Abstract
We characterized a novel Holospora-like bacterium (HLB) (Alphaproteobacteria, Holosporales) living in the macronucleus of the brackish water ciliate Frontonia salmastra. This bacterium was morphologically and ultrastructurally investigated, and its life cycle and infection capabilities were described. We also obtained its 16S rRNA gene sequence and performed in situ hybridization experiments with a specifically-designed probe. A new taxon, “Candidatus Hafkinia simulans”, was established for this HLB. The phylogeny of the family Holosporaceae based on 16S rRNA gene sequences was inferred, adding to the already available data both the sequence of the novel bacterium and those of other Holospora and HLB species recently characterized. Our phylogenetic analysis provided molecular support for the monophyly of HLBs and placed the new endosymbiont as the sister genus of Holospora. Additionally, the host ciliate F. salmastra, recorded in Europe for the first time, was concurrently described through a multidisciplinary study. Frontonia salmastra’s phylogenetic position in the subclass Peniculia and the genus Frontonia was assessed according to 18S rRNA gene sequencing. Comments on the biodiversity of this genus were added according to past and recent literature.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Symbiosis between bacteria and ciliated protists was frequently reported [1,2,3,4,5,6,7,8,9,10]. Investigations on Holospora bacteria, highly infectious endonuclear symbionts belonging to Alphaproteobacteria, have monopolized the field. The complex life cycle of these bacteria and their specificity for host and site of infection suggest that there is a system of diverse interactions with the host, both at the stage of infection and during infection maintenance, either in the macronucleus (MA) or in the micronucleus (MI) [11,12,13,14,15,16]. Research on Holospora officially started in 1890, when Waldemar Mordechai Wolff A. Hafkine (1860–1930) found and described three intranuclear bacteria in the ciliate Paramecium caudatum. That was the first accurate description of Holospora, with the species Holospora obtusa infecting the MA, and both Holospora undulata and Holospora elegans infecting the MI of that ciliate [17].
Holospora and Holospora-like bacteria (HLB) (e.g., Preeria caryophila [18] and “Candidatus Gortzia” [19, 20]) are endonuclear symbionts of different Paramecium species. This has traditionally been considered a basic feature of the clade [12, 21,22,23]. HLB have been characterized as immobile, host- and compartment-specific, infectious bacteria manifesting reproductive forms (RF) and infectious forms (IF) in their life cycle. They have been traditionally defined as bacteria able only to invade nuclei of certain Paramecium species.
To date, at least 13 symbionts assignable to HLB were characterized from Paramecium species [3, 14, 16, 19, 20, 24, 25], although only part of them have been investigated from a molecular point of view [4, 16, 19, 20, 26, 27]. Recently, HLBs were described from P. jenningsi [19], P. multimicronucleatum [20], and P. chlorelligerum [16]. None have been reported in other Paramecium species (such as P. duboscqui, P. woodruffi, P. polycaryum); they remain yet to be discovered.
Despite being traditionally associated with Paramecium, several endosymbionts assignable to HLBs according to their morphology and life cycle have been discovered in ciliates phylogenetically distant from Paramecium. In detail, they have been observed as macronuclear symbionts of Metopus caudatus [28], Metopus spinosus [29], Prorodon teres [3, 30], Stentor multiformis [31], Stentor polymorphus [3, 32], Stentor roeselii [31], Trithigmostoma cucullulus [4, 33], Vorticella sp. [34], Vorticella convallaria [35], Zoothamnium pelagicum [36], Spirostomum minus [4], and Trichodina pediculus [37]. Unfortunately, none of them have ever been molecularly investigated, or even properly morphologically/ultrastructurally described, also due to their occasional retrieval.
Nevertheless, these findings suggest that besides Paramecium, a great variety of other Ciliophora could harbor HLB as well: this is the case of Frontonia salmastra, where a macronuclear endosymbiont assignable to a novel clade of HLB was observed. This reports the morphological and ultrastructural investigation as well as the analysis of the life cycle and infection capabilities of this endosymbiont. We sequenced its 16S rRNA gene, and the phylogenetic analysis based on it suggested a close affinity with Holospora, which resulted in being a sister genus of the novel HLB. We named this bacterium “Candidatus Hafkinia simulans” in honor of Waldemar M.W.A. Hafkine, for his fundamental contribution in the study of the HLB. Additionally, herein we present the host (F. salmastra) redescription according to a multidisciplinary study approach and discuss issues surrounding Frontonia systematics.
Materials and Methods
Host Isolation, Cultivation, and Identification
Frontonia salmastra was sampled between November 2005 and May 2006 in the permanent brackish water pond (salinity: 5–14 g/L) located along the Ligurian sea coastline close to the mouth of Serchio river (Parco Naturale di Migliarino–San Rossore–Massaciuccoli, Migliarino, Pisa district, Tuscany, Italy; N 43° 78′ 73.9″, E 10° 26′ 81.4″). Several ciliates were collected from original samples to obtain some populations maintained in laboratory conditions with the diatom Phaeodactylum tricornutum as a food source (salinity: 5 g/L). During lab cultivation, ciliate culture medium salinity was slightly modified to roughly test the ciliate tolerance to different salinity values. Freshwater condition was never reached due to ciliate intolerance. As the same ciliate have also been retrieved in similar habitats from several other water bodies in Tuscany with salinity between 4 and 20 g/L, some trials of adaptation to higher salinity conditions were also performed, and some subcultures survived up to 40 g/L salinity.
About 85% of cells belonging to the populations obtained from original samples (referred to as ITM-5F, ITM-7F, and ITM-9F) manifested HLB infection of the MA. From the population ITM-5F, the infected monoclonal strain ITM-5F-2 was obtained and used for the following analysis. For experimental infection, the aposymbiotic monoclonal culture ITM-5F-6, isolated from the same population, was used.
Ciliate identification was performed through the observation of living and silver-impregnated specimens; characters were examined according to the original description of F. salmastra [38].
Light and Transmission Electron Microscopy
Living F. salmastra cells were observed for morphological details with a Leitz (Germany) Differential Interference Contrast (DIC) microscope at a magnification of × 300–1250 with the help of a compression device [39]. For silver impregnation, ciliates were fixed with Champy’s solution and silver nitrate-stained after Corliss [40]. As for the investigation on the macronuclear bacteria, specimens were fixed for electron microscopy according to a well-established protocol [41] and examined with a SX 100S JEOL transmission electron microscope (JEOL Ltd., Tokyo, Japan).
Experimental infection was carried out using a homogenate prepared from infected cells according to Preer [21]. Frontonia salmastra aposymbiotic cells (monoclonal strain ITM-5F-6) were infected by mixing this cell culture to an equal volume of the homogenate, in a 3-mL depression slide, and maintained at 18–24 °C. To check the infection status, a set of living cells (n = 10) was observed under the DIC microscope 24 and 48 h after the mixing with homogenate, and then rechecked after 7, 9, and 14 days.
