Abstract
Synergistetes strain MFA1 is an asaccharolytic ruminal bacterium isolated based on its ability to degrade fluoroacetate, a plant toxin. The amino acid and peptide requirements of the bacterium were investigated under different culturing conditions. The growth of strain MFA1 and its fluoroacetate degradation rate were enhanced by peptide-rich protein hydrolysates (tryptone and yeast extract) compared to casamino acid, an amino acid-rich protein hydrolysate. Complete utilization and preference for arginine, asparagine, glutamate, glycine, and histidine as free amino acids from yeast extract were observed, while the utilization of serine, threonine, and lysine in free form and peptide-bound glutamate was stimulated during growth on fluoroacetate. A predominant peptide in yeast extract preferentially utilized by strain MFA1 was partially characterized by high-liquid performance chromatography-mass spectrometry as a hepta-glutamate oligopeptide. Similar utilization profiles of amino acids were observed between the co-culture of strain MFA1 with Methanobrevibacter smithii without fluoroacetate and pure strain MFA1 culture with fluoroacetate. This suggests that growth of strain MFA1 could be enhanced by a reduction of hydrogen partial pressure as a result of hydrogen removal by a methanogen or reduction of fluoroacetate.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The recently classified phylum Synergistetes consists of amino acid-fermenting anaerobes from a wide variety of anaerobic habitats including wastewater sludge, human oral cavity, and animal gastrointestinal tracts [1–5]. Fermenting amino acids is thought to be the primary ecological role of Synergistetes in nature [6], with some species using peptides as their preferred energy source [3, 5, 7–11].
Similar to other fermentative bacteria, members of the Synergistetes produce hydrogen and organic compounds (acetate, propionate, and butyrate) from amino acid fermentation [12, 13]. However, the increase in hydrogen partial pressure as a result of amino acid fermentation can lead to inhibition of further fermentation due to thermodynamic constraints [14, 15]. In the presence of suitable electron acceptors, some Synergistetes are capable of switching from fermentative pathways to anaerobic respiration to prevent the metabolic constraints resulting from an increased hydrogen partial pressure. Species from the genera Dethiosulfovibrio and Thermanaerovibrio have the ability to use elemental sulfur and thiosulfate as hydrogen sinks [5, 16–18], whilst other species from the genus Anaerobaculum reduces crotonate [11], sulfite [9], or cystine [16].
The Synergistetes bacterium strain MFA1 is most closely related to members of the genera Synergistes and Cloacibacillus [19]. Strain MFA1 was isolated from the bovine rumen and appears to be a low-abundance ubiquitous member of the gastrointestinal tracts of various herbivores [19]. This bacterium is asaccharolytic and ferments amino acids for growth [19], which are characteristics shared across the Synergistetes phylum [6]. Moreover, strain MFA1 is capable of anaerobically degrading fluoroacetate, a toxic compound found in some Acacia plant species and heartleaf (Gastrolobium spp.) shrubs. These fluoroacetate-bearing plants are periodically associated with poisoning of ruminant livestock, particularly in southern hemisphere countries [20, 21]. Livestock that accidentally graze these plants usually die within 30 hours of ingestion of the shrubs [22]. Therefore, understanding the fluoroacetate metabolism of strain MFA1 and its growth has potential for the development of antidotal therapy for fluoroacetate poisoning in cattle.
Preliminary biochemical characterization of strain MFA1 was reported by Davis et al. [19], but further investigation of the factors that influence its survival and growth in gut ecosystems is required. The primary aim of this study was to investigate the nutritional requirement of strain MFA1 with an emphasis on amino acid and peptide utilization under various culturing conditions, which may facilitate the establishment and maintenance of strain MFA1 in the rumen. Different types of protein hydrolysates (casamino acids, phytone peptone, tryptone, and yeast extract) added at 0.4 % (w/v) were initially used to examine their effects on the growth of strain MFA1. Amino acid and peptide utilization by strain MFA1 from a yeast extract-rich medium were then examined by comparing utilization profiles in the presence of fluoroacetate. Additionally, the metabolic versatility of strain MFA1 was explored by investigating its amino acid utilization and fluoroacetate-degrading capacity in a methanogenic environment which is analogous to conditions in the rumen. Co-cultivation of MFA1 with the hydrogen-scavenging methanogen, Methanobrevibacter smithii, revealed a competition for hydrogen between the two in the presence of fluoroacetate. Our results provide insight into the nutritional ecology of strain MFA1 and its potential to degrade fluoroacetate under methanogenic conditions, as found in the rumen.
Methods
Strain and Culture Conditions
Synergistetes bacterium strain MFA1 was isolated from a bovine rumen as described by Davis and colleagues [19], and a pure culture was stored at −80 °C in 20 % (v/v) glycerol solution. Strain MFA1 was grown at 39 °C and maintained by daily subculture on an artificial rumen fluid basal medium described by Davis et al. [19], which contains mineral salts and 10 % (v/v) clarified rumen fluid. Casamino acids (0.8 %, w/v) and yeast extract (0.2 %, w/v) were supplemented as carbon/energy sources. The autotrophic methanogen, M. smithii ATCC35061, was purchased from American Type Culture Centre and grown in medium recommended by the ATCC (Medium 1340) under a H2/CO2 (80:20) atmosphere at 150-kPa pressure.
