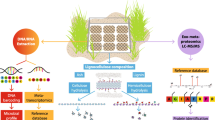
Abstract
The conversion of lignocellulosic biomass into biofuels can potentially be improved by employing robust microorganisms and enzymes that efficiently deconstruct plant polysaccharides at elevated temperatures. Many of the geothermal features of Yellowstone National Park (YNP) are surrounded by vegetation providing a source of allochthonic material to support heterotrophic microbial communities adapted to utilize plant biomass as a primary carbon and energy source. In this study, a well-known hot spring environment, Obsidian Pool (OBP), was examined for potential biomass-active microorganisms using cultivation-independent and enrichment techniques. Analysis of 33,684 archaeal and 43,784 bacterial quality-filtered 16S rRNA gene pyrosequences revealed that archaeal diversity in the main pool was higher than bacterial; however, in the vegetated area, overall bacterial diversity was significantly higher. Of notable interest was a flooded depression adjacent to OBP supporting a stand of Juncus tweedyi, a heat-tolerant rush commonly found growing near geothermal features in YNP. The microbial community from heated sediments surrounding the plants was enriched in members of the Firmicutes including potentially (hemi)cellulolytic bacteria from the genera Clostridium, Anaerobacter, Caloramator, Caldicellulosiruptor, and Thermoanaerobacter. Enrichment cultures containing model and real biomass substrates were established at a wide range of temperatures (55–85 °C). Microbial activity was observed up to 80 °C on all substrates including Avicel, xylan, switchgrass, and Populus sp. Independent of substrate, Caloramator was enriched at lower (<65 °C) temperatures while highly active cellulolytic bacteria Caldicellulosiruptor were dominant at high (>65 °C) temperatures.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Microorganisms that are able to enzymatically deconstruct the complex, heteropolymeric structure of lignocellulosic biomass are of particular interest since these strains or their enzymes can potentially be developed for biofuel production from renewable plant material [1, 2]. The decomposition of lignocellulosic biomass in nature occurs through the action of complex microbial communities organized into specific metabolic guilds with complementary or synergistic activities. Studies employing metagenomics and/or pyrosequencing of 16S rRNA tags have provided new insights into the diversity and organization of biomass-degrading communities such as within wood-feeding insects [3, 4]; rumen environments [5–7], composting systems [8, 9], and laboratory microcosms [8, 10]. Thermophilic microorganisms with (hemi)cellulolytic capabilities provide additional advantages over mesophiles for utilizing plant biomass such as faster reaction kinetics and greater enzyme stabilities under process conditions [11]; therefore, several model thermophiles are being developed for cost-effective, second-generation biofuel production [12–18]. Despite the intense interest in thermophilic biomass decomposition, microbial ecology-based studies examining the community-wide (hemi)cellulolytic potential of certain terrestrial hot springs have only recently emerged [19]. While many carbohydrate-active isolates have been obtained from hot spring environments, their study in isolation provides little information regarding the potential interactions between other microbes within their niche.
With an interest in rapid thermophilic deconstruction of plant biomass, we examined a well-known terrestrial hot spring environment known for its broad diversity in both cultivated and uncultivated Bacteria [20–24] and Archaea [25–27]. Previous cultivation-based studies from samples collected at Obsidian Pool (OBP), Yellowstone National Park, WY produced highly cellulolytic and xylanolytic strains from the genus Caldicellulosiruptor [24, 28] and the isolate Caldicellulosiruptor obsidiansis has been shown to effectively colonize cellulose surfaces [29] and deconstruct biomass through a suite of carbohydrate-active enzymes [30–32].
The aim of this study was to get a broader view of the diversity of biomass-active microorganisms inhabiting OBP using both high-throughput (HT) pyrosequencing and laboratory enrichment cultures containing both model and real-world biomass substrates. Enrichment cultures were employed in order to collect metabolite data to verify that active fermentation was occurring on individual insoluble substrates at various temperatures, which would be difficult to control using substrates deployed in situ (“bugtraps”). Enrichments established at elevated temperatures (55–85 °C) were designed to select for organisms that were most active on crystalline cellulose (Avicel), xylan, acid-pretreated switchgrass, and dilute acid-pretreated Populus biomass. A broad community analyses were performed using 454 FLX pyrosequencing on samples collected from the primary fluid source in the spring itself as well as a flooded depression adjacent to OBP supporting dense ground vegetation (Juncus tweedyi). The full-length 16S rRNA gene sequences from clone libraries derived from enriched samples were obtained to provide higher-resolution taxonomic identification of biomass-active organisms at the species or strain level. As expected, much higher overall diversity was encountered in sediment samples collected from the vegetated area where temperatures varied from 57 to 74 °C. Likewise, this environment produced active biomass-fermenting communities dominated by bacteria up to ∼80 °C, but activity was inhibited at the higher temperatures tested. Numerous uncultivated organisms were detected in the enrichment samples providing a source of isolation targets for future studies aimed at cooperative biomass deconstruction in the extremely thermophilic temperature range.
Materials and Methods
Sampling Site and Sample Collection
Obsidian Pool is a hot spring in the Mud Volcano Area, which is located within the Yellowstone Caldera (N44°36.603′ W110°26.331′) of Yellowstone National Park (YNP). Sediment and water samples were collected from the main pool (OBP5, OBP3) and a flooded depression (OBP10) on the eastern edge of the spring, which contains both living and decaying J. tweedyi (Tweedy’s rush) as a source of lignocellulosic material. The temperature and pH of each collection site was measured before sampling. Replicate samples were collected as described earlier [24]. The samples were kept in tightly sealed bottles and placed at 4 °C until they were transported in a cooler (∼10 h) to the Oak Ridge National Laboratory (ORNL), Oak Ridge, TN. The samples were stored in the dark at 4 °C until processed within 1 week.
Elemental Analysis
Immediately upon arrival to ORNL, a total of 25 metals were quantified in OBP3 and OBP10 sediments via inductively-coupled plasma mass spectrometry (ICP-MS, PerkinElmer Elan 600) including Li, Be, Na, Mg, Al, K, Ca, Cr, Fe, Mn, Ni, Co, Cu, Zn, Ga, As, Se, Sr, Ag, Cd, Cs, Ba, Pb, Bi, and U. Previous studies have provided water chemistry and hydrogen concentration for OBP: sulfide, 17.6 μM; sulfate, 0.33–0.52 μM; H2, 133.2–325.3 nM; CH4, 0.1 μM; CO2, 12.8–14.9 mM; C n H n , 21–63 ng l−1; and Cl, 25–305 mg l−1 [33, 34].
