Abstract
Wolbachia are inherited intracellular bacteria that cause male-specific death in some arthropods, called male-killing. To date, three Wolbachia strains have been identified in the oriental tea tortrix Homona magnanima (Tortricidae, Lepidoptera); however, none of these caused male-killing in the Japanese population. Here, we describe a male-killing Wolbachia strain in Taiwanese H. magnanima. From field-collected H. magnanima, two female-biased host lines were established, and antibiotic treatments revealed Wolbachia (wHm-t) as the causative agent of male-killing. The wsp and MLST genes in wHm-t are identical to corresponding genes in the nonmale-killing strain wHm-c from the Japanese population, implying a close relationship of the two strains. Crossing the Japanese and Taiwanese H. magnanima revealed that Wolbachia genotype rather than the host genetic background was responsible for the presence of the male-killing phenotype. Quantitative PCR analyses revealed that the density of wHm-t was higher than that of other Wolbachia strains in H. magnanima, including wHm-c. The densities of wHm-t were also heterogeneous between host lines. Notably, wHm-t in the low-density and high-density lines carried identical wsp and MLST genes but had distinct lethal patterns. Furthermore, over 90% of field-collected lines of H. magnanima in Taiwan were infected with wHm-t, although not all host lines harboring wHm-t showed male-killing. The host lines that showed male-killing harbored a high density of Wolbachia compared to the host lines that did not show male-killing. Thus, the differences in the phenotypes appear to be dependent on biological and genetic characteristics of closely related Wolbachia strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Various microbes, including viruses, fungi, and bacteria, are symbiont of insects [1,2,3]. Some of these microbes are inherited from mothers to offspring via transovarian transmission [4]. Wolbachia pipientis, an alpha-proteobacteria hereafter referred to as Wolbachia, is one such intracellular bacteria found in various arthropods [5, 6]. Since its survival mostly relies on female hosts, Wolbachia is considered to have unique adaptive strategies to manipulate the host, especially host reproduction [5, 7]. These strategies include male-killing, which directly produces female-biased sex ratios to promote Wolbachia inheritance [5, 8]. Other intracellular bacteria, microsporidia, and viruses have also been reported to induce similar male-killing at early embryonic stages (early male-killing) or at larval and pupal stages (late male-killing) [5, 8,9,10].
Diverse species of intracellular microbes are known to induce male-killing, and the phenomenon has been investigated for many decades in terms of ecological and evolutionary significance and molecular mechanism [5, 11]. The mechanism of male-killing has attracted a great deal of attention and has been well studied in two intracellular bacteria, Spiroplasma and Wolbachia. Both bacteria cause defects in dosage compensation and sex determination cascade in host insects [12,13,14]. Recently, a gene encoding the male-killing toxin was identified in Spiroplasma poulsonii [12]. A candidate gene has also been postulated for Wolbachia male-killing [15]. It has been reported that the density of Wolbachia in the host line affects the strength of both the male-killing and cytoplasmic incompatibility phenotypes [16,17,18,19]. Furthermore, identical Wolbachia strains have been observed to exhibit different phenotypes after interspecific transfer to a new host [20,21,22] or intraspecific introgression to a new host population [23], indicating that host genetic background interacts with the bacteria to generate the different outcomes observed in Wolbachia hosts.
Homona magnanima (Tortricidae, Lepidoptera) is a serious pest of tea plants in East Asia. In the Japanese populations of the insect, Spiroplasma [24] and a presumed RNA virus [9, 25, 26] were shown to kill males at the embryonic and larval stages, respectively. In addition, three Wolbachia strains have also been identified and characterized in the Japanese host population—wHm-a had no effect on the host, wHm-b caused cytoplasmic incompatibility in the host, and wHm-c compensated host reproduction; however, none of them distorted the sex ratio [19]. In this study, we identified a male-killing Wolbachia strain from H. magnanima in Taiwan. To characterize this male-killing strain, phylogenetic analyses were conducted and the effects of the strain on host development were investigated. A series of crossing experiments were conducted using the Taiwanese and Japanese H. magnanima lines to determine whether the male-killing phenotype was dependent on host genetic background. The effects of Wolbachia density on host mortality or fecundity were also investigated, along with the prevalence and density of Wolbachia in Taiwanese field populations.
Materials and Methods
Insects and Establishment of H. magnanima Lines
In 2015, 72 egg-masses of H. magnanima were collected from tea plantations at Tea Research and Extension Station (Taoyuan City, Taiwan) and imported with permission from the Ministry of Agriculture, Forestry and Fisheries (No. 27 - Yokohama Shokubou 891). Each field-collected egg-mass was placed on a plastic box (23 × 16 × 8 cm) containing artificial diet INSECTA LF (Nosan, Yokohama, Japan). Larvae hatching from each egg-mass were reared collectively in the box until adult eclosion. The adults that emerged were mated, and offspring maintained in the laboratory as described previously by Tsugeno et al. [24] and Arai et al. [19].
From the 72 egg-masses, three host lines were successfully established and maintained—two female-biased lines named WT12 and WT24 along with one line with normal sex ratio (NSR). One female adult from each line was subjected to microbe detection assays as mentioned below. Every generation, Wolbachia infection was checked using three to five adult females. The female-biased lines were mated with males from the NSR line and maintained in the laboratory as mentioned above. Three lines of Japanese H. magnanima Wa, Wb, and Wc lines were also maintained in the same manner. These lines are singly infected with the Wolbachia strains wHm-a, wHm-b, and wHm-c [19].