Endosymbiont 16S rRNA Gene Sequencing and Phylogeny
Infected ciliates were isolated in sterile artificial seawater (salinity: 5 g/L) and starved for 24–48 h. Single specimens were collected by micropipette and quickly washed in sterile distilled water, in order to minimize the presence of contaminants; then, they were fixed in 70% ethanol and stored at −20 °C. DNA was extracted from 50 infected cells; extraction was performed with Nucleospin® Plant Extraction Kit (Macherey-Nagel). Amplification of 16S rRNA gene, through polymerase chain reaction (PCR) was performed using the primer combination 16S alpha F19a and 16S alpha R1517, both specifically designed for the class Alphaproteobacteria [42]. A “touchdown” PCR cycle [43] was used in order to increase the specificity of the reaction: after a preliminary denaturation (94 °C-3′), the annealing temperature was set at 63 °C for the first 5 cycles, then shifted to 57 °C for 15 cycles, and reduced to 50 °C for the remaining 25 cycles. TaKaRa Ex Taq polymerase was used for the reaction. Positive and negative controls were included in the experiment to exclude false results. The reaction was done in a Primus 96 plus (MWG Biotech.) thermocycler. The amount of PCR product was assessed by electrophoresis on agarose gel containing ethidium bromide. The PCR product was purified using Quantum Prep® PCR Kleen Spin Columns (Bio-Rad). Amplified gene fragment was directly full-lengths sequenced in both directions with the internal primers 16S F343 ND, 16S R515 ND, and 16S F785 ND [42]. The sequencing reaction was performed by MWG Customer Service (MWG Biotech.). Electropherograms were manually checked and assembled using Chromas Lite software.
The 16S rRNA gene sequence was aligned using the automated aligner ARB software package [44], automatically checked against more than approximately 600,000 bacterial sequences on the SSU rRNA SILVA 123 Ref NR 99 database [45], and further manually checked, against those belonging to the Holosporales and to the selected outgroup (about 100 sequences), to obtain a proper alignment.
As for phylogenetic analysis, 79 sequences were employed in total, 68 selected sequences belonging to order Holosporales and 10 belonging to order Rickettsiales as outgroup. The alignment was reduced in length according to the shortest sequence on each end, obtaining a 1288 character matrix. Both maximum likelihood (ML) analysis (PHYML 5.3.2) [46] and Bayesian inference (BI) analysis (MrBayes 3.2) [47] were performed, with the GTR + I + G substitution model, as indicated by the AIC (Akaike’s information criterion), calculated by jModelTest 2.2 [48]. ML analysis was applied with 1000 pseudoreplicates, while for BI analysis three different Markov Chain Monte Carlo runs were employed, with one cold chain and three heated chains each, running for 1000,000 generations.
Fluorescence In situ Hybridization
Fluorescence in situ hybridization (FISH) was performed both on ciliates of the population ITM-5F and of the monoclonal strain ITM-5F-2. Frontonia salmastra cells were starved for 2 days and, after a thrice washing in distillated water, fixed for 5–10 min in depression slides with 4% (v/v) formaldehyde in PBS. Experiments were performed as previously described [49], with (30%) or without formamide in the hybridization buffer. The eubacterial universal probe EUB338 [50] was employed in preliminary FISH experiments to highlight the presence of bacteria associated with the ciliate MA. Moreover, we tested the following probes: the alphaprotobacterial probes Alf1b [51] and Hafk_sim_147 (5′-TGA AGT TTC CTC CAG TTA TTC-3′). The probe Hafk_sim_147 was specifically designed on the 16S rRNA gene sequence of F. salmastra symbiont.
This probe was tested in silico on the RDP (ribosomal database project) [52] and SILVA database using TestProbe 3.0 [45], allowing 0 mismatches. The sequence of probe Hafk_sim_147 matched three sequences belonging to the class Acidobacteria (namely, Telmatobacter bradus AM887760, uncultured bacterium EU284344, uncultured bacterium HG325035).
Double hybridizations, using probe Hafk_sim_147 or Alf1b in competition with the probe EUB338, were performed, in order to check the possible presence of other symbiotic bacteria inside the host. Cells were observed with a Leica DMR light microscope, equipped for epifluorescence.
Frontonia salmastra 18S rRNA Gene Sequencing and Phylogeny
Starting from 20 fixed cells of ITM-5F-2 ciliate strain, total DNA extraction, amplification, and 18S rRNA gene sequencing were carried out using the same reagents, instruments, and sequencing company cited above. Primers employed for PCR amplification were 18S ITM-5F-2 (Forward, 5′-GAA ACT GCG AAT GGC TC-3′) and 18S R1513 Karyo [53], with the following settings: 3 min at 94 °C, 35 cycles × [(94 °C (30 s), 57 °C (30 s) and 72 °C (120 s)], plus 7 min at 72 °C, cooling at 8 °C (forever). For direct sequencing, three internal primers, namely 18S F919, 18S R536, and 18S R1052 [54] were used.
To reconstruct the phylogeny of F. salmastra we relied on the same softwares and parameters cited above. A total of 59 sequences of other Oligohymenophoreans, 46 of which belonging to the subclass Peniculia, were selected. The alignment was manually checked, sequence length was reduced according to the shortest sequence, and a 1664-character matrix was obtained.
Results
Endosymbiont Morphology and Its Life Cycle
The HLB retrieved in the MA of F. salmastra manifested the classical Holospora’s polymorphism. Two distinctive forms of straight non-motile bacteria with different size and structure were observed in the infected MA (Figs. 1a, 2, 3, 4, 5, and 6). There were RFs, 2–4 × 1.0–1.5 μm in size, and IFs, which could reach 8–30 × 2.0–3.5 μm (Figs. 2, 3, 4, 5, and 6). The IFs presented a very peculiar spindle-shape or even skittles-like form (Figs. 2a–f, 3g, 5b). They looked extremely long if compared to the RFs (Figs. 3g and 5b). However, a large spectrum of intermediate morphological forms (IMFs) was present. IMFs were 5–10 × 1.5–2.5 μm in size, thus bigger than RFs. In many cases, IMFs had not yet manifested compartmentalization of cytoplasmic and periplasmic portions or of the recognition tip (Figs. 4b and 5a). Nevertheless, under the DIC microscope, some IMFs showed a number of bright stripes in the cytoplasm, which probably corresponded to periplasmic parts (Fig. 2d). Moreover, some IFs manifested many periplasmic zones without the formation of the recognition tip, which is a distinctive trait of the IF. Conversely, in some cases, IFs were observed to generate two recognition tips, located at the opposite ends of the bacterial cell (Fig. 2e, f). Bacteria were randomly distributed in the MA karyoplasm, but sometimes few IFs were observed in the center of the MA. Generally, in terms of shape and size, the mature IFs looked like the diatom P. tricornutum (Figs. 2e–g, 3c, g) (laboratory food source of our F. salmastra).