Enrichment Culture Experiments
The enrichment culture medium consisted of 4.5 mM KCl, 4.7 mM NH4Cl, 2.5 mM MgSO4, 1.0 mM NaCl, 0.7 mM CaCl2·2H2O, 0.25 mg/ml Resazurin, 1.4 mM cysteine-HCl·H2O, 1 mM Na2S·9H2O, 3.0 mM NaHCO3, 1 mM phosphate buffer, 10 mM 3-(N-morpholino)propanesulfonic acid (MOPS) pH 6.8, 1× ATCC trace minerals (Manassas, VA), 1× ATCC vitamin supplement (Manassas, VA), 0.02 % (w/v) yeast extract (Fisher Scientific, Pittsburgh, PA), and 1.0 g/l carbon source. Carbon and energy sources included (1) high-purity microcrystalline cellulose with ∼50-μm particle size (Avicel® PH-101) (Sigma-Aldrich Co., St. Louis, MO), (2) xylan (≥90 % HPLC) from beechwood (Sigma-Aldrich Co., St. Louis, MO), (3) dilute sulfuric acid-pretreated Alamo switchgrass (Panicum virgatum), and (4) dilute sulfuric acid-pretreated shavings of Populus deltoides. Pretreated biomass substrates were provided by the National Renewable Energy Laboratory, Golden, CO and were processed under the following conditions: 190 °C, 0.050 g sulfuric acid g−1 biomass, residence time of 1 min, and 25 % (w/w) total solids. Media were prepared anaerobically using a modified Hungate technique [35] and sterilized by autoclaving. Enrichment cultures were established by inoculating seven 125-ml serum bottles containing 50 ml of medium with 0.5 ml from environmental sample and incubating at 55, 60, 65, 70, 75, 80, and 85 °C with shaking at 90-rpm. Incubation times varied from 16 to 24 h, after which samples were transferred to a new serum bottle of the same substrate composition. Cultures were transferred three times to ensure enrichment on specified substrates. Growth was monitored by observing direct microscopic cell count, measuring changes in pH, and end-product analyses. Cell densities were determined by using a Thoma cell counting chamber (Blaubrand, Wertheim, Germany) with a Zeiss Axioskop2 Plus microscope with phase-contrast illumination.
Metabolite Measurements
Low Molecular Weight Fatty Acid and Alcohol Analysis
Acetate, lactate, and ethanol production was analyzed via high-performance liquid chromatography (HPLC) by filtering (0.2 μm) 1 ml of supernatant samples and acidifying with 200 mM sulfuric acid (final concentration of 5 mM) before injection into a Waters Breeze 2 HPLC system (Waters Corp., Milford, MA). Metabolites were separated on an Aminex HPX-87H column (BioRad Laboratories, Hercules, CA) at 60 °C using 5 mM sulfuric acid as the mobile phase at 0.6 ml min−1 and then passed through a refractive index (RI) detector (Waters 2414). Identification was performed by comparison of retention times with known standards. Quantitation of the metabolites was calculated against linear standard curves. All standards were prepared in uninoculated culture media to account for interference of salts in the RI detector. Samples were run in triplicate.
Gas Analysis
Gases were sampled from the headspace of the serum vial and analyzed immediately on an Agilent 6850 gas chromatograph (Agilent Technologies, USA) equipped with a thermal conductivity detector (TCD). All gas analytes were separated on an HP-PLOT U column (30 m × 0.32 mm × 0.10 μm film; J&W Scientific, Agilent Technologies, USA). Two HP-PLOT U columns were joined together for a total length of 60 m for optimized separation. Samples for carbon dioxide measurements were injected into a 185 °C split-splitless injector with a split ratio set to 3:1 and isocratic oven (70 °C) and helium carrier flow (5.1 ml min−1). The detector had a helium makeup flow of 10 ml min−1 at 185 °C, with the detector filament set for positive polarity. Samples to detect hydrogen concentrations were injected into a 185 °C split-splitless injector with a split ratio of 3:1 and isocratic oven (180 °C) and nitrogen carrier flow (3.5 ml min−1). The detector had a nitrogen makeup flow of 10 ml min−1 at 185 °C with the detector filament at negative polarity. Peak identifications were performed by comparison with known standards. Quantification of each compound was calculated against individual linear standard curves.
DNA Extraction from Environmental Samples, 454 Pyrosequencing of 16S rRNA Genes, and Analyses
Total community genomic DNA (gDNA), which contained both prokaryotic and eukaryotic DNA, was extracted from 5 g of original OBP sediment slurry samples using the PowerMax® Soil DNA Isolation Kit (MO BIO Laboratories, Inc., Carlsbad, CA). The theoretical size of the microbial community was estimated based on the total DNA recovered and the assumption that the predicted effective genome size of the individual cell of soil/sediment bacteria or archaea is 4.7 Mb as estimated from metagenomics data [36, 37], and a genome of this size would weigh 4.05 fg [38].
The hypervariable V4 regions of the 16S rRNA gene for Bacteria (∼290 bp) and Archaea (∼340 bp) were amplified from the total gDNA using the high-fidelity AccuPrime Pfx DNA polymerase (Invitrogen, Carlsbad, CA) as described earlier [39, 40]. Sequencing reactions were performed on a 454 Life Sciences Genome Sequencer FLX (Roche Diagnostics, Indianapolis, IN). FASTA files (raw 454 reads) were initially processed through the Ribosomal Database Project (RDP) pyrosequencing pipeline [41], where they were sorted by sample identifiers; the tag and 16S primer sequences were trimmed off, and low-quality sequences (using minimum average quality score of 20, maximum number of N = 0, minimum sequence length = 100) were removed. For each sample, 9,040 to 18,354 high-quality sequences of size 200–210 bp were obtained. Chimeric sequences were detected by using the Chimera Slayer [42] and were removed from the analyses. The 454 pyrosequences were analyzed using the RDP pyrosequencing pipeline tools for alignment, clustering, rarefaction, and determining Shannon and Chao1 diversity indices from a single sample file. Operational taxonomic units (OTUs) were defined at 97 % identity, and representative sequences were selected using the RDP pipeline. The 16S rRNA gene sequences were assigned to a set of hierarchical taxa using the RDP Naïve Bayesian rRNA Classifier v.2.0 [43] with a bootstrap cutoff of 50 % for short pyrosequences [44]. Additionally, the 454 pyrosequences processed through the RDP pyrosequencing pipeline were uploaded to MG-RAST [45]. The MG-RAST webserver was used to compare the sequencing data using the “best hit classification” function with RDP as the annotation source, a maximum E value cutoff of 10−5, a minimum percent identity of 97 %, and a minimum alignment length of 50 bp. Phylogenetic data downloaded from MG-RAST were imported into STAMP v2.0.0 (release candidate 6; [46] for additional statistical analyses using ANOVA, and P < 0.05 was considered significant.