In 2017, a survey of the prevalence of Wolbachia, Spiroplasma, and RNA virus was completed using 139 egg-masses and 82 larvae of H. magnanima collected and imported from the same Taiwanese tea field with permission from the Ministry of Agriculture, Forestry and Fisheries (No. 29 - Yokohama Shokubou 1326). Egg-masses were reared as mentioned above until adult eclosion. Field-collected larvae were reared individually on INSECTA LF in a 1/2-oz cup until adult eclosion. After eclosion, the sex of the individual was confirmed by morphology, and the sex ratio of adults from each egg-mass was calculated. These adults were subjected to microbe detection assays as described below.
Detection of Microbes and Molecular Typing of Wolbachia
Total DNA was extracted from individual H. magnanima abdomen, as described in Arai et al. [19]. Total RNA was extracted from the abdomen, using ISOGEN (Nippon Gene, Tokyo, Japan) and following the manufacturer’s protocol. Briefly, samples were individually placed in a new plastic tube and briefly homogenized in 1000 μL ISOGEN using a sterilized pestle, mixed with 200 μL chloroform, and centrifuged. The resulting supernatant (500 μL) was transferred to a new tube and precipitated with 500 μL isopropanol. The precipitated RNA was then washed with 1000 μL 70% ethanol [v/v], dissolved in 50 μL distilled water, treated with 2 μL DNase I (Nippon Gene), re-extracted with ISOGEN as described in the manufacturer’s protocol, and stored at − 80 °C.
To detect Wolbachia and Spiroplasma, DNA extracted from adults were PCR-amplified as previously reported [19], using TaKaRa Ex Taq or EmeraldAmp MAX PCR Master Mix (TaKaRa Bio, Shiga, Japan) and primer combinations listed in Table S1. DNA extracted from Wa [19] and IN12 [24] lines were used as positive controls for Wolbachia and Spiroplasma, respectively. To detect male-killing virus, RNA samples were diluted with Milli-Q water to 50–100 ng/μL, reverse transcribed with AMV Reverse Transcriptase XL (TaKaRa), and amplified using primers C3-F and C3-R against MK 1241 [25]. RNA extracted from the LMK line [25] was used as the positive control for male-killing RNA virus. β-Actin of the host insect was amplified as control [19, 24]. PCR products were separated electrophoretically on 1.5% w/v agarose, stained with ethidium bromide, and visualized on a transilluminator. Infection with each microbe was checked once per sample.
For molecular typing of Wolbachia in WT12 and WT24 lines, wsp genes and Wolbachia multilocus sequence typing (MLST) genes (gatB, coxA, hcpA, ftsZ, and fbpA) were amplified using primer sets designed by Baldo et al. [27]. The PCR products of wsp and MLST genes from WT12 and WT24 lines were cloned and sequenced as described previously [19].
Antibiotic Treatments
Each egg-mass from WT12, WT24, and NSR lines was reared for one generation on the artificial diet SilkMate 2S (Nosan) supplemented with 0.1% w/w tetracycline, with each line treatment in three replicates. The sex ratio at treated generation (G0) was calculated based on the number of males and females in adult stage, and Wolbachia infection was confirmed by PCR amplification of wsp gene using one individual from each treatment. G0 females from each line were mated with NSR males, and G1 offspring were reared on SilkMate 2S without tetracycline to assess Wolbachia infection and the sex ratio.
Crossing Between Taiwanese and Japanese Host Lines
Five females from WT12 and WT24 lines were mated with five males from the Japanese Wc line, and similarly, five Wc females were mated with five NSR males from Taiwan. Crossing was replicated five times each, and five F1 egg-masses were selected from each cross. Hatched larvae were individually reared on SilkMate 2S in 1/2-oz cups. After eclosion, the sex ratio was calculated. F1 females were also backcrossed with Wc or NSR males, respectively, over five generations, and the sex ratio in the offspring was determined.
Determination of Wolbachia Density in Wolbachia-Infected Host Lines
To estimate Wolbachia density, five or six newly emerged females from each Wolbachia-infected line (WT12, WT24, Wa, Wb, and Wc) were randomly sampled and individually subjected to DNA extraction as mentioned above. Each DNA template was diluted to 10 ng/μL. A series of qPCR assays were performed using StepOnePlus real-time PCR system (Life Technologies, Carlsbad, CA, USA). Wolbachia wsp gene copy number in each sample was quantified using a reaction mixture containing 10 ng DNA, 30 μM primers wHm-uni_qpcrF and wHm-uni_qpcrR (Table S1), and 5 μL FastStart Universal SYBR Green master mix (Roche, Basel, Switzerland) [19]. A standard curve was generated using 10−4, 10−3, 10−2, 10−1, 1, and 10 ng plasmid DNA harboring a wsp fragment, and the number of wsp copies in the plasmid DNA was estimated from the molecular weight of the plasmid and wsp sequence. Finally, the total amount of Wolbachia wsp copies per 10 ng DNA was calculated.
