacteria, regardless of the total or sub-communities (Fig. S5). Thus, microeukaryotic communities are more structured by species sorting relative to nestedness than bacterial communities. These findings collectively support the “size-plasticity” hypothesis: small-size bacteria are less environment filtered than larger size microeukaryotes [19]. However, the importance of body size does not rule out the influence of other factors in triggering this pattern, such as dormancy, a strategy that is uncommon in microeukaryotes [28]. Notably, both rearing environment and temporal processes (host age) significantly shaped the gut sub-communities. However, which of these processes is more important depends on the degree of habitat specialization (Table 1). At first, it seems surprising that specialists were much less influenced by pure environmental component compared with pure temporal component for both microeukaryotes and bacteria, given previous evidence that environmental specialists with narrower niche breadths were more governed by local conditions [19, 49]. However, considerations of co-evolution hypothesis may explain this apparent paradox [57]. From a host perspective, host should tightly select and reserve suitable commensals, thereby providing immense potential for adaptation to changing conditions, such as diet and external environment [8, 57, 58]. Thus, it is expected that specialists were more strongly governed by temporal process (Table 1). Our recent work shows that gut age-discriminatory taxa could accurately predict shrimp age [29], reinforcing the prominent imprint of host age. By this logic, variations in rearing water along shrimp development could reasonably be expected to affect gut microbiotas (Table 1).
To compare the different facets of community structure, we further partitioned biodiversity into local (α), regional (γ), and the overall difference between local communities (β) [21]. We found that the temporal dynamics of both gut bacteria and microeukaryotes were primarily ascribed to the high microbial dissimilarities along time points (\( \overline{\beta} \)IntraTimes) (Fig. 4), which reinforce again that host age governs the gut microbial assembly. It is known that gut microbiotas exhibit asynchronous alterations in response to host ages and external environmental conditions [6, 17, 30]. To improve host fitness, nestedness between communities ensures that the recruited commensals can thrive and replace less adapted species, as suggested by the hologenome theory of evolution [58]. Along these same lines, and given the higher relative contribution of \( \overline{\beta} \)IntraTimes in microeukaryotes compared with bacteria (Fig. 4), microeukaryotes display less nestedness from horizontally inherited taxa (source from preceding younger host). Owing to the low functional redundancy of microeukaryotes [28], it will be interesting to explore how and to what extent this low nestedness of microeukaryotic components affects host health and lifespan. Conversely, bacterial communities exhibited a significantly higher\( \overline{\upalpha} \)LocalCommunities compared with microeukaryotic ones (Fig. 4), which indicates a highly diversified potential of bacteria for conferring ecological insurance. Consistent with this assertion, it has been shown that the gut bacterial communities have high functional redundancy, with compositional difference but functional similarity in fish [22, 59], and shrimp [60].
Numerous studies have depicted contrasting ecological processes involved in generating the spatial distribution patterns of free-living bacterial and microeukaryotic communities [49, 61]. Whether the successions of host-associated microbial communities are structured by similar rules is an ignored question in microbial ecology. Overall, we found comparable relative importances of ecological processes in governing the two gut microbial communities (Fig. 5), which were distinct from the underlying processes for the biogeography of free-living microorganisms. Homogeneous selection was the primary process governing both the bacterial and microeukaryotic communities, though a higher importance was observed for microeukaryotes (Fig. 5). These results are in line with ones from Wu et al. [49] who showed that protist communities are structured more strongly by species sorting than bacterial counterparts. A possible explanation for this pattern is that the colonization and/or nestedness of gut commensals are directionally selected as the host aged, such as immune maturity [29, 30]. As a consequence, there is a covariation between gut microbiotas and shrimp age, with a given age exerting convergent selection (i.e., homogeneous selection) leading to similar microbial assemblages (Fig. 1). Accordingly, we found a relatively low dissimilarity among communities within each shrimp age (\( \overline{\beta} \)IntraTimes) (Fig. 4). It may seem surprising that variable selection contributed a minor effect on the gut microbiotas (Fig. 5), given previous evidence that suggests rearing conditions markedly affect the gut bacterial communities in aquatic animals [6, 30, 60, 62]. Nevertheless, our recent work has shown that life stage exerts overriding effects on the succession of gut microbiota, instead of an inadequate constraint by water geochemical variables [8]. However, due to the unequal tolerance of gut bacteria and microeukaryotes to host selection, a lower dispersal limitation (here analogue of nestedness) was observed for microeukaryotes (Fig. 5). Together, these findings highlight the importance of shrimp age in contributing divergence of both domains, with eukaryotes being more affected than bacteria.