DNA Extraction from Enrichment Cultures, Cloning, Sanger Sequencing, and Analyses
The total gDNA was extracted from each out of 24 combinations of growth temperature and substrate using 3 ml of the enrichment culture and the PowerSoil™ DNA Isolation Kit (MoBio Laboratories, Inc., Carlsbad, CA). The genomic DNA was amplified using GoTaq Flexi DNA polymerase (Promega, Madison, WI) and universal bacterial primers targeted to Escherichia coli 16S rRNA positions 8–27 (5′-AGA GTT TGA TCC TGG CTC AG-3′) and 1510 to 1492 (5′-GGT TAC CTT TTA CGA CTT-3′). The resulting PCR product of approximately 1.5 kb contained essentially the complete 16S rRNA gene allowing better taxonomic resolution. PCR products were purified from UltraPure™ Agarose (Invitrogen, Carlsbad, CA) using QIAquick Gel Extraction Kit (Qiagen Inc., Valencia, CA). PCR products were ligated in pCR 2.1-TOPO vectors (Invitrogen), transformed into TOP10 chemically competent E. coli, and plated onto LB agar containing 50 μg ml−1 kanamycin and X-gal. Transformants were incubated overnight at 37 °C, and white colonies were selected. The clones were sequenced in both directions at The Genome Center of Washington University using Applied Biosystems 3730 Platform (unfortunately clones from 70 °C were lost during transportation). The Sanger raw sequences were run via the Joint Genome Institute’s 16S GeneLib pipeline yielding 1,509 high-quality sequences. The 16S rRNA gene sequences were assigned to a set of hierarchical taxa using the RDP Naïve Bayesian rRNA Classifier v.2.0 [43] with confidence threshold of 80 %. The sequences were aligned using the Needleman alignment method [47] and the greengenes alignment database within the program Mothur [48]. To determine variations between combinations of substrate and temperature, the enrichments were compared at the sequence level using an unweighted UniFrac algorithm [49, 50] and visualized through principal component analysis (PCA). For UniFrac analysis, the neighbor-joining tree generated using MEGA4 software [51] was rooted with the Methanosarcina mazei 16S rRNA gene sequence (EF452664). The environmental input file for the UniFrac analysis contained the count of how many times the representative sequences appeared in the clone library. P values were corrected for multiple comparisons by multiplying the calculated P value by the number of comparisons made (Bonferroni correction) [49, 50].
Nucleotide Accession Numbers
The Sanger sequences obtained during this study were deposited in GenBank under accession numbers KJ479277–KJ480727. The 454 pyrosequences were uploaded to MG-RAST and made publically available with the following accession numbers: 4535428.3, 4535429.3, 4535430.3, 4535431.3, 4535432.3, and 4535433.3.
Results
Sampling Site Description
The Mud Volcano area (encompassing OBP) is generally categorized as a vapor-dominated (acid-sulfate) system, which is fed via fractures by reduced gases (e.g., H2 and H2S) from the 3–10-km-deep magma body [52, 53]. Approaching the surface, the sulfides are oxidized resulting in acidic fluids of elevated sulfate concentrations capable of considerable chemical weathering, which releases cations into solution [54]. Elevated concentrations of metals (except Al, Be, K, Se, Ag, Cs, Bi) were found in the peripheral area in comparison to the main pool (Table S1). Obsidian Pool is relatively shallow with a maximum depth of ∼1 m and varies in size throughout the year [34]. Algal mats and both living and decaying J. tweedyi (Tweedy’s rush) plants were present, especially at the eastern and southern edge of OBP, which is flooded with heated water (Fig. 1).
Relative abundance of Bacteria and Archaea in Obsidian Pool (OBP5, OBP3) and adjacent vegetated area (OBP10). Samples were collected in October 2007 (OBP5) and July 2008 (OBP3, OBP10). Increased steam due to low air temperatures partially obscures the OBP5 image. Collection sites (arrows) and water temperatures are indicated. The yellow box indicates the OBP10 sample area in relation to Obsidian Pool. Clones that could not be assigned with the >50 % confidence threshold to any phylum were referred as “unclassified.” The bacterial group “Other” includes the phyla Actinobacteria, Bacteroidetes, Chlorobi, Chloroflexi, Gemmatimonadetes, Nitrospirae, Synergistetes, and Tenericutes, which were present at <0.4 %
Microbial Community Size Estimation
The gDNA yield ranged from 0.6 to 2.0 μg g−1 of water-sediment slurry. The theoretical size of the microbial community based on the yield of gDNA ranged from 1.5 × 108 to 5.0 × 108 cells g−1 (Table 1). However, the theoretical estimation of microbial cells was significantly higher than DAPI whole cell counts of 2.3 × 106 cells g−1 determined previously from OBP sediments [55]. This overestimation is potentially due to the eukaryotic component of the total extracted gDNA.
Bacterial and archaeal diversity in the main OBP varied somewhat between the October and July sampling times. Archaeal diversity in the main pool was higher than bacterial; however, in the vegetated area, bacteria were more diverse (Table 1). Rarefaction analyses show that a number of individuals sampled in the main pool reached saturation and more intensive sampling was not likely to yield new additional species (Fig. S1, Table 1). The steep slope of the curve for the OBP10 Bacteria (Fig. S1) reflects significantly higher phylotype diversity in the bordering area supporting J. tweedyi (OBP10) than in the main pool (OBP5 and OBP3). The OBP10 sample containing live and dead plant material was selected as an inoculum for enrichment cultures to further investigate the presence of putative cellulolytic and xylanolytic microorganisms.
Changes in Microbial Community Structure with Varying Growth Substrate and Incubation Temperature
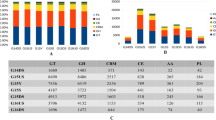
Based on the diversity analysis results, the OBP10 sample was selected as an inoculum for laboratory enrichment culture experiments with crystalline cellulose (Avicel), xylan, acid pretreated switchgrass, or Populus as the dominant carbon and energy source. Incubation temperatures ranged from 55 to 85 °C in 5 °C increments to examine temperature-dependent changes in microbial community activity and structure. Regardless of the substrate used, cell counts after the third passage on the respective substrate were higher at 55–65 °C (ca. 108 cells ml−1) than at 75–80 °C (ca. 107 cells ml−1), and no cells were detected at 85 °C. An increase in incubation temperature caused a shift in bacterial community structure from a Caloramator-dominated community at 55 °C to a Caldicellulosiruptor-dominated community at higher temperatures (Fig. 2).
Changes in bacterial community structure when grown on Avicel, switchgrass, Populus, and xylan with increasing incubation temperature. The data are derived from a total of 1,509 high-quality, full-length 16S rRNA gene sequences obtained by the Sanger method. The sequences were assigned to taxa using the Ribosomal Database Project Naïve Bayesian rRNA Classifier v.2.0 [43] with a confidence threshold of 80 %
The metabolite measurements showed that the primary product regardless of substrate and temperature combinations was acetate (Table 2). A maximum acetate concentration of 7.6 mM was detected on Populus at 65 °C; low acetate concentrations (1.2–1.3 mM) were measured on xylan at 55–70 °C, and no acetate was detected at 85 °C. The production of lactate was observed at temperatures of 75 and 80 °C on Avicel (2.3–2.6 mM) or Populus (1.8–2.0 mM), whereas ethanol at a concentration of 1.1 mM was detected only on Avicel at 80 °C (Table 2). Gases including H2 (0.4–8.24 mM) and CO2 (0.5–3.6 mM) were detected in all enrichments (Table 2).
A relationship between bacterial communities derived from individual enrichments was analyzed using an unweigthed UniFrac algorithm [49]. PCA revealed that 36.5 % of the total variance could be explained by the first two principal coordinates (Fig. 3). Both factors correlated with temperature. Axis 1 accounted for 24.6 % of the variation between communities grown at lower (55–65 °C) temperatures and communities from higher (75–80 °C) temperatures. Axis 2 explained 11.9 % of the variation and corresponded with changes in lignocellulosic substrate within any given temperature set. Regardless of the substrate, 16S rRNA gene sequences from 55 °C enrichments were placed in close proximity to each other and separately from others indicating that temperature, not substrate, was the main parameter affecting community development.