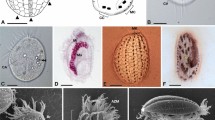
General view and morphological details of Frontonia salmastra. a, d living cells, by differential interference contrast (DIC) microscopy; b, c silver stained cells. a lateral view of the ciliate. CV, contractile vacuole; OA, oral apparatus; MA, macronucleus. b, ventral side of the stained ciliate. c, oral ciliature. VK, vestibular kineties; PK, postoral kineties; PO, paraoral membrane; P1–P3, peniculi; PS, postoral suture line. d nuclear apparatus. MI, micronuclei, e trichocysts (arrows), viewed from above. Scale bars represent 20 μm (a, b) and 5 μm (c–e)
Different stages of “Candidatus Hafkinia simulans” in the macronucleus of Frontonia salmastra. All micrographs were made from living cells by DIC microscopy. a Infectious form (IF) in the macronucleus (MA) after experimental infection. b the infected MA with numerous IFs. Micronuclei (MI) are visible. c crushed MA presenting both forms of bacteria: reproductive form (RF) and IF. Arrows indicating trichocysts. d intermediate forms (IMF) of the symbiont. e, f mature IF of the symbiont. g the main food source of F. salmastra, the diatom Phaeodactylum tricornutum. Scale bars represent 25 μm
Results of FISH experiments on Frontonia salmastra cells infected by “Candidatus Hafkinia simulans”. a–c infected macronucleus (MA) of F. salmastra; d–g infected MA in division. a, b the same MA, showing positive signal to EUB338 probe (emitting in green) and to Hafk_sim_147 probe (emitting in red), with infectious forms (IF) well visible; c higher magnification of MA showing positivity to EUB338 probe with reproductive forms (RF) together with IFs well visible; d, e the same dividing MA showing positivity to EUB338 (emitting in green) and Alf1b (emitting in red), with almost exclusively RFs present in MA, and some bacteria present in the cytoplasm (arrowhead); f, g the same dividing MA showing positivity to EUB338 (emitting in green) and Hafk_sim_147 (emitting in red), with both IFs and RFs well visible. Scale bars represent 20 μm
Ultrastructural morphology of “Candidatus Hafkinia simulans” harbored in the macronucleus of Frontonia salmastra. a reproductive and infectious forms (RF and IF) of the bacterium in the host macronucleus (MA); the arrowhead indicates the periplasmic portion of IF, double arrowhead indicates the cytoplasmic part of IF. b RF, IF, and intermediate form (IMF) of the bacterium. Arrows indicate polyribosome concentration in IMF. Scale bars represent 1.5 μm (a) and 1.0 μm (b)
Ultrastructural peculiarities of Frontonia salmastra infected by “Candidatus Hafkinia simulans”. a nuclear apparatus of the ciliate (MA, macronucleus; MI, micronucleus) and unusual type of mitochondria (MT); note the cristae inside. MA harboring reproductive and intermediate forms (RF and IMF) of the endosymbiont. b nucleolus (N) and different forms (RF, IMF, and IF – infectious form) of the bacterium in the matrix of infected MA. Arrowhead indicates the periplasmic portion of IF. Scale bars represent 3 μm
Ultrastructural peculiarities of the infectious form of “Candidatus Hafkinia simulans”. a longitudinal and cross sections of infectious form (IF); arrowheads point to the fine fibrillar layer on IF’s surface, b IF of the bacteria in the cytoplasm of Frontonia salmastra; arrows indicate the classical type of mitochondria. Scale bars represent 3 μm
The endosymbiotic forms RF, IMF, and IF could also be discriminated at ultrastructural level (Figs. 4 and 5). RFs resembled RFs of previously investigated HLB, showing the typical, homogeneous, and relatively electron-transparent prokaryotic cytoplasm. IMFs displayed a more heterogeneous and denser cytoplasm in comparison with that of RFs. Sometimes a darker zone was observed located at IMF poles (Fig. 4b): it could be interpreted either as a starting developmental stage of periplasm, or as a kind of polyribosome concentration. IFs always showed the periplasmic portion (sometimes with invaginations into the cytoplasm) and a fine fibrous layer on their surface (Figs. 4a and 6a).
The karyoplasm of the infected MA presented relatively large areas almost entirely occupied by HLB. In those parts, very few chromatin bodies were observed (Figs. 4a and 5). Moreover, the nucleolus of the infected MA was detected (Fig. 5b). The MI showed the classical “compact-type” micronuclear structure (Fig. 5a). Few IFs were recorded in the ciliate cytoplasm (Fig. 6b): probably they were released from the infected MA.
During the division process of the infected MA, HLB were randomly distributed in daughter nuclei, without producing any structure similar to the connecting piece of “classical” Holospora (Fig. 3d–f). The number of HLB in F. salmastra infected MA greatly varied from cell to cell, and sometimes the majority of them were represented by the RFs only (Fig. 3d, e).
Experimental infection of the aposymbiotic monoclonal strain ITM-5F-6 highlighted the typical Holospora’s life cycle for the studied HLB. After 24 h from the mixing, IFs were detected in the MA, and then RFs were produced (Fig. 2a, b). The first IF generation was recorded 9 days after the beginning of experimental infection. A new generation of IFs was established in 2 weeks on average.
Endosymbiont Molecular Characterization
A 1442-bp long 16S rRNA gene sequence was obtained for the HLB endosymbiont of F. salmastra; it has been deposited on the NCBI database (accession number: MH319377). From similarity matrix calculation, the average identity values of 94.2% with Holospora sequences (93.9–94.5%) and 91.9% with “Ca. Gortzia” sequences (91.6–92.1%) were recorded respectively (Table 1). In phylogenetic trees, this endosymbiont fell inside the Holosporaceae, as a sister taxon to Holospora clade, with strong statistical supports (100/1.00), (Fig. 7). Phylogenetic trees obtained with ML and BI analyses showed a coherent topology between each other and high statistical value supports.
The HLB from F. salmastra showed bright positive signals by the tested probes in FISH experiments (Fig. 3). Double staining with EUB338 and Alf1b probes revealed the presence of few bacterial cells also in the cytoplasm of some F. salmastra cells (Fig. 3d, e). Using a combination of Eubacteria probe and species-specific probe for the HLB symbiont (Hafk_sim_147) overlapping signals were recorded (Fig. 3a, b): no other bacterial species were detected inside the ciliate host.