Conclusions
Our study provides important insights for explaining the succession of gut bacteria and microeukaryotes as host aged. Gut microeukaryotes are governed to a greater extent by host homogeneous selection than bacterial communities, which support the “size-plasticity” hypothesis. This differed patterns could be attributed to differences in \( \overline{\beta} \)IntraTimes (bacteria < microeukaryotes), proportion of temporal generalists, habitat niche breadth, and nestedness (bacteria > microeukaryotes in the three cases) between the two microbial domains. In contrast to free-living microorganisms, different underlying ecological processes but similar successional patterns were detected between gut bacteria and microeukaryotes. Under this scenario, to attain a comprehensive understanding of the processes that govern microbial successions, free-living and host-associated meta-communities should be analyzed and interpreted independently in future ecological studies.
References
Zhou J, Ning D (2017) Stochastic community assembly: does it matter in microbial ecology? Microbiol Mol Biol Rev 81:e00002–e00017
Zhu J, Dai W, Qiu Q, Dong C, Zhang J, Xiong J (2016) Contrasting ecological processes and functional compositions between intestinal bacterial community in healthy and diseased shrimp. Microb Ecol 72:975–985
Huang Z, Zeng S, Xiong J, Hou D, He J (2020) Microecological Koch’s postulates reveal that intestinal microbiota dysbiosis contributes to shrimp white feces syndrome. Microbiome 8:32
Lu J, Zhang X, Qiu Q, Chen J, Xiong J (2020) Identifying potential polymicrobial pathogens: moving beyond differential abundance to driver taxa. Microb Ecol. https://doi.org/10.1007/s00248-020-01511-y
Hou D, Huang Z, Zeng S, Liu J, Wei D, Deng X, Weng S, Yan Q, He J (2018) Intestinal bacterial signatures of white feces syndrome in shrimp. Appl Microbiol Biotechnol 102:3701–3709
Holt CC, van der Giezen M, Daniels CL, Stentiford GD, Bass D (2020) Spatial and temporal axes impact ecology of the gut microbiome in juvenile European lobster (Homarus gammarus). ISME J 14:531–543
Smith P, Willemsen D, Popkes M, Metge F, Gandiwa E, Reichard M, Valenzano DR (2017) Regulation of life span by the gut microbiota in the short-lived African turquoise killifish. eLife 6:e27014
Xiong J, Xuan L, Yu W, Zhu J, Qiu Q, Chen J (2019) Spatiotemporal successions of shrimp gut microbial colonization: high consistency despite distinct species pool. Environ Microbiol 21:1383–1394
Greenhalgh K, Meyer KM, Aagaard KM, Wilmes P (2016) The human gut microbiome in health: establishment and resilience of microbiota over a lifetime. Environ Microbiol 18:2103–2116
Yan Q, Li J, Yu Y, Wang J, He Z, Van Nostrand JD, Kempher ML, Wu L, Wang Y, Liao L, Li X, Wu S, Ni J, Wang C, Zhou J (2016) Environmental filtering decreases with fish development for the assembly of gut microbiota. Environ Microbiol 18:4739–4754
Zeng S, Huang Z, Hou D, Liu J, He J (2017) Composition, diversity and function of intestinal microbiota in pacific white shrimp (Litopenaeus vannamei) at different culture stages. PeerJ 5:e3986
Yu W, Wu J, Zhang J, Yang W, Chen J, Xiong J (2018) A meta-analysis reveals universal gut bacterial signatures for diagnosing the incidence of shrimp disease. FEMS Microbiol Ecol 94:fiy147
Björk JR, Hui F, O’Hara RB, Montoya JM (2018) Uncovering the drivers of host-associated microbiota with joint species distribution modeling. Mol Ecol 27:2714–2724
Stegen JC, Lin X, Fredrickson JK, Chen X, Konopka A (2013) Quantifying community assembly processes and identifying features that impose them. ISME J 7:2069–2079
Baselga A (2010) Partitioning the turnover and nestedness components of beta diversity. Glob Ecol Biogeogr 19:134–143
Si X, Baselga A, Leprieur F, Song X, Ding P, Meiri S (2016) Selective extinction drives taxonomic and functional alpha and beta diversities in island bird assemblages. J Anim Ecol 85:409–418
Xiong J, Dai W, Qiu Q, Zhu J, Yang W, Li C (2018) Response of host–bacterial colonization in shrimp to developmental stage, environment and disease. Mol Ecol 27:3686–3699
Zhang Z, Li D, Refaey MM, Xu W, Tang R, Li L (2018) Host age affects the development of southern catfish gut bacterial community divergent from that in the food and rearing water. Front Microbiol 9:495