PCA analysis of 16S rRNA gene sequences derived from different enrichments with unweighted UniFrac. Neighbor-joining tree was rooted with an archaeal outgroup. The ends of the alignment were trimmed prior to the analysis so that the aligned sequences were all the same length. The enrichment names contain incubation temperature from 55 to 80 °C with interval of 5 °C, and letter stands for substrate: A Avicel, P Populus, S switchgrass, X xylan
Characterization of Bacterial and Archaeal 16S rRNA Pyrosequences
A total of 40,134 and 28,508 high-quality sequences were obtained after processing of 454 pyrosequencing data generated for the V4 region of bacterial and archaeal 16S rRNA genes, respectively. A small fraction of sequences (0.4–2.1 %) amplified with Bacteria-specific primers were affiliated with Archaea, and 1.1–4.4 % of sequences amplified with Archaea-specific primers were affiliated with Bacteria (Table 1). These sequences were removed from further analyses.
In the main OBP, the phylum Aquificae accounted for up to 98.2 and 87.2 % of the bacterial population in the October and July samples (P < 1e−15), respectively (Figs. 1 and S2). The relative abundance of other bacteria in the October sample was as low as 1.8 % of the total sequences with Proteobacteria (0.6 %) being the highest. While the phylum Crenarchaeota represented the majority of Archaea in both October (100.0 %) and July (99.3 %) main pool samples (Fig. 1), a change in sulfur-metabolizing members of the community from sulfur-reducing Acidianus dominating in October to organotrophic Thermofilum dominating in July was detected. In July, the temperature in the main pool was 6 °C higher and elemental sulfur deposits were observed on the surrounding banks. An increase of sulfur-dependent Crenarchaeota (Thermofilum, Pyrobaculum, Caldisphaera, Staphylothermus; P < 1e−15; Fig. S3), sulfate-reducing bacteria Thermodesulfobacteria (P < 1e−15), organotrophs such as Deinococcus-Thermus (P < 1e−15) and Dictyoglomi (P < 1e−15; Fig. S2), plus organotrophic archaea including Fervidicoccus (P < 1e−15, Fig. S3) was observed. The phyla Thaumarchaeota and Korarchaeota were found in July main pool samples in low (<0.6 %) abundance.
The flooded marsh-like area contained high bacterial diversity with 24.2 % of the sequences assigned to unclassified bacteria. Between the vegetated sediments and the main pool, a decrease in the relative abundance of Aquificae (P < 1e−15), and Thermodesulfobacteria (P = 3.06e−5) was observed, while changes in Deinococcus-Thermus (P = 0.789) and Dictyoglomi (P = 3.949) were not significant (Figs. 1 and S4). The members of the phyla Proteobacteria, Firmicutes, Verrucomicrobia, Acidobacteria, Spirochaetes, and Thermotogae were significantly higher (P < 1e−15) in abundance in OBP10 (Figs. 1 and S4). The Firmicutes comprised 20.4 %, the majority of which belonged to ten families (21 genera) of the class Clostridia including potentially cellulolytic bacteria from the genera Clostridium, Anaerobacter, Caloramator, Thermoanaerobacter, and Caldicellulosiruptor. The phyla Cyanobacteria, Chloroflexi, Gemmatimonadetes, Chlorobi, and Nitrospirae were only found in the peripheral sample (OBP10). With respect to Archaea, the peripheral area showed a decrease in the total abundance of Crenarchaeota (P < 1e−15), especially Caldisphaera (P < 1e−15), Pyrobaculum (P < 1e−15), and Staphylothermus (P = 0.019). Nevertheless, increases in crenarchaeal representatives such as Fervidicoccus (P = 2.08e−5) and Thermosphaera (P < 1e−15) were observed. A relative abundance of Candidatus Nitrosocaldus, a member of the phylum Thaumarchaeota, increased substantially up to 16.2 % (P < 1e−15) in OBP10. The methanogenic euryarchaeote, Methanocella, was also abundant (P = 0.04) in the peripheral area sediments (Figs. 1 and S5).
Phylogenetic Comparison of Bacterial Communities from Hot Spring Samples and Lignocellulose Enrichments
Bacterial sequences from sample OBP10 obtained through 454 pyrosequencing upon amplification with bacterial primers (9,014 sequences), archaeal primers (290 sequences), and Sanger sequences derived from enrichments (1,545 sequences) comprised a total of 10,848 sequences. These sequences were aligned and clustered using the RDP pyrosequencing pipeline resulting in 740 OTUs at 0.03 distance cutoff with 333 OTUs containing a single sequence. The remaining 407 OTUs were divided into five groups: 350 (86.0 %) OTUs contained only 454 pyrosequences obtained with bacterial primers, 30 (7.4 %) OTUs contained only Sanger sequences derived from enrichments, 21 (5.2 %) OTUs contained bacterial 454 pyrosequences obtained with both bacterial and archaeal primers, 5 (1.2 %) OTUs contained 454 pyrosequences and Sanger sequences, and 1 (0.2 %) OTU contained bacterial 454 pyrosequences obtained only with archaeal primers. Taxonomic identification showed that 108 OTUs were affiliated with potential cellulolytic bacteria such as Clostridia (103 OTUs), Dictyoglomi (3 OTUs), and Thermotogae (2 OTUs). In spite of the fact that bacteria from the genus Caldicellulosiruptor were abundant in the enrichments (668 sequences, 43.2 %) and there were no priming mismatches found, only 9.8 % of the total pyrosequences were annotated in the MG-RAST analysis as belonging to Caldicellulosiruptor saccharolyticus at an average identity of 98 % over 50 % of the alignment length. These Caldicellulosiruptor-like sequences were detected in the peripheral OBP10 area but not in water and sediment samples from the primary pool. A member of the genus Thermoanaerobacter, which could potentially utilize cellodextrins or xylan, was detected in laboratory enrichments (43 sequences, 2.8 %) with 37 sequences detected in xylan enrichments at 60, 65, and 75 °C. In the peripheral OBP10 area, the pyrosequences affiliated with the family Thermoanaerobacteraceae comprised only 0.7 % and were related to Carboxydothermus sp., Thermanaeromonas sp., and Thermoanaerobacter sp. Bacteria involved in the breakdown of lignocellulosic material such as Dictyoglomus, Fervidobacterium (Thermotogae), Caloramator, and Clostridium were detected in environmental OBP samples and in enrichments (Fig. 4).
