Abstract
Industrial wasteland soils with aged PAH and heavy metal contaminations are environments where pollutant toxicity has been maintained for decades. Although the communities may be well adapted to the presence of stressors, knowledge about microbial diversity in such soils is scarce. Soil microbial community dynamics can be driven by the presence of plants, but the impact of plant development on selection or diversification of microorganisms in these soils has not been established yet. To test these hypotheses, aged-contaminated soil samples from a field trial were collected. Plots planted with alfalfa were compared to bare soil plots, and bacterial and fungal diversity and abundance were assessed after 2 and 6 years. Using pyrosequencing of 16S rRNA gene and ITS amplicons, we showed that the bacterial community was dominated by Proteobacteria, Actinobacteria, and Bacteroidetes and was characterized by low Acidobacteria abundance, while the fungal community was mainly represented by members of the Ascomycota. The short-term toxic impact of pollutants usually reduces the microbial diversity, yet in our samples bacterial and fungal species richness and diversity was high suggesting that the community structure and diversity adapted to the contaminated soil over decades. The presence of plants induced higher bacterial and fungal diversity than in bare soil. It also increased the relative abundance of bacterial members of the Actinomycetales, Rhizobiales, and Xanthomonadales orders and of most fungal orders. Multivariate analysis showed correlations between microbial community structure and heavy metal and PAH concentrations over time, but also with edaphic parameters (C/N, pH, phosphorus, and nitrogen concentrations).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The decline of coal-mining activities has left behind large areas of industrial wasteland soils that display high levels of multi-contamination by polycyclic aromatic hydrocarbons (PAHs) and heavy metals (HMs) [1]. During the twentieth century, industrial facilities such as gas or coking plants caused chronic pollution of soils. After an aging period, both PAHs and HMs persisted in the environment due to their low availability/mobility and low degradability/reactivity. Thus, the high concentrations of pollutants and their toxicity have been exerting a selection pressure on microorganisms for decades. These particular environmental conditions potentially enabled the microbial community to adapt, and favored microorganisms able of using pollutant as carbon source or tolerate pollutant by detoxification mechanisms [2, 3]. This long-term process should lead to the development of a new microbial community diversity after a few decades [4, 5]. Yet, little is known about the impact of ancient contamination on microbial diversity in such aged multi-polluted soils. Most studies focus on the short-term modifications of microbial diversity following recent contamination events, mainly in laboratory experiments (i.e., spiking of unpolluted soils) [6–8]. Environmental conditions and edaphic parameters are components that shape the soil microbial communities [9]. Similarly, chronic pollution could also be an important parameter that can select soil microbial diversity. While the short-term impact of contamination tends to decrease microbial abundance, richness and diversity, we hypothesized that in an aged multi-contaminated soil the microbial community would adapt over time toward a unique and rich diversity pattern [4, 5].
Some studies show that vegetation can help to restore contaminated environments. Phytoremediation strategies have been developed based on plants’ contribution to the stabilization, extraction, and degradation of pollutants [10]. While shaping microbial diversity by stimulating some microbial populations, plant root exudates could favor PAH degradation through rhizodegradation processes [11–13] and modify HM speciation, mobility, and availability [10] in the rhizosphere. Among different plant species used in rhizoremediation assays, alfalfa is described as tolerant to pollution and interesting for PAH remediation [14, 15], similarly to other legumes such as red lentil [16]. A contribution of mycorrhizal colonization of plant roots to PAH dissipation [17] and HM tolerance [18] has also been shown. There is an effect of plants on soil microbial biomass and diversity because vegetal litter or residues bring complex organic matter into soils [19] and also pose the question of selection or diversification of microorganisms. As most studies focus on one particular microbial function or population, little is known about the impact of plants and the effect of the rhizosphere on bacterial and fungal richness and diversity in multi-contaminated soils. A dramatic shift in the bacterial community structure in the rhizosphere of sunflower in a creosote-contaminated soil was demonstrated in a pot experiment after 90 days in the greenhouse during PAH degradation [20]. Although fungal community dynamic is mainly driven by plants [21], the diversity of this microbial community needs to be explored. Overall, as the rhizosphere effect could modify soil parameters and fate of pollutants, there is still a need to study the long-term impact of plants in comparison to bare soil, on shaping the microbial communities during natural attenuation processes of aged multi-contaminated soils.
It is well established that environmental parameters impact microbial community composition [9]. Although bacteria and fungi are in constant interaction in soils [22], these two components of the microbial community have a different ecology (e.g., nutrient and carbon source requirement, growth rate…) and respond differently to environmental conditions and stressors. To study the system as a whole, bacterial and fungal diversity must be studied together. To our knowledge, no study has yet evaluated how plants and edaphic parameters drive both fungal and bacterial diversity of a multi-contaminated soil on the long term.
In this context, the aims of our study were to (1) describe the diversity of bacterial and fungal communities in an aged multi-contaminated soil from a former coking plant site (Neuves-Maisons, France), (2) determine the impact of plant development on microbial richness and diversity, (3) characterize the temporal modifications of microbial diversity according to evolution of edaphic parameters over 6 years. To address this issue, we collected soil samples from a field trial where a long-term plant-assisted natural attenuation experiment was being conducted [23]. We studied three replicate plots of planted and non-planted soil, sampled at three time-points (0, after 2 and 6 years). We analyzed bacterial and fungal community abundance using real-time quantitative PCR, and diversity using tag-encoded pyrosequencing of the 16S rRNA gene and ITS amplicons, respectively.
Materials and Methods
Site Description and Sample Collection
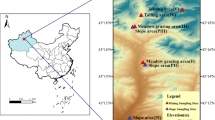
The field trial, previously described by Ouvrard et al. [23], was set up in September 2005. Among the 24 plots filled with an aged PAH- and HM-contaminated soil (NM) from a coking plant wasteland (Neuves-Maisons, Meurthe et Moselle, France), only three replicates of two treatments were studied. The bare soil plots (NM-BS) were kept clear of vegetation by hand weeding and the planted plots (NM-Msm) were sown with alfalfa (40 g of seeds of Medicago sativa var. Europe per plot) in September 2005 and co-inoculated with two mycorrhizal fungal strains (Glomus mosseae and Glomus intra-radices, recently renamed Funneliformis mosseae and Rhizophagus intraradices [24], commercial inoculum from the Institüt für Pflanzenkultur, Germany). The plots were monitored twice a year, in May and September, for 6 years. In September, the alfalfa biomass was cut and dry weight was measured (Table 1). Mycorrhizal colonization of roots was determined by Trypan blue staining as described by [25] and notation was performed using the method described in [26]. Six soil subsamples were collected per plot and mixed to obtain one composite soil sample per plot. After sieving to 5 mm, 500 g were dried and dedicated to soil characterization, and 50 g were stored at −80 °C until DNA extraction. Three sampling dates were studied: samples T0 (September 2005), T4 (September 2007), and T12 (September 2011), i.e., 18 soil samples altogether. Sample characteristics (humidity level, physico-chemical properties, total and extractable heavy metals, and PAH concentrations) were measured at the LAS-INRA laboratory (Arras, France) and are listed in Table 1.
DNA Extraction
Total genomic DNA was extracted from 0.5 g wet weight of each of the 18 soil samples with the FastDNA SPIN Kit for Soil (MP Biomedicals, Illkirch, France), and resuspended in 100 μl of DES (Dnase-Free water). DNA concentrations and quality (purity ratio A 260/A 280) were measured using a spectrophotometer (UV1800, Shimadzu) equipped with a TrayCell unit (Hellma).
