Abstract
Methanogenic archaea survive under aerated soil conditions in paddy fields, and their community is stable under these conditions. Changes in the abundance and composition of an active community of methanogenic archaea were assessed by analyzing mcrA gene (encoding α subunit of methyl-coenzyme M reductase) and transcripts during a prolonged drained period in a paddy-upland rotational field. Paddy rice (Oryza sativa L.) was planted in the flooded field and rotated with soybean (Glycine max [L.] Merr.) under upland soil conditions. Soil samples were collected from the rotational plot in the first year, with paddy rice, and in the two successive years, with soybean, at six time points, before seeding, during cultivation, and after harvest as well as from a consecutive paddy (control) plot. By the time that soybean was grown in the second year, the methanogenic archaeal community in the rotational plot maintained high mcrA transcript levels, comparable with those of the control plot community, but the levels drastically decreased by over three orders of magnitude after 2 years of upland conversion. The composition of active methanogenic archaeal communities that survived upland conversion in the rotational plot was similar to that of the active community in the control plot. These results revealed that mcrA gene transcription of methanogenic archaeal community in the rotational field was affected by a prolonged non-flooding period, longer than 1 year, indicating that unknown mechanisms maintain the stability of methanogenic archaeal community in paddy fields last up to 1 year after the onset of drainage.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
CH4 is the second most important greenhouse gas, after CO2, and paddy fields are one of its major emission sources [1]. Methanogenic archaea are strict anaerobes, favor highly reduced, anoxic growth conditions [2], and produce CH4 in the paddy field soil [3]. However, methanogenic archaea survive well under aerated conditions in the paddy fields and their community is stable under these conditions [4]. The population and community composition did not change in the drained periods during wheat (Triticum aestivum L.) cultivation in winter after the paddy rice (Oryza sativa L.) was cropped with flooding in the summer [4,5,6,7,8]. On the other hand, the composition and abundance of methanogenic archaeal community were affected by the upland conversion of paddy fields for one or more years in paddy-upland rotational fields where soybean (Glycine max [L.] Merr.) was cultivated under drained conditions in the summer instead of paddy rice. In the rotational fields, the copy number of mcrA gene that encodes the α subunit of methyl-coenzyme M reductase decreased to about one tenth in comparison with that in the consecutive paddy (control) field. In addition, some members of the order Methanosarcinales were fatally affected, as assessed by the analysis of 16S ribosomal RNA (rRNA) gene by polymerase chain reaction (PCR)-denaturing gradient gel electrophoresis (DGGE) [9]. In contrast, Scavino et al. [10] showed that 16S rRNA and mcrA gene copy numbers, and the composition of methanogenic archaeal community estimated by terminal restriction fragment length polymorphism (T-RFLP) of 16S rRNA genes, were not significantly affected by the pasture following the irrigated rice cultivation. Watanabe et al. [7, 8] observed fluctuations in the ratios of 16S rRNA/16S rRNA genes and mcrA transcripts/mcrA genes and thus variations in the composition of the active communities passing from flooded to drained conditions in a double cropping paddy field. However, almost no changes in the DGGE band patterns of 16S rRNA were found [7]. In addition, mcrA transcript levels fluctuated within one order of magnitude during a year [8] or during rice cultivation [11]. Recently, Breidenbach and Conrad [12] and Breidenbach et al. [13] also reported that 16S rRNA copy numbers of methanogenic archaea were similar by comparing fields with only rice and fields with rice-maize rotation. Nevertheless, the response of an active methanogenic archaeal community to upland conversion in paddy-upland rotational fields where a prolonged drained period lasts over 1 year has not been investigated thus far. Under those conditions, the composition and abundance of methanogenic archaeal community may be more affected than under double cropping conditions.
Here, we investigate the effect of upland conversion of a more than 1-year duration on an active community of methanogenic archaea in a paddy-upland rotational field. The abundance and composition of the active methanogenic archaeal community were estimated by analyzing mcrA transcripts as already reported [8, 11, 14,15,16]. The present study aims to clarify to what degree the abundance of the active methanogenic archaeal community decreases and which members of the active community are affected by the long duration of upland conversion compared with its impact on the extant community assessed by DNA analyses. The composition and abundance of active methanogenic archaeal community were examined by T-RFLP, clone library analyses, and quantitative PCR (qPCR) methods by targeting mcrA genes and their transcripts over 3 years in a paddy-upland rotational field.
Materials and Methods
Sampling Sites and Soil Samples
Soil samples were collected from an experimental paddy field located in NARO Tohoku Agriculture Research Center (Daisen [Omagari], Akita, Japan; 39° 29′ N, 140° 29′ E) in 2012, 2013, and 2014, designated as “E” field in a previous study [9]. The paddy field soil was Gray Lowland soil (Typic Fluvaquents) [17]. The paddy field consisted of a rotational plot and a consecutive paddy rice plot (as a control), with two replicates each. The rotational field history since 1982 is shown in Table 1. The information pertaining to the plots and the physical and chemical properties of the soil have already been reported by Shirato et al. [17] and Liu [9]. Soil samples for messenger RNA (mRNA) and DNA analyses of the methanogenic archaea were collected from the plow layer, at 5–10 cm depth, at four locations/plot, on July 10, 2012; April 23, July 30 and November 1, 2013; and on April 10 and July 31, 2014. The soils collected at four points were mixed well to obtain a composite sample for each of two replicate plots. Soil samples for RNA extraction were dispensed into extraction tubes, and immediately frozen in liquid nitrogen. The tubes were transported to the laboratory in a styrene foam box filled with dry ice and stored at − 80 °C until use. The remaining soil samples were stored at 4 °C, passed through a 2-mm mesh sieve and used for measurements of CH4 production potential and moisture content. Soil moisture was determined at six sampling time points by placing in an oven at 105 °C for 24 h and is shown in Table S1.