Phylogenetic analysis of 16S rRNA gene sequences obtained by FLX 454 pyrosequencing of the V4 region amplified from OBP10 environmental gDNA and full-length Sanger sequences derived from OBP10 batch enrichments. The clusters with >10 sequences and those affiliated with potential cellulolytic bacteria are presented. The number of clones in each cluster is indicated in brackets. Symbols indicate clusters with sequences detected by 454 pyrosequencing using bacterial primers (open circles), or bacterial and archaeal primers (open triangles), clusters with Sanger sequences only (filled diamonds), and clusters which contain sequences obtained by 454 pyrosequencing and Sanger sequencing (filled squares). Additionally, the clusters with 454 pyrosequences and Sanger sequences are in bold, and the number of sequences obtained by each approach is indicated. The tree was produced by a neighbor-joining method. Bootstrap values were based on 1,000 replicates. The scale bar represents 0.05 changes per nucleotide position
Discussion
This study demonstrated that (1) bacterial diversity was significantly higher in the adjacent vegetated area of OBP versus the main spring likely in response to temperature gradients and higher organic nutrients, (2) microbial community structure and activity changed with respect to incubation temperature and biomass substrate in enrichment culture experiments but was limited by temperatures above 80 °C, and (3) the most active cellulolytic microorganisms were detected through enrichment culture techniques.
In OBP, members of the thermophilic phyla Aquificae and Crenarchaeota, which are capable of using carbon dioxide as the sole carbon source and gain energy by the oxidation of inorganic substances like hydrogen and sulfur [34], dominated over other microorganisms. Both bacteria and archaea in OBP showed variations in abundance, which may reflect changes in geochemical activity. Previous studies of OBP showed fluctuations in H2 (133.2–325.3 nM), CO2 (12.8–14.9 mM), and SO4 2− (0.33–0.52 μM) [33]; the SO4 2− concentration can range from 51.8 μM to greater than 10.4 mM [55]. Increases in elemental sulfur and SO4 2− concentrations potentially led to increases in the abundance of potentially sulfur-dependent Crenarchaeota and sulfate-reducing bacteria Thermodesulfobacteria and Thermodesulforhabdus. The abundance of Aquificae, which exclusively or preferentially use molecular H2 as an energy source [34], changed between 98.2 % in October and 87.2 % in July. Crenarchaeota of the family Sulfolobaceae was detected only in the October sample with Acidianus spp. being dominant. High abundance of the Aquificae and Crenarchaeota suggested that primary production in OBP is likely driven by chemoautotrophic hydrogen or sulfur oxidation. Cole et al. [56] used pyrosequencing tags to examine the microbial diversity from a number of samples collected from Great Boiling Spring (GBS) in Nevada. They compared communities from the spring water to those inhabiting sediments, some of which were collected near the edge of the spring, which is also surrounded by grasses. Similar to OBP, the main source water was dominated by Aquificae and crenarachaeotes (Thermoprotei); however, sediment samples were much more diverse with archaea dominating sediments at higher temperatures (82–87 °C) and bacterial groups representing the most abundant OTUs between 80 and 62 °C [56]. While this study did not examine as many samples as Cole et al., the overall observations were similar in that saturated OBP sediments occurring at lower temperatures than the hydrothermal source also harbor a much greater diversity of thermophilic microorganisms which are predominantly bacterial.
Cellulosic detritus from heat-tolerant J. tweedyi [57] is a possible source of carbon for heterotrophic microorganisms at least in localized niches, where the dilute acid-sulfate water of OBP [55] and elevated temperatures likely aid in solubilizing C5 sugars and possibly lignin providing a mild pretreatment of plant material for further enzymatic decomposition. The phylogenetic analysis of 16S rRNA gene pyrosequences from the OBP10 sample showed a diverse community represented by 3 archaeal and 18 bacterial phyla and many unclassified bacteria. High taxonomical and functional diversity driven by the abundance of micronutrients and organic material creates the potential for discovering novel bacterial species with robust cellulolytic capabilities. The OBP10 sample contained a high abundance of Firmicutes, for example Clostridium, Anaerobacter, Caloramator, Thermoanaerobacter, and Caldicellulosiruptor, which could participate in different stages of cellulose degradation. Interestingly, Caldicellulosiruptor-like sequences detected by 16S rRNA gene pyrosequencing had low percent identity to Caldicellulosiruptor spp. detected in the OBP10 sample enrichments, while Caldicellulosiruptor sequences from the OBP10 enrichments were 96.6 % identical to C. obsidiansis OB47 which was previously isolated from OBP [24]. This possibly suggests that in the natural environment, Caldicellulosiruptor spp. are highly diverse with only a small portion (below 0.005 %) of Caldicellulosiruptor population able to grow under laboratory enrichment conditions tested in this study. The same conclusion may be true for another potentially hemicellulose-degrading bacterium, Thermoanaerobacter, which was detected at low abundance (<0.7 %) in the OBP10 sample, and these sequences had an average of 80 % identity to Thermoanaerobacter spp. detected in xylan enrichments.
Different combinations of substrate and temperature stimulated the formation of distinct microbial communities with varying levels of metabolic activity. Metabolite measurements showed that acetate was the main product with any combination of substrate and temperature, whereas lactate and ethanol were produced at 75–80 °C on Avicel and Populus. Phylogenetic analyses of each enriched microbial community detected significant changes bound to temperature-dependent distribution of Caloramator and Caldicellulosiruptor. Caloramator spp. dominated Avicel, switchgrass, and Populus enrichments at 55 and 60 °C. Crystalline cellulose utilization is not widespread in the genus Caloramator; however, the ability to grow on cellulose has been reported for Caloramator australicus [58] and other strains have dominated cellulose enrichments under thermophilic conditions [59]. At higher incubation temperatures, Caldicellulosiruptor spp. clearly outcompeted other organisms, especially for Avicel, and pretreated switchgrass and Populus were crystalline and amorphous cellulose would be the primary carbon source. Many Caldicellulosiruptor spp. express a diverse set of glycoside hydrolases and can utilize polymeric components of plant cell walls including cellulose and hemicellulose [15, 60, 61], plus the 70–80 °C temperature range is optimal for their growth [24, 62]. A Dictyoglomus sp., represented by a single OTU, was also detected in 80 °C enrichments, especially when xylan was added as the carbon and energy source which is consistent with the known ability of these organisms to efficiently breakdown C5 polymers [63, 64]. Fervidobacterium spp. were also present in cultures with xylan incubated at 75–80 °C, and members of this genus have been reported to readily ferment xylan at temperatures up to 82 °C [65].
Other recent attempts to characterize lignocellulose degraders at extreme to hyperthermophilic temperature ranges in situ at GBS [19] produced quite different results compared to our laboratory enrichments. Thermotoga spp., while present in low abundance in the natural background, were highly enriched on deployed aspen and corn stover samples at 85 °C while our enrichments produced virtually no growth at these temperatures. However, enrichment of Caldicellulosiruptor spp. was not detected by Peacock et al. [19] at lower temperatures (77 °C), which contrasts our results suggesting that these organisms are not ubiquitous in freshwater hot spring environments. The temperature limit for cultivable plant biomass-degrading communities inhabiting Obsidian Pool appears to be ∼85 °C, where no production of acetate, lactate, or ethanol was observed with the exception of a trace amount of acetate (1 mM) in 85 °C xylan enrichments. Interestingly, no lactate was produced from enrichments with pretreated switchgrass or xylan at 75 and 80 °C, while lactate was produced from Avicel and Populus at these temperatures. Similar end-product profiles were also observed with the isolate C. obsidiansis when grown on these substrates [24], and highly related organisms dominated the Avicel and Populus enrichments at 75 and 80 °C. While this potentially provides an explanation for the difference in end-products at the higher temperature enrichments between substrates, factors controlling substrate-dependent (i.e., Avicel vs. switchgrass) carbon flux to either acetate or lactate are not well understood in Caldicellulosiruptor spp. Although acetate production was observed across all temperatures and substrates, the highest combined concentrations of acetate, lactate, and ethanol were always found to be at temperatures of 75 and 80 °C.