Assessment of Fungal and Bacterial Abundance by Real-Time Quantitative PCR
Real-time quantitative PCR was performed as previously described in Cébron et al. [27] and Thion et al. [21]. Primer sets FF390R (5′-GAGGTCTCGTTCGTTATCG-3′)/Fung5F (5′-GGGAACCAGGACTTTTAC-3′) [28] and 968 F (5′-AACGCGAAGAACCTTAC-3′)/1401R (5′-CGGTGTGTACAAGACCC-3′) [29] were used to quantify fungi and bacteria by targeting the 18S rRNA and 16S rRNA genes, respectively. Amplification reactions (20 μl) were carried out using 1 μl of ten-fold diluted DNA (corresponding to 2.8 to 6.5 ng), 10 μl of iQ SYBR Green SuperMix (Bio-Rad), 0.8 μl of each primer (10 μM), 0.4 μl of bovine serum albumin (3 %), 0.2 μl of dimethylsulfoxide, and 0.08 μl of T4gp32 (500 μg/ml) (MP Biomedicals). Quantification was performed using a CFX96 Real-Time PCR detection system (Bio-Rad) and standard plasmid dilution series from 108 to 101 copies μl−1. Data were then expressed as gene copy numbers per gram of dry weight soil after data correction using soil sample humidity values.
PCR and Pyrosequencing
The primer set 515 F (5′-GTG CCA GCM GCC GCG GTA A-3′)/907R (5′-CCG TCA ATT CMT TTR AGT TT-3′) [30] was used to amplify the V4–V5 region of bacterial 16S rRNA genes and the primer set ITS1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′) [31]/ITS2 (5′-GCTGCGTTCTTCATCGATGC-3′) [32] was used to amplify the internal transcribed spacer (ITS1) region of fungal rRNA genes. Primers were combined at the 5′ end with a multiplex identifier (MID, Roche) corresponding to a sequence of 10 sample-specific nucleotides. The PCR mixture (50 μl) contained 5× Taq-&Go™ ready to use PCR Mix (MP Biomedicals), 10 pM of each primer, and 2 μl of ten-fold diluted template DNA (or 2 μl of sterile water for the negative control). Five independent PCR reactions were performed for each sample. The PCR conditions were 94 °C for 5 min; 30 cycles of 94 °C for 30 s, 55 °C or 50 °C for 1 min, 72 °C for 1 min, followed by 72 °C for 10 min for primers 515 F/907R or ITS1F/ITS2, respectively. PCR efficiency was checked by agarose gel electrophoresis (1 %) and visualized after ethidium bromide staining under UV light with a GelDoc XR transilluminator (Bio-Rad).
The five independent PCR products were pooled, purified using the QIAquick PCR Purification Kit (Qiagen, France) and DNA was quantified by using a spectrophotometer (UV1800, Shimadzu) equipped with a TrayCell unit (Hellma) and by visualization and comparison with a mass ladder (Low DNA Mass Ladder, Invitrogen) on 1 % agarose gels as described above. Two equimolar mixes (multiplexed samples) from the 18 samples, one for 16S rRNA genes and one for ITS amplicons, were prepared in 50 μl to reach a total concentration of 40 ng/μl. Pyrosequencing was performed by Beckman Coulter Genomics (Danvers, MA, USA) using a 454/Roche GS-FLX Titanium system.
Analysis of 16S rRNA Genes and ITS Pyrosequencing Data
Bacterial Community Analysis
Ribosomal Database Project (RDP, [33]) pyrosequencing pipeline was used for filtering and trimming of 16S rRNA gene reads based on sff-files. The presence of both forward and reverse primers (max. edit distance of 2) was checked to recover sense and anti-sense sequences. At the end, 482,163 reads passed the length (longer than 300 bp) and quality criteria (exclusion a reads with unresolved nucleotides and a low average quality score below 20) before being distributed into 18 samples according to their MIDs, the 16S rRNA gene reads were filtered and trimmed according to different criteria. Mothur 1.29.2 program [34] was used following previously described analyses steps [35]. Alignment of bacterial sequences used the SILVA database as a template. Based on distance matrix generation, the sequences were clustered and operational taxonomic unit (OTU) was formed to 97 % sequence similarity. To compare all sample conditions, a subsampling to 18,100 sequences per sample was performed. Finally, the taxonomic affiliation was obtained using the RDP database as a template.
Fungal Community Analysis
For the fungal community, 360,477 quality reads of ITS were obtained from the Mothur-based pipeline previously described by Coince et al. [36]. The fungal ITS sequences were distributed into 18 samples according to their MIDs, the reads were cleaned in Mothur 1.20.1 with the default parameters [34]. ITS1 was extracted by using Fungal ITS extractor version 2 [37] and a filtration step by length (>100 bp) was performed. Quality ITS reads were aligned and clustered into OTUs (97 % similarity) and consensus sequences were generated by using Uclust version 3.0 (parameter iddef = 2.97 % similarity). Then, subsampling to 9100 sequences per sample was performed. Consensus sequences were assigned against the UNITE database by using the Basic Local Alignment search Tool algorithm Blastn version 2.2.2.3 [38].
Diversity Analysis
Rarefaction curves were obtained using the Analytic Rarefaction 2.0 tool (Hunt Mountain Software); richness and diversity indices were calculated using EstimateS 9.1.0 [39]. The sequence data generated in this study were deposited in the NCBI Sequence Read Archive (SRA) and are available under the BioProject ID: PRJNA269565 and PRJNA269566 for bacteria and fungi, respectively.
Statistical Analyses
Statistical analyses were performed using XLStat2012 software (Addinsoft). A two-way analysis of variance (ANOVA) followed by a Newman-Keuls post-hoc test was performed to determine statistical differences (p < 0.05) in the microbial communities among the six conditions, depending on time and the presence of a plant cover. Canonical correspondence analysis (CCA) followed by a Monte Carlo permutation test (1000 permutations) in XLStat-ADA 2015 (Addinsoft) was used to evaluate the relationship between environmental factors (plant biomass, Log10 of bacterial and fungal abundances obtained through qPCR and edaphic characteristics that were significantly different (ANOVA, p < 0.05) between samples). Two-dimensional hierarchical clustering was performed on standardized data set (z score) of the most abundant (>2 % in at least one sample) bacterial and fungal OTUs (affiliation at 97 % similarity; i.e. 24 bacterial and 37 fungal OTUs) in the three replicates of the six conditions. This was performed within the XLStat2011-Omics package (Addinsoft). Both samples and OTUs were clustered with Ascending Hierarchical Classification based on distance metric (euclidian), generating dendrograms. Heat maps were reordered by dendrogram clustering to relocate covarying OTUs and samples.
Results
Evolution of Soil Characteristics with Time and Plant Colonization
The edaphic parameters of the NM soil evolved over time as described in Table 1. Over the 6 years, pH values, C/N, available phosphorous, and humidity increased (or were higher at T12), while PAH concentrations, extractable Mn and Zn decreased with time, and total metal concentration remained unchanged.
Bacterial and Fungal Abundance
Over time, bacterial abundance (mean value of 1.69 × 109 ± 0.58 × 109 16S rRNA gene copies g dw soil−1) was about ten-fold higher than fungal abundance (mean value of 2.14 × 108 ± 0.15 × 108 18S rRNA gene copies g dw soil−1; Fig. 1). Bacterial abundance was significantly higher (p < 0.05) at T0 and then decreased with time. Conversely, fungal abundance remained constant over time. In the Msm plots, no colonization of roots by mycorrhizal fungi was observed at T4, while the frequency of mycorrhizal roots reached 24 % at T12 and arbuscules were observed (Table 1). The presence of vegetation had no impact on bacterial abundance, while it enhanced fungal abundance: it was significantly higher (p < 0.05) in MsmT4 samples when the biomass of alfalfa colonizing the Msm plots reached its highest value (1693 ± 156 g plot−1; Fig. 1). Consequently, the ratio fungi to bacteria was higher in MsmT4 soil compared to the other samples.