Morphology, Biology, and Phylogenetic Position of the Host Ciliate F. salmastra Dragesco & Dragesco-Kernéis, 1986
Considering that the ability to maintain the ciliate Frontonia in a permanent culture was rarely mentioned in literature to our best knowledge [55,56,57], the first notable result of the present study was the success in maintaining F. salmastra in culture for about 5 months. As for this achievement, we consider critical the choice of the diatom P. tricornutum as the food source [58].
Before our investigation, this frontoniid was found once in brackish water in Benin (West Africa) and briefly described by French protozoologists [38]. Then, it was mentioned from the littoral zone of the Caspian Sea [59].
We performed the morphological study for the ciliate population ITM-5F infected by the novel endosymbiont HLB sampled from a pond nearby the mouth of Serchio river (Fig. 1). Frontonia salmastra ITM-5F was a middle-sized ciliate: body length was 100–190 μm (Х ± SD = 145.2 ± 12.5 μm) and body width 40–60 (50.4 ± 4.9 μm); it presented 75–95 somatic kineties; 2–3, relatively small MI (usually with diameter about 3–4 μm) of the “compact” type,” one contractile vacuole (CV), located in the equatorial part of the cell, with one pore (PCV) were present. A number of collecting canals (6–8) were observed starting from the main reservoir of CV and producing a distinctive network beneath the cortex, not always well visible; this feature has never been reported in other frontoniids. The ciliature connected with the oral region presented 4–5 vestibular, 4–5 postoral kineties, and a double paroral membrane. The cytostome was situated anteriorly some 1/5 to 1/6 of body length. Peniculi 1, 2, and 3 had the following composition (number of rows of cilia): 5 + 5 (1, 2); the third peniculus started with 4 ciliary rows, but at its posterior end they were reduced to 2–3 rows. Vestibular kineties extended posteriorly, reaching roughly double the length of the oral aperture size (Fig. 1c). Unlike the majority of Frontonia species, F. salmastra showed rhomb-like trichocysts, in cross-view (Fig. 1e). The ciliate always swam in a clockwise direction with respect to the longitudinal body axis when viewed from the posterior end.
As for the ultrastructure, the most relevant feature in the cytoplasm of F. salmastra was the presence of two different types of mitochondria: one had a classical aspect, while the second had an unusual appearance, showing an irregular shape and a very electron-dense, congested matrix, but still clean-cut cristae (Figs. 5a and 6b). Some intermediate forms of mitochondria were detected in TEM (data not shown).
Frontonia salmastra 18S rRNA gene sequence (1669 bp) was deposited in the NCBI database under the following accession number: MH319376. Identity values with other SSU rRNA encoding gene sequences belonging to other Frontonia species ranged from 98.9% (F. subtropica, FJ868202) to 91.9% (F. pusilla, FJ868201) (Table 2). Topology of BI tree suggested F. salmastra a sister taxon of F. subtropica—F. magna—F. sinica group although with limited statistical support (0.90) (Fig. 8). The genus Frontonia is not monophyletic, but forms three distinguishable clades composed of 1) F. canadensis, F. salmastra, F. subtropica, F. magna, F. sinica, F. tchibisovae, and F. mengi; 2) F. paramagna, “F. vernalis,” and F. leucas; 3) F. didieris, “F. ocularis,” F. elegans, and F. pusilla. Clade 3 Frontonia appeared sister of Apofrontonia dohrni although with limited statistical support, and together (Clade 3 Frontonia + Apofrontonia) resulted sister of Paramecium. Peniculia group members (Frontonia, Apofrontonia, Paramecium, Paranassula, Stokesia, and Lembadion) clustered together showing high values of statistical support (1.00/100).
Discussion
Characterization of the Novel HLB
The morphology of the new HLB herein described as an endosymbiont of F. salmastra, especially the IF (Figs. 2, 3, 4, 5, and 6), is quite different from HLBs investigated to date, i.e., true holosporas, Preeria caryophila, and members of the “Ca. Gortzia” genus. Indeed, the IF of the novel HLB exceeds in size the largest representative of the other HLB (i.e. Holospora obtusa), and it is also different in shape and inner structure, showing many periplasmic zones; as for the last feature, somewhat reminiscent of “Ca. Gortzia shahrazadis” [20]. The new HLB is unable to produce the “connecting piece,” the formation used by most of the representatives of Holospora genus to segregate the majority of IFs during cell division (Fig. 3f). To date, the absence of this mechanism of the IFs releasing from the host cell was recorded only for H. parva among Holospora representatives [16].
Some IFs were detected in the cytoplasm of F. salmastra (Fig. 6b) as previously reported in “Ca. Gortzia shahrazadis” [20]. This feature could be a sort of back docking process, which allows the release of IFs from the infected MA through a strategy already observed in some other HLB [24, 60, 61]. We also speculate that the fine fibrillar layer on the IFs envelope observed would be helpful in this case (Fig. 6a). Concerning the infection of new specimens of F. salmastra, the morphology of IFs of this new HLB is undeniably similar to the shape of diatoms, thus it could possibly be hypothesized that this is an adaptation by IF to be better accepted by the ciliate host. Both the above suggestions, although intriguing, require further investigation.
Our phylogenetic trees were coherent with previously published works [18, 25, 62,63,64,65] and we decided to follow the systematic organization which considers Holosporales as an order [64, 65]. Molecular data suggested a close affinity of the new HLB endosymbiont from F. salmastra with Holospora; but low identity values between their 16S rRNA gene sequences indicated that it could be considered more likely a sister to the Holospora clade. Given that the average identity value with respect to members of Holospora (94.2%; Table 1) exceeds the proposed genus-level threshold of 94.5% [66, 67], we can conclude that the new clade corresponds to a new genus. We propose this new clade as “Candidatus Hafkinia simulans”, named after the scientist who first described Holospora clade members, i.e. Waldemar Mordecai Wolff A. Hafkine. The specific name refers to the resemblance of its IF to the shape of a diatom, with diatoms, in our experience being a reliable food source for F. salmastra. The description of this novel taxon of HLB is presented below.
Frontonia salmastra and Other Frontoniids
Frontonia salmastra should be considered as a quite rare representative of brackish water frontoniids. After the first record in Benin in 1986 [38], it has been reported from the littoral zone of the Caspian Sea [59] and in a brief report from Italy [68, 69]. Few findings so far, despite the intense work of Chinese colleagues, who described several brackish and marine-water frontoniids during the last decade [70,71,72,73,74,75,76]. Those authors succeed to retrieve along the Chinese coastline several frontoniids previously described from Europe, USA, and Canada [75, 76]; thus, probably, at least some brackish and marine-water Frontonia species do not have a restricted distribution. According to our results, this could be the case of F. salmastra as well.