Pandit SN, Kolasa J, Cottenie K (2009) Contrasts between habitat generalists and specialists: an empirical extension to the basic metacommunity framework. Ecology 90:2253–2262
Hu A, Wang H, Cao M, Rashid A, Yu CP (2019) Environmental filtering drives the assembly of habitat generalists and specialists in the coastal sand microbial communities of Southern China. Microorganisms 7:598
Escalas A, Bouvier T, Mouchet MA, Leprieur F, Bouvier C, Troussellier M, Mouillot D (2013) A unifying quantitative framework for exploring the multiple facets of microbial biodiversity across diverse scales. Environ Microbiol 15:2642–2657
Escalas A, Troussellier M, Tong Y, Bouvier T, Bouvier C, Mouchet MA, Flores Hernandez D, Ramos Miranda J, Zhou J, Mouillot D (2017) Functional diversity and redundancy across fish gut, sediment, and water bacterial communities. Environ Microbiol 19:3268–3282
Laforest-Lapointe I, Arrieta MC (2018) Microbial eukaryotes: a missing link in gut microbiome studies. mSystems 3:e00201–e00217
Andersen LO, Vedel NH, Stensvold CR (2013) Waiting for the human intestinal Eukaryotome. ISME J 7:1253–1255
Dai W, Yu W, Zhang J, Zhu J, Tao Z, Xiong J (2017) The gut eukaryotic microbiota influences the growth performance among cohabitating shrimp. Appl Microbiol Biotechnol 101:6447–6457
Parfrey LW, Walters WA, Knight R (2011) Microbial eukaryotes in the human microbiome: ecology, evolution, and future directions. Front Microbiol 2:153
Xiong J, Yu W, Dai W, Zhang J, Ou C (2018) Quantitative prediction of shrimp disease incidence via the profiles of gut eukaryotic microbiota. Appl Microbiol Biotechnol 102:3315–3326
Massana R, Logares R (2013) Eukaryotic versus prokaryotic marine picoplankton ecology. Environ Microbiol 15:1254–1261
Xiong J, Zhu J, Dai W, Dong C, Qiu Q, Li C (2017) Integrating gut microbiota immaturity and disease-discriminatory taxa to diagnose the initiation and severity of shrimp disease. Environ Microbiol 19:1490–1501
Burns AR, Zac Stephens W, Stagaman K, Wong S, Rawls JF, Guillemin K, Bohannan BJ (2016) Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J 10:655–664
Xiong J (2018) Progress in the gut microbiota in exploring shrimp disease pathogenesis and incidence. Appl Microbiol Biotechnol 102:7343–7350
Farjalla VF, Srivastava DS, Marino NAC, Azevedo FD, Dib V, Lopes PM, Rosado AS, Bozell RL, Esteves FA (2012) Ecological determinism increases with organism size. Ecology 93:1752–7959
AQSIQ (2007) The specification for marine monitoring of China-Part 4: seawater analysis (GB 17378.4-2007). General administration of quality supervision, inspection and quarantine (AQSIQ) of the People’s Republic of China. (in Chinese).
Herlemann DPR, Lundin D, Jürgens K, Bertilsson S, Waniek JJ, Andersson A, F. (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571–1579
Hugerth LW, Muller EE, Hu YO, Lebrun LA, Roume H, Lundin D, Wilmes P, Andersson AF (2014) Systematic design of 18S rRNA gene primers for determining eukaryotic diversity in microbial consortia. PLoS One 9:e95567
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963
Caporaso JG, Kuczynski J, Stombaugh J (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: A versatile open source tool for metagenomics. PeerJ 4:e2584
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Christian Q, Elmar P, Pelin Y, Jan G, Timmy S, Pablo Y, Jörg P, Oliver GF (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596
R Development Core Team (2013) R: a language and environment for statistical computing. The R foundation for statistical computing, Vienna ISBN: 3-900051-07-0. http://www.R-project.org/
Peres-Neto PR, Jackson DA (2001) How well do multivariate data sets match? The advantages of a Procrustean superimposition approach over the Mantel test. Oecologia 129:169–178
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Aust Ecol 26:32–46
Xiong J, Zhu J, Wang K, Wang X, Ye X, Liu L, Zhao Q, Hou M, Qiuqian L, Zhang D (2014) The temporal scaling of bacterioplankton composition: High turnover and predictability during shrimp cultivation. Microb Ecol 67:256–264
Székely AJ, Langenheder S (2014) The importance of species sorting differs between habitat generalists and specialists in bacterial communities. FEMS Microbiol Ecol 87:102–112
McArdle BH, Anderson MJ (2001) Fitting multivariate models to community data: a comment on distance-based redundancy analysis. Ecology 82:290–297