Growth on Different Sources of Amino Acids and Peptides
A minimal concentration of the protein hydrolysates at 0.4 % (w/v) was used to clearly elucidate the effect of each hydrolysate on strain MFA1’s growth and its fluoroacetate degradation capability. Filter-sterilized protein hydrolysates (casamino acids, acid hydrolysate of casein; phytone peptone, enzymatic digest of soy peptone; tryptone, enzymatic digest of casein; and yeast extract, water soluble portion of autolyzed Saccharomyces cerevisiae cells) (BD Biosciences, CA, USA) were added separately to a final concentration of 0.4 % (w/v) in the artificial rumen fluid PA basal medium with 0.002 % yeast extract. Similar concentration of the protein hydrolysates was also added separately into a medium containing 20 mM fluoroacetate and 0.002 % yeast extract. All experiments were performed in triplicate in anaerobic 27 ml Balch tubes with a headspace of 17 ml that consisted of 95 % CO2 and 5 % H2. Strain MFA1 cultures were inoculated and incubated at 39 °C for 120 h. Growth and fluoroacetate degradation by strain MFA1 were monitored by measuring optical density (OD) at 600 nm and fluoride production using a Thermo Scientific fluoride ion-selective electrode (Thermo Fisher Scientific, MA, USA), respectively, as described by Davis et al. [19].
Utilization of “Free” and “Peptide-Bound” Amino Acids
Amino acids and peptides utilized by strain MFA1 in a nutrient-rich yeast extract medium (0.8 %, w/v) were investigated to determine the growth factors for the bacterium. The utilization profiles of strain MFA1 at its exponential phase of growth were performed using the medium with either 0.8 % yeast extract (BYE medium) or 0.8 % yeast extract and 20 mM fluoroacetate (FYE medium). Strain MFA1 was inoculated and incubated for 120 h. Cultures were centrifuged at 13,000 g for 10 min, and supernatants were collected for amino acid and peptide analyses.
Free and total amino acid compositions of culture samples and uninoculated sterile BYE medium (blank) were analyzed by the Australian Proteome Analysis Facility (Macquarie University, Sydney, Australia) on a Waters ACQUITY™ Ultra Performance Liquid Chromatography (UPLC) system (BEH RP C18, 2.1 × 100 mm, 1.7-μm column) (Waters Corporation, MA, USA). During the amino acid analysis, two independent UPLC runs were performed for each sample. Peptide-bound amino acids were estimated as the difference between total and free amino acids. The analysis was performed by firstly adding internal standards α-aminobutyric acid and norvaline to each sample for free amino acid analysis. Samples were filtered through 10-kDa molecular weight cutoff (MWCO) centrifugal units (Merck Millipore, Victoria, Australia). The filtrates were then derivatized using a Waters AccQ-Tag Ultra derivatization kit [23] prior to amino acid analysis using the UPLC system. Amino acids were separated by a gradient elution with eluent A following 20-fold dilution and eluent B provided by the Waters AccQ-Tag Ultra Chemistry kit, with a flow rate of 0.7 ml min−1 at 57 °C and a 10.2-min analysis time per sample. The amino acid peak intensities were measured using UV absorption at 260 nm.
For total amino acid analysis, samples were treated for 24 h to vapor-phase hydrolysis using 6 M HCl at 110 °C [24], followed by pretreatment and analysis with the UPLC systems as described above. Due to the deamination of glutamine to glutamic acid and asparagine to aspartic acid under acidic conditions, the amounts of these acids are reported as the sum of those respective components.
HPLC Peptide Isolation and Mass Spectrometry Identification
In order to characterize the peptides utilized by strain MFA1 from yeast extract, culture medium was analyzed for peptide composition using high-performance liquid chromatography (HPLC) and fractions representing peptide peaks that had decreased were collected and sequenced using mass spectrometry by the following methods. Initially, strain MFA1 culture medium samples (after 120 h of incubation) and uncultured sterile medium (blank media) were collected after centrifugation at 13,000 g for 10 min. The supernatant was further filtered through a 10-kDa MWCO centrifugal unit by centrifugation at 14,000 g. The peptides were separated using a Grace Vydac C18 HPLC column (Polymeric 300 Å, 5 μm, 2.1 mm × 250 mm) (Grace Davison Discovery Sciences, IL, USA) using a gradient elution with a mixture of two solvents; solvent A was 0.1 % (v/v) trifluoroacetic acid (TFA) in water, and solvent B was 0.1 % TFA in 90 % (v/v) aqueous acetonitrile. The gradient elution was performed as follows: 0–5 min, 0 % B; 5–45 min, 0–45 % B; 45–48 min, 45–80 % B; 48–58 min, 80 % B; and 58–68 min, 0 % B. The flow rate was 0.2 ml min−1, and the column was used at 30 °C and detected at 215 nm.
The peptide fractions separated by HPLC were collected manually and dried in a SpeedVac concentrator (Savant Inc., MI, USA). The samples were resuspended in 0.1 % TFA and 45 % acetonitrile. The matrix (α-cyano-4-hydroxycinnamic acid (CHCA)) (Sigma-Aldrich Co., MO, USA) was dissolved in 2:1 (water/acetonitrile) with 0.1 % TFA. The concentrated peptide samples were added to the CHCA matrix in 1:1 ratio onto a matrix-assisted laser desorption/ionization (MALDI) plate. Peptide de novo sequencing was performed by an Applied Biosystems 4700 Proteomics Analyzer (Life Technologies, CA, USA) with time of flight/time of flight (TOF/TOF) ion optics. The MS spectra and tandem MS spectra were carried out in reflector mode with external calibration using the 4700 calibration mixture kit (Life Technologies). MS/MS data was interpreted using MASCOT and PARAGON algorithms within the ProteinPilot 2.0 software.