Characterizing and understanding natural microbial communities that are able to deconstruct plant biomass at elevated temperatures is highly relevant for improving future lignocellulosic bioconversion technologies. Obsidian Pool in YNP remains a “hot spot” for revealing diverse microbial communities with a wide range of phenotypic characteristics. In this study, we have pinpointed an area of the spring that harbors an excellent source of cellulolytic and xylanolytic thermophiles due to the unique ecology of the OBP10 sample environment. Deep-dive metagenomics/proteomics and identification of the specific glycoside hydrolases involved in plant cell wall deconstruction (studies in progress) will shed more light on the microbial ecology of thermophilic biomass decomposition in nature with future applications for biotechnology.
References
Carroll A, Somerville C (2009) Cellulosic biofuels. Annu Rev Plant Biol 60:165–182. doi:10.1146/annurev.arplant.043008.092125
Bayer E, Shoham Y, Lamed R (2006) Cellulose-decomposing bacteria and their enzyme systems. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes. Springer, New York, pp 578–617
Reid NM, Addison SL, Macdonald LJ, Lloyd-Jones G (2011) Biodiversity of active and inactive bacteria in the gut flora of wood-feeding huhu beetle larvae (Prionoplus reticularis). Appl Environ Microbiol 77:7000–7006
Warnecke F, Luginbuhl P, Ivanova N, Ghassemian M, Richardson TH, Stege JT, Cayouette M, McHardy AC, Djordjevic G, Aboushadi N, Sorek R, Tringe SG, Podar M, Martin HG, Kunin V, Dalevi D, Madejska J, Kirton E, Platt D, Szeto E, Salamov A, Barry K, Mikhailova N, Kyrpides NC, Matson EG, Ottesen EA, Zhang XN, Hernandez M, Murillo C, Acosta LG, Rigoutsos I, Tamayo G, Green BD, Chang C, Rubin EM, Mathur EJ, Robertson DE, Hugenholtz P, Leadbetter JR (2007) Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450:560–565
Fouts DE, Szpakowski S, Purushe J, Torralba M, Waterman RC, MacNeil MD, Alexander LJ, Nelson KE (2012) Next generation sequencing to define prokaryotic and fungal diversity in the bovine rumen. PLoS One 7. doi:10.1371/journal.pone.0048289
Hess M, Sczyrba A, Egan R, Kim TW, Chokhawala H, Schroth G, Luo SJ, Clark DS, Chen F, Zhang T, Mackie RI, Pennacchio LA, Tringe SG, Visel A, Woyke T, Wang Z, Rubin EM (2011) Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331:463–467. doi:10.1126/science.1200387
Brulc JM, Antonopoulos DA, Miller MEB, Wilson MK, Yannarell AC, Dinsdale EA, Edwards RE, Frank ED, Emerson JB, Wacklin P, Coutinho PM, Henrissat B, Nelson KE, White BA (2009) Gene-centric metagenomics of the fiber-adherent bovine rumen microbiome reveals forage specific glycoside hydrolases. Proc Natl Acad Sci U S A 106:1948–1953. doi:10.1073/pnas.0806191105
Allgaier M, Reddy A, Park JI, Ivanova N, D’Haeseleer P, Lowry S, Sapra R, Hazen TC, Simmons BA, VanderGheynst JS, Hugenholtz P (2010) Targeted discovery of glycoside hydrolases from a switchgrass-adapted compost community. PLoS One 5:e8812. doi:10.1371/journal.pone.0008812
van der Lelie D, Taghavi S, McCorkle SM, Li LL, Malfatti SA, Monteleone D, Donohoe BS, Ding SY, Adney WS, Himmel ME, Tringe SG (2012) The metagenome of an anaerobic microbial community decomposing poplar wood chips. PLoS One 7. doi:10.1371/journal.pone.0036740
Reddy AP, Allgaier M, Singer SW, Hazen TC, Simmons BA, Hugenholtz P, VanderGheynst JS (2011) Bioenergy feedstock-specific enrichment of microbial populations during high-solids thermophilic deconstruction. Biotechnol Bioeng 108:2088–2098. doi:10.1002/bit.23176
Taylor MP, Eley KL, Martin S, Tuffin MI, Burton SG, Cowan DA (2009) Thermophilic ethanologenesis: future prospects for second-generation bioethanol production. Trends Biotechnol 27:398–405. doi:10.1016/j.tibtech.2009.03.006
Shaw AJ, Podkaminer KK, Desai SG, Bardsley JS, Rogers SR, Thorne PG, Hogsett DA, Lynd LR (2008) Metabolic engineering of a thermophilic bacterium to produce ethanol at high yield. Proc Natl Acad Sci U S A 105:13769–13774. doi:10.1073/pnas.0801266105
Olson DG, McBride JE, Shaw AJ, Lynd LR (2012) Recent progress in consolidated bioprocessing. Curr Opin Biotechnol 23:396–405. doi:10.1016/j.copbio.2011.11.026
Blumer-Schuette SE, Kataeva I, Westpheling J, Adams MW, Kelly RM (2008) Extremely thermophilic microorganisms for biomass conversion: status and prospects. Curr Opin Biotechnol 19:210–217. doi:10.1016/j.copbio.2008.04.007
Yang SJ, Kataeva I, Hamilton-Brehm SD, Engle NL, Tschaplinski TJ, Doeppke C, Davis M, Westpheling J, Adams MWW (2009) Efficient degradation of lignocellulosic plant biomass, without pretreatment, by the thermophilic anaerobe Anaerocellum thermophilum DSM 6725. Appl Environ Microbiol 75:4762–4769
Cha M, Chung D, Elkins JG, Guss AM, Westpheling J (2013) Metabolic engineering of Caldicellulosiruptor bescii yields increased hydrogen production from lignocellulosic biomass. Biotechnol Biofuels 6:85. doi:10.1186/1754-6834-6-85
Svetlitchnyi VA, Kensch O, Falkenhan DA, Korseska SG, Lippert N, Prinz M, Sassi J, Schickor A, Curvers S (2013) Single-step ethanol production from lignocellulose using novel extremely thermophilic bacteria. Biotechnol Biofuels 6:31. doi:10.1186/1754-6834-6-31
Yao S, Mikkelsen MJ (2010) Metabolic engineering to improve ethanol production in Thermoanaerobacter mathranii. Appl Microbiol Biotechnol 88:199–208. doi:10.1007/s00253-010-2703-3
Peacock JP, Cole JK, Murugapiran SK, Dodsworth JA, Fisher JC, Moser DP, Hedlund BP (2013) Pyrosequencing reveals high-temperature cellulolytic microbial consortia in Great Boiling Spring after in situ lignocellulose enrichment. PLoS One 8:e59927. doi:10.1371/journal.pone.0059927
Hugenholtz P, Pitulle C, Hershberger KL, Pace NR (1998) Novel division level bacterial diversity in a Yellowstone hot spring. J Bacteriol 180:366–376
Kashefi K, Holmes DE, Reysenbach AL, Lovley DR (2002) Use of Fe(III) as an electron acceptor to recover previously uncultured hyperthermophiles: isolation and characterization of Geothermobacterium ferrireducens gen. nov., sp. nov. Appl Environ Microbiol 68:1735–1742