Xuan L, Sheng Z, Lu J, Qiu Q, Chen J, Xiong J (2019) Bacterioplankton community responses and the potential ecological thresholds along disturbance gradients. Sci Total Environ 696. https://doi.org/10.1016/j.scitotenv.2019.134015
Borcard D, Legendre P (2002) All-scale spatial analysis of ecological data by means of principal coordinates of neighbour matrices. Ecol Model 153:51–68
Wu W, Lu HP, Sastri A, Yeh YC, Gong GC, Chou WC, Hsieh CH (2018) Contrasting the relative importance of species sorting and dispersal limitation in shaping marine bacterial versus protist communities. ISME J 12:485–494
Lande R (1996) Statistics and partitioning of species diversity, and similarity among multiple communities. Oikos 76:5–13
Logares R, Tesson SVM, Canbäck B, Pontarp M, Rengefors K (2018) Contrasting prevalence of selection and drift in the community structuring of bacteria and microbial eukaryotes. Environ Microbiol 20:1–15
Luis P, Saint-Genis G, Vallon L, Bourgeois C, Bruto M, Marchand C, Record E, Hugoni M (2019) Contrasted ecological niches shape fungal and prokaryotic community structure in mangroves sediments. Environ Microbiol 21:1407–1424
Bogitsh B, Carter C, Oeltmann T (2005) Human parasitology3rd edn. Elsevier Academic Press, Burlington
Linda W, Heintz-Buschart A, Hogan A, Muller EEL, Narayanasamy S, Laczny CC, Hugerth LW, Bindl L, Bottu J, Andersson AF (2017) Colonization and succession within the human gut microbiome by archaea, bacteria, and microeukaryotes during the first year of life. Front Microbiol 8:738
Astorga A, Oksanen, Luoto J, Soininen M, Virtanen R, Muotka T (2012) Distance decay of similarity in freshwater communities: do macro- and microorganisms follow the same rules? Glob Ecol Biogeogr 21:365–375
Soininen J, Heino J, Wang J (2018) A meta-analysis of nestedness and turnover components of beta diversity across organisms and ecosystems. Glob Ecol Biogeogr 27:96–109
Mcfall-Ngai M, Hadfield MG, Bosch TCG, Carey HV, Wernegreen JJ (2013) Animals in a bacterial world, a new imperative for the life sciences. Proc Natl Acad Sci U S A 110:3229–3236
Rosenberg E, Koren O, Reshef L, Efrony R, Zilber-Rosenberg I (2007) The role of microorganisms in coral health, disease and evolution. Nat Rev Microbiol 5:355–362
Mouchet MA, Bouvier C, Bouvier T, Troussellier M, Escalas A, Mouillot D (2012) Genetic difference but functional similarity among fish gut bacterial communities through molecular and biochemical fingerprints. FEMS Microbiol Ecol 79:568–580
Tzeng TD, Pao YY, Chen PC, Weng FC, Jean WD, Wang D (2015) Effects of host phylogeny and habitats on gut microbiomes of oriental river prawn (Macrobrachium nipponense). PLoS One 10:e0132860
Isabwe A, Ren K, Wang Y, Peng F, Chen H, Yang J (2019) Community assembly mechanisms underlying the core and random bacterioplankton and microeukaryotes in a river–reservoir system. Water 11:1127
Hou D, Huang Z, Zeng S, Liu J, He J (2018) Comparative analysis of the bacterial community compositions of the shrimp intestine, surrounding water and sediment. J Appl Microbiol 125:792–799
Acknowledgments
We wish to acknowledge Prof. SG Tringe from DOE Joint Genome Institute for her excellent comments on the manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (31872693), the Natural Science Fund for Distinguished Young Scholars of Zhejiang Province (LR19C030001), the Technology Innovation Team of Ningbo (2015C110018), and the K.C. Wong Magna Fund in Ningbo University.
Author information
Authors and Affiliations
Contributions
JC and JX conceived and designed research. XL, MY, and QQ conducted experiments. JX contributed analytical tools. JX and JL analyzed data. JX wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
This article does not contain any studies with human participants performed by any of the authors. The use of shrimp were followed the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 766 kb)
Rights and permissions
About this article
Cite this article
Xiong, J., Li, X., Yan, M. et al. Comparable Ecological Processes Govern the Temporal Succession of Gut Bacteria and Microeukaryotes as Shrimp Aged. Microb Ecol 80, 935–945 (2020). https://doi.org/10.1007/s00248-020-01533-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-020-01533-6