Nucleic Acid Extraction and RT
Nucleic acids (DNA/RNA) were extracted from 0.4 to 0.7 g soil samples according to modified protocols of Lueders et al. [18] and Murase et al. [19]. Briefly, prior to nucleic acid extraction, stored soil samples were treated with 300 μL RNAlater-ICE (Invitrogen Life Technologies, Carlsbad, CA, USA) and the preparations were left at − 20 °C for 12 h. DNA and RNA (110 μL) were coextracted and purified on filtration columns (Zymo-Spin™ IV-HRC column, Zymo Research, Irvine, CA, USA). An aliquot from each nucleic acid extract (50 μL) was further treated with DNase I (Promega, Madison, WI, USA) to remove DNA. RNA was suspended in RNase-free TE buffer and stored at − 80 °C. DNA removal was confirmed by PCR using mcrA primer set described below and the RNA template. Complementary DNA (cDNA) was synthesized using PrimeScirpt®RT reagent Kit (TaKaRa, Otsu, Japan) and random hexamer primers. Nucleic acid extracts not treated with DNase I were directly used as DNA templates for further DNA-based analyses.
Quantification of mcrA Gene Copy Numbers and Transcripts
The abundance of methanogenic archaea was determined by quantitative PCR (qPCR)-targeting of mcrA gene and its transcripts. qPCR was performed with Thermal Cycler Dice Real Time System (TaKaRa) and a specific primer set (mcrA-f: 5′-GGT GGT GTM GGA TTC ACA CAR TAY GCWACA GC-3′ and mcrA-r: 5′-TTC ATT GCRTAG TTW GGR TAG TT-3′) [20] following the procedure of Liu et al. [21]. Mixtures of known quantities of mcrA gene fragments (101–107 copies) or mcrA transcripts (103–109 copies) from three methanogenic archaeal strains were used as a reference, according to Watanabe et al. [22]. Approximately 3 ng DNA and 20 ng RNA were used as templates in a reaction. Blanks (negative controls) were always run, with water as template. Standard curve was obtained, with an average slope of − 3.491, 95.5% efficiency, average y intercept of 47.99, and R 2 = 0.999. Amplification of non-specific DNA fragments was not detected during dissociation curve analysis indicating good specificity of the primer set in the reactions with DNA and cDNA samples.
Measurements of CH4 Production Potential
To measure CH4 production, three replicates of each soil sample were statically incubated for 2 months. Briefly, 20 g fresh soil was placed into a 120-mL glass vial. The vials were capped with butyl stoppers and aluminum seals. The headspace was flushed with N2 for 3 min, pressurized to 111 kPa, and then incubated at 25 °C in the dark. CH4 concentration in the headspace was measured every 5–7 days using a gas chromatograph (GC-14B, Shimadzu, Kyoto, Japan).
T-RFLP Analysis of Methanogenic Archaeal Community
T-RFLP analysis of the methanogenic archaeal mcrA gene and its transcripts was conducted using the primer set mcrA-f/mcrA-r, where the forward primer was labeled with 6-carboxyfluorescein. The PCR reactions were carried out twice for DNAs and cDNAs from the soil samples for each of two replicate plots with a TaKaRa PCR Thermal Cycler (TaKaRa). The 50 μL mixture contains 1 μL of template DNA or cDNA, 2.5 μmol of forward and reverse primers, 5.0 U of Ex-Taq polymerase (TaKaRa), 25 μmol of Ex-Taq buffer (TaKaRa), 25 μmol of dNTP mixture (TaKaRa), and was filled up with Milli-Q water. The PCR amplification program was as follows: initial denaturation at 95 °C for 90 s, followed by 35 cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s and extension at 72 °C for 90 s, and a final extension at 72 °C for 6 min. PCR amplicons were purified with NucleoSpin Gel and PCR Cleanup Purification Kit (Macherey-Nagel, Duren, Germany), and approximately 30 ng amplicons were subjected to a Klenow fragment treatment to eliminate PCR bias caused by partially single-stranded amplicons [23]. After purification, all the PCR amplicons were digested with MspI (TaKaRa) at 37 °C for 2 h [24] and re-purified by ethanol precipitation. Digestion reaction products were mixed with 0.25 μL size standard LIZ600 (Applied Biosystems, Foster City, CA, USA) and 9.75 μL Hi-Di formamide (Applied Biosystems). Fragments were separated according to size using ABI Prism 3130 Genetic Analyzer (Applied Biosystems) in GeneScan mode. T-RFLP profiles were obtained and analyzed using Peak Scanner Software v 1.0 (Applied Biosystems). The relative abundance of all terminal restriction fragments (T-RFs) in each sample was calculated by dividing the individual peak height by total peak height. T-RFs with the average proportion of two replicates < 1% were eliminated from the profile of each sample.