Effects of Wolbachia on Host Survival and Fitness
To survey the effects of Wolbachia on host development, egg-masses from WT12, WT24, and NSR lines were obtained as mentioned previously [19]. Survival rate throughout the embryonic stage was quantified as hatchability (number of hatched neonate/number of eggs). The number of eggs per egg-mass was estimated based on a regression line between the area of egg-masses and the number of eggs derived from 50 egg-masses from the NSR line. We note that in H. magnanima, matured embryos (pharate) form black head capsules after 4 days post oviposition (dpo), indicating successful development to late embryonic stage (Fig. S1). Healthy pharate hatch within 1 day, while damaged pharate larvae will not. To verify the lethal stage during embryogenesis of H. magnanima, both survival rate until late embryonic stage (number of pharate larvae/number of eggs) and survival rate of pharate larvae (number of hatched neonate/number of pharate larvae) were calculated. Hatched neonates and remaining pharate larvae were counted under a microscope.
To determine sex in pharate and neonates, the W chromosome was stained using lactic acetic orcein as previously reported [28]. Briefly, pharates and neonates were dissected on slide glass with forceps, followed by fixation with methanol and acetic acid and staining with lactic acetic orcein. Finally, mortality in female and male pharate larvae was calculated as the number of remaining male or female pharate larvae 7 days post oviposition per total number of pharate larvae in the egg-mass.
Larvae from the WT12, WT24, and NSR lines were individually reared in 1/2-oz cups, using SilkMate 2S as described above. The larval development time and pupal weight (within 1 day post pupation) was measured as mentioned previously [19]. The longevity and number of egg-mass of adult females were measured in a 120-cc plastic cup with cotton supplied with water. Ten adult females per line were individually subjected to the experiment.
Transmission of Wolbachia Strains in Female-Biased Lines
One female each from the WT12 and WT24 lines was mated with males from the NSR line. After oviposition and hatching, 50 neonate larvae were chosen randomly and reared in 1/2-oz cups until adult eclosion, using the artificial diet INSECTA LF. Wolbachia infection was tested in F1 adults by PCR assay as described above. Transmission rate was calculated as the number of infected F1 adults divided by the total number of F1 adults tested.
H. magnanima Sex Ratio in the Field and Prevalence and Density of Wolbachia
Larvae (n = 82) and egg-masses (n = 96) collected from the field in 2017 were reared until adult eclosion as outlined above. Total DNA and RNA were separately extracted from abdomen of the 82 samples collected at the larval stage, after dividing the abdomen into two pieces using a pair of forceps, and the template was used to detect Wolbachia, Spiroplasma, and RNA virus simultaneously. The Wolbachia-infected samples were then genotyped using diagnostic primer sets for wHm-a, wHm-b, and wHm-c [19] (Table S1). Three newly emerged females from each egg-mass were individually subjected to DNA extraction, detection of Wolbachia, and determination of Wolbachia density as mentioned above.
Data Analysis and Statistics
A χ2 test was used to assess bias in the sex ratio of the host lines. The correlation between egg-mass size and number of eggs per egg-mass was investigated by regression analysis using a general linear model. Hatchability, larval developmental time, pupal weight, and Wolbachia density in each host line were analyzed by Steel–Dwass test. Wolbachia densities in male-killing egg-masses and nonmale-killing egg-masses were compared using Wilcoxon test. All statistical analyses were performed using JMP v9 (SAS, Cary, NC, USA). The MLST and wsp sequences of wHm-t were deposited in GenBank under accession numbers LC427375 to LC427380. The sequences were aligned with known Wolbachia sequences obtained from the Wolbachia MLST database [27] (Table S2) and wHm-a, wHm-b, and wHm-c (GenBank accession numbers LC363921 to LC363938) using ClustalW [29]. A phylogenetic tree was estimated using MEGA6 [30] by maximum likelihood with bootstrap resampling of 1000 replicates.
Results
Wolbachia Strain wHm-t Is a Causative Agent for the Female-Biased Sex Ratio in Taiwanese H. magnanima
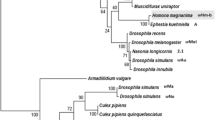
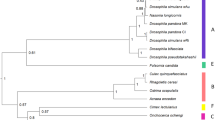
Two female-biased lines (WT12 and WT24) were established from 72 egg-masses collected in 2015, alongside one line with a NSR (sex ratios 98.6 ± 0.5% female (χ2 test, χ2 = 886.5, p < 0.001), 99.6 ± 0.2% (χ2 = 927.1, p < 0.001), and 50.4 ± 1.2% (χ2 = 1.837, p > 0.05), respectively (Fig. 1). In the two female-biased lines, Wolbachia was detected by diagnostic PCR, and Spiroplasma or the male-killing virus was not detected. NSR adults were free of intracellular bacteria and male-killing virus (Fig. 2). To survey multiple Wolbachia infections, 16 wsp fragments cloned from DNA derived from an adult female from the WT12 and WT24 lines were sequenced and were found to be identical. Furthermore, sequences of wsp and MLST genes were completely identical between wHm-c from Japanese Wc line (normal sex ratio) and Wolbachia derived from both WT12 and WT24 lines (female-biased sex ratio) (Fig. 3). From now on, the Taiwanese wHm-c type Wolbachia will be referred as wHm-t.