Co-Cultivation of Strain MFA1 and M. smithii
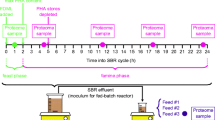
Co-cultivation with M. smithii in a medium containing a non-growth-limiting amount of yeast extract (0.8 %, w/v) was investigated to determine the effect of hydrogen on strain MFA1. The methanogen was initially grown under a H2/CO2 (80:20) atmosphere at 150-kPa pressure in BYE medium at 39 °C. Strain MFA1 and M. smithii were then separately transferred to BYE and FYE media. Both organisms were subcultured twice separately under the same culture conditions before inoculating each culture with similar cell densities (0.1 OD; 600 nm) into serum bottles containing 100 ml of BYE or FYE medium with 150 ml of headspace in triplicate. Pure cultures of each microorganism from the two different media served as controls. The headspace of the co-culture study contained an atmosphere of 95 % CO2 and 5 % H2 with no additional hydrogen, but pure M. smithii control cultures were further supplemented with 50 kPa of hydrogen. Two-milliliter culture samples were removed from fermentations at 0, 15, 27, 39, 51, 63, and 74 h after inoculation for determination of OD, fluoride ion production as an indicator of fluoroacetate degradation, and amino acid and short-chain fatty acid (SCFA) analyses.
Chemical Analyses
Headspace gas analysis for hydrogen and methane was performed throughout the co-culture study by withdrawing 2 ml of the headspace gas from each bottle at 0, 15, 27, 39, 51, 63, and 74 h after inoculation. Culture supernatants for SCFA (acetic, propionic, n-butyric, isobutyric, n-valeric, and branched C5 acid) determination were obtained at 0 and 74 h from cultures during co-culture study. Gas and SCFAs were analyzed by a gas chromatography (GC-2014, Shimadzu, Japan) using previously described methods [19].
Statistical Analysis
Sample means from amino acid and peptide utilization data were analyzed using one-way analysis of variance. Differences were considered significant at p < 0.05. Error bars shown in graphs represent standard error of the mean.
Results
Growth of Strain MFA1 on Different Protein Hydrolysates
Growth of strain MFA1 was enhanced when 20 mM fluoroacetate was added to cultures (Fig. 1a). During the exponential phase, the fluoroacetate-supplemented cultures exhibited a significantly shorter doubling time (p < 0.05, two-tailed paired t test) than the non-fluoroacetate-supplemented cultures. Strain MFA1 in fluoroacetate medium with individually supplemented casamino acids, phytone peptone, tryptone, and yeast extract had a doubling time of approximately 5 to 6 h. Growth in the absence of fluoroacetate resulted in longer doubling times in media supplemented with only casamino acids (10 h), phytone peptone (12 h), tryptone (10 h), and yeast extract (7 h) (Fig. 1a). Variations in apparent cell densities with different types of protein hydrolysates in fluoroacetate-supplemented media were observed, with yeast extract or tryptone both reaching an OD600 of 0.38, followed by phytone peptone with an OD600 of 0.25, and casamino acids demonstrating the least growth with an OD600 of 0.13.
Effect of different peptide sources on a the bacterial growth of Synergistetes isolate strain MFA1 and b fluoroacetate degradation. Solid markers represent media containing 20 mM fluoroacetate and 0.4 % of the respective protein digests (casamino acids (circles); phytone peptone (squares); tryptone (triangles); yeast extract (diamonds)) and 0.002 % yeast extract; open markers represent media containing only 0.4 % of the protein digests and 0.002 % yeast extract
The extent of fluoroacetate degradation was strongly correlated to MFA1 growth on different types of protein substrates (r 2 value of 0.979). Strain MFA1 was able to completely degrade 20 mM fluoroacetate in cultures containing either yeast extract or tryptone after 77 h of incubation (Fig. 1b). However, only 14 mM fluoride ions were produced from fluoroacetate degradation in the culture containing phytone peptone and 20 mM fluoroacetate. When the fluoroacetate medium was supplemented with casamino acids, strain MFA1 only degraded 10 mM fluoroacetate.
Amino Acid and Peptide Utilization from Yeast Extract
The yeast extract-rich BYE medium contained 49.6 mM of total amino acids (Table 1). Glutamate was the most abundant amino acid in yeast extract, with approximately 7.4 mM of free and peptide-bound glutamate in the sterile medium.
During growth on high amounts of yeast extract, strain MFA1 preferentially used hydrophilic amino acids, metabolizing 11.4 mM (39 %) of hydrophilic amino acids compared to 1.4 mM (7 %) of hydrophobic amino acids (Table 1). Arginine, asparagine, glutamate, glycine, and histidine from yeast extract were completely utilized. Leucine was the most utilized hydrophobic free amino acid (508 μM), followed by 247 μM of proline and 180 μM of phenylalanine.
When the growth medium was supplemented with 20 mM fluoroacetate, a significantly higher amount of free amino acids from yeast extract was degraded by strain MFA1 (Table 1). Elevated utilization for cysteine, lysine, serine, and threonine was observed, with percentage increases of 9.8, 30.0, 40.1, and 19.7 %, respectively. Glutamate, glycine, histidine, and asparagine as free amino acids were completely used similar to the non-fluoroacetate-containing medium. Hydrophobic free amino acids that were utilized to a greater extent (p < 0.05) during improved growth in the presence of fluoroacetate compared to basal medium were leucine, methionine, phenylalanine, and tyrosine, with 28.9, 45.9, 29.0, and 18.6 % increase, respectively.
Peptide-bound lysine was the most metabolized peptide containing amino acids from yeast extract and was the preferred form compared to the utilization of the free form (1264 μM compared to 176 μM, respectively). The addition of fluoroacetate did not significantly alter the utilization profile of peptide-bound lysine. Conversely, greater quantities (p < 0.05) of peptide-bound glutamate were metabolized from the fluoroacetate-containing medium, with a further increase of 628 μM glutamate peptides from 162 μM utilized during its growth in the basal yeast extract medium.