Dunfield PF, Tamas I, Lee KC, Morgan XC, McDonald IR, Stott MB (2012) Electing a candidate: a speculative history of the bacterial phylum OP10. Environ Microbiol 14:3069–3080. doi:10.1111/j.1462-2920.2012.02742.x
Hamilton-Brehm SD, Gibson RA, Green SJ, Hopmans EC, Schouten S, van der Meer MT, Shields JP, Damste JS, Elkins JG (2013) Thermodesulfobacterium geofontis sp. nov., a hyperthermophilic, sulfate-reducing bacterium isolated from Obsidian Pool, Yellowstone National Park. Extremophiles 17:251–263. doi:10.1007/s00792-013-0512-1
Hamilton-Brehm SD, Mosher JJ, Vishnivetskaya T, Podar M, Carroll S, Allman S, Phelps TJ, Keller M, Elkins JG (2010) Caldicellulosiruptor obsidiansis sp. nov., an anaerobic, extremely thermophilic, cellulolytic bacterium isolated from Obsidian Pool, Yellowstone National Park. Appl Environ Microbiol 76:1014–1020. doi:10.1128/aem.01903-09
Barns SM, Fundyga RE, Jeffries MW, Pace NR (1994) Remarkable archaeal diversity detected in a Yellowstone National Park hot spring environment. Proc Natl Acad Sci U S A 91:1609–1613
Elkins JG, Podar M, Graham DE, Makarova KS, Wolf Y, Randau L, Hedlund BP, Brochier-Armanet C, Kunin V, Anderson I, Lapidus A, Goltsman E, Barry K, Koonin EV, Hugenholtz P, Kyrpides N, Wanner G, Richardson P, Keller M, Stetter KO (2008) A korarchaeal genome reveals insights into the evolution of the Archaea. Proc Natl Acad Sci U S A 105:8102–8107. doi:10.1073/pnas.0801980105
Podar M, Makarova KS, Graham DE, Wolf YI, Koonin EV, Reysenbach AL (2013) Insights into archaeal evolution and symbiosis from the genomes of a nanoarchaeon and its inferred crenarchaeal host from Obsidian Pool, Yellowstone National Park. Biol Direct 8:9. doi:10.1186/1745-6150-8-9
Vishnivetskaya TA, Raman B, Phelps TJ, Podar M, Elkins JG (2012) Cellulolytic microorganisms from thermal environments. In: Anitori RP (ed) Extremophiles: microbiology and biotechnology. Caister Academic Press, Norfolk, pp 131–158
Wang ZW, Hamilton-Brehm SD, Lochner A, Elkins JG, Morrell-Falvey JL (2011) Mathematical modeling of hydrolysate diffusion and utilization in cellulolytic biofilms of the extreme thermophile Caldicellulosiruptor obsidiansis. Bioresour Technol 102:3155–3162. doi:10.1016/j.biortech.2010.10.104
Lochner A, Giannone RJ, Keller M, Antranikian G, Graham DE, Hettich RL (2011) Label-free quantitative proteomics for the extremely thermophilic bacterium Caldicellulosiruptor obsidiansis reveal distinct abundance patterns upon growth on cellobiose, crystalline cellulose, and switchgrass. J Proteome Res 10:5302–5314. doi:10.1021/pr200536j
Lochner A, Giannone RJ, Rodriguez M Jr, Shah MB, Mielenz JR, Keller M, Antranikian G, Graham DE, Hettich RL (2011) Use of label-free quantitative proteomics to distinguish the secreted cellulolytic systems of Caldicellulosiruptor bescii and Caldicellulosiruptor obsidiansis. Appl Environ Microbiol 77:4042–4054. doi:10.1128/AEM.02811-10
Blumer-Schuette SE, Giannone RJ, Zurawski JV, Ozdemir I, Ma Q, Yin Y, Xu Y, Kataeva I, Poole FL 2nd, Adams MW, Hamilton-Brehm SD, Elkins JG, Larimer FW, Land ML, Hauser LJ, Cottingham RW, Hettich RL, Kelly RM (2012) Caldicellulosiruptor core and pangenomes reveal determinants for noncellulosomal thermophilic deconstruction of plant biomass. J Bacteriol 194:4015–4028. doi:10.1128/JB.00266-12
Spear JR, Walker JJ, McCollom TM, Pace NR (2005) Hydrogen and bioenergetics in the Yellowstone geothermal ecosystem. Proc Natl Acad Sci U S A 102:2555–2560. doi:10.1073/pnas.0409574102
Shock EL, Holland M, Meyer-Dombard DAR, Amend JP (2005) Geochemical sources of energy for microbial metabolism in hydrothermal ecosystems: Obsidian Pool, Yellowstone National Park, USA. In: Inskeep W, McDermott T (eds) Geothermal biology and geochemistry in Yellowstone National Park. Thermal Biology Institute, Montana State University, Bozeman, pp 95–112
Miller TL, Wolin MJ (1974) A serum bottle modification of the Hungate technique for cultivating obligate anaerobes. Appl Microbiol 27:985–987
Raes J, Korbel JO, Lercher MJ, von Mering C, Bork P (2007) Prediction of effective genome size in metagenomic samples. Genome Biol 8:R10. doi:10.1186/gb-2007-8-1-r10
Angly FE, Willner D, Prieto-Davà A, Edwards RA, Schmieder R, Vega-Thurber R, Antonopoulos DA, Barott K, Cottrell MT, Desnues C, Dinsdale EA, Furlan M, Haynes M, Henn MR, Hu Y, Kirchman DL, McDole T, McPherson JD, Meyer F, Miller RM, Mundt E, Naviaux RK, Rodriguez-Mueller B, Stevens R, Wegley L, Zhang L, Zhu B, Rohwer F (2009) The GAAS metagenomic tool and its estimations of viral and microbial average genome size in four major biomes. PLoS Comput Biol 5:e1000593
Ellenbroek FM, Cappenberg TE (1991) DNA-synthesis and tritiated-thymidine incorporation by heterotrophic fresh-water bacteria in continuous culture. Appl Environ Microbiol 57:1675–1682
Mackelprang R, Waldrop MP, DeAngelis KM, David MM, Chavarria KL, Blazewicz SJ, Rubin EM, Jansson JK (2011) Metagenomic analysis of a permafrost microbial community reveals a rapid response to thaw. Nature 480:368–371
Porat I, Vishnivetskaya TA, Mosher JJ, Brandt CC, Yang ZK, Brooks SC, Liang L, Drake MM, Podar M, Brown SD, Palumbo AV (2009) Characterization of archaeal community in contaminated and uncontaminated surface stream sediments. Microb Ecol 60:784–795. doi:10.1007/s00248-010-9734-2
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarrell DM, Marsh T, Garrity GM, Tiedje JM (2009) The ribosomal database project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37:D141–D145. doi:10.1093/nar/gkn879
Haas BJ, Gevers D, Earl AM, Feldgarden M, Ward DV, Giannoukos G, Ciulla D, Tabbaa D, Highlander SK, Sodergren E, Methe B, DeSantis TZ, Petrosino JF, Knight R, Birren BW (2011) Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res 21:494–504. doi:10.1101/gr.112730.110