Phylogenetic analysis of wsp and MLST genes in Wolbachia infecting Homona magnanima. The Wolbachia strains quoted are listed in Table S2. The tree was constructed by maximum likelihood based on the Tamura–Nei model. Bootstrap values exceeding 50% in 1000 replicates are indicated
To investigate whether wHm-t is responsible for the female-biased sex ratio in Taiwanese H. magnanima lines, wHm-t was eliminated from these lines by antibiotic treatment. In tetracycline-treated G0, the sex ratio was still female-biased in WT12 and WT24 (Fig. 4). The sex ratio at G1 was not female-biased in any antibiotic-treated lines, implying that a tetracycline-sensitive microbe caused the sex ratio distortion in Taiwanese H. magnanima. Furthermore, Wolbachia was absent from tetracycline-treated G0 and G1 adults of WT12 and WT24 lines (Fig. 4). Taken together, these results confirm that wHm-t infection distorts the sex ratio in Taiwanese H. magnanima.
Effect of tetracycline on Wolbachia infection and host sex ratio at G0 and G1. Wolbachia infection was tested by diagnostic PCR. Black circles marked W+ indicate lines infected with Wolbachia, while gray circles marked W− are uninfected. Black and white boxes indicate lines with distorted and normal sex ratio, respectively
Effect of Host Genetic Background on the Sex Ratio in Japanese and Taiwanese H. magnanima Infected with Closely Related Wolbachia Strains
Taiwanese (WT12, WT24, and NSR) and Japanese (Wc) H. magnanima adults were successfully mated, generating viable, fertile offspring. The sex ratio was near 1:1 (48.5% females, n = 297) in the F1 generation of Wc (female) × NSR (male), but female-biased in the F1 generation of WT12 × Wc (100% females, n = 205) and F1 generation of WT24 × Wc (100% females, n = 186). After backcrossing, the sex ratio was normal in the offspring of F1 (Wc × NSR) × NSR (back cross: BC1) (48.6% females, n = 251), but female-biased in the offspring of F1 (WT12 × Wc) × Wc (100% females, n = 140) and F1 (WT24 × Wc) × Wc (100% females, n = 152). H. magnanima females infected with wHm-t produced only females over five generations of backcrossing with Wc males (Fig. S2). On the other hand, H. magnanima infected with wHm-c showed normal sex ratio over the same generations of backcrossing with NSR males (Fig. S2). These results indicate that the host genotype is not a key driver of sex ratio distortion.
Wolbachia Density Within Strains, Effects for Host Survival and Fitness, and Transmission of wHm-T Within Taiwanese Host
Wolbachia density was significantly higher in WT12 than in any other lines tested, including WT24 (Steel–Dwass test, Z = − 4.128, p < 0.001), Wa (Z = 3.594, p < 0.001), Wb (Z = 3.923, p < 0.001), and Wc (Z = 3.923, p < 0.001) (Fig. 5). Wolbachia density was also significantly higher in WT24 than in Wb (Z = 3.923, p < 0.001) and Wc (Z = 3.923, p < 0.001), but was comparable between WT24 and Wa (Z = 0.692, p = 0.958).
The area of an egg-mass (x, mm2) was significantly correlated with the number of eggs in an egg-mass (y) (general linear model, y = 2.9273x + 3.9248 with R2 = 0.8556 and p < 0.01). This regression line was then used to estimate survival rate and hatchability. The mean hatchability was significantly higher in NSR than in WT12 (Steel–Dwass test, Z = − 5.287, p < 0.01, Table 1) and in WT24 (Z = − 3.146, p < 0.01), and higher in WT24 than in WT12 (Z = 4.273, p < 0.01), indicating that wHm-t reduced the viability of embryos. The survival rate until late embryonic stage was significantly lower in WT12 than in NSR (Z = − 4.249, p < 0.01) and WT24 (Z = 4.610, p < 0.01), but did not differ between NSR and WT24 (Z = 0.929, p = 0.62). The survival rate of pharate larvae was significantly higher in NSR than in WT12 (Z = − 6.057, p < 0.01) and in WT24 (Z = − 5.047, p < 0.01) and significantly higher in WT12 than in WT24 (Z = 5.002, p < 0.01). These results indicate that wHm-t caused male-killing at different stages in the WT12 and WT24 lines.
Pharate larvae were female-biased in WT12 (χ2 test, χ2 = 23.6, p < 0.001), but not in NSR (χ2 = 0.011, p > 0.05) and WT24 (χ2 = 0.137, p > 0.05) based on acetic lactic orcein staining (Fig. S3). On the other hand, neonates were female-biased both in WT12 (χ2 = 69.1, p < 0.001) and WT24 (χ2 = 45.0, p < 0.001) but not in NSR (χ2 = 0.011, p > 0.05, Fig. S4). Furthermore, mortality in female pharate larvae was significantly higher in WT12 than in WT24 (Steel–Dwass test, Z = − 4.594, p < 0.01) and NSR (Z = − 4.635, p < 0.01, Table S4). These results indicate that wHm-t caused male-killing in WT12 during early embryogenesis and in WT24 lines at late embryogenesis and also caused defects in females in the WT12 line.