Our phylogenetical analyses agree with previous studies regarding Frontonia and Peniculia [72,73,74,75,76,77,78,79,80]. As the first consideration, the monophyly of Frontonia was not supported by the 18S rRNA gene based phylogeny [72, 78,79,80,81,82]. Frontonia salmastra clustered with F. subtropica, F. magna, and F. sinica, but branched off basally with respect to the rest of the subgroup. This is a quite coherent result as all members of this group are brackish water species (although sampled at various salinities) and have, at least, one synapomorphic feature: the double paraoral membrane. Such a composition of the oral structure is absent only from F. sinica, which however shows “several argentophilic lines to left of paroral membrane” [74].
The brackish water F. subtropica [75] is the species showing the highest 18S rRNA gene identity value with F. salmastra (98.9%). The former species can be distinguished from the latter based on the following morphological features: cell dimensions (200 × 70 μm vs. 145 × 50 μm); the presence of a single MI (vs. 2–3 MI); the composition of the peniculi (4 + 4 + 4 vs. 5 + 5 + 4 rows of cilia). Also the CV structure, being likely without canals (vs. a network of canals), could be a character useful for separation. Nevertheless, given the fact that the original description of F. subtropica is unclear in some points (some values reported in the text conflict with those reported in descriptive tables) [75], every such consideration should be cautious.
Unlike the previous studies on brackish water Frontonia species, one of the aims of the present research was to investigate F. salmastra at the ultrastructural level. We observed in its cytoplasm the presence of two morphologically different structures which in our opinion can be attributable to mitochondria: one is a classical mitochondrion, while the second appears rather unusual in aspect. Although the latter should definitely deserve further investigation with methods other than electron microscopy, at present we only speculate that it could be a kind of transient morphology of the classical mitochondrion, especially considering the intermediate forms concurrently observed. As in F. leucas (freshwater species), only classical organelles were observed [57], it might be also possible that this feature is connected to living in a particular habitat (e.g. brackish water). In literature on ciliates, only once to our knowledge have two different mitochondria been reported in the same cell, namely in Euplotes minuta [83].
Finally, during a survey of recent and past literature on the genus Frontonia, we realized that there are some misinterpretations regarding several species, and that these have caused some confusion. For example, F. ocularis, first described by Bullington [55] from the USA and recently redescribed from China [76], should be very likely attributed to the European F. fusca [84,85,86] since they share the same morphological key features. Unfortunately, in the redescription by Pan and colleagues [76] some former species of the genus (i.e. F. salmastra, F. marina, F. frigida [38, 84, 86, 87]) have been neglected and some features we consider important are not mentioned at all (such as number and type of micronuclei, structure of CV, and type of swimming rotation).
Moreover, in all published molecular trees concerning Frontonia the18S rRNA gene sequence with accession number U97110, which is shown in GenBank database as F. vernalis, is present. This sequence is the only one available for that species and was deposited by Hirt and colleagues in 1997, not supported by any publication. Indeed, it was used in a publication only in 2000 by Strüder-Kypke and colleagues [88], but again without providing any morphological data. It is known that Hirt and colleagues worked with freshwater frontoniids hosting Chlorella-like cytoplasmic symbionts, see various publications [56, 89, 90]; these frontoniids do not match the morphotype of F. vernalis described by Ehrenberg [91], Kahl [85], or Bullington [55]. However, the species described by Bullington also does not fit previous descriptions of F. vernalis [85, 91], but rather resembles F. fusca described by Quennerstedt [84] and recently redescribed by Fokin [86]. In this context, we suggest that the sequence U97110 should be only prudently associated with F. vernalis, at least until this ciliate will be redescribed according to a multidisciplinary study approach [92]. Consequently, in the phylogenetic tree herein presented (Fig. 8), the sequence of F. vernalis has been indicated in quotes.
Description of “Candidatus Hafkinia” gen. nov.
Hafkinia (Haf’kini.a; N.L. fem. n. Hafkinia, to the discoverer of Holospora, the bacteriologist Waldemar Mordecai Wolff A. Hafkine): Gram-negative, Alphaproteobacteria, Holosporales, belonging to the Holosporaceae family, sister genus of Holospora. Macronuclear endosymbiont of Frontonia salmastra. Monospecific genus. The type species is “Candidatus Hafkinia simulans” (this study).
Description of “Candidatus Hafkinia simulans” sp. nov.
Hafkinia simulans (Haf’kini.a si.mu.lans; N.L. fem. n. Hafkinia, to the discoverer of Holospora, the bacteriologist Waldemar Mordecai Wolff A. Hafkine; N.L. adj. simulans, imitating, because the infectious form of this endosymbiont resembles the diatoms eaten by its host ciliate). Macronuclear endosymbiont of the free-living ciliate F. salmastra, sampled in a brackish water pond of the “Parco Naturale di Migliarino, San Rossore, Massaciuccoli” (Migliarino, Pisa district, Italy). Infectious form (IF) spindle-shaped or skittles-like, straight form, 8–30 × 2.0–3.5 μm in size. IF shows cellular subcompartments: cytoplasm, periplasm, and recognition tip. IFs with extensive periplasmic space often produce several stripes and dots along the cell body. Rod-shaped reproductive forms (RF) 2–4 × 1.0–1.5 μm in size, with homogeneous cytoplasm.
IFs with shape and size similar to the diatom Pheodactylum tricornutum, the organism eaten by the host ciliate. According to this observed apparent imitation of the food, it has been hypothesized that the endosymbiont developed its morphology to better colonize particular ciliate hosts. No “connecting piece” or killer traits detected. Endosymbiont can perform horizontal transmission between hosts of the same species. Uncultured. Other characteristics are present in generic description. Basis of assignment: 16S rRNA gene sequence (MH319377) and probe Hafk_sim_147 (5′-TGA AGT TTC CTC CAG TTA TTC-3′). Type species of the genus.