Utilization of Glutamate-Containing Peptides from Yeast Extract
HPLC separation of yeast extract peptides from sterile FYE medium identified two dominant peptide peaks at 17- and 23-min retention times (fractions 1 and 4, respectively; Fig. 2). The peptide profile of FYE medium following 120 h of incubation with strain MFA1 shows a reduction of these peaks and the appearance of two major peaks at 29 and 32 min (fractions 5 and 6, respectively; Fig. 2). The most depleted peptide, fraction 4, was characterized using MALDI-TOF/TOF. A clean mass spectrum consisting of three major peptide ions of m/z 1230.4, 1101.4, and 714.2 was generated (Fig. S1a). The difference between the first and second peaks was 387.1 Da, followed by 129.0-Da difference between the second and last peaks. These differences represent the loss of three and one glutamate, respectively. From the MS/MS ladder sequencing, the spectrum demonstrated fragment y ions separated by a mass difference of 129 Da, indicating that the peptide sequence is composed of mainly glutamate (Fig. S1b). The amino (N) terminal amino acid was predicted to be a glutamate due to the mass change of the first fragment peak (m/z 1101.2) at exactly 129.0 Da., resulting in the prediction of the peptide to be composed of a hepta-glutamate sequence at the N-terminus (Fig. 2).
Separation and identification of peptides from FYE medium. C18 reverse-phase HPLC reveals chromatograms of FYE uninoculated medium (black line) and after 120-h incubation with Synergistetes isolate strain MFA1 (grey line). The major peptide peak (fraction 4) consists of an oligopeptide with N-terminal hepta-glutamate
Co-Culture Experiment with M. smithii
Growth, Fluoroacetate Degradation, and Gas Production
Maximum growths of pure cultures of strain MFA1 and M. smithii in BYE medium were OD600 of 0.33 and 0.06, respectively (Fig. 3a). Co-cultivation of strain MFA1 with M. smithii reached a maximum OD600 of 0.50. In the presence of 20 mM fluoroacetate, cell densities of strain MFA1 alone and the co-culture of strain MFA1 with M. smithii reached an OD600 of 0.80. Fluoroacetate degradation through the detection of fluoride ions as an end product showed no difference in the rates or extent of strain MFA1 growth on its own or in co-culture with M. smithii (Fig. 3b)
Time course of co-culture of Synergistetes isolate strain MFA1 and Methanobrevibacter smithii: a microbial cell density measured at 600 nm; b fluoroacetate degradation; c headspace hydrogen concentration; and d headspace methane gas production of pure culture strain MFA1 (diamonds), pure culture M. smithii (triangles), and co-culture of strain MFA1 and M. smithii (squares). The study was carried out in BYE medium (open markers), and solid markers represent the FYE medium
Changes in the composition of the culture headspace gases were observed throughout the study. In BYE medium, there was a net accumulation of approximately 200 μmol of hydrogen by strain MFA1 (Fig. 3c), while no accumulation of hydrogen was observed by strain MFA1 in FYE medium, where approximately 40 μmol of hydrogen was maintained throughout the study. M. smithii grew in both BYE and FYE media with the addition of 50 kPa of hydrogen (approximately 3500 μmol). Hydrogen metabolism was continuously coupled with methane production in both media by the pure culture of M. smithii.
Co-cultures of strain MFA1 and M. smithii (without the addition of 50 kPa of hydrogen) in BYE medium generated no excess hydrogen and were continuously consumed to an undetectable level by 74 h. Similarly, in co-culture experiments with fluoroacetate, hydrogen levels were detected at relatively low levels throughout the study, at approximately 20 μmol. However, methane production during co-culture with fluoroacetate (FYE medium) was significantly lower compared to the co-culture in BYE medium (Fig. 3d). A total of 37 μmol of methane was detected from the headspace of the co-culture grown on fluoroacetate at 74 h, while approximately 470 μmol of methane was produced by M. smithii in co-culture grown on BYE medium.
Amino Acid Utilization
There was an overall utilization of approximately 10.9 mM of total amino acids by strain MFA1 when cultured alone in BYE medium, with the production of 574 μM ornithine and 200 μM alanine (Table 2). In co-culture with M. smithii, there was a 27 % increase in the utilization of total amino acids (13.8 mM), which includes utilization of 213 μM ornithine. In addition, the co-culture utilized a substantial amount of lysine (1338 μM) and serine (1565 μM) compared to strain MFA1 alone. Glutamate utilization was not significantly different between the pure culture of strain MFA1 and co-culture. An actively growing M. smithii in pure culture supplemented with 50 kPa hydrogen only consumed 0.4 mM of total amino acids.
SCFA Production
In BYE medium, strain MFA1 produced approximately 2 mM acetate and 4 mM propionate (Table 3) as the metabolic end products of fermentation. The co-culture in BYE medium produced higher concentrations of acetate (7.4 mM) and propionate (6.1 mM), while the actively growing pure M. smithii culture assimilated approximately 2 mM of acetate from the BYE medium. The production of branched C5 acid was undetectable in the pure culture of strain MFA1 culture, but trace amounts of branched C5 acid were produced in the co-culture (0.2 mM).
With the addition of 20 mM fluoroacetate to the media, strain MFA1 in pure culture produced approximately 23 mM of acetate, 6 mM propionate, and 0.4 mM branched C5 acid. Similar values of acetate and propionate were detected in the FYE medium after 74 h of incubation with strain MFA1 and M. smithii in co-culture. SCFAs were undetectable from pure M. smithii culture in FYE medium. Production or assimilation of n-butyric, isobutyric, and n-valeric acid was not detected in any of the cultures.