Mean mortality of larvae and pupae were significantly higher in WT24 than in WT12 (Steel–Dwass test, Z = − 2.653, p < 0.05) and NSR (Z = 2.647, p < 0.05) (Table S5), but no significance was detected between WT12 and NSR (Z = 2.207, p > 0.05). Mortality at the first and second instar was higher in WT24 than in NSR (first instar: Z = − 2.653, p < 0.05; second instar: Z = − 2.389, p < 0.05) (Table S5).
There were no significant differences in larval development time, adult longevity, pupal weight, number of egg-mass per female, and egg-mass size between lines with or without Wolbachia (Table 2). Transmission rates of wHm-t to the next generation were 97.9% (49/50) in WT12 and 100% (50/50) in WT24, respectively. These results suggested that wHm-t does not compensate host development and reproduction but can be successfully inherited to host offspring with high transmissibility.
Wolbachia Prevalence and Correlation Between Male-Killing Phenotype and Wolbachia Density
A total of 82 larvae collected from the field in 2017 were reared until adult eclosion, producing 57 female and 25 male adults. Larvae hatched from 96 of the 139 egg-masses collected in field were also successfully reared until adult eclosion. Of these, 46 egg-masses were female-biased, while the sex ratio was normal in the other 50 egg-masses.
In adults reared from field-collected larvae, the prevalence of Wolbachia, Spiroplasma, and male-killing virus was 89.0% (n = 73/82), 2.4% (n = 2/82), and 14.6% (n = 12/82), respectively (Table S3). For Wolbachia, wHm-a and wHm-b were not detected (based on strain-specific PCR), but wHm-c was detected at the rate of 89.0% (n = 73/82). Similarly, the prevalence of Wolbachia in adults reared from field-collected egg-masses was 90.6% (n = 87/96).
Although Wolbachia prevalence was around 90% in the field, only 46 of 96 egg-masses were female-biased. Since 9 of the 96 egg-masses were not infected with Wolbachia as determined by diagnostic PCR, Wolbachia densities were quantified by qPCR in newly ecloded adult females derived from the remaining 87 egg-masses. Figure 6 shows the relationship between the proportion of female and Wolbachia density. The mean Wolbachia density (wsp copies/10 ng total DNA) was significantly higher in the female-biased lines (402,955 ± 50,288, n = 46) than that in the NSR lines (511 ± 64, n = 41) (Wilcoxon test, χ2 = 194.4, degree of freedom = 1, p < 0.001). Collectively, the data suggest that not only Wolbachia infection but also its density is correlated to the occurrence of male-killing.
Relationship between proportion of females and Wolbachia density in field-collected Homona magnanima. The Wolbachia density in established male-killing host lines WT12 and WT24 are marked with arrows. Wolbachia density (x-axis) is plotted in Fig. 1 against the proportion of females (y-axis, %)
Discussion
In this study, we observed that a Wolbachia strain, wHm-t, causes male-killing in Taiwanese H. magnanima. Remarkably, the male-killing strain wHm-t and the nonmale-killing strain wHm-c, isolated from Japanese H. magnanima [19], are identical based on phylogenetic analysis of wsp and MLST genes. Host-switching experiments confirmed that host genotype did not modify Wolbachia phenotypes. However, lethality patterns and mortality in H. magnanima differed in response to density of wHm-t—mortality was higher in the host lines with high wHm-t titers (WT12), than in the host line with low wHm-t titers (WT24) (Table 1). Furthermore, pharate mortality in females of WT12 line was higher than that in WT24 line. Lastly, wHm-t prevalence in the field exceeds 90%, and only host lines infected at high titers undergo male-killing.
The phenotypes of Wolbachia are thought to be outcomes of interactions between host and Wolbachia. In this study, we found that wHm-t and wHm-c are of the same MLST genotype [27], but the corresponding phenotypes are completely different. The strain wHm-c was previously shown to have no effect on the sex ratio in H. magnanima, to shorten larval development time, and to increase pupal weight [19], whereas wHm-t induces female-biased sex ratio but does not affect larval and pupal development (Table 2). Wolbachia phenotypes are well known to be modified by host genetic factors. For instance, the Wolbachia strain wBol1 caused male-killing in Hypolimnas bolina in Polynesia [31], but not in Southeast Asia [23]. Crossing experiments between Polynesian and Southeast Asian populations revealed that the host genetic background influenced the occurrence of male-killing caused by wBol1 in H. bolina [23] and was dependent on a single region of the butterfly chromosome 25 [32, 33]. In H. magnanima, Japanese and Taiwanese populations were confirmed to be the same biological species based on morphology (U Jinbo, personal communication), and the offspring of the two populations were fertile. The results of crossing experiments showed that wHm-t distorted the sex ratio in the Taiwanese population and in hybrids between Taiwanese and Japanese populations, whereas wHm-c did not distort the sex ratio in the Japanese population nor in the hybrids. This result indicates that wHm-t phenotype did not differ between geographically distinct host populations. Likewise, WT12 and WT24 lines showed different lethality patterns and wHm-t titers; however, these two host lines appear to have identical genetic backgrounds as they were collected from the same tea field and also mated to the NSR line for over 25 generations in the laboratory. Thus, the male-killing phenotype in wHm-t is due exclusively to its own biological and genetic factors, independent of host genetic background.