Abundance of fungal and bacterial (18S and 16S rRNA gene copy numbers) communities and alfalfa biomass over time (T0: September 2005, T4: September 2007 and T12: September 2011) for the two BS (bare soil) and Msm (planted with Medicago sativa and inoculated with mycorrhizal fungi) treatments. Different letters indicate significant differences among plots (p < 0.05) resulting from a two-way analysis of variance (ANOVA) followed by Neuman-Keuls multiple comparison test; abc, AB, and A′B′ refer respectively to statistics on 16S rRNA gene copy numbers. g of dry weight soil−1, 18S rRNA gene copy numbers. g of dry weight soil−1 and alfalfa biomass
Bacterial and Fungal Richness and Diversity
After filtering steps, 88 and 82 % of 16S rRNA genes and ITS reads were kept for further analyses (Table 2). The rarefaction analysis revealed that plateau levels were not reached in any of the samples (Supplementary Figure S1). Based on complete data, we found ten times more OTUs for the bacterial (1963 to 3057 OTUs g dw soil−1) than for the fungal community (284 to 592 OTUs g dw soil−1; Table 2). Similarly, the Chao1 estimator of OTU richness predicted a ten-fold higher richness for the bacterial community (mean of 6350 estimated species at T0) as compared to the fungal community (mean of 463 estimated species at T0; Table 2). Bacterial richness was neither influenced by the presence of a plant cover or by time (Table 2 and Supplementary Figure S1). By contrast, fungal richness increased significantly (p < 0.05) with time and was significantly higher (p < 0.05) under the MsmT4 condition (Table 2). The Shannon and inverse of Simpson indices of bacterial and fungal diversity increased significantly (p < 0.05) with time and with the presence of plants at MsmT4, as well as at T12 for fungi (Table 2).
Taxonomic assignment of OTUs allowed us to identify at least 17 bacterial phyla and 5 fungal phyla (Fig. 2). The bacterial community was mainly composed of Proteobacteria (57.4 ± 13.4 %), Actinobacteria (13.2 ± 7.7 %), Bacteroidetes (6.0 ± 1.9 %) and unclassified bacteria (2.4 ± 1.0 %). Proteobacteria were dominated by members of the Gamma- (21.3 ± 8.8 %), Alpha- (19.0 ± 1.5 %) and Beta-Proteobacteria (15.0 ± 6.4 %) classes. The fungal community was mainly composed of Ascomycota (53.6 ± 9.4 %), unclassified fungi (41.3 ± 8.2 %) and Basidiomycota (4.2 ± 5.2 %).
We found 24 and 37 dominant bacterial and fungal OTUs, represented by at least 2 % of the sequences for at least one sample (Fig. 3). They accounted for 28.6 ± 7.5 % and 67.9 ± 3.3 % of the bacterial 16S rRNA genes and fungal ITS sequences, respectively. At T0, the bacterial community was dominated by OTU11 affiliated to the Thiobacillus genus (Beta-Proteobacteria) and the fungal community was dominated by two OTUs affiliated to the Talaromyces and Scedosporium genera (Ascomycota) and by a third OTU affiliated to unclassified fungi.
Clustering analysis generating double dendrograms (vertically OTUs dendrogram and horizontally sample conditions dendrogram) and heat map representation reflecting data values of the most abundant (>2 % in at least one sample) bacterial and fungal OTUs in the six conditions (affiliation at 97 % similarity). The relative abundance of 24 bacterial and 37 fungal OTUs were normalized (z score) to compare studied conditions (three replicates per condition)
Modification of Bacterial and Fungal Diversity Over Time
The relative abundance of some bacterial phyla significantly (p < 0.05) increased (Bacteroidetes, Firmicutes, Nitrospirae, OD1, Gemmatimonadetes, Verrucomicrobia) or decreased (Gamma-Proteobacteria, Beta-Proteobacteria) over time, while others were specifically increased at T4 (Actinobacteria, Acidobacteria, Alpha-Proteobacteria) (Fig. 2). The relative abundance of Glomeromycota phylum increased significantly (p = 0.007) at T12 (Fig. 2). Moreover, the relative abundance of two orders belonging to the Ascomycota (Hypocreales and Pleosporales) increased significantly (p = 0.062 and p = 0.009) over time. On the contrary, the relative abundance of Eurotiales decreased significantly (p < 0.001) over time.
Clustering of samples according to the dominant OTUs (horizontal dendrogram, Fig. 3) showed higher similarity between samples at T0 than between samples at the two others times. This clustering of T0 samples is mainly due to 19 OTUs, mostly belonging to bacteria (Fig. 3). The relative abundance of 12 bacterial and 10 fungal OTUs increased significantly (p < 0.05) over time (Fig. 3 and Table S1). The 12 bacterial OTUs were affiliated to one Actinobacteria (Arthrobacter), one Bacteroidetes (Ohtaekwangia), one unclassified Firmicute, one Gemmatimonadetes (Gemmatimonas), one Nitrospirae (Nitrospira), two Alpha-Proteobacteria (unclassified Erythrobacteraceae and unclassified Bradyrhizobiaceae), three Beta-Proteobacteria (Rhodocyclaceae, Methyloversatilis and unclassified Burkholderiales), and two Gamma-Proteobacteria (unclassified and Thiohalophilus). The ten fungal OTUs were affiliated to five different genera: three were affiliated to the Ascomycota phylum, the Pezizomycotina subphylum, and the Sordariomycetes class (Acremonium, Bionectria, Fusarium), one belonged to unclassified Basidiomycota, and one was affiliated to unclassified fungi. The relative abundance of two bacterial OTUs affiliated to the Acidobacteria Gp7 and Ohtaekwangia genera (Bacteroidetes) and of seven fungal OTUs affiliated to Phoma, unclassified Plectosphaerellaceae and unclassified fungi, increased significantly at T4 (Fig. 3 and Table S1). The relative abundance of nine bacterial and three fungal OTUs decreased significantly (p < 0.05) over time (Table S1). Seven of these bacterial OTUs were affiliated to Gamma-Proteobacteria and belonged to the orders Acidithiobacillales (Thermithiobacillus), Chromatiales (Thiohalophilus, Halothiobacillus), Xanthomonadales (unclassified Xanthomonadaceae, Dyella, Rhodanobacter), Methylohalomonas or unclassified Gamma-Proteobacteria, one was affiliated to Chloroflexi (Sphaerobacter) and one to Beta-Proteobacteria (Thiobacillus). The three fungal OTUs belonged to the Ascomycota phylum, Pezizomycotina subphylum and Eurotiomycetes (Talaromyces) or Sordariomycetes (Pseudallescheria) class, and one was affiliated to unclassified fungi (Fig. 3).