Discussion
We sought to gain a better understanding of the metabolic capabilities of Synergistetes isolate strain MFA1 with the ultimate goal of promoting its growth in the rumen and providing a microbial strategy to protect cattle from fluoroacetate poisoning. This study revealed that protein hydrolysates exert a stimulatory effect on strain MFA1 growth, regardless of the animal or plant origin and nitrogenous composition of the protein hydrolysates. The growth of strain MFA1 was significantly (p < 0.01, two-tailed t test) higher in tryptone, a pancreatic digest of casein with a higher peptide content, as compared to its growth in casamino acids, an amino acid-rich hydrolysate of casein. Yeast extract, a protein hydrolysate from autolyzed yeast, resulted in similar growth promotion as tryptone. Moreover, the growth of strain MFA1 was moderately induced by phytone peptone, an enzymatic digest of plant protein, likely due to its high peptide content despite its low total nitrogen content [25].
The differences in growth of strain MFA1 on the respective protein hydrolysates are primarily due to their total nitrogen composition and secondarily to their peptide abundance. The amino acid utilization profile of strain MFA1 from this study on yeast extract and from a previous study on casamino acids [19] demonstrated a complete utilization of arginine, asparagine, glutamic acid, glycine, and histidine amino acids regardless of the type of protein hydrolysate. These amino acids may represent the fundamental energy source for the growth of strain MFA1. However, certain peptides potentially promote the growth of strain MFA1. This notion is supported by the enhanced growth of strain MFA1 in protein hydrolysates that consist of a higher proportion of peptide than free amino acids. Furthermore, it was noted that glutamate- and lysine-containing peptides were preferentially utilized by strain MFA1 from yeast extract in the presence of fluoroacetate. Therefore, peptides, particularly the glutamate-containing peptides, promote the growth of strain MFA1, over the addition of just essential amino acids. Both tryptone and yeast extract have a higher abundance of glutamate peptides than phyptone peptone, while casamino acids consist of mainly free glutamate [26]. It is also worth noting that free glutamate is metabolized extensively by the predominant non-cellulolytic ruminal bacteria (Prevotella bryantii B14, Selenomonas ruminantium HD4, and Streptococcus bovis ES1) [27]. In addition, hydrophilic peptides containing a large proportion of arginine, aspartate, lysine, and glutamate are preferentially used by mixed ruminal bacteria in vitro [28]. Many factors other than peptide concentration may also influence their uptake, and therefore, a growth competition study of strain MFA1 with other predominant proteolytic ruminal bacteria may reveal the competitive attributes of strain MFA1.
Fluoroacetate results in a marked increase in the growth of strain MFA1, as demonstrated from this study and previous work [19]. Interestingly, the rate of fluoroacetate degradation by strain MFA1 in the four different protein hydrolysates strongly correlated to its growth rate in the respective protein sources, suggesting that the fluoroacetate degradation is linked to bacterial cell density. This is in accordance with other Synergistetes bacteria, where despite the increase in growth of these bacteria due to the presence of a suitable electron acceptor (e.g., crotonate and elemental sulfur), their reduction could be influenced by the type of nitrogen source [5, 29].
The degradation of fluoroacetate by strain MFA1 appeared to be consistent with other anaerobic respiratory mechanisms, and this bacterium may potentially use the intermediate organic acids and reducing agents generated from the fermentation of these protein hydrolysates such as formate, pyruvate, and hydrogen as the electron donor for the reduction of fluoroacetate. From the amino acid and peptide analyses, it is noted that more amino acids (e.g., serine, threonine, and lysine) were fermented in the presence of fluoroacetate. This observation is commonly associated with many organotrophic Synergistetes, where the presence of a suitable electron acceptor stimulates the growth of these bacteria on fermentable substrates [5, 11, 17, 18]. Moreover, the lack of a measureable production of fermentative hydrogen in the strain MFA1 culture during its growth with fluoroacetate is consistent with other Synergistetes bacteria during anaerobic respiration processes with inorganic/organic compounds [5, 12, 18].
Co-cultivation with a hydrogen-scavenging methanogen in the absence of fluoroacetate showed that the growth of strain MFA1 can be enhanced when fermentative hydrogen is removed by the methanogen. Many methanogenic archaea, including M. smithii, derive their energy from autotrophic growth through the coupling of hydrogen oxidation to carbon dioxide reduction, leading to exclusive production of methane [30]. Accordingly, previous studies have estimated the growth of methanogens in batch culture from the accumulation of methane in the system [30–33]. The final growth of M. smithii in co-culture with strain MFA1 in basal yeast extract medium was predicted to be relatively similar to the pure M. smithii culture at an OD (600 nm) of approximately 0.05 based on the comparable methane gas production by the methanogen in pure and co-culture. Therefore, the major increase in OD in the co-culture is likely due to strain MFA1.
Moreover, the increase in SCFA production from the co-culture in BYE medium is consistent with an increase in strain MFA1. M. smithii assimilates acetate during its growth in non-fluoroacetate-supplemented medium. This is in agreement with the genomic and metabolomic study conducted by Samuel et al. [34] that demonstrated the assimilation of acetate by M. smithii and up-regulation of genes involved in assimilation through an incomplete reductive tricarboxylic acid (TCA) cycle. However, no assimilation of acetate by M. smithii was observed during growth on fluoroacetate-containing medium. Early studies have shown that fluoroacetate is a powerful inhibitor of acetate oxidation in animals, yeast, and bacteria [35, 36] and completely inhibits acetoclastic methanogenesis [37]. M. smithii is not an acetolastic methanogen, but it has been demonstrated that acetate is a crucial growth-inducing compound for this hydrogen scavenger. Therefore, the presence of fluoroacetate potentially inhibits the growth of M. smithii. Conversely, the growth of strain MFA1 was not inhibited by fluoroacetate because this bacterium does not naturally assimilate acetate.