Clone Library Analysis of mcrA Genes and Their Transcripts
For soil samples collected on July 10, 2012 and July 31, 2014, mcrA gene fragments were PCR-amplified twice from DNAs and cDNAs for each of two replicate plots using the same primer set, mcrA-f/mcrA-r, and the amplification procedure as mentioned above. PCR products were purified using NucleoSpin Extract II (Macherey-Nagel, Duren, Germany) and the preparations from the two replicate plots were combined. Then, the mixture was subjected to a ligation reaction into pT7-Blue-T-Vector (Novagen, San Diego, USA) using Ligation Solution I (Takara) and used to transform competent cells of Escherichia coli XLI-Blue (Toyobo, Osaka, Japan), according to the manufacturer’s instructions. Positive clones were verified by blue/white selection and by colony PCR with the primer set and the amplification procedure specified above. Plasmid DNA was extracted using NucleoSpin Plasmid EasyPure kit (Macherey-Nagel). Clones (96) were randomly selected and sequenced using BigDye Terminator v 3.1 Cycle Sequencing Kit with ABI 3530 Genetic Analyzer (Applied Biosystems). The obtained sequences were virtually translated into amino acid sequences using EMBOSS Transeq program (http://www.ebi.ac.uk/Tools/st/emboss_transeq/). The closest relatives of the translated amino acid sequences were searched using BLAST algorithm on NCBI website (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The sequences were aligned using ClustalW v 1.83 on DDBJ website (http://clustalw.ddbj.nig.ac.jp), with default parameters. Operational taxonomic units (OTUs) and Chao1 and Shannon indices were determined at 90% sequence similarity threshold by MOTHUR [25], according to a previous study [8]. Eight clone libraries constructed from mcrA gene and transcripts in 2012 and 2014 were compared by non-metric multidimensional scaling (NMDS) using the relative proportion of OTUs of clones in each library. Phylogenetic tree of amino acid sequences of the representative clones from DNAs, cDNAs, and related mcrA gene fragments was constructed using neighbor-joining method in MEGA6 [26].
Statistical Analysis
Logarithmically transformed mcrA gene copy numbers and transcript abundances, as well as CH4 production measurements were subjected to one-way ANOVA and Tukey’s test to determine the significance of differences between groups using Microsoft Excel 2013 for Windows (Microsoft Corp., Tokyo, Japan) and Ekuseru-Toukei 2012 (Social Survey Research Information Co., Ltd., Tokyo, Japan) at P < 0.05. NMDS analysis on the relative T-RF abundance and relative OTU proportion data and the Wilcoxon signed-rank test for the Shannon index were performed using PAST3 [27].
Results
Effect of Upland Conversion on Abundance of mcrA Genes and Transcripts
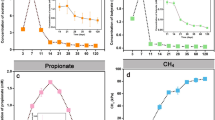
Copy numbers of methanogenic archaeal mcrA genes ranged from 7.2 × 107 to 4.0 × 108 and from 6.7 × 108 to 1.8 × 109 copies g−1 dry soil in the rotational and control plots, respectively (Fig. 1a). Levels of mcrA transcripts in methanogenic archaea ranged from 2.4 × 105 to 1.1 × 109 and from 4.5 × 108 to 6.9 × 109 copies g−1 dry soil in the rotational and control plots, respectively. Abundance of mcrA transcripts ranged from 1.1 × 109 to 3.0 × 108 in the rice growing period of 2012 to the soybean harvesting period of 2013 after 1 year of upland conversion but drastically decreased to 3.2 × 105 copies g−1 dry soil in the rotational plot afterwards (Fig. 1b). Copy numbers of the methanogenic archaeal mcrA genes were significantly lower in the rotational plot than in the control plot (P < 0.01), with some fluctuations among the sampling occasions in the both plots. The abundance of methanogenic archaeal mcrA transcripts was significantly lower in the rotational plot than in the control plot (P < 0.01), with the decrease by about three orders of magnitude, after 1 year of upland conversion in the rotational plot. The abundance of mcrA transcripts in the rotational plot until July of 2013 was not significantly different from that in the control plot.
mcrA gene copy numbers (a) and transcript abundances (b) in methanogenic archaeal communities from six sampling points in 2012, 2013, and 2014. The numbers are presented as log10 values. BF/BS, before paddy rice flooding or before soybean seeding; CG, crop (rice/soybean) growth period; AH, after crop (rice/soybean) harvesting. Bars represent mean value range from duplicate measurements from two replicate plots. Different letters indicate statistically significant differences (P < 0.05)
CH4 Production Potential
In soil samples collected during drained conditions, CH4 was produced at very low levels in both the control and rotational plots. CH4 was produced under flooded conditions in the control as well as rotational plots in 2012 (Fig. 2) when paddy rice was cultivated in the second year of conversion from soybean to paddy rice (Table 1). Indeed, CH4 production potential of the rotational plot was almost same as that in the control plot when paddy rice was cultivated under flooded conditions. In contrast, only small amounts of CH4 were produced from the soil under drained conditions in the rotational plot during soybean growth in 2013 and 2014. The CH4 production potential showed significantly low values during soybean growing periods in the rotational plot compared with the control plot (P < 0.01). A positive linear correlation was observed between the mcrA transcript/gene abundance ratio (mRNA/DNA) in methanogenic archaeal community and the CH4 production potential in the control plot (Fig. S1).