Modification of Bacterial and Fungal Diversity with the Presence of Plant
Clustering of samples according to the dominant OTUs (horizontal dendrogram, Fig. 3) showed high similarities in planted soils and between bare soils at T4 and T12. Clustering of the dominant OTUs (vertical dendrogram, Fig. 3) showed that plant development increased the relative abundance of 15 and 11 OTUs, at T4 and T12 respectively, mainly belonging to fungi. The relative abundance of other bacterial OTUs was increased in bare soil (Fig. 3 and Table S1). The relative abundance of some bacterial phyla increased significantly (p < 0.05) (Actinobacteria) or decreased (Bacteroidetes, Nitrospira, Planctomycetes, Gamma-Proteobacteria, OD1, Gemmatimonadetes) with the presence of plants. Concerning fungal phyla, the relative abundance of Zygomycota and Glomeromycota increased significantly in alfalfa-planted plots (p < 0.05). Moreover, the fungal order Microascales was significantly (p < 0.001) less represented in planted soils (Fig. 3).
The relative abundance of five bacterial and ten fungal OTUs increased significantly (p < 0.05) with the presence of plants (Fig. 3 and Table S1). These bacterial OTUs were affiliated to one Actinobacteria (Arthrobacter), one unclassified Firmicute, one Alpha-Proteobacteria (unclassified Bradyrhizobiaceae) and two Gamma-Proteobacteria belonging to the Xanthomonadaceae family (unclassified and Dyella). Among the fungal OTUs, four belonged to the Sordariomycetes class among the Pezizomycotina subphylum (Acremonium, Bionectria, and Fusarium and unclassified Plectosphaerellaceae), and the other six were unclassified. The relative abundance of 6 bacterial and 5 fungal OTUs decreased significantly (p < 0.05) with the presence of plants (Table S1). These bacterial OTUs were affiliated to one Bacteroidetes (Ohtaekwangia), one Gemmatimonadetes (Gemmatimonas), one Nitrospiraea (Nitrospira), one Alpha-Proteobacteria (unclassified Erythrobacteraceae), and two Beta-Proteobacteria (unclassified Rhodocyclaceae and Burkholderiales). Three fungal OTUs belonged to the Sordariomycetes (Phoma, Pseudallescheria, Scedosporium) and two to unclassified fungi (Fig. 3).
Multivariate Analysis
A canonical correspondence analysis (CCA) was used to evaluate which variables best explained the microbial community compositions (Fig. 4). CCA ordination of bacterial and fungal taxons (relative abundance at the order level) and environmental variables (plant biomass, microbial abundance, and edaphic characteristics of the samples) depict relationships between the different plots and sampling dates. The first two CCA axes explained 69.63 % of the total variance in the microbial diversity data (Fig. 4). The six T0 samples grouped together and were separated from the other samples on the first axis (F1) discriminating the samples according to the effect of time. These T0 samples were discriminated according to higher bacterial abundance (16S), higher total calcium carbonate concentration, and higher extractable zinc concentration. At T4, the three samples from the planted plots MsmT4 were clearly separated from the BST4 samples on the second axis (F2) discriminating the samples according to the presence of plants, higher total N content and fungal abundance (18S). At T12, although the samples were still discriminated, the difference between the Msm and BS plots was thinner. T12 samples were discriminated according to higher total metal contents, extractable Fe and Cu, higher pH, C/N ratio and available phosphorous.
Canonical correspondence analysis (CCA) ordination of bacterial (in green) and fungal (in blue) taxons (relative abundance of the dominant orders represented by more than 10 sequences), and environmental variables (abundance of fungal 18S and bacterial 16S rRNA gene copy numbers, biomass of dry alfalfa and edaphic characteristics (in red), showing the relationships between the different plots and sampling dates (three replicates of the six conditions, in black bold type)). Numbers in italic correspond to the quantity of OTUs that belong to the taxon. Fungal and bacterial orders: Acidi.: Acidimicrobiales, Acidithio.: Acidithiobacillales, Actinomyc.: Actinomycetales, Altero.: Alteromonadales, Bacill.: Bacillales, Bdello.: Bdellovibrionales, Burkhol.: Burkholderiales, Caldi.: Caldilineale, Capno : Capnodiales, Caulo.: Caulobacterales, Chaetos.: Chaetosphaeriales, Chlamy.: Chlamydiales, Chroma.: Chromatiales, Clostri.: Clostridiales, Dehalo.: Dehalococcoidales, Desulfur.: Desulfuromonadales, Elusi.: Elusimicrobiales, Euro.: Eurotiales, Euzeb.: Euzebyales, Flavo.: Flavobacteriales, Gemma.: Gemmatimonadales, Helot.: Helotiales, Hydrogeno.: Hydrogenophilales, Hypo.: Hypocreales, Lactob.: Lactobacillales, Legion.: Legionellales, Methylo.: Methylococcales, Microasc.: Microascales, Mortier.: Mortierellales, Myxoc.: Myxococcales, Nitro.: Nitrospirales, Nitroso.: Nitrosomonadales, Oceano.: Oceanospirillales, Onyg : Onygenales, Opitu.: Opitutales, Parvu.: Parvularculales, Pseudo.: Pseudomonadales, Punicei.: Puniceicoccales, Rhizo.: Rhizobiales, Rhodo.: Rhodobacterales, Rhodocyc.: Rhodocyclales, Rhodospiri.: Rhodospirillales, Rickett.: Rickettsiales, Soliru.: Solirubrobacterales, Sphaero.: Sphaerobacterales, Sphingo.: Sphingomonadales, Sphingobacter.: Sphingobacteriales, Sporidio.: Sporidiobolales, Syntro.: Syntrophobacterales, Theleb.: Thelebolales, Thermo.: Thermoanaerobacterales, Thermoleo.: Thermoleophilales, Thiotric.: Thiotrichales, Treme.: Tremellales, Un. Acido_Gp1: Unclassified Acidobacteria_GP1, Un. Acido_Gp10: Unclassified Acidobacteria_GP10, Un. Acido_Gp16: Unclassified Acidobacteria_GP16, Un. Acido_Gp17: Unclassified Acidobacteria_GP17, Un. Acido_Gp3: Unclassified Acidobacteria_GP3, Un. Acido_Gp6: Unclassified Acidobacteria GP6, Un. Acido_Gp7: Unclassified Acidobacteria_GP7, Un. Actino.: Unclassified Actinobacteria, Un. Bacteria.: Unclassified Bacteria, Un. Bacteroid.: Unclassified Bacteroidetes, Un. Basidio.: Unclassified Basidiomycota, Un. Chloro.: Unclassified Chloroflexi, Un. Firmi. Unclassified Firmicutes, Un. Fungi: Unclassified Fungi, Un. Glomero.: Unclassified Glomeromycota, Un. Hypocreo.: Unclassified Hypocreomycetidae, Un. OD1: Unclassified OD1, Un. Plancto.: Unclassified Planctomycetes, Un. Strepto.: Unclassified Streptophyta, Un. Thermo.: Unclassified Thermomicrobia, Un. α-proteo.: Unclassified Alphaproteobacteria, Un. γ-proteo.: Unclassified Gammaproteobacteria, Un. δ-proteo.: Unclassified Deltaproteobacteria, Xantho.: Xanthomonadales. Edaphic characteristics: extractable copper, iron, and zinc (Cu ext, Fe ext, and Zn ext, respectively), 16 PAHs (mg/kg of dry weight soil)
At T0, four bacterial orders belonging to Gamma-proteobacteria (Chromatiales), Beta-proteobacteria (Acidithiobacillales and Hydrogenophilales) and Elusimicrobia phyla (Elusimicrobiales) and one fungal order (Sporidiobolales) belonging to Basidiomycota were the most correlated with high extractable zinc concentration, high CaCO3 and high bacterial abundance. Over 6 years, part of the microbial community was modified and more affected by changes in edaphic parameters. At T12, five bacterial orders belonging to five different phyla: Nitrospirae (Nitrospirales), Verrucomicrobia (Puniceicoccales), Actinobacteria (Euzebyales), Acidobacteria (Unclassified Acidobacteria_GP16 ), and Chloroflexi (Unclassified Thermomicrobia), and two fungal orders: Unclassified Glomeromycota, and Unclassified Basidiomycota, were highly correlated with higher C/N ratio, higher pH, higher P2O5, higher extractable iron and copper content and higher zinc and chrome concentrations. Moreover, seven bacterial orders belonging to five different phyla: Actinobacteria (Solirubrobacterales Thermoleophilales), Chloroflexi (Unclassified Chloroflexi), Acidobacteria (Unclassified Acidobacteria_GP10), Planctomycetes (Unclassified), and Firmicutes (Unclassified), were highly correlated with higher copper and lead concentrations. At T4, most fungal orders were positively correlated to alfalfa biomass and to the higher fungal abundance. Among these seven orders, six belong to the Ascomycota phylum and Pezizomycotina subphylum and belong to four different classes: Dothideomycetes (Capnodiales), Eurotiomycetes (Onygenales), Leotiomycetes (Helotiales, Thelebolales), and Sordariomycetes (Chaetosphaeriales, and Unclassified Hypocreomycetidae), and one to the Basidiomycota phylum (Tremellales).