References
Preer JR, Preer LB, Jurand A (1974) Kappa and other endosymbionts in Paramecium aurelia. Bacteriol. Rev. 38:113
Görtz HD (1988) Endocytobiosis. In: Görtz HD (ed) Paramecium. Springer Verlag, Berlin, pp 393–405
Fokin SI (2004) Bacterial endocytobionts of Ciliophora and their interactions with the host cell. Int. Rev. Cytol. 236:181–249
Fokin SI (2012) Frequency and biodiversity of symbionts in representatives of the main classes of Ciliophora. Eur. J. Protistol. 48:138–148
Fujishima M (2009) Endosymbionts in Paramecium. In: Fujishima M (ed) Münster: microbiology monograph, vol 12. Springer-Verlag, Berlin, p 252
Bright M, Bulgheresi S (2010) A complex journey: transmission of microbial symbionts. Nat Rev Microbiol 8:218–230
Modeo L, Petroni G, Lobban CS, Verni F, Vannini C (2013) Morphological, ultrastructural, and molecular characterization of Euplotidium rosati n. sp. (Ciliophora, Euplotida) from Guam. J. Eukaryot. Microbiol. 60:25–36
Castelli M, Sassera D, Petroni G (2016) Biodiversity of “non-model” Rickettsiales and their association with aquatic organisms. In: Thomas S (ed) Rickettsiales. Springer, Cham, pp 59–91
Castelli M, Serra V, Senra MVX, Basuri CK, Soares CAG, Fokin SI, Modeo L, Petroni G (2018) The hidden world of Rickettsiales symbionts: “Candidatus Spectririckettsia obscura,” a novel bacterium found in Brazilian and Indian Paramecium caudatum. Microb. Ecol. https://doi.org/10.1007/s00248-018-1243-8
Nitla V, Serra V, Fokin SI, Modeo L, Verni F, Sandeep BV, Kalavati C, Petroni G (2018) Critical revision of the family Plagiopylidae (Ciliophora: Plagiopylea), including the description of two novel species, Plagiopyla ramani and Plagiopyla narasimhamurthii, and redescription of Plagiopyla nasuta Stein, 1860 from India. Zool J Linnean Soc XX, p 1–45. https://doi.org/10.1093/zoolinnean/zly041
Görtz HD (1996) Symbiosis in ciliates. In: Hausmann K, Bradbury PS (eds) Ciliates. Cells as organisms. Fischer, Stuttgart, pp 441–462
Görtz HD, Schmidt HJ (2005) Genus Holospora. In: Garrity et al (eds) Bergey’s manual of systematic bacteriology, vol 2, part C2nd edn. Springer, New York, pp 149–151
Görtz HD, Fokin SI (2009) Diversity of endosymbiotic bacteria in Paramecium. In: Fujishima M (ed) Endosymbionts in Paramecium. microbiology monographs Vol. 12, chapter 6. Springer-Verlag, Heidelberg, pp 132–160
Fokin SI, Görtz HD (2009) Diversity of Holospora bacteria in Paramecium and their characterization. In: Fujishima M (ed) Endosymbionts in Paramecium. Microbiology monograph Vol. 12, chapter 7. Springer-Verlag, Heidelberg, pp 161–199
Sabaneyeva EV, Derkacheva ME, Benken KA, Fokin SI, Vainio S, Skovorodkin IN (2009) Actin-based mechanism of Holospora obtusa trafficking in Paramecium caudatum. Protist 160:205–219
Lanzoni O, Fokin S, Lebedeva N, Migunova A, Petroni G, Potekhin A (2016) Rare freshwater ciliate Paramecium chlorelligerum Kahl, 1935 and its macronuclear symbiont “Candidatus Holospora parva”. PLoS ONE 11(12):e0167928. https://doi.org/10.1371/journal.pone.0167928
Hafkine MW (1890) Maladies infectieuses des paramécies. Ann Inst Pasteur 4:363–379
Potekhin A, Schweikert M, Nekrasova I, Vitali V, Schwarzer S, Anikina A, Kaltz O, Petroni G, Schrallhammer M (2018) Complex life cycle, broad host range and adaptation strategy of the intranuclear Paramecium symbiont Preeria caryophila comb. nov. FEMS Microbiol. Ecol. 94(7):fiy076
Boscaro V, Fokin S, Schralhammer M, Schweikert M, Petroni G (2013) Revised systematics of Holospora-like bacteria and characterization of “Candidatus Goertzia infectiva”, a novel macronuclear symbiont of Paramecium jenningsi. Microb. Ecol. 65:255–267
Serra V, Fokin SI, Castelli M, Basuri CK, Nitla VM, Verni F, Sandeep BV, Kalavathi C, Petroni G (2016) “Candidatus Gortzia shahrazadis”, a novel endosymbiont of Paramecium multimicronucleatum and revision of the biogeographical distribution of Holospora-like bacteria. Front. Microbiol. 7:1704. https://doi.org/10.3389/fmicb.2016.01704
Preer LB (1969) Alpha, an infectious macronuclear symbiont of Paramecium aurelia. J Protozool 16:570–578
Gromov BV, Ossipov DV (1981) Holospora (ex Hafkine 1890) nom. rev., a genus of bacteria inhabiting the nuclei of paramecia. Int. J. Syst. Bacteriol. 31:348–352
Preer JR, Preer LB (1982) Revival of names of protozoan endosymbionts and proposal of Holospora caryophila nom. nov. Int. J. Syst. Bacteriol. 32:140–141
Fokin SI, Brigge T, Brenner J, Görtz H-D (1996) Holospora species infecting the nuclei of Paramecium appear to belong into two groups of bacteria. Eur. J. Protistol. 32(Supp l1):19–24
Fokin SI (2000) Host specificity of Holospora and its relationships with Paramecium phylogeny. Japan J Protozool 33:94
Rautian MS, Wackerow-Kouzova ND (2013) Phylogenetic placement of two previously described intranuclear bacteria from the ciliate Paramecium bursaria (Protozoa, Ciliophora): “Holospora acuminata” and “Holospora curviuscula”. Int. J. Syst. Evol. Microbiol. 63:1930–1933. https://doi.org/10.1099/ijs.0.046631-0
Dohra H, Tanaka K, Suzuki T, Fujishima M, Suzuki H (2014) Draft genome sequences of three Holospora species (Holospora obtusa, Holospora undulata, and Holospora elegans), endonuclear symbiotic bacteria of the ciliate Paramecium caudatum. FEMS Microbiol. Lett. 359:16–18. https://doi.org/10.1111/1574-6968.12577
Jankowski AW (1964) Morphology and evolution of ciliophora. III. Diagnoses and phylogenesis of 53 sapropelebionts, mainly of the order Heterotrichida. Arch. Protistenkd. 107:185–294
Esteban G, Fenchel T, Finlay B (1995) Diversity of free-living morphospecies in the ciliates genus Metopus. Arch. Protistenkd. 146:137–164
von Stein F (1867) Der Organismus der Infusionsthiere. II . Abth 2 Naturgeschichte der heterotrichen Infusorien. Wilhelm Engelmann, Leipzig
Görtz HD, Wiemann M (1987) Colonization of the ciliate Stentor multiformis by three different endocytobionts. Endocyt Cell Res 4:177–184
Balbiani EG (1892) Nouvelles recherches expérimentales sur la mérotomie des infusoires ciliés. Ann Microgr 5:1–25