Cumulative CH4 production potential of the soil in control and rotational plots for 37 days of incubation. BF/BS, before paddy rice flooding or before soybean seeding; CG, crop (rice/soybean) growth period; AH, after crop (rice/soybean) harvesting. Bars represent mean value ranges from triplicate measurements from two replicate plots. Different letters indicate statistically significant differences (P < 0.05)
Composition of Methanogenic Archaeal Community Determined by T-RFLP Analysis of mcrA Gene and Transcripts
Changes in methanogenic archaeal community composition in the rotational and control plots under flooded and drained conditions are shown in NMDS scatter plot of the relative T-RFs of mcrA gene and its transcripts (Fig. 3). Methanogenic archaeal community delineated by mcrA gene analyses showed similarity with subgrouping by the rotational and control plots in the NMDS scatter plot. Based on mcrA transcript analyses, the community in the control plot showed dissimilarity with the community characterized by mcrA gene analyses. In contrast, the community in the rotational plot was more distantly scattered, with a tendency to divide into two subgroups (before and after the soybean cultivation in 2013), which was not as clearly observed in the community delineated by mcrA gene analyses. Overall, the archaeal community composition of the rotational plot was different from the control plot, but the community described by mcrA transcript analyses under drained conditions in the rotational plot showed some similarities with the community based on mcrA gene analysis in the control plot though the stress value (0.193) indicated that the separation/grouping should be interpreted with caution.
Non-metric multidimensional scaling (NMDS) analysis of T-RFLP profiles of mcrA gene and transcripts using relative T-RF abundance at all the sampling points in control and rotational plots. BF/BS, before paddy rice flooding or before soybean seeding; CG, crop (rice/soybean) growth period; AH, after crop (rice/soybean) harvesting. Relative T-RF abundance represents mean value from duplicate measurements for each of two replicate plots
Community Composition of Methanogenic Archaea Examined by Clone Library Analysis of mcrA Genes and Transcripts
In total, eight clone libraries were constructed, and 83–85 and 71–84 clone sequences were retrieved for mcrA genes and transcripts, respectively (Table S2). OTU numbers and diversity indices are shown in Table S2. Based on the Chao1 estimator, which indicates species richness in samples, the clones covered 73–93 and 77–86% of methanogenic archaeal communities in mcrA gene and transcript clone libraries, respectively. As a whole, 19 OTUs were recognized and they were phylogenetically affiliated with Methanomicrobiales, Methanosarcinales, Methanocellales, Methanobacteriales, Candidatus Methanoperedenaceae, and Methanomassiliicoccales (Fig. 4). The Shannon index did not significantly differ between the control and rotational plots for both mcrA gene and transcripts (Wilcoxon signed-rank test; P = 0.63). NMDS scatter plots of the relative OTU proportion of mcrA genes and transcripts in the clone libraries revealed that the proportion of OTUs was different between the rotational and control plots and was also different between the 2012 and 2014 libraries from the rotational plot (Fig. S2). It is interesting that the library of mcrA transcripts from 2014 (the second year of upland conversion) in the rotational plot showed some similarities phylogenetically with that of mcrA transcripts library from 2012 control plot.
Neighbor-joining phylogenetic tree constructed from the deduced amino acid sequences of proteins encoded by mcrA gene and transcripts of representative OTUs and related sequences. Methanopyrus kandleri was used as outgroup. The numbers in parentheses after the OTU number represent ratio of clones obtained from the libraries in the order of mcrA gene in control and rotational plots in 2012, mcrA transcripts in control and rotational plots in 2012, mcrA gene in control and rotational plots in 2014, and mcrA transcripts in control and rotational plots in 2014
The predominant groups in mcrA gene and transcript values libraries were Methanomicrobiales, Methanosarcinales (Methanosarcinaceae and Methanosaetaceae), and Methanocellales (Fig. 5). The community composition in the rotational plot was different from that in the control plot based on both mcrA genes and transcripts. The relative abundance of Methanomicrobiales decreased and that of Methanosarcinales and Methanocellales increased in the rotational plot compared with the control plot.
In mcrA gene analyses, OTU1 (Methanosarcinaceae), OTUs 2 and 8 (Methanosaetaceae), and OTU5 (Methanocellales) were dominant groups in the rotational plot, whereas OTUs 3 and 4 (Methanomicrobiales) and OTU5 (Methanocellales) were dominant groups in the control plot (Fig. S3a). In addition, some OTUs were present only in the control (OTUs 9, 10, 11, 12, and 16) and rotational (OTU 8, 13, and 15) plots. In the transcript analyses, OTU3 (Methanomicrobiales) and OTU2 (Methanosaetaceae) were dominant groups in the rotational plot, whereas OTUs 3, 4, and 6 (Methanomicrobiales), as well as OTU2 (Methanosaetaceae) were dominant groups in the control plot (Fig. S3b). OTU8 was present only in the rotational plot. Furthermore, in the rotational plot, OTUs 5 and 8 were mainly dominant groups in 2012 during rice cultivation, and OTUs 4 and 6 were mainly dominant groups in 2014 (the second year of upland conversion).