However, along with other fermentative bacteria, the growth of strain MFA1 may be inhibited with an increase in hydrogen partial pressure. Accumulation of hydrogen, unlike other fermentation products such as ammonia, causes feedback inhibition to fermentation processes, and the disposal of hydrogen using interspecies hydrogen transfer or anaerobic respiration leads to better growth [11, 12, 17, 38–42]. Ornithine is usually produced by strain MFA1 as a fermentative product in the basal medium [19]. But, ornithine was consumed by strain MFA1 when M. smithii (this study) or fluoroacetate [19] was used as an electron acceptor. The metabolism of ornithine and other amino acids such as serine by strain MFA1 is likely to account for the increase in acetate and propionate production in the co-culture. Similar observations have been made in Clostridium sporogenes and a phylogenetically distant Synergistetes bacterium, Thermanaerovibrio acidaminovorans [12, 43]. From these studies, it was proposed that hydrogen is a strict barrier for ornithine metabolism, which could be overcome by the addition of an electron acceptor [12, 43]. The significance of hydrogen disposal for growth of fermentative bacteria is that the oxidation process of reducing agents such as NADH and FADH2 coupled to proton reduction is only thermodynamically feasible under low-hydrogen conditions [44, 45]. In the rumen ecosystem, where hydrogen partial pressures are maintained at low levels due to the presence of methanogenic archaea, strain MFA1 would most likely metabolize ornithine from arginine catabolism to ammonia and SCFAs.
Similar to ornithine utilization during syntrophic growth, strain MFA1 also metabolized this non-proteinogenic amino acid in the presence of fluoroacetate [19]. Although both approaches for the disposal of hydrogen resulted in relatively similar amino acid utilization profiles, fluoroacetate degradation may have advantages over interspecies hydrogen transfer. This is clearly demonstrated from the enhanced bacterial growth in the presence of fluoroacetate compared to those in the syntrophic association. The degradation of fluoroacetate may be coupled with the oxidation of organic compounds during a respiratory process similar to halorespiration, which creates an electrochemical gradient that provides the chemical energy required for bacterial growth [46–48]. These results demonstrate that the syntrophic association may have resulted in the enhanced growth of strain MFA1 by promoting fermentation, but halorespiration of fluoroacetate may be more advantageous by facilitating ATP synthesis through electrochemical gradients in addition to amino acid fermentation.
During the growth of strain MFA1 in medium containing only amino acids, conditions may not be favorable for the fermentation of amino acids through Stickland reactions, which usually involve one amino acid acting as an electron donor for the reduction of another amino acid (electron acceptor) [49]. This is apparent from the production of hydrogen [19], which causes feedback inhibition to amino acid fermentation processes at higher concentrations. Whilst the introduction of peptides provides better growth due to a higher rate of amino acid fermentation, strain MFA1 can potentially benefit from both fermentative- and respiratory-type metabolism in the presence of an appropriate electron acceptor (e.g., fluoroacetate). This is indicated by the absence of measurable hydrogen production or detectable consumption of hydrogen by M. smithii in the co-culture with fluoroacetate. Strain MFA1 has the metabolic flexibility for energy conservation through fermentation, facultative syntrophy, or anaerobic respiration. This characteristic is shared with some other Synergistetes bacteria [5, 11, 18, 50]. For example, Anaerobaculum mobile can grow in syntrophy with a methanogen or through anaerobic respiration using either thiosulfate, sulfur, cystine, or crotonate as its electron acceptor [11].
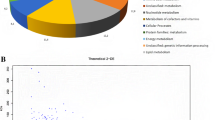
In summary, strain MFA1 predominately used five different hydrophilic amino acids for its growth, and use of peptides containing lysine or glutamate from yeast extract was stimulated during its syntrophic and respiratory growth. Fermentation of these amino acids and peptides during syntrophic growth with other hydrogen-scavenging methanogens in the rumen produces acetate, propionate, and hydrogen. However, in the presence of an electron acceptor, strain MFA1 readily switches to a respiratory-type metabolism using amino acids and fermentation intermediates as electron donors for the synthesis of ATP. We have begun to unravel the molecular underpinnings of these metabolic observations by sequencing the 3.5 Mb genome of strain MFA1 (data not shown). Preliminary analyses indicate that MFA1 encodes a high percentage of amino acid metabolism and transport genes, corresponding to 13 % of the total COG functional categories, which is a characteristic of the Synergistetes phylum and higher than the bacterial average (~8 %) [6]. Furthermore, multiple complete pathways exist for the metabolism of glutamate to carboxylic acid and other amino acids such as glutamine and γ-aminobutyric acid (data not shown).
Further studies to promote the growth of strain MFA1 in vitro and in vivo using a range of peptides consisting of glutamate or lysine in conjunction with an appropriate electron acceptor should be conducted. In addition, this study has provided us with an opportunity to investigate an approach to alter fermentative pathways resulting in the reduction of hydrogen availability for methane formation in the rumen.