Other than the variations in wsp and MLST genes, the differences in phenotypes between wHm-c- and wHm-t-related strains are likely due to variations in Wolbachia genes that are related to density and/or male-killing. It is well known that closely related bacteria strains may exhibit different pathogenicity due to mutations in specific genomic regions [12, 34, 35]. For example, the pathogenic bacterium Salmonella typhimurium harbors various virulence factors in its genome, and mutations in these genes alter pathogenicity and antibiotic resistance [35]. Since wsp and MLST genes only represent a tiny fraction of the Wolbachia genome, it is necessary to compare genomes of those Wolbachia to identify the underlying genomic regions for male-killing and density (as seen in Ellegaard et al. [36] and LePage et al. [37]).
Wolbachia density was previously demonstrated to be crucial in phenotype development [16,17,18,19]. For example, in Drosophila melanogaster, a Wolbachia strain wMelpop proliferates to higher densities and exhibit higher virulence than its related strain wMel [38, 39]. In H. bolina, a series of antibiotic treatments can artificially modify the timing of male-killing caused by Wolbachia, and it was hypothesized that Wolbachia represents the ability to kill males during embryogenesis and larval stages based on Wolbachia amount and virulence [40]. This is in line with our findings that low-density wHm-t (in WT24) killed insects in late developing stages, but high-density wHm-t (in WT12) killed early developing stages. Furthermore, male-killing by Wolbachia in Drosophila bifasciata requires a threshold Wolbachia density in eggs [16]. In the current study, male-killing of H. magnanima is always accompanied by high titer of wHm-t infection, but the nonmale-killing strain wHm-c shows low titer in its host (Figs. 1 and 5). Some insect intracellular bacteria are known to alter transcriptional levels of pathogenic genes depending on the bacterial titer, resulting in changes in pathogenicity, called “phenotypic plasticity” [41,42,43]. If wHm-c- and wHm-t-related strains have an identical genome, phenotypic plasticity based on bacterial density could be another explanation for differences in phenotypes among Wolbachia strains in H. magnanima.
Male-killing has been defined as male-specific death [5, 7]. In this study, mortality was also high in female as well as in male of WT12 pharate larvae (Table S4). We call this phenomenon “host lethal phenotype” caused by wHm-t infection. Some male-killing bacteria are known to enhance female fitness in terms of fertility or resistance to natural enemies [44, 45] or degrade female activity and longevity [46]; however, “female-killing” as host lethal phenotype would seem counterproductive for an endosymbiont that is mainly transmitted from female hosts to offspring. In fact, there are not many insects infected with wHm-t at high titers, as in WT12, in the field. Possibly, the host lethal phenotype caused by wHm-t in H. magnanima may be driven either by (1) male-killing through a molecule that targets only males, with female-killing achieved via other factors; or by (2) differences between males and females in “tolerance” to a specific substance, so that males are killed at lower titer while females are killed at higher titer. So far, a symbiont-encoded male-killing gene has been determined only in Spiroplasma infecting D. melanogaster, and expression of a single Spiroplasma gene reproduces male-killing phenotypes [12], although Spiroplasma additionally harbor genes encoding toxins that modulate host biology [47, 48]. Wolbachia also have multiple virulence factors in their genomes [49,50,51]; however, whether a single molecule or multiple molecules cause male-killing phenotype is yet to be elucidated. The virulence factors might affect only males or both males and females, which eventually lead to host lethal phenotype of wHm-t in H. magnanima. Further genomic and mechanistic studies may facilitate the understanding of the mechanism of male-killing and interactions between Wolbachia and their host.
References
Duron O, Bouchon D, Boutin S, Bellamy L, Zhou L, Engelstädter J, Hurst GDD (2008) The diversity of reproductive parasites among arthropods: Wolbachia do not walk alone. BMC Biol 6:27
Gibson CM, Hunter MS (2010) Extraordinarily widespread and fantastically complex: comparative biology of endosymbiotic bacterial and fungal mutualists of insects. Ecol Lett 13:223–234
Roossinck MJ (2015) Plants, viruses and the environment: ecology and mutualism. Virology 479:271–277
Bright M, Bulgheresi S (2010) A complex journey: transmission of microbial symbionts. Nat Rev Microbiol 8:218–230
Werren JH, O’Neill SL (1997) The evolution of heritable symbionts. In: O’Neill SL, Hoffmann AA, Werren JH (eds) Influential passengers: inherited microorganisms and arthropod reproduction. Oxford University Press, Oxford, pp 1–41