Discussion
Changes in mcrA Gene and Transcript Abundance and CH4 Production Potential Caused by Upland Conversion
The abundance of mcrA genes was ca. one order of magnitude lower in the rotational plot than in the control plot (Fig. 1a), confirming the findings in the same field in the years 2008–2011 in a previous study [9]. The abundance of mcrA transcripts in the control plot was 108–109 copies g−1 dry soil, with fluctuations within one order of magnitude (Fig. 1b). The fluctuation range coincided with data on the fluctuations of mcrA transcript abundance during the flooding and drainage of paddy fields [8, 11, 15]. Indeed, there was a positive correlation between the ratio of mcrA transcript/gene numbers (mRNA/DNA) and CH4 production potential in the control plot (Fig. S1). The abundance of mcrA transcripts in the rotational plot was comparable with that in the control plot from 2012 (CG-12) when paddy rice was cultivated on both fields under flooded conditions. It remained at 108 copies g−1 dry soil, slightly lower, in the rotational plot than in the control plot for the two following sampling time points (BF/BS-13 and CG-13) in the subsequent year though the difference was not significant, when soybean was grown under drained conditions. However, the abundance of mcrA transcripts drastically decreased after harvest (AH-13) and was at 105 copies g−1 dry soil in the second year of upland conversion (BF/BS-14 and CG-14) (Fig. 1b). This was in contrast to that mcrA gene abundance was not conspicuously reduced at the time (Fig. 1a) and CH4 production potential of the soil remained very low but still positive in the rotational plot during the first and second years of upland conversion. The drastic decrease in mcrA transcript abundance implies that the transcriptional activity of mcrA gene in the methanogenic archaeal community was greatly reduced in the rotational plot on account of > 1 year upland conversion, indicating a detrimental effect of prolonged dry and oxic soil conditions on the active community of methanogenic archaea. The transcriptional activity of mcrA gene in methanogenic archaea is probably decreased by oxygen exposure, low moisture content, and no substrate availability in the soil under upland conditions, which has been in part demonstrated by a moderate (within one order of magnitude) decrease in the mcrA transcript levels in previous studies [8, 11, 14, 15]. The decrease in mcrA transcript number in the rotational plot by more than three orders of magnitude suggested a much more severe impact of the soil conditions, which was probably potentiated by a long, 2 year, duration of upland conversion. The large difference between mcrA gene and transcript abundances may indicate that majority of the extant community of methanogenic archaea was dormant, with almost no activity, e.g., “maintenance or survival mode” [28], under these conditions although resistant forms of cells (e.g., spores) have not been found in methanogenic archaea.
Interestingly, at least by the soybean growth period (CG-13) in the first year of upland conversion, mcrA transcript level was almost the same as that during flooding before upland conversion (CG-12) (Fig. 1b). How could methanogenic archaea sustain a high abundance of mcrA transcripts within 1 year after the onset of drainage? Firstly, some methanogenic archaea possess oxidative stress defense mechanisms, based on enzymes, such as superoxide dismutase and catalase, which alleviate the damage caused by oxygen exposure [29, 30]. In addition, protection afforded by minerals such as pyrite [31] and the encystment by soil protozoa [32] might play a role in tolerance of dry/oxic conditions. Secondly, methanogenic archaea may resist unfavorable soil conditions by maintaining the abundance of mRNAs or ribosomal RNAs. In fact, the number of 16S rRNAs was also measured in our soil samples. The number was sustained at 2.7 × 1010, 4.9 × 1010, and 6.2 × 1010 copies g−1 dry soil in CG-12, BS-13, and CG-13 samples, respectively, whereas it decreased to about one tenth, 5.6 × 109 copies g−1 dry soil, in AH-13. Maintenance of a high 16S rRNA number was also reported for drained conditions in rice-maize rotational fields under double cropping system [12, 13]. However, it is not clear how methanogenic archaea maintain their transcriptional activities or prevent mRNA/rRNA degradation under drained soil conditions up to 1 year after the onset of drainage. The synthesis and maintenance of long-lived rRNAs/mRNAs under extreme energy limitation [28] might in part explain these findings. CH4 production potential remained positive, but very low, in the soil samples of the rotational plot during upland conversion (Fig. 2), suggesting that methanogenic archaea obtained only small quantities of maintenance energy from methanogenesis, the sole energy source in methanogenic archaea, under those circumstances. The universal distribution of fermentative hydrogen production in terrestrial ecosystems [33] may comprise, in part, energy supply for hydrogenotrophic methanogenic archaea in the soil. In addition, hydrogen production by nitrogen-fixing bradyrhizobia in soybean nodules [34] might have also made some contributions to the energy supply in the rotational plot. Suitable micro-niches might exist in the drained upland soil allowing anaerobic decomposition, providing methanogenic substrates [35, 36]. Indeed, acetate was found in upland soils [37] and degradation of organic matter, such as lignin and monosaccharides, in anaerobic micro-sites by facultative aerobes was reported in aerobic forest soils [38, 39]. In the future study, further investigation should be conducted to elucidate the mechanisms of this phenomenon by monitoring oxygen concentrations and redox potentials in the soil or the micro-niches. Additional possible factor that may have facilitated retardation of degradation of mRNAs/rRNAs in the soil could be the temperature. Low temperature in the area of the investigated field, from about − 3 to 7 °C (the monthly average air temperature from November 2012 to April 2013 at Omagari observatory [39° 29′ N, 140° 30′ E], Japan Meteorological Agency; http://www.jma.go.jp/jp/amedas), was associated with the snowfall in the winter season, from the crop harvesting period to the flooding of the field or soybean seeding. Although higher temperature, from 14 to 24 °C, from May to October during the crop-growing period should have counteracted the retarding effect, mcrA transcript level was nonetheless still sustained. Since Methanosarcina barkeri exhibited high resistance to exposure to air and high temperature after desiccation [40], the drying caused by drainage in the paddy rice harvesting period (Table S1) may have induced some archaeal mechanism to survive the soil conditions after the onset of drainage. Further studies need to be conducted to characterize the mechanisms by which methanogenic archaea maintain high transcript levels even 1 year after the onset of drainage. These mechanisms would be responsible for the high stability of methanogenic archaeal community in paddy fields used for the cultivation of paddy rice under flooded conditions at least once a year, including fields with double cropping systems with upland crop cultivation during the annual winter/dry season.