References
Vartoukian SR, Palmer RM, Wade WG (2007) The division “Synergistes”. Anaerobe 13:99–106
Godon JJ, Moriniere J, Moletta M, Gaillac M, Bru V, Delgenes JP (2005) Rarity associated with specific ecology niches in the bacterial world: the ‘Synergistes’ example. Environ Microbiol 7:213–224
Baena S, Fradeau ML, Ollivier B, Labat M, Thomas P, Garcia JL, Patel BKC (1999) Aminomonas paucivorans gen. nov., sp. nov., a mesophilic, anaerobic, amino-acid-utilizing bacterium. Int J Syst Evol Microbiol 49:975–982
Ganesan A, Sebastien C, Tarrade A, Dauga C, Bouchez T, Pelletier E, Le Paslier D, Sghir A (2008) Cloacibacillus evryensis gen. nov., sp. nov., a novel asaccharolytic, mesophilic, amino-acid-degrading bacterium within the phylum ‘Synergistetes’, isolated from an anaerobic sludge digester. Int J Syst Evol Microbiol 58:2003–2012
Zavarzina DG, Zhilina TN, Tourova TP, Kuznetsov BB, Kostrikina NA, Bonch-Osmolovskaya EA (2000) Thermanaerovibrio velox sp. nov., a new anaerobic, thermophilic, organotrophic bacterium that reduces elemental sulfur, and emended description of the genus Thermanaerovibrio. Int J Syst Evol Microbiol 50:1287–1295
Hugenholtz P, Hooper SD, Kyrpides NC (2009) Synergistetes. Environ Microbiol 11:1327–1329
Allison MJ, Mayberry WR, McSweeney CS, Stahl DA (1992) Synergistes jonesii, gen. nov., sp. nov.: a rumen bacterium that degrades toxic pyridinediols. Syst Appl Microbiol 15:522–529
Rees GN, Patel BKC, Grassia GS, Sheeny AJ (1997) Anaerobaculum thermoterrenum gen. nov., sp. nov., a novel, thermophilic bacterium which ferments citrate. Int J Syst Evol Microbiol 47:150–154
Jumas-Bilak E, Carlier JP, Jean-Pierre J, Citron D, Bernard K, Damay A, Gay B, Teyssier C, Campos J, Marchandin H (2007) Jonquetella anthropi gen. nov., sp. nov., the first member of the candidate phylum ‘Synergistetes’ isolated from man. Int J Syst Evol Microbiol 57:2743–2748
Looft T, Levine UY, Stanton TB (2013) Cloacibacillus porcurum sp. nov., a mucin-degrading bacterium from the swine intestinal tract and emended description of the genus Cloacibacillus. Int J Syst Evol Microbiol 63:1960–1966
Menes RJ, Muxi L (2002) Anaerobaculum mobile sp. nov., a novel anaerobic, moderately thermophilic, peptide-fermenting bacterium that uses crotonate as an electron acceptor, and emended description of the genus Anaerobaculum. Int J Syst Evol Microbiol 52:157–164
Plugge CM, Stams AJM (2001) Arginine catabolism by Thermanaerovibrio acidaminovorans. FEMS Microbiol Lett 195:259–262
McSweeney CS, Allison MJ, Mackie RI (1993) Amino acid utilization by the ruminal bacterium Synergistes jonesii strain 78-1. Arch Microbiol 159:131–135
Adams CJ, Redmond MC, Valentine DL (2006) Pure-culture growth of fermentative bacteria, facilitated by H2 removal: bioenergetics and H2 production. Appl Environ Microbiol 72:1079–1085
Soboh B, Linder D, Hedderich R (2004) A multisubunit membrane-bound [NiFe] hydrogenase and an NADH-dependent Fe-only hydrogenase in the fermenting bacterium Thermoanaerobacter tengcongenesis. Microbiology 150:2451–2463
Dahle H, Birkeland NK (2006) Thermovirga lienii gen. nov., sp. nov., a novel moderately thermophilic, anaerobic, amino-acid-degrading bacterium isolated from a North Sea oil well. Int J Syst Evol Microbiol 56:1539–1545
Magot M, Ravot G, Campaignolle X, Ollivier B, Patel BKC, Fardeau ML, Thomas P, Crolet JL, Garcia JL (1997) Dethiosulfovibrio peptidovorans gen. nov., sp. nov., a new anaerobic, slightly halophilic thiosulfate-reducing bacterium from corroding offshore oil wells. Int J Syst Evol Microbiol 47:818–824
Surkov AV, Dubinina GA, Lysenko AM, Glockner FO, Kuever J (2001) Dethiosulfovibrio russensis sp. nov., Dethiosulfovibrio marinus sp. nov. and Dethiosulfovibrio acidaminovorans sp. nov., novel anaerobic, thiosulfate- and sulfur-reducing bacteria isolated from ‘Thiodendron’ sulfur mats in different saline environments. Int J Syst Evol Microbiol 51:327–337
Davis CK, Webb RI, Sly LI, Denman SE, McSweeney CS (2012) Isolation and survey of novel fluoroacetate-degrading bacteria belonging to the phylum Synergistetes. FEMS Microbiol Ecol 80:671–684
Twigg LE, Wright GR, Potts MD (1999) Fluoroacetate content of Gastrolobium brevipes in Central Australia. Aust J Bot 47:877–880
Twigg LE, King DR (1991) The impact of fluoroacetate-bearing vegetation on native Australian fauna: a review. Oikos 61:412–430
Robison WH (1970) Acute toxicity of sodium monofluoroacetate to cattle. J Wildl Manag 34:647–648
Cohen SA, Michaud DP (1993) Synthesis of a fluorescent derivatizing reagent, 6-aminoquinolyl-N-hydroxysuccinimidyl carbamate, and its application for the analysis of hydrolysates amino acids via high performance liquid chromatography. Anal Biochem 211:279–287
Meltzer NM, Tous GI, Gruber S, Stein S (1987) Gas-phase hydrolysis of proteins and peptides. Anal Biochem 160:356–361
Rada V, Petr J (2000) A new selective medium for the isolation of glucose non-fermenting bifidobacteria from hen caeca. J Microbiol Methods 43:127–132
BD Biosciences (2006) Advanced bioprocessing. BD Bionutrients technical manual, 3rd edn. BD Biosciences, Sparks