Discussion
Microbial Diversity of the Aged Multi-Contaminated NM Soil
The NM soil had a high carbon concentration, neutral pH, low nutrient concentrations and high concentrations of PAHs and heavy metals that explain its low fertility and high toxicity [23]. This soil represented a unique ecosystem: contamination dated back to ca. 100 years, so selection pressure on soil microorganisms had lasted for decades. While the short-term impact of contamination tends to decrease microbial abundance, richness and diversity, we hypothesized that in such aged multi-contaminated soil the microbial community would adapt over time toward a unique diversity as compared to unpolluted soil.
In NM soil, bacteria were approximately ten-fold more abundant than fungi regardless of time or of the presence of vegetation. The same trend was previously observed in the same soil [40], and in many uncontaminated soils but with a higher difference between bacterial and fungal abundance levels. In fact, the environments harboring such high fungal abundance and a high fungal to bacterial ratio are usually rich in carbon, like the litter horizon in forest soils [41]. The Chao1 index evidenced higher bacterial than fungal species richness in our NM soil samples, as previously shown in other soils [42]. Freshly contaminated soils usually harbor low bacterial richness, evenness, and diversity after spiking [6]. Yet bacterial richness and diversity in our NM soil was relatively high and comparable to the levels found in a diesel-contaminated soil in Poland [43] and in an aged PAH-contaminated soil from a coking plant wasteland in Beijing, China [44], using a similar sequencing effort. The result from our NM soil suggests a long-term adaptation of the microbial community toward a diversified and pollutant-resistant community. In addition, fungal richness was close to the richness found in forest soils [36], but fungal diversity was relatively low and close to the level found in a soil highly contaminated by hydrocarbons [45].
The dominance of three bacterial phyla (Proteobacteria, Actinobacteria, and Bacteroidetes) and Ascomycota and unclassified fungi with poor representation of Basidiomycota is in accordance with fingerprinting analyses of dominant bacterial and fungal species inhabiting the NM soil [21, 46]. Overall, at the phylum level, the bacterial and fungal community diversities were close to other soil types. However, the relative proportion of each bacterial phylum seems to be specific to polluted environments, contrary to fungal ones. Proteobacteria are usually the dominant phylum in contaminated soils [47]. The co-occurrence of Alpha-, Beta-, and Gamma-Proteobacteria in similar relative proportions (15–21 %) seems typical of polluted soils: Alpha- are usually in higher proportions in non-contaminated soils, while Beta- and Gamma- subdivisions are favored in hydrocarbon-contaminated soils [48]. Furthermore, Actinobacteria was the second most abundant phylum in our NM soil as found in other heavy metal-contaminated soils [47, 49]. Actinobacteria were previously described in extremely arid environments [50], with similar properties (low nutrient and low carbon availability) to aged multi-contaminated soil. Finally, some genera belonging to the Actinobacteria are able to degrade PAH compounds [51]. Members of Acidobacteria were found scarce in our NM soil as found in other PAH-contaminated soil [44], but they were largely less represented than in non-polluted soils [52, 53]. Overall, similar fungal phylum diversity was also found in non-polluted soils [54].
Impact of Edaphic Parameters on Soil Microbial Diversity Over Time
Although this result could be specific to this particular site under specific climatic condition, the NM soil monitoring over 6 years revealed interesting relationship between bacterial and fungal diversity and edaphic parameters. Our canonical correspondence analysis showed that a temporal shift occurred in the composition of the soil microbial community, with some taxa correlated with modifications in edaphic parameters.
Among the different soil parameters that increased simultaneously over time (C/N ratio, available phosphorous, extractable iron, copper, chrome, zinc concentrations), the pH can be an important driver of the bacterial community but not of the fungal one [55]. Most orders belonging to Acidobacteria were favored over time, with subgroup-16 positively correlated with pH values. Unlike the other subgroups, the relative abundance of Acidobacteria subgroup 1 was greater at T0 and was negatively correlated with the pH level, as previously found [55].
Despite a low availability and mobility of PAH and HM, and significant decrease of these fractions over time [23], we observed a significant decrease in total PAH (c.a. 40 % dissipation) and extractable Zn concentrations over the 6 years of monitoring that could also contribute to explain changes in microbial diversity. Xu et al. [44] suggested that a decrease in pollutant concentration allowed for the development of members of the Acidobacteria phylum in a PAH-contaminated soil. The PAH decrease could also explain the emergence of PAH-degrading microorganisms that benefited from the biodegradation of the pollutant. As previously observed [21, 46], OTUs affiliated to the Arthrobacter and Fusarium genera increased over time in NM soil samples. Members of the Arthrobacter and Fusarium genera have been found in hydrocarbon-contaminated environments [56, 57] and could be potential PAH degraders [51, 58]. The relative abundance of the Nitrospirae phylum, one Nitrospira OTU and the Nitrospirales order increased over time. On CCA it was positively correlated to the C/N ratio, and negatively correlated to total nitrogen content. This bacterial group is more competitive with other nitrite-oxidizing bacteria (i.e. Nitrobacter) in soils with low N availability [59]. Relative abundance of the Gemmatimonadetes phylum and one OTU affiliated to the Gemmatimonas genus also increased over time. Although members of this phylum are frequently detected in soils and more prevalent in dry soils with poor nutrient availability and low microbial activity [60], their ecological properties are still unclear.
On the contrary, the relative abundance of other microbial taxa decreased over time. The fungal Eurotiales order and one OTU affiliated to Talaromyces (Ascomycota; Eurotiales) were more abundant at T0. This fungal taxon seems to be well adapted to high PAH contamination levels, as similar strains were isolated from an oil-contaminated soil and were able to grow at high PAH concentrations [61]. Within the bacterial community, the relative abundance of the Gamma-Proteobacteria phylum and of seven OTUs affiliated to Gamma-Proteobacteria decreased over time. Among these, Thiohalophilus, Halothiobacillus, Methylohalomonas, known as halophilic bacteria, could be repressed by the leaching of extractable metals and sulfate salts over time. The relative abundance of Thiobacillus (Beta-proteobacteria; Hydrogenophilales) was the most abundant bacterial OTU at T0 (ca. 15 %), but also decreased over time. Thiobacillus was previously detected in heavy metal-contaminated soils [47]. The implication of the Thiobacillus genus in PAH degradation has also been described [62] although our results indicate a drastic decrease over time.