Robust and Thriving Members of the Methanogenic Archaeal Community During Upland Conversion
Almost all methanogenic archaeal members identified by mcrA transcript analysis in the rotational plot during paddy rice cultivation under flooded conditions (CG-12) thrived after 2 years of upland conversion (CG-14), as indicated by the order/family and OTU levels (Fig. 5 and Fig. S3). The dominant members of methanogenic archaeal community were Methanomicrobiales, Methanosarcinales, and Methanocellales (Fig. 5). These have also been identified in a subtropical aerated soil [41] and subtropical estuarine marsh soils [42] by DNA-based analyses and in other paddy fields [8, 11, 14, 15] by mcrA transcript analyses. The community compositions constructed from mcrA gene and transcript levels were different in the control and rotational plots, confirming DNA-based findings from the same field [9]. We noted that the relative abundance of methanogenic archaea varied with respect to composition, depending on mcrA gene or transcript analyses, especially Methanosarcinaceae, Methanosaetaceae, and Methanocellales (Fig. 5), and also on the OTU level (Fig. S3).
In addition, some changes in the community composition of methanogenic archaea in the rotational plot brought about by 2-year upland conversion were observed at the OTU level. However, it was not the case of specific groups disappearing during the 2-year upland conversion, indicating that almost all the members of methanogenic archaeal community survived and withstood the prolonged 2-year drainage. It is interesting that the composition of the community surviving during upland conversion in the rotational plot, as assessed by mcrA transcript analysis, was rather similar to the composition of the active community in the control plot (Fig. S2). The surviving community may have represented autochthonous members of the soil, as a seed community for community recovery under probable favorable conditions in the future.
Resilience of a Methanogenic Archaeal Active Community After Plot Re-conversion to Paddy Field
In paddy-upland rotational fields, CH4 emission in the second year, or later, of paddy rice cultivation after upland conversion was comparable with that from fields with consecutively cultivated paddy rice, but the emission in the first year of paddy rice cultivation following upland conversion decreased to 20–50% of that in the consecutive paddy plot [13, 43,44,45]. In the present study, the rotational plot in 2012 was in the second year of paddy rice cultivation under flooding, after 3-year upland conversion (Table 1). The recovery of CH4 emission in the second year of paddy rice cultivation and also the comparable CH4 production potential from both the control and rotational plots (Fig. 2) were correlated well with the recovery of mcrA transcript levels (Fig. 1). Should this relationship between CH4 emission/production and mcrA transcript abundance be also valid in the first year of paddy rice cultivation after upland conversion, mcrA transcript levels cannot be expected to fully recover. This may be a cause of low CH4 emission from rotational fields in the first year of re-conversion to paddy rice cultivation. Once the soil conditions become favorable for methanogenic archaeal growth, an active archaeal community and CH4 production potential can gradually recover in the rotational plot, but full recovery may take ~ 2 years. Further investigation of the rotational plot is needed to verify this hypothesis by monitoring long-term methanogenic archaeal dynamics and CH4 emission/production at more sampling time points under various kinds of rotational intervals in the future study.
Conclusions
High mcrA transcript levels were maintained by the methanogenic archaeal community in the rotational plot and were comparable to those of archaeal community in the control plot by the time of soybean growth in the first year of upland conversion. This finding was probably associated with the high stability of methanogenic archaeal community in paddy fields where paddy rice is at least once a year cultivated under flooded conditions. However, the levels drastically decreased, by more than three orders of magnitude, after 2 years of upland conversion. Almost all members of the active methanogenic archaeal community, as assessed by mcrA transcript analyses, survived and withstood the prolonged 2-year drainage despite some changes in the community composition of methanogenic archaea. The composition of active community in the rotational plot surviving during upland conversion was similar to the composition of active community in the control plot. Community members surviving in the drained soil may have played a role of a seed community, awaiting favorable conditions in the rotational plot. Elucidation of the mechanisms of their survival, together with the ability of methanogenic archaeal community to recuperate after the return to the flooded paddy field in a paddy-upland rotational field in the future study, may provide pivotal information about the control of CH4 emission/production in paddy fields.