Atasoglu C, Valdes C, Walker ND, Newbold JC, Wallace JR (1998) De novo synthesis of amino acids by the ruminal bacteria Prevotella bryantii B14, Selenomonas ruminantium HD4, and Streptococcus bovis ES1. Appl Environ Microbiol 64:2836–2843
Chen G, Strobel HJ, Russell JB, Sniffen CJ (1987) Effect of hydrophobicity on utilization of peptides by ruminal bacteria in vitro. Appl Environ Microbiol 53:2021–2025
Baena S, Fardeau ML, Labat M, Ollivier B, Garcia JL, Patel BKC (2000) Aminobacterium mobile sp. nov., a new anaerobic amino-acid-degrading bacterium. Int J Syst Evol Microbiol 50:259–264
Garcia JL (1990) Taxonomy and ecology of methanogens. FEMS Microbiol Rev 87:297–308
Savant DV, Shouche YS, Prakash S, Ranade DR (2002) Methanobrevibacter acididurans sp. nov., a novel methanogen from a sour anaerobic digester. Int J Syst Evol Microbiol 52:1081–1087
Lai M-C, Shu C-M, Chen S-C, Lai L-J, Chiou M-S, Hua JJ (2000) Methanosarcina mazei strain O1M9704, methanogen with novel tubule isolated from estuarine environment. Curr Microbiol 41:15–20
Maestrojuan GM, Boone DR (1991) Characterization of Methanosarcina barkeri MST and 227, Methanosarcina mazei S-6T, and Methanosarcina vacuolata Z-761T. Int J Syst Evol Microbiol 41:267–274
Samuel BS, Hansen EE, Mancester JK, Coutinho PM, Henrissat B, Fulton R, Latreille P, Kim K, Wilson RK, Gordon JI (2007) Genomic and metabolic adaptations of Methanobrevibacter smithii to the human gut. Proc Natl Acad Sci U S A 104:10643–10648
Bartlett GR, Barron ESG (1947) The effect of fluoroacetate on enzymes and on tissue metabolism. Its use for the study of the oxidation pathway of pyruvate metabolism. J Biol Chem 170:67–82
Kalnitsky G, Guzman Barron ES (1947) The effect of fluoroacetate on the metabolism of yeast and bacteria. J Biol Chem 170:83–95
Winfrey MR, Zeikus JG (1979) Anaerobic metabolism of immediate methane precursors in Lake Mendota. Appl Environ Microbiol 37:244–253
Goodwin S, Giraldo-Gomez E, Mobarry B, Switzenbaum MS (1991) Comparison of diffusion and reaction rates in anaerobic microbial aggregates. Microb Ecol 22:161–174
Iannotti EL, Kafhewitz D, Wolin MJ, Bryant MP (1973) Glucose fermentation products of Ruminococcus albus grown in continuous culture with Vibrio succinogenes: changes caused by interspecies transfer of H2. J Bacteriol 114:1231–1240
Robert C, Del’Homme C, Bernalier-Donadille A (2001) Interspecies H2 transfer in cellulose degradation between fibrolytic bacteria and H2-utilizing microorganisms from the human colon. FEMS Microbiol Ecol 205:209–214
Rychlik JL, May T (2000) The effect of a methanogen, Methanobrevibacter smithii, on the growth rate, organic acid production, and specific ATP activity of three predominant ruminal cellulolytic bacteria. Curr Microbiol 40:176–180
Scheifinger CC, Lineham B, Wolin MJ (1975) H2 production of Selenomonas ruminatium in the absence and presence of methanogenic bacteria. Appl Microbiol 29:480–483
Wildenauer FX, Winter J (1986) Fermentation of isoleucine and arginine by pure and syntrophic cultures of Clostridium sporenges. FEMS Microbiol Lett 38:373–379
Stams AJM (1994) Metabolic interactions between anaerobic bacteria in methanogenic environments. Antonie Van Leeuwenhoek 66:271–294
Thauer RK, Jungermann K, Decker K (1977) Energy conservation in chemotrophic anaerobic bacteria. Bacteriol Rev 41:100–180
van de Pas BA, Jansen S, Dijkema C, Schraa G, De Vos WM, Stams AJM (2001) Energy yield of respiration on chloroaromatic compounds in Desulfitobacterium dehalogenans. Appl Environ Microbiol 67:3958–3963
Mohn WW, Tiedje JM (1991) Evidence for chemiosmotic coupling of reductive dechlorination and ATP synthesis in Desulfomonile tiedjei. Arch Microbiol 157:1–6
Schumacher W, Holliger C (1996) The proton electron ratio of the menaquinone-dependent electron transport from dihydrogen to tetrachloroethene in ‘Dehalobacter restrictus’. J Bacteriol 178:2328–2333
Stickland L (1934) Studies in the metabolism of the strict anaerobes (genus Clostridium): the chemical reactions by which Cl. sporogenes obtains its energy. Biochem J 28:1746–1759
Diaz-Cardenas C, Lopez G, Patel BKC, Baena S (2010) Dethiosulfovibrio salsuginis sp. nov., an anaerobic, slightly halophilic bacterium isolated from a saline spring. Int J Syst Evol Microbiol 60:850–853
Acknowledgments
The authors thank E. J. Gagen for comments on the manuscripts. The research was partly funded by the Commonwealth Science and Industrial Research Organization (CSIRO) Office of Chief Executive postgraduate scholarship program and Meat and Livestock Australia. PH was supported by an ARC Discovery Outstanding Research Award (ARC-DP120103498).
Conflict of Interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 40 kb)
Rights and permissions
About this article
Cite this article
Leong, L.E.X., Denman, S.E., Hugenholtz, P. et al. Amino Acid and Peptide Utilization Profiles of the Fluoroacetate-Degrading Bacterium Synergistetes Strain MFA1 Under Varying Conditions. Microb Ecol 71, 494–504 (2016). https://doi.org/10.1007/s00248-015-0641-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-015-0641-4