Impact of Vegetation on Soil Microbial Abundance and Diversity
Our results show that the plant cover influenced microbial abundance in the NM soil, especially at T4 when the plant cover was the thickest. Although the role of root exudates in regulating soil fungal community composition and diversity is well known [63], the impact of the rhizosphere on increasing soil fungal abundance, as found here, is less documented as compared to bacteria [63].
The plant cover had an unexpected impact on fungal and bacterial community structure in the NM soil. At T4, PAH dissipation was significantly lower in presence of plants than in bare soil, probably explained by the release of more easily degradable carbon compounds in root exudates [64]. At T4, the community diversity is contrasted between bare and alfalfa-planted soils, and the multivariate analysis showed that plant biomass is the dominant explicative factor compared to PAH content. Bacterial and fungal diversity indices increased with the presence of vegetation, as previously shown [21, 46]. Although the impact of plants varies depending on seasons, usually fungal and bacterial diversity usually decrease in the rhizospheric or planted soils as compared to the bulk or unplanted controls [65, 66]. Lower microbial diversity in the rhizosphere could be due to selective pressure from root exudates. In aged hydrocarbon- and heavy metal-contaminated soils that are extremely oligotrophic environments, microbial activity is limited by nutrient availability [67]. Most studies describe the impact of plants on microbial diversity in non-contaminated and/or nutrient-rich soils (e.g., agricultural or forest soils). The presence of plants on a nutrient-poor soil could increase diversity due to the input of diversified carbon and nutrient sources, such as carbohydrates from exudates and plant debris preferentially degraded by bacteria and fungi, respectively. Moreover, the development of plants and roots creates numerous novel niches favoring the establishment of more diverse microbial communities. As previously shown, the functional community was also impacted by plant covers because enzymatic activity measured (FDase, arylamidase, aryl sulfatase, cellulose, urease…) were higher in vegetalized NM soil than in bare soil [40].
Our analyses allowed us to discriminate between planted soil and bare soil plots and to identify microbial taxa whose relative abundance levels were differently impacted by the presence of plants. Actinomycetales and Arthrobacter, the most represented order and most abundant OTU of the bacterial community, were positively correlated to alfalfa biomass. Increased numbers of Actinobacteria in the rhizosphere of different plant species such as maize [66] and strawberry [68] are documented. In addition, the Bradyrhizobiaceae family and the Rhizobiales order were among the dominant bacterial OTUs favored by the development of alfalfa. These nitrogen-fixing bacteria are known to be abundant in the rhizosphere of legumes due to their symbiotic association [69]. However, we never observed nodules throughout the monitoring period. Finally, the relative abundance of the Nitrospirae phylum and Nitrospira OTUs, a well-known nitrite-oxidizing group [70], decreased in alfalfa-planted plots. Competition for nitrogen between plants and nitrifying bacteria has already been reported in unfertilized nutrient-poor soil [71].
Plants have a higher impact on fungal than on bacterial community structure. According to our multivariate analysis, most fungal orders were positively correlated to the presence of plants (alfalfa biomass). The relative abundance of Ascomycota belonging to the Pezizomycotina sub-division and affiliated to various classes, i.e. Sordariomycetes, Leotiomycetes, Dothideomycetes, and Eurotiomycetes, increased. Among the Sordariomycetes, the Chaetosphaeriales, and the unclassified Hypocreales were the most important orders allowing for discrimination between treatments. Among these orders, the relative abundance of some dominant fungal OTUs affiliated to the Acremonium, Bionectria, and Fusarium genera significantly increased with the presence of alfalfa. While the Acremonium genus is well known as a fungal endophyte of plants that improves mineral nutrition [72], Bionectria and Fusarium species are often plant pathogens [73, 74]. Conversely, in bare soil, the Microascales order was the only one whose relative abundance increased. Although alfalfa was co-inoculated with two mycorrhizal Glomus strains, Glomeromycota was not a dominant phylum, but its relative abundance was higher in planted plots. Our results show that unclassified Glomeromycota were more correlated with planted soil after 6 years than after 2 years. Colonization of roots by mycorrhizal fungi was detected only after 6 years, suggesting potential initial toxicity of the contaminated soil toward mycorrhizal fungi, as suggested previously [23].
Conclusion
To conclude, the high fungal and bacterial abundance levels in the NM soil probably reflect its high carbon content due to contamination. Microbial diversity and richness in the aged-contaminated soil were surprisingly high and closer to that found in an unstressed soil than in freshly contaminated soil. A specific microbial community settled over time through modifications of edaphic parameters, pollution content and due to the presence of plants. Over time the C/N increased, due to a decrease of soil nitrogen content (no fertilization), favoring Nitrospirae phylum, and nitrogen-fixing bacteria in planted soil. Over time the pH increased, probably due to sulfate leaching that favored some Acidobacteria groups and repressed halophilic bacteria. In 6 years, the decrease of PAH content by c.a. 40 % favored the appearance of few taxa previously described as PAH degraders. The modification of pollution content and bioavailability contributed probably to reduce environment toxicity and led to adaptation in microbial community composition. The alfalfa cover structured the microbial community, probably due to a rhizosphere effect while this partly inhibited the PAH dissipation. Indeed vegetation (i) selected a specific community by releasing root exudates or plant debris that input nutrient sources more easily degraded than PAHs in the oligotrophic soil; (ii) increased bacterial diversity, and (iii) favored most fungal species compared to bare soil.