References
Ciais P, Sabine C, Bala G, Bopp L, Brovkin V, Canadell J, Chhabra A, DeFries R, Galloway J, Heimann M, Jones C, Le Quéré C, Myneni RB, Piao S, Thornton P (2013) Carbon and other biogeochemical cycles. In: Stocker TF, Qin D, Plattner GK, Tignor M, Allen SK, Boschung J, Nauels A, Xia Y, Bex V, Midgley PM (eds) Climate change 2013: the physical science basis. Contribution of Working Group I to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change. Cambridge University Press, Cambridge, pp. 465–570
Thauer RK, Kaster AK, Seedorf H, Buckel W, Hedderich R (2008) Methanogenic archaea: ecologically relevant differences in energy conservation. Nat Rev Microbiol 6:579–591
Conrad R (2007) Microbial ecology of methanogens and methanotrophs. Adv. Agron. 96:1–63
Watanabe T, Kimura M, Asakawa S (2006) Community structure of methanogenic archaea in paddy field soil under double cropping (rice-wheat). Soil Biol. Biochem. 38:1264–1274
Asakawa S, Hayano K (1995) Populations of methanogenic bacteria in paddy field soil under double cropping conditions (rice-wheat). Biol. Fertil. Soils 20:113–117
Asakawa S, Akagawa-Matsushita M, Koga Y, Hayano K (1998) Communities of methanogenic bacteria in paddy field soils with long-term application of organic matter. Soil Biol. Biochem. 30:299–303
Watanabe T, Kimura M, Asakawa S (2007) Dynamics of methanogenic archaeal communities based on rRNA analysis and their relation to methanogenic activity in Japanese paddy field soils. Soil Biol. Biochem. 39:2877–2887
Watanabe T, Kimura M, Asakawa S (2009) Distinct members of a stable methanogenic archaeal community transcribe mcrA genes under flooded and drained conditions in Japanese paddy field soil. Soil Biol. Biochem. 41:276–285
Liu D, Ishikawa H, Nishida M, Tsuchiya K, Takahashi T, Kimura M, Asakawa S (2015) Effect of paddy-upland rotation on methanogenic archaeal community structure in paddy field soil. Microb. Ecol. 69:160–168
Scavino AF, Ji Y, Pump J, Klose M, Claus P, Conrad R (2013) Structure and function of the methanogenic microbial communities in Uruguayan soils shifted between pasture and irrigated rice fields. Environ. Microbiol. 15:2588–2602
Lee HJ, Kim SY, Kim PJ, Madsen EL, Jeon CO (2014) Methane emission and dynamics of methanotrophic and methanogenic communities in a flooded rice field ecosystem. FEMS Microbiol. Ecol. 88:195–212
Breidenbach B, Conrad R (2014) Seasonal dynamics of bacterial and archaeal methanogenic communities in flooded rice fields and effect of drainage. Front. Microbiol. 5:752–752
Breidenbach B, Blaser MB, Klose M, Conrad R (2016) Crop rotation of flooded rice with upland maize impacts the resident and active methanogenic microbial community. Environ. Microbiol. 18:2868–2885
Yuan Y, Conrad R, Lu Y (2011) Transcriptional response of methanogen mcrA genes to oxygen exposure of rice field soil. Environ. Microbiol. Rep. 3:320–328
Ma K, Conrad R, Lu Y (2012) Responses of methanogen mcrA genes and their transcripts to an alternate dry/wet cycle of paddy field soil. Appl. Environ. Microbiol. 78:445–454
Bridgham SD, Cadillo-Quiroz H, Keller JK, Zhuang Q (2013) Methane emissions from wetlands: biogeochemical, microbial, and modeling perspectives from local to global scales. Glob Change Biol 19:1325–1346
Shirato Y, Yagasaki Y, Nishida M (2011) Using different versions of the rothamsted carbon model to simulate soil carbon in long-term experimental plots subjected to paddy–upland rotation in Japan. Soil Sci. Plant Nutr. 57:597–606
Lueders T, Manefield M, Friedrich MW (2004) Enhanced sensitivity of DNA- and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ. Microbiol. 6:73–78
Murase J, Hida A, Ogawa K, Nonoyama T, Yoshikawa N, Imai K (2015) Impact of long-term fertilizer treatment on the microeukaryotic community structure of a rice field soil. Soil Biol. Biochem. 80:237–243
Luton PE, Wayne JM, Sharp RJ, Riley PW (2002) The mcrA gene as an alternative to 16S rRNA in the phylogenetic analysis of methanogen populations in landfill. Microbiology 148:3521–3530
Liu D, Suekuni C, Akita K, Ito T, Saito M, Watanabe T, Kimura M, Asakawa S (2012) Effect of winter-flooding on methanogenic archaeal community structure in paddy field under organic farming. Soil Sci. Plant Nutr. 58:553–561
Watanabe T, Wang G, Taki K, Ohashi Y, Kimura M, Asakawa S (2010) Vertical changes in bacterial and archaeal communities with soil depth in Japanese paddy fields. Soil Sci. Plant Nutr. 56:705–715
Egert M, Friedrich MW (2005) Post-amplification Klenow fragment treatment alleviates PCR bias caused by partially single-stranded amplicons. J Microbiol Meth 61:69–75
Barbier BA, Dziduch I, Liebner S, Ganzert L, Lantuit H, Pollard W, Wagner D (2012) Methane-cycling communities in a permafrost-affected soil on Herschel Island, Western Canadian Arctic: active layer profiling of mcrA and pmoA genes. FEMS Microbiol. Ecol. 82:287–302