Görtz HD, Maier G (1991) A bacterial infection in a ciliate from sewage sludge. Endocyt Cell Res 8:45–52
Kirby H (1941) A parasite of the macronucleus of Vorticella. J Parasit 27:311–314
Platt-Rohloff L (1996) Untersuchungen an bakteriellen Endosymbiosen in neu isolierten Ciliaten aus marinen und aus Brackwasser-Biotopen. Inauguraldissertion zur Erlangung des Doktorgrades der Fachbereichs Biologie der Freien Universitat Berlin
Laval M (1970) Presence de bacteries intranucleaires chez Zoothamnium pelagicum (Cilie Peritriche) leur role dans la formation des pigments intracytoplasmiques des zoides. Septième Congrès International la Microscopie Électronique, Grenoble, 403–404
Hausmann K, Hausmann E (1981) Structural studies on Trichodina pediculus (Ciliophora, Peritricha). II. The adhesive disc. J Ultrastrust Res 74:144–155
Dragesco J, Dragesco-Kernéis A (1986) Cilies libres de l’Afrique intertropicale. Introduction a la connaissance et a l’etude des cilies. Faune Tropicale 26:1–559
Skovorodkin IN (1990) A device for immobilization of biological objects in light microscope studies. Cytology (Sankt-Petersburg) 32:301–302
Corliss JO (1953) Silver impregnation of ciliated protozoa by the Chatton-Lwoff technique. Stain. Technol. 28:97–100
Fokin SI, Gortz HD (1993) Caedibacter macronucleorum sp. nov., a bacterium inhabiting the macronucleus of Paramecium duboscqui. Arch. Protistenkd. 143:319–324
Vannini C, Rosati G, Verni F, Petroni G (2004) Identification of the bacterial endosymbionts of the marine ciliate Euplotes magnicirratus (Ciliophora, Hypotrichia) and proposal of “Candidatus Devosia euplotis”. Int. J. Syst. Evol. Microbiol. 54:1151–1156
Don RH, Cox PT, Wainwright BJ, Baker K, Mattick JS (1991) Touchdown’PCR to circumvent spurious priming during gene amplification. Nucleic Acids Res. 19:4008
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Kumar Y, Buchner A, Lai T, Steppi S, Jobb G, Förster W, Brettske I, Gerber S, Ginhart AW, Gross O, Grumann S, Hermann S, Jost R, König A, Liss T, Lüßmann R, May M, Nonhoff B, Reichel B, Strehlow R, Stamatakis A, Stuckmann N, Vilbig A, Lenke M, Ludwig T, Bode A, Schleifer KH (2004) ARB: a software environment for sequence data. Nucleic Acids Res. 32:1363–1371
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41:590–596. https://doi.org/10.1093/nar/gks1219
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52:696–704
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna D, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61:539–542. https://doi.org/10.1093/sysbio/sys029
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods 9:772–772
Fokin SI, Schweikert M, Brummer F, Gortz HD (2005) Spirostomum spp. (Ciliophora, Protista), a suitable system for endocytobiosis research. Protoplasma 225:93–102
Amann RI, Binder BJ, Olson RJ, Chisholm SW, Devereux R, Stahl DA (1990) Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl. Environ. Microbiol. 56:1919–1925
Manz W, Amann R, Ludwig W, Wagner M, Schleifer KH (1992) Phylogenetic oligodeoxynucleotide probes for the major subclasses of proteobacteria: problems and solutions. Syst. Appl. Microbiol. 15:593–600
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarrell DM, Marsh T, Garrity GM, Tiedje JM (2009) The ribosomal database project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 37:141–145. https://doi.org/10.1093/nar/gkn879
Andreoli I, Mangini L, Ferrantini F, Santangelo G, Verni F, Petroni G (2009) Molecular phylogeny of unculturable Karyorelictea (Alveolata, Ciliophora). Zool Scripta 38:651–662
Modeo L, Fokin SI, Boscaro V, Andreoli I, Ferrantini F, Rosati G, Verni F, Petroni G (2013) Morphology, ultrastructure and molecular phylogeny of the ciliate Sonderia vorax with insights into the systematics of order Plagiopylida. BMC Microbiol. 13:40. https://doi.org/10.1186/1471-2180-13-40
Bullington WE (1939) A study of spiraling in the ciliate Frontonia with a review of the genus and a description of two new species. Arch. Protistenkd. 92:10–66
Berninger UG, Finlay BJ, Canter HM (1986) The spatial distribution and ecology of zoochlorellae-bearing ciliate in a productive pond. J Protozool 33:557–563
Fokin SI, Schweikert M (2003) Bacterial endobionts of Frontonia leucas (Ciliophora, Peniculida). Europ J Protistol 39:311–318
Rossi A, Boscaro V, Carducci D, Serra V, Modeo L, Verni F, Fokin SI, Petroni G (2016) Ciliate communities and hidden biodiversity in freshwater biotopes of the Pistoia province (Tuscany, Italy). Europ J Protistol 53:11–19
Alekperov IH (2005) Atlas of freeliving ciliates (classes Kinetofragminophora, Colpodea, Oligohymenophorea, Polyhymenophora). Borcali NPM, Baku 309 p. (in Russian)
Fokin SI, Sabaneyeva EV (1997) Release of endonucleobiotic bacteria Holospora bacillata and Holospora curvata from the macronucleus of their host cells Paramecium woodruffi and Paramecium calkinsi. Endocytobiosis Cell Res 12:49–55
Fokin SI (2015) Release of Holospora-like bacteria in different ciliate species. Abstracts of VII ECOP-ISOP joint meeting. Sevilla, Spain, 279
Hess S, Suthaus A, Melkonian M (2016) Characterisation of “Candidatus Finniella” (Rickettsiales, Alphaproteobacteria), novel endosymbionts of Viridiraptorid amoeboflagellates (Cercozoa, Rhizaria). Appl. Environ. Microbiol. 82:659–670
Szokoli F, Castelli M, Sabaneyeva E, Schrallhammer M, Krenek S, Doak TG, Berendonk TU, Petroni G (2016) Disentangling the taxonomy of Rickettsiales and description of two novel symbionts (“Candidatus Bealeia paramacronuclearis” and “Candidatus Fokinia cryptica”) sharing the cytoplasm of the ciliate protist Paramecium biaurelia. Appl Environ Microb 82:7236–7247
Schrallhammer M, Castelli M, Petroni G (2018) Phylogenetic relationships among endosymbiotic R-body producer: bacteria providing their host the killer trait. Syst. Appl. Microbiol. 41:213–220
Tashyreva D, Prokopchuk G, Votýpka J, Yabuki A, Horák A, Lukeš J (2018) Life cycle, ultrastructure, and phylogeny of new diplonemids and their endosymbiotic bacteria. mBio 9(2):e02447–e02417