References
Sandrin TR, Maier RM (2003) Impact of metals on the biodegradation of organic pollutants. Environ Health Perspect 111:1093
Cerniglia CE, Heitkamp MA (1989) Microbial degradation of polycyclic aromatic hydrocarbons (PAH) in the aquatic environment. Metab Polycycl Aromat Hydrocarb Aquat Environ 41–68
Peng R-H, Xiong A-S, Xue Y et al (2008) Microbial biodegradation of polyaromatic hydrocarbons. FEMS Microbiol Rev 32:927–955
Schimel J, Balser TC, Wallenstein M (2007) Microbial stress-response physiology and its implications for ecosystem function. Ecology 88:1386–1394. doi:10.2307/27651246
Paissé S, Coulon F, Goñi-Urriza M et al (2008) Structure of bacterial communities along a hydrocarbon contamination gradient in a coastal sediment. FEMS Microbiol Ecol 66:295–305. doi:10.1111/j.1574-6941.2008.00589.x
Ding GC, Heuer H, Smalla K (2012) Dynamics of bacterial communities in two unpolluted soils after spiking with phenanthrene: soil type specific and common responders. Front Microbiol 3:290
Zhou H, Wong AY, Yu RK et al (2009) Polycyclic aromatic hydrocarbon-induced structural shift of bacterial communities in mangrove sediment. Microb Ecol 58:153–160. doi:10.1007/s00248-008-9456-x
Gremion F, Chatzinotas A, Kaufmann K, Von Sigler W, Harms H (2004) Impacts of heavy metal contamination and phytoremediation on a microbial community during a twelve-month microcosm experiment. FEMS Microbiol Ecol 48:273–283, Wiley
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364. doi:10.1890/05-1839
Wenzel W (2009) Rhizosphere processes and management in plant-assisted bioremediation (phytoremediation) of soils. Plant Soil 321:385–408. doi:10.1007/s11104-008-9686-1
Binet P, Portal JM, Leyval C (2000) Fate of polycyclic aromatic hydrocarbons (PAH) in the rhizosphere and mycorrhizosphere of ryegrass. Plant Soil 227:207–213. doi:10.1023/A:1026587418611
Corgié SC, Joner EJ, Leyval C (2003) Rhizospheric degradation of phenanthrene is a function of proximity to roots. Plant Soil 257:143–150. doi:10.1023/A:1026278424871
Haritash AK, Kaushik CP (2009) Biodegradation aspects of polycyclic aromatic hydrocarbons (PAHs): a review. J Hazard Mater 169:1–15. doi:10.1016/j.jhazmat.2009.03.137
Sun M, Fu D, Teng Y et al (2011) In situ phytoremediation of PAH-contaminated soil by intercropping alfalfa (Medicago sativa L.) with tall fescue (Festuca arundinacea Schreb.) and associated soil microbial activity. J Soils Sediments 11:980–989. doi:10.1007/s11368-011-0382-z
Muratova AY, Turkovskaya OV, Hübner T, Kuschk P (2003) Studies of the efficacy of alfalfa and reed in the phytoremediation of hydrocarbon-polluted soil. Appl Biochem Microbiol 39:599–605. doi:10.1023/A:1026238720268
Kawasaki A, Watson E, Kertesz M (2012) Indirect effects of polycyclic aromatic hydrocarbon contamination on microbial communities in legume and grass rhizospheres. Plant Soil 358:169–182. doi:10.1007/s11104-011-1089-z
Joner EJ, Leyval C (2001) Influence of arbuscular mycorrhiza on clover and ryegrass grown together in a soil spiked with polycyclic aromatic hydrocarbons. Mycorrhiza 10:155–159. doi:10.1007/s005720000071
Leyval C, Turnau K, Haselwandter K (1997) Effect of heavy metal pollution on mycorrhizal colonization and function: physiological, ecological and applied aspects. Mycorrhiza 7:139–153. doi:10.1007/s005720050174
Cébron A, Beguiristain T, Bongoua-Devisme J, et al. (2015) Impact of clay mineral, wood sawdust or root organic matter on the bacterial and fungal community structures in two aged PAH-contaminated soils. Environ Sci Pollut Res 1–15
Tejeda-Agredano MC, Gallego S, Vila J et al (2013) Influence of the sunflower rhizosphere on the biodegradation of PAHs in soil. Soil Biol Biochem 57:830–840. doi:10.1016/j.soilbio.2012.08.008
Thion C, Cébron A, Beguiristain T, Leyval C (2012) Long-term in situ dynamics of the fungal communities in a multi-contaminated soil are mainly driven by plants. FEMS Microbiol Ecol 82:169–181. doi:10.1111/j.1574-6941.2012.01414.x
Buée M, De Boer W, Martin F et al (2009) The rhizosphere zoo: an overview of plant-associated communities of microorganisms, including phages, bacteria, archaea, and fungi, and of some of their structuring factors. Plant Soil 321:189–212. doi:10.1007/s11104-009-9991-3
Ouvrard S, Barnier C, Bauda P et al (2011) In situ assessment of phytotechnologies for multicontaminated soil management. Int J Phytoremediation 13:245–263. doi:10.1080/15226514.2011.568546
Schüßler A, Walker C (2010) The Glomeromycota: a species list with new families and new genera.
Koske RE, Gemma JN (1989) A modified procedure for staining roots to detect VA mycorrhizas. Mycol Res 92:486–488. doi:10.1016/S0953-7562(89)80195-9
Trouvelot A, Kough JL, Gianinazzi-Pearson V (1986) Mesure du taux de mycorhization VA d’un système radiculaire. Recherche de méthodes d’estimation ayant une signification fonctionnelle. 217–221
Cébron A, Norini M-P, Beguiristain T, Leyval C (2008) Real-time PCR quantification of PAH-ring hydroxylating dioxygenase (PAH-RHDα) genes from Gram positive and Gram negative bacteria in soil and sediment samples. J Microbiol Methods 73:148–159. doi:10.1016/j.mimet.2008.01.009
Lueders T, Wagner B, Claus P, Friedrich MW (2004) Stable isotope probing of rRNA and DNA reveals a dynamic methylotroph community and trophic interactions with fungi and protozoa in oxic rice field soil. Environ Microbiol 6:60–72. doi:10.1046/j.1462-2920.2003.00535.x
Felske A, Akkermans ADL, De Vos WM (1998) Quantification of 16S rRNAs in complex bacterial communities by multiple competitive reverse transcription-PCR in temperature gradient gel electrophoresis fingerprints. Appl Environ Microbiol 64:4581–4587
Turner S, Pryer KM, Miao VP, Palmer JD (1999) Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis1. J Eukaryot Microbiol 46:327–338
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes‐application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, White JJSJ (eds) PCR Protoc. Academic, San Diego, pp 315–322
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Schloss PD, Westcott SL, Ryabin T et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Sogin ML, Morrison HG, Huber JA et al (2006) Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc Natl Acad Sci 103:12115–12120
Coince A, Caël O, Bach C et al (2013) Below-ground fine-scale distribution and soil versus fine root detection of fungal and soil oomycete communities in a French beech forest. Fungal Ecol 6:223–235. doi:10.1016/j.funeco.2013.01.002
Nilsson RH, Veldre V, Hartmann M et al (2010) An open source software package for automated extraction of ITS1 and ITS2 from fungal ITS sequences for use in high-throughput community assays and molecular ecology. Fungal Ecol 3:284–287. doi:10.1016/j.funeco.2010.05.002
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. doi:10.1016/S0022-2836(05)80360-2
Colwell R (2013) EstimateS: Statistical estimation of species richness and shared species from samples. 2006. Consult. En Httpviceroy Eeb Uconn Eduestimates
Cébron A, Cortet J, Criquet S et al (2011) Biological functioning of PAH-polluted and thermal desorption-treated soils assessed by fauna and microbial bioindicators. Spec Issue Environ Microbiol 162:896–907. doi:10.1016/j.resmic.2011.02.011
Baldrian P, Kolarik M, Stursova M et al (2012) Active and total microbial communities in forest soil are largely different and highly stratified during decomposition. ISME J 6:248–258
Gottel NR, Castro HF, Kerley M et al (2011) Distinct microbial communities within the endosphere and rhizosphere of Populus deltoides roots across contrasting soil types. Appl Environ Microbiol 77:5934–5944. doi:10.1128/AEM.05255-11
Sutton NB, Maphosa F, Morillo JA et al (2013) Impact of long-term diesel contamination on soil microbial community structure. Appl Environ Microbiol 79:619–630. doi:10.1128/AEM.02747-12