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Claesson MJ, O’Sullivan O, Wang Q, Nikkila J, Marchesi JR, Smidt H, de Vos WM, Ross RP, O’Toole PW (2009) Comparative analysis of pyrosequencing and a phylogenetic microarray for exploring microbial community structures in the human distal intestine. PLoS One 4:e6669. doi:10.1371/journal.pone.0006669
Meyer F, Paarmann D, D’Souza M, Olson R, Glass EM, Kubal M, Paczian T, Rodriguez A, Stevens R, Wilke A, Wilkening J, Edwards RA (2008) The metagenomics RAST server - a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9. doi:10.1186/1471-2105-9-386
Parks DH, Beiko RG (2010) Identifying biologically relevant differences between metagenomic communities. Bioinformatics 26:715–721. doi:10.1093/bioinformatics/btq041
Needleman SB, Wunsch CD (1970) A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol 48:443–453
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. doi:10.1128/aem.01541-09
Lozupone C, Hamady M, Knight R (2006) UniFrac - an online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinforma 7:371–385. doi:10.1186/1471-2105-7-371
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235. doi:10.1128/aem.71.12.8228-8235.2005
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Fournier RO (1989) Geochemistry and dynamics of the Yellowstone National Park hydrothermal system. Annu Rev Earth Planet Sci 17:13–53. doi:10.1146/annurev.earth.17.1.13
Kennedy BM, Lynch MA, Reynolds JH, Smith SP (1985) Intensive sampling of noble-gases in fluids at Yellowstone. 1. Early overview of the data - regional patterns. Geochim Cosmochim Acta 49:1251–1261. doi:10.1016/0016-7037(85)90014-6
White DE (1981) Active geothermal systems and hydrothermal ore deposits. Economic geology 75th anniversary volume
Meyer-Dombard DR, Shock EL, Amend JP (2005) Archaeal and bacterial communities in geochemically diverse hot springs of Yellowstone National Park, USA. Geobiology 3:211–227. doi:10.1111/j.1472-4669.2005.00052.x
Cole JK, Peacock JP, Dodsworth JA, Williams AJ, Thompson DB, Dong HL, Wu G, Hedlund BP (2013) Sediment microbial communities in Great Boiling Spring are controlled by temperature and distinct from water communities. ISME J 7:718–729. doi:10.1038/ismej.2012.157
Stout RG, Al-Niemi TS (2002) Heat-tolerant flowering plants of active geothermal areas in Yellowstone National Park. Ann Bot 90:259–267
Ogg CD, Patel BKC (2009) Caloramator australicus sp nov., a thermophilic, anaerobic bacterium from the Great Artesian Basin of Australia. Int J Syst Evol Microbiol 59:95–101. doi:10.1099/ijs.0.000802-0
Orlygsson J, Sigurbjornsdottir MA, Bakken HE (2010) Bioprospecting thermophilic ethanol and hydrogen producing bacteria from hot springs in Iceland. Icel Agric Sci 23:73–85
Vanfossen AL, Verhaart MR, Kengen SM, Kelly RM (2009) Carbohydrate utilization patterns for the extremely thermophilic bacterium Caldicellulosiruptor saccharolyticus reveal broad growth substrate preferences. Appl Environ Microbiol 75:7718–7724
Blumer-Schuette SE, Giannone RJ, Zurawski JV, Ozdemir I, Ma Q, Yin YB, Xu Y, Kataeva I, Poole FL, Adams MWW, Hamilton-Brehm SD, Elkins JG, Larimer FW, Land ML, Hauser LJ, Cottingham RW, Hettich RL, Kelly RM (2012) Caldicellulosiruptor core and pangenomes reveal determinants for noncellulosomal thermophilic deconstruction of plant biomass. J Bacteriol 194:4015–4028. doi:10.1128/jb.00266-12
Yang S-J, Kataeva I, Wiegel J, Yin Y, Dam P, Xu Y, Westpheling J, Adams MWW (2009) eclassification of ‘Anaerocellum thermophilum’ as Caldicellulosiruptor bescii strain DSM 6725T sp. nov. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.017731-0, ijs.0.017731-017730
Mathrani IM, Ahring BK (1992) Thermophilic and alkalophilic xylanases from several Dictyoglomus isolates. Appl Microbiol Biotechnol 38:23–27
Mathrani IM, Ahring BK (1991) Isolation and characterization of a strictly xylan-degrading Dictyoglomus from a man-made, thermophilic anaerobic environment. Arch Microbiol 157:13–17
Podosokorskaya OA, Merkel AY, Kolganova TV, Chernyh NA, Miroshnichenko ML, Bonch-Osmolovskaya EA, Kublanov IV (2011) Fervidobacterium reparium sp. nov., a thermophilic anaerobic cellulolytic bacterium isolated from a hot spring. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.026070-0
Acknowledgments
We thank the National Park Service and especially Christie Hendrix for coordinating and allowing sampling under permit #YELL-2008-SCI-5714. Heidi Anderson at the Yellowstone Center for Resources helped with plant species identification. We kindly thank Zamin Koo Yang for the assistance with pyrosequencing and Xiangping Yin for ICP analysis. Christopher W. Schadt provided helpful comments on the manuscript. This work was supported by the BioEnergy Science Center (BESC), which is a U.S. Department of Energy Bioenergy Research Center supported by the Office of Biological and Environmental Research in the DOE Office of Science, Oak Ridge National Laboratory. Oak Ridge National Laboratory is managed by UT-Battelle, LLC, for the U.S. Department of Energy under contract DE-AC05-00OR22725.
Author information
Authors and Affiliations
Corresponding author
Additional information
The submitted manuscript has been authored by a contractor of the U.S. Government under contract DE-AC05-00OR22725. Accordingly, the U.S. Government retains a nonexclusive, royalty-free license to publish or reproduce the published form of this contribution, or allow others to do so, for U.S. Government purposes.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table S1
(DOCX 37 kb)
Supplementary Fig. S1
(DOCX 131 kb)
Supplementary Fig. S2
(DOCX 239 kb)
Supplementary Fig. S3
(DOCX 503 kb)
Supplementary Fig. S4
(DOCX 454 kb)
Supplementary Fig. S5
(DOCX 295 kb)
Rights and permissions
About this article
Cite this article
Vishnivetskaya, T.A., Hamilton-Brehm, S.D., Podar, M. et al. Community Analysis of Plant Biomass-Degrading Microorganisms from Obsidian Pool, Yellowstone National Park. Microb Ecol 69, 333–345 (2015). https://doi.org/10.1007/s00248-014-0500-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-014-0500-8