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer KH, Whitman WB, Euzéby J, Amann R, Rosselló-Móra R (2014) Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 12:635–645. https://doi.org/10.1038/nrmicro3330
Rosselló-Móra R, Amann R (2015) Past and future species definitions for Bacteria and archaea. Syst. Appl. Microbiol. 38:209–216
Fokin SI, Schrallhammer M, Vannini C, Ferrantini F, Petroni G, Görtz HD (2006b) New Holospora endocytobionts in some common ciliates. In: 25th Wiss Tag Deut Gesel Protozool Liebenwalde (Berlin), 28
Ferrantini F, Fokin SI, Vannini C, Verni F, Petroni G (2007) Characterization of a novel Holospora-like symbiont from Frontonia (Ciliophora, Oligohymenophorea). J. Eukaryot. Microbiol. 54:29S
Long H, Song W, Gong J, Hu X, Ma H, Zhu M, Wang M (2005) Frontonia lynni n. sp., a new marine ciliate (Protozoa, Ciliophora, Hymenostomatida) from Qingdao, China. Zootaxa 1003:57–64
Long H, Song W, Al-Rasheid KAS, Wang Y, Yi Z, Al-Quraishy SA, Lin X, Al-Farraj SA (2008) Taxonomic studies on three marine species of Frontonia from northern China: F. didieri n. sp., F. multinucleata n. sp. and F. tchibisovae Burkovsky 1970 (Ciliophora: Peniculida). Zootaxa 1687:35–50
Gao S, Chen ZG, Shao C, Long HA, Al-Rasheid KA, Song WB (2008) Reconsideration of the phylogenetic position of Frontonia-related Peniculia (Ciliophora, Protozoa) inferred from the small subunit ribosomal RNA gene sequences. Acta Protozool. 47:47–54
Fan X, Chen X, Song W, Al-Rasheid KA, Warren A (2011) Two novel marine Frontonia species, Frontonia mengi spec. nov. and Frontonia magna spec. nov. (Protozoa; Ciliophora), with notes on their phylogeny based on small-subunit rRNA gene sequence data. Int J Syst Evol Microb 61:1476–1486
Fan X, Lin X, Liu W, Xu Y, Al-Farra SA, Al-Rasheid KA, Warren A (2013) Morphology of three new marine Frontonia species (Ciliophora; Peniculida) with note on the phylogeny of this genus. Europ J Protist 49:312–323
Pan X, Gao F, Liu W, Fan X, Warren A, Song W (2013a) Morphology and SSU rRNA gene sequences of three Frontonia species, including a description of F. subtropica spec. nov. (Ciliophora, Peniculida). Europ J Protistol 49:67–77
Pan X, Liu W, Yi Z, Fan X, Al-Rashid K, Lin X (2013b) Studies on three diverse Frontonia species (Ciliophora, Peniculida), with brif notes on 14 marine or brackish congeners. Acta Protozool. 52:35–49
Fokin SI, Andreoli I, Verni F, Petroni G (2006a) Apofrontonia dohrni sp. n. and the phylogenetic relationships within Peniculia (Protozoa, Ciliophora, Oligohymenophorea). Zool Scripta 35:289–300
Andreoli I, Fokin SI, Verni F, Petroni G (2007) Phylogenetic relationships within Frontoniids. J. Eukaryot. Microbiol. 54:28S
Cai X, Wang C, Pan X, El-Serehy HA, Mu W, Gao F, Qiu Z (2018) Morphology and systematics of two freshwater Frontonia species (Ciliophora, Peniculida) from northeastern China, with comparisons among the freshwater Frontonia spp. Europ J Protistolo 63:105–116
Xu Y, Gao F, Fan X (2018) Reconsideration of the systematics of Peniculida (Protista, Ciliophora) based on SSU rRNA gene sequences and new morphological features of Marituja and Disematostoma. Hydrobiologia 806:313–331
Chen Y, Zhao Y, Pan X, Ding W, Al-Rasheid KA, Qiu Z (2014) Morphology and phylogeny of a new Frontonia ciliate, F. paramagna spec. nov. (Ciliophora, Peniculida) from Harbin, Northeast China. Zootaxa 3827:375–386
Zhao Y, Yi Z, Gentekaki E, Zhan A, Al-Farraj SA, Song W (2016) Utility of combining morphological characters, nuclear and mitochondrial genes: an attempt to resolve the conflicts of species identification for ciliated protists. Mol Phylogen Evol 94:718–729
Jurand A, Lipps HJ (1973) Two types of mitochondria in Euplotes minuta. Arch. Protistenkd. 115:133–136
Quennerstedt A (1869) Bidrag till Sveriges Infusorie-fauna III. Acta Univ Lund 6:1–35
Kahl A (1931) Holotricha. In: Dahl F (ed) Die Tierwelt Deutschlands and der angrenzenden Meeresteile. 21 Teil. Urtiere oder Protozoa. G. Fisher, Jena, pp S.181–S.398
Fokin SI (2008) Rediscovery and characterisation of Frontonia fusca (Quennerstedt, 1869) Kahl, 1931 (Ciliphora, Peniculia). Denisia 23:251–259
Petz W, Song W, Wilbert N (1995) Taxonomy and ecology of the ciliate fauna (Protozoa, Ciliphora) in the endopagial and pelagial of the Weddell Sea, Antarctica. Stapfia 40:1–223
Strüder-Kypke MC, Wright ADG, Fokin SI, Lynn DH (2000) Phylogenetic relationships of the subclass Peniculia (Oligohymenophorea, Ciliophora) inferred from small subunit rRNA gene sequences. J Euk Microbiol 47:419–429
Finlay BJ, Berninger UG, Stewart LJ, Hindle RM, Davidson W (1987) Some factors controlling the distribution of two pond-dwelling ciliates with algal symbionts (Frontonia vernalis and Euplotes daidaleos). J Protozool 34:349–356
Esteban GF, Fenchel T, Finlay BJ (2010) Mixotrophy in ciliates. Protist 161:621–641. https://doi.org/10.1016/j.protis.2010.08.002
Ehrenberg CG (1838) Die infusionsthierchen als vollkommene organismen. Voss, Leipzig
Fokin SI (2017) The genus Frontonia Ehrenberg, 1833. Unfinished story. Inter Congr Protistol 30th July–4th August 2017. Prague, Czech Republic. Book of abstracts:77
Acknowledgments
The authors are grateful to Simone Gabrielli for his patient and precious assistance in tree editing.
Funding
This research was supported by a grant from Italian Research Ministry (MIUR – Ministero Italiano dell’ Università e della Ricerca) to S.I. Fokin; and by the PRA_2018_63 project from University of Pisa to G. Petroni.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Fokin, S.I., Serra, V., Ferrantini, F. et al. “Candidatus Hafkinia simulans” gen. nov., sp. nov., a Novel Holospora-Like Bacterium from the Macronucleus of the Rare Brackish Water Ciliate Frontonia salmastra (Oligohymenophorea, Ciliophora): Multidisciplinary Characterization of the New Endosymbiont and Its Host. Microb Ecol 77, 1092–1106 (2019). https://doi.org/10.1007/s00248-018-1311-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-018-1311-0