Xu Y, Sun G-D, Jin J-H et al (2014) Successful bioremediation of an aged and heavily contaminated soil using a microbial/plant combination strategy. J Hazard Mater 264:430–438. doi:10.1016/j.jhazmat.2013.10.071
Bell TH, El-Din Hassan S, Lauron-Moreau A et al (2014) Linkage between bacterial and fungal rhizosphere communities in hydrocarbon-contaminated soils is related to plant phylogeny. ISME J 8:331–343
Cébron A, Beguiristain T, Faure P et al (2009) Influence of vegetation on the in situ bacterial community and polycyclic aromatic hydrocarbon (PAH) degraders in aged PAH-contaminated or thermal-desorption-treated soil. Appl Environ Microbiol 75:6322–6330
Gołębiewski M, Deja-Sikora E, Cichosz M et al (2014) 16S rDNA pyrosequencing analysis of bacterial community in heavy metals polluted soils. Microb Ecol 67:635–647. doi:10.1007/s00248-013-0344-7
Labbé D, Margesin R, Schinner F, Whyte LG, Greer CW (2007) Comparative phylogenetic analysis of microbial communities in pristine and hydrocarbon-contaminated Alpine soils. FEMS Microbiol Ecol 59:466–475, Wiley
Gremion F, Chatzinotas A, Harms H (2003) Comparative 16S rDNA and 16S rRNA sequence analysis indicates that Actinobacteria might be a dominant part of the metabolically active bacteria in heavy metal-contaminated bulk and rhizosphere soil. Environ Microbiol 5:896–907. doi:10.1046/j.1462-2920.2003.00484.x
Köberl M, Müller H, Ramadan EM, Berg G (2011) Desert farming benefits from microbial potential in arid soils and promotes diversity and plant health. PLoS ONE 6:e24452. doi:10.1371/journal.pone.0024452
Kallimanis A, Frillingos S, Drainas C, Koukkou A (2007) Taxonomic identification, phenanthrene uptake activity, and membrane lipid alterations of the PAH degrading Arthrobacter sp. strain Sphe3. Appl Microbiol Biotechnol 76:709–717. doi:10.1007/s00253-007-1036-3
Roesch LFW, Fulthorpe RR, Riva A et al (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1:283–290
Nacke H, Thürmer A, Wollherr A et al (2011) Pyrosequencing-based assessment of bacterial community structure along different management types in German forest and grassland soils. PLoS ONE 6:e17000. doi:10.1371/journal.pone.0017000
Buée M, Reich M, Murat C et al (2009) 454 Pyrosequencing analyses of forest soils reveal an unexpectedly high fungal diversity. New Phytol 184:449–456. doi:10.1111/j.1469-8137.2009.03003.x
Rousk J, Baath E, Brookes PC et al (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4:1340–1351
Panov AV, Esikova TZ, Sokolov SL et al (2013) Influence of soil pollution on the composition of a microbial community. Microbiology 82:241–248. doi:10.1134/S0026261713010116
Lladó S, Gràcia E, Solanas AM, Viñas M (2013) Fungal and bacterial microbial community assessment during bioremediation assays in an aged creosote-polluted soil. Soil Biol Biochem 67:114–123. doi:10.1016/j.soilbio.2013.08.010
Hong JW, Park JY, Gadd GM (2010) Pyrene degradation and copper and zinc uptake by Fusarium solani and Hypocrea lixii isolated from petrol station soil. J Appl Microbiol 108:2030–2040. doi:10.1111/j.1365-2672.2009.04613.x
Attard E, Poly F, Commeaux C et al (2010) Shifts between Nitrospira‐ and Nitrobacter‐like nitrite oxidizers underlie the response of soil potential nitrite oxidation to changes in tillage practices. Environ Microbiol 12:315–326
DeBruyn JM, Nixon LT, Fawaz MN et al (2011) Global biogeography and quantitative seasonal dynamics of Gemmatimonadetes in soil. Appl Environ Microbiol 77:6295–6300
Reyes-César A, Absalón Á, Fernández F et al (2014) Biodegradation of a mixture of PAHs by non-ligninolytic fungal strains isolated from crude oil-contaminated soil. World J Microbiol Biotechnol 30:999–1009. doi:10.1007/s11274-013-1518-7
Singleton D, Jones M, Richardson S, Aitken M (2013) Pyrosequence analyses of bacterial communities during simulated in situ bioremediation of polycyclic aromatic hydrocarbon-contaminated soil. Appl Microbiol Biotechnol 97:8381–8391. doi:10.1007/s00253-012-4531-0
Berg G, Smalla K (2009) Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Microbiol Ecol 68:1–13. doi:10.1111/j.1574-6941.2009.00654.x
Tian L, Ma P, Zhong J-J (2003) Impact of the presence of salicylate or glucose on enzyme activity and phenanthrene degradation by Pseudomonas mendocina. Process Biochem 38:1125–1132
Shahsavari E, Adetutu E, Anderson P, Ball A (2013) Tolerance of selected plant species to petrogenic hydrocarbons and effect of plant rhizosphere on the microbial removal of hydrocarbons in contaminated soil. Water Air Soil Pollut 224:1–14. doi:10.1007/s11270-013-1495-3
Li X, Rui J, Mao Y et al (2014) Dynamics of the bacterial community structure in the rhizosphere of a maize cultivar. Soil Biol Biochem 68:392–401. doi:10.1016/j.soilbio.2013.10.017
Breedveld GD, Karlsen DA (2000) Estimating the availability of polycyclic aromatic hydrocarbons for bioremediation of creosote contaminated soils. Appl Microbiol Biotechnol 54:255–261. doi:10.1007/s002530000362
Smalla K, Wieland G, Buchner A et al (2001) Bulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: plant-dependent enrichment and seasonal shifts revealed. Appl Environ Microbiol 67:4742–4751
McCaig AE, Glover LA, Prosser JI (1999) Molecular analysis of bacterial community structure and diversity in unimproved and improved upland grass pastures. Appl Environ Microbiol 65:1721–1730
Koops H-P, Purkhold U, Pommerening-Röser A, Dworkin M, Falkow S, Rosenberg E et al (2006) The lithoautotrophic ammonia-oxidizing bacteria. In: The Prokaryotes. Springer, New York, pp 778–811
Stienstra AW, Klein Gunnewiek P, Laanbroek HJ (1994) Repression of nitrification in soils under a climax grassland vegetation. FEMS Microbiol Ecol 14:45–52. doi:10.1111/j.1574-6941.1994.tb00089.x
Rahman MH, Saiga S (2007) Endophyte effects on nutrient acquisition in tall fescue grown in andisols. J Plant Nutr 30:2141–2158. doi:10.1080/01904160701700632
Luo J, Zhuang W-Y (2010) New species and new Chinese records of Bionectriaceae (Hypocreales, Ascomycota). Mycol Prog 9:17–25
Pegg GF, Parry DW (1983) Infection of lucerne (Medicago sativa) by Fusarium species. Ann Appl Biol 103:45–55. doi:10.1111/j.1744-7348.1983.tb02739.x
Acknowledgments
This study was funded by OTELo (Observatoire Terre et Environnement de Lorraine) and the Agence Nationale de la Recherche (ANR-2008-CESA-010 and ANR-13-JSV7-0007_01). The authors thank the GISFI (www.gisfi.fr) for the access to the experimental site at Homécourt and Annie Buchwalter for English editing.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Rarefaction curve representing the number of operational taxonomic units (OTUs) depending on the number of 16S rRNA genes (A) and ITS (B) sequences amplified from the six conditions. After OTU formation to 97 % similarity, subsampling was performed to 18,100 and 9,100 sequences per sample for bacterial and fungal data, respectively. (GIF 322 kb)
Table S1
Affiliation at 97 % similarity of the most abundant (>2 % in at least one sample) bacterial and fungal OTUs in the six conditions (mean and standard deviation, n = 3). Two-way analysis of variance (ANOVA) followed by Neuman-Keuls multiple comparison test to study the effect of plant cover and time on the most abundant bacterial (16S) and fungal (ITS) OTUs. Sections colored in dark grey / grey / light grey and white mean respectively that OTUs were significantly (p < 0.05) favored at all time-points / favored at T4 / inhibited or not influenced by the plant cover or by time. (DOC 169 kb)
Rights and permissions
About this article
Cite this article
Bourceret, A., Cébron, A., Tisserant, E. et al. The Bacterial and Fungal Diversity of an Aged PAH- and Heavy Metal-Contaminated Soil is Affected by Plant Cover and Edaphic Parameters. Microb Ecol 71, 711–724 (2016). https://doi.org/10.1007/s00248-015-0682-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-015-0682-8