Zug R, Hammerstein P (2012) Still a host of hosts for Wolbachia: analysis of recent data suggests that 40% of terrestrial arthropod species are infected. PLoS One 7:e38544
Werren JH, Baldo L, Clark ME (2008) Wolbachia: master manipulators of invertebrate biology. Nat Rev Microbiol 6:741–751
Hurst GDD, Bandi C, Sacchi L, Cochrane AG, Bertrand D, Karaca I, Majerus MEN (1999) Adonia variegata (Coleoptera: Coccinellidae) bears maternally inherited Flavobacteria that kill males only. Parasitology 118:125–134
Morimoto S, Nakai M, Ono A, Kunimi Y (2001) Late male-killing phenomenon found in a Japanese population of the oriental tea tortrix, Homona magnanima (Lepidoptera: Tortricidae). Heredity 87:435–440
Jaenike J (2007) Spontaneous emergence of a new Wolbachia phenotype. Evolution 61:2244–2252
Engelstädter J, Hurst GDD (2009) The ecology and evolution of microbes that manipulate host reproduction. Annu Rev Ecol Evol Syst 40:127–149
Harumoto T, Lemaitre B (2018) Male-killing toxin in a bacterial symbiont of Drosophila. Nature 557:252–255
Fukui T, Kawamoto M, Shoji K, Kiuchi T, Sugano S, Shimada T, Suzuki Y, Katsuma S (2015) The endosymbiotic bacterium Wolbachia selectively kills male hosts by targeting the masculinizing gene. PLoS Pathog 11:e1005048
Harumoto T, Anbutsu H, Lemaitre B, Fukatsu T (2016) Male-killing symbiont damages host’s dosage-compensated sex chromosome to induce embryonic apoptosis. Nat Commun 7:12781
Perlmutter JI, Bordenstein SR, Unckless RL, LePage DP, Metcalf JA, Hill T, Martinez J, Jiggins FM, Bordenstein SR (2019) The phage gene wmk is a candidate for male killing by a bacterial endosymbiont. PLoS Pathog 15:e1007936
Hurst GDD, Johnson AP, Schulenburg JHGVD, Fuyama Y (2000) Male-killing Wolbachia in Drosophila: a temperature-sensitive trait with a threshold bacterial density. Genetics 156:699–709
Kondo N, Shimada M, Fukatsu T (2005) Infection density of Wolbachia endosymbiont affected by co-infection and host genotype. Biol Lett 1:488–491
Watanabe M, Miura K, Hunter MS, Wajnberg E (2011) Superinfection of cytoplasmic incompatibility-inducing Wolbachia is not additive in Orius strigicollis (Hemiptera: Anthocoridae). Heredity 106:642–648
Arai H, Hirano T, Akizuki N, Abe A, Nakai M, Kunimi Y, Inoue MN (2019) Multiple infection and reproductive manipulations of Wolbachia in Homona magnanima (Lepidoptera: Tortricidae). Microb Eco 77:257–266
Sasaki T, Ishikawa H (1999) Wolbachia infections and cytoplasmic incompatibility in the almond moth and the Mediterranean flour moth. Zool Sci 16:739–744
Sasaki T, Kubo T, Ishikawa H (2002) Interspecific transfer of Wolbachia between two lepidopteran insects expressing cytoplasmic incompatibility: a Wolbachia variant naturally infecting Cadra cautella causes male killing in Ephestia kuehniella. Genetics 162:1313–1319
Kageyama D, Wang CH, Hatakeyama M (2017) Wolbachia infections of the butterfly Eurema mandarina interfere with embryonic development of the sawfly Athalia rosae. J Invertebr Pathol 150:76–81
Hornett EA, Charlat S, Duplouy AMR, Davies N, Roderick GK, Wedell N, Hurst GDD (2006) Evolution of male-killer suppression in a natural population. PLoS Biol 4:e283
Tsugeno Y, Koyama H, Takamatsu T, Nakai M, Kunimi Y, Inoue MN (2017) Identification of an early male-killing agent in the oriental tea tortrix, Homona magnanima. J Hered 108:553–560
Nakanishi K, Hoshino M, Nakai M, Kunimi Y (2008) Novel RNA sequences associated with late male killing in Homona magnanima. P Roy Soc B-Biol Sci 275:1249–1254
Hoshino M, Nakanishi K, Nakai M, Kunimi Y (2008) Gross morphology and histopathology of male-killing strain larvae in the oriental tea tortrix, Homona magnanima (Lepidoptera: Tortricidae). Appl Entomol Zool 43:119–125
Baldo L, Hotopp JCD, Jolley KA, Bordenstein SR, Biber SA, Choudhury RR, Hayashi C, Maiden MCJ, Tettelin H, Werren JH (2006) Multilocus sequence typing system for the endosymbiont Wolbachia pipientis. Appl Environ Microbiol 72:7098–7110
Kageyama D, Traut W (2004) Opposite sex–specific effects of Wolbachia and interference with the sex determination of its host Ostrinia scapulalis. P Roy Soc B-Biol Sci 271:251–258
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Dyson EA, Kamath MK, Hurst GDD (2002) Wolbachia infection associated with all-female broods in Hypolimnas bolina (Lepidoptera: Nymphalidae): evidence for horizontal transmission of a butterfly male killer. Heredity 88:166–171
Hornett EA, Moran B, Reynolds LA, Charlat S, Tazzyman S, Wedell N, Jiggins CD, Hurst GD (2014) The evolution of sex ratio distorter suppression affects a 25 cM genomic region in the butterfly Hypolimnas bolina. PLoS Gen 10:e1004822