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75:7537–7541
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30:2725–2729
Hammer Ø, Harper DAT, Ryan P D (2001) Paleontological statistics software: package for education and data analysis. Palaeontol Electron 4:1–9
Lever MA, Rogers KL, Lloyd KG, Overmann J, Schink B, Thauer RK, Hoehler TM, Jorgensen BB (2015) Life under extreme energy limitation: a synthesis of laboratory- and field-based investigations. FEMS Microbiol. Rev. 39:688–728
Shima S, Sordel-Klippert M, Brioukhanov A, Netrusov A, Linder D, Thauer RK (2001) Characterization of a heme-dependent catalase from Methanobrevibacter arboriphilus. Appl Environ Microbiol 67:3041–3045
Brioukhanov AL, Thauer RK, Netrusov AI (2002) Catalase and superoxide dismutase in the cells of strictly anaerobic microorganisms. Microbiology 71:281–285
Fetzer S, Bak F, Conrad R (1993) Sensitivity of methanogenic bacteria from paddy soil to oxygen and desiccation. FEMS Microbiol Ecol 12:107–115
Miyaki T, Oyaizu H, Matsumoto S (1996) Methanogenesis from dried and heat treated paddy soil. Microbes Environ 11:51–55
Greening C, Biswas A, Carere CR, Jackson CJ, Taylor MC, Stott MB, Cook GM, Morales SE (2016) Genomic and metagenomic surveys of hydrogenase distribution indicate H2 is a widely utilised energy source for microbial growth and survival. ISME J 10:761–777
Conrad R, Seiler W (1980) Contribution of hydrogen production by biological nitrogen fixation to the global hydrogen budget. J Geophys Res 85:5493–5498
von Fischer JC, Hedin LO (2002) Separating methane production and consumption with a field-based isotope pool dilution technique. Global Biogeochem Cycles 16:8–13
Megonigal JP, Guenther AB (2008) Methane emissions from upland forest soils and vegetation. Tree Physiol 28:491–498
Kusel K, Drake HL (1995) Effects of environmental parameters on the formation and turnover of acetate by forest soils. Appl Environ Microbiol 61:3667–3675
Degelmann DM, Kolb S, Dumont M, Murrell JC, Drake HL (2009) Enterobacteriaceae facilitate the anaerobic degradation of glucose by a forest soil. FEMS Microbiol Ecol 68:312–319
DeAngelis KM, Allgaier M, Chavarria Y, Fortney JL, Hugenholtz P, Simmons B, Sublette K, Silver WL, Hazen TC (2011) Characterization of trapped lignin-degrading microbes in tropical forest soil. PLoS One 6: e19306
Anderson KL, Apolinario EE, Sowers KR (2012) Desiccation as a long-term survival mechanism for the archaeon Methanosarcina barkeri. Appl Environ Microbiol 78:1473–1479
Xie W, Jiao N, Ma CL, Fang S, Phelps JT, Zhu RX, Zhang CL (2017) The response of archaeal species to seasonal variables in a subtropical aerated soil: insight into the low abundant methanogens. Appl Microbiol Biotechnol 101:6505–6515
Tong C, Cadillo-Quiroz H, Zeng ZH, She CX, Yang P, Huang JF (2017) Changes of community structure and abundance of methanogens in soils along a freshwater–brackish water gradient in subtropical estuarine marshes. Geoderma 299:101–110
Kumagai K, Konno Y (1998) Methane emission from rice paddy fields after upland farming. Jpn J Soil Sci Plant Nutr 69: 333–339 (in Japanese with English summary)
Eusufzai MK, Tokida T, Okada M, Sugiyama S, Liu GC, Nakajima M, Sameshima R (2010) Methane emission from rice fields as affected by land use change. Agr Ecosyst Environ 139:742–748
Shiono H, Saito H, Nakagawa F, Kumagai K (2014) Effects of crop rotation and rice straw incorporation in spring on methane and nitrous oxide emissions from an upland paddy field in a cold region of Japan. Jpn J Soil Sci Plant Nutr 85:420–430 (in Japanese with English summary)
Acknowledgements
We thank Professor Emeritus Makoto Kimura from the Graduate School of Bioagricultural Sciences, Nagoya University, for his helpful suggestions and comments.
Funding Information
This work was supported in part by the Environmental Research Projects from the Sumitomo Foundation [Grant number: 143218] and by the Soil e-DNA Project from the Ministry of Agriculture, Forestry, and Fisheries of Japan [Grant number: eDNA-07-101-1].
Author information
Authors and Affiliations
Corresponding author
Additional information
Accession number: Sequences of mcrA genes and transcripts from methanogenic archaea obtained in the course of the present study have been deposited in GenBank/EMBL/DDBJ under accession numbers LC154114-LC154792.
Electronic Supplementary Material
Supplemental Fig. S1
(PPTX 177 kb)
Supplemental Fig. S2
(PPTX 219 kb)
Supplemental Fig. S3
(PPTX 404 kb)
Supplemental Table 1
(DOCX 20 kb)
Supplemental Table 2
(DOCX 24 kb)
Rights and permissions
About this article
Cite this article
Liu, D., Nishida, M., Takahashi, T. et al. Transcription of mcrA Gene Decreases Upon Prolonged Non-flooding Period in a Methanogenic Archaeal Community of a Paddy-Upland Rotational Field Soil. Microb Ecol 75, 751–760 (2018). https://doi.org/10.1007/s00248-017-1063-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-1063-2