Reynolds LA, Hornett EA, Jiggins CD, Hurst GD (2019) Suppression of Wolbachia-mediated male-killing in the butterfly Hypolimnas bolina involves a single genomic region. PeerJ 7:e7677
Hensel M (2000) Salmonella pathogenicity island 2. Mol Microbiol 36:1015–1023
Deiwick J, Nikolaus T, Shea JE, Gleeson C, Holden DW, Hensel M (1998) Mutations in Salmonella pathogenicity island 2 (SPI2) genes affecting transcription of SPI1 genes and resistance to antimicrobial agents. J Bacteriol 180:4775–4780
Ellegaard KM, Klasson L, Näslund K, Bourtzis K, Andersson SG (2013) Comparative genomics of Wolbachia and the bacterial species concept. PLoS Genet 9:e1003381
LePage DP, Metcalf JA, Bordenstein SR, On J, Perlmutter JI, Shropshire JD, Layton EM, Funkhouser-Jones LJ, Beckmann JF, Bordenstein SR (2017) Prophage WO genes recapitulate and enhance Wolbachia-induced cytoplasmic incompatibility. Nature 543:243–247
Min KT, Benzer S (1997) Wolbachia, normally a symbiont of Drosophila, can be virulent, causing degeneration and early death. Proc Natl Acad Sci U S A 94:10792–10796
Chrostek E, Teixeira L (2015) Mutualism breakdown by amplification of Wolbachia genes. PLoS Biol 13:e1002065
Charlat S, Davies N, Roderick GK, Hurst GDD (2007) Disrupting the timing of Wolbachia-induced male-killing. Biol Lett 3:154–156
Casadevall A, Pirofski LA (1999) Host-pathogen interactions: redefining the basic concepts of virulence and pathogenicity. Infect Immun 67:3703–3713
Login FH, Balmand S, Vallier A, Vincent-Monégat C, Vigneron A, Weiss-Gayet M, Rochat D, Heddi A (2011) Antimicrobial peptides keep insect endosymbionts under control. Science 334:362–365
Enomoto S, Chari A, Clayton AL, Dale C (2017) Quorum sensing attenuates virulence in Sodalis praecaptivus. Cell Host Microbe 21:629–636
Ebbert MA (1991) The interaction phenotype in the Drosophila willistoni-Spiroplasma symbiosis. Evolution 45:971–988
Xie J, Butler S, Sanchez G, Mateos M (2014) Male killing Spiroplasma protects Drosophila melanogaster against two parasitoid wasps. Heredity 112:399
Hurst GDD, Jiggins FM (2000) Male-killing bacteria in insects: mechanisms, incidence, and implications. Emerg Infect Dis 6:329–336
Ballinger MJ, Perlman SJ (2017) Generality of toxins in defensive symbiosis: ribosome-inactivating proteins and defense against parasitic wasps in Drosophila. PLoS Pathog 13:e1006431
Masson F, Copete SC, Schüpfer F, Garcia-Arraez G, Lemaitre B (2018) In vitro culture of the insect endosymbiont Spiroplasma poulsonii highlights bacterial genes involved in host-symbiont interaction. mBio 9:e00024-18
Pichon S, Bouchon D, Cordaux R, Chen L, Garrett RA, Grève P (2009) Conservation of the type IV secretion system throughout Wolbachia evolution. Biochem Bioph Res Commun 385:557–562
Sheehan KB, Martin M, Lesser CF, Isberg RR, Newton ILG (2016) Identification and characterization of a candidate Wolbachia pipientis type IV effector that interacts with the actin cytoskeleton. mBio 7:e00622-16
Rice DW, Sheehan KB, Newton ILG (2017) Large-scale identification of Wolbachia pipientis effectors. Genome Biol Evol 9:1925–1937
Acknowledgments
We thank Dr. Utugi Jimbo (National Museum of Nature and Science, Tokyo, Japan) for morphological identification of Taiwanese population of H. magnanima and Dr. Katsuhiko Ito (Tokyo University of Agriculture and Technology, Tokyo, Japan) for lending us the StepOnePlus™ real-time PCR system (Applied Biosystems, Tokyo, Japan). We also thank Dr. Hisashi Anbutsu (National Institute of Advanced Industrial Science and Technology, Tsukuba, Japan) and Professor Greg Hurst (Institute of Integrative Biology, University of Liverpool, Liverpool, UK) for revising the manuscript.
Data Archiving
The sequence data of wsp and MLST genes of wHm-t were deposited in GenBank under accession numbers LC427375 to LC427380.
Author information
Authors and Affiliations
Contributions
In this work, HA conducted field surveys, all experiments, and data analysis. SRL organized to collect insects in Taiwan and contributed to the discussion. MN supported the entire experiments and contributed to the discussion. YK sampled insects in Taiwanese tea field and contributed to the entire discussions of this study. Lastly, MNI had full access to all data and had responsibility for the decision to submit for publication.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Electronic Supplementary Material
Supplementary Figure 1
(PNG 275 kb)
Supplementary Figure 2
(PNG 145 kb)
Supplementary Figure 3
(PNG 630 kb)
Supplementary Figure 4
(PNG 250 kb)
ESM 1
(DOCX 41.2 kb)
Rights and permissions
About this article
Cite this article
Arai, H., Lin, S.R., Nakai, M. et al. Closely Related Male-Killing and Nonmale-Killing Wolbachia Strains in the Oriental Tea Tortrix Homona magnanima. Microb Ecol 79, 1011–1020 (2020). https://doi.org/10.1007/s00248-019-01469-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-019-01469-6