Abstract
Bacterial resistance to antibiotics has become a public health issue. Over the years, pathogenic organisms with resistance traits have been studied due to the threat they pose to human well-being. However, several studies raised awareness to the often disregarded importance of environmental bacteria as sources of resistance mechanisms. In this work, we analyze the diversity of antibiotic-resistant bacteria occurring in aquatic environments of the state of Rio de Janeiro, Brazil, that are subjected to distinct degrees of anthropogenic impacts. We access the diversity of aquatic bacteria capable of growing in increasing ampicillin concentrations through 16S rRNA gene libraries. This analysis is complemented by the characterization of antibiotic resistance profiles of isolates obtained from urban aquatic environments. We detect communities capable of tolerating antibiotic concentrations up to 600 times higher than the clinical levels. Among the resistant organisms are included potentially pathogenic species, some of them classified as multiresistant. Our results extend the knowledge of the diversity of antibiotic resistance among environmental microorganisms and provide evidence that the diversity of drug-resistant bacteria in aquatic habitats can be influenced by pollution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Antibiotic-resistant bacteria represent a burden to the health of millions of individuals worldwide. This matter has become a public health issue globally [64, 36, 18]. However, this problem is not only of medical concern but also of ecological relevance. Drug resistance is not a trait restricted to pathogenic organisms, but is largely disseminated among environmental bacteria as well [14, 24, 39]. Even so, antibiotic resistance has mostly been investigated under the medical perspective, and the important role of environmental bacteria as sources of antibiotic resistance traits has frequently been disregarded [60]. Resistance genes originated from environmental bacteria can be mobilized into the genomes of pathogenic organisms through horizontal gene transfer [9, 32, 33]. Thus, to fully understand how resistance emerges and propagates, it is necessary to investigate not only medical scenarios but also the ecological and evolutionary processes of resistant microorganisms and their genomes [25, 42, 66].
Antibiotic molecules have been present among Earth’s microbiome long before the use of these substances for medical purposes [8, 10, 27]. Microbial communities from soil and water commonly produce antibiotics, but not necessarily for their use as bactericidal substances. Instead, these compounds are used as intercellular signaling molecules [16, 37]. In this context, antibiotics can be considered fundamental tools for maintenance of community homeostasis, regulating the complex interactions and metabolic pathways that take place within microbial consortia [1, 41, 42]. This hypothesis is supported by evidences that sub-inhibitory concentrations of these compounds can induce significant shifts in bacterial gene expression patterns [28, 37, 67].
Resistance genes also originated long before the use of antibiotics as drugs. Vancomycin and tetracycline resistance genes were shown to be at least 30,000 years old [20], while β-lactamases may have appeared on Earth millions of years ago [65]. These proteins are thought to have originated from genes associated with physiological functions such as detoxification, secretion, and signaling [6, 51]. Upon the selective pressure imposed by the extensive use of antibiotics, they evolved to have drug resistance as their primary function [40].
In the presence of a selective pressure imposed by antibiotics, resistance tends to rapidly spread among bacterial populations [68]. However, in the absence of such drugs, resistant bacteria can present reduced fitness when compared to the antibiotic susceptible strains [3, 26]. This would favor the elimination of resistance genes from bacteria that are not subjected to antibiotics as a relevant selective pressure. However, the presence of these genes in a bacterial genome may carry little to no fitness costs at all [41, 65] or can be associated with other physiological functions [40], which favors the maintenance, and potential dissemination of those features. Therefore, environmental bacteria are often multiresistant and, in some cases, capable, of tolerating extremely high doses these molecules [8, 15, 21]. Some soil microbiomes are capable of subsisting on antibiotics as their sole carbon source [15].
Habitats submitted to varying degrees of anthropogenic impacts were shown to be important sources of resistance genes [69] and resistant bacteria [6]. Environments submitted to very little anthropogenic impacts like deep terrestrial subsurface [8], glaciers [5, 49], and Antarctic waters [17] are rich in such genes, suggesting that their distribution is not restricted to sites subjected to human intervention. Water environments are crucial agents for the mixing and counteracting of resistant organisms originated from highly selective sites (e.g., hospitals and farms), with environmental bacteria, playing a role as reservoirs of resistance genes [6]. A significant body of evidence suggests that pollution promotes the spread of resistance in aquatic sites: Wastewater discharges influence the diversity of resistant microorganisms [13, 56, 59] and of resistance genes [33, 54]. Additionally, hospital effluents were shown to be an important source of resistant bacteria to water habitats [46, 48], a consequence of the extensive use of antimicrobials made in such places [34]. Nevertheless, we do not fully understand the factors controlling the distribution of antibiotic resistance in aquatic ecosystems.
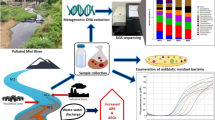
Here, we investigate how pollution affects antibiotic resistance among cultivable bacterial communities from water bodies located in the state of Rio de Janeiro, each subjected to distinct degrees of domestic and industrial sewage pollution (Fig. 1a). We selected communities of environmental bacteria based on their capacity to grow on ampicillin-supplemented Luria–Bertani (LB) medium. This drug is a semi-synthetic beta-lactam antibiotic. Beta-lactams inactivate bacterial transpeptidases, which are proteins responsible for performing cross-links between peptidoglycan strands during cell wall formation. This results in a weak cell wall that is also prone to ruptures, consequently impairing bacterial growth [71]. Beta-lactams are widely used throughout the world for tackling bacterial infections and have a well-established history of resistance. DNA extracted from these cultures was later used for construction of 16S rRNA gene libraries. In addition, we performed isolation of bacteria from the aquatic environments sampled, followed by characterization of the antibiotic resistance profiles of these isolates, so to identify multiresistant and potentially pathogenic organisms.
a Map of sampling sites. Sites located in Ilha grande: Parnaioca River (PR), Parnaioca Beach (PB), and Mangrove system (MS). Sites from Rio de Janeiro city: Barra da Tijuca (BT), Guanabara Bay (GB), and Cunha Channel (CC). The white line delimits the Jacarepaguá lagoon systems, and the stars represent sampling sites within this location. b An overview of our experimental design
Methods
Sampling
Six 50-mL water samples were retrieved from distinct sites throughout Rio de Janeiro state (white dots, Fig. 1a). Three samples were obtained from aquatic environments of Ilha Grande, a relatively preserved island within the Atlantic rain forest biome: freshwater from Parnaioca river (PR) (23°11′21″ S/44°15′11″ W), seawater from Parnaioca beach (PB) (23°11′24″ S/44°15′15″ W), and brackish water from a nearby mangrove system (MS) (23°10′26″ S/44°17′08″W).
In parallel, three samples were collected from impacted sites of Rio de Janeiro city, subjected to distinct levels of wastewater discharges: seawater from Barra da Tijuca beach (BT) (23°00′43″ S/43°21′58″ W), Guanabara Bay (GB) (22°50′18″ S/43°14′00″W), and Cunha Channel (CC) (22°51′14″ S/43°14′13″ W). BT is an urban coastal environment subjected to low impact; this site is used by the local population for recreational purposes. Barra da Tijuca beach is connected to the coastal, mildly impacted, Jacarepaguá lagoons, which discharge its waters in the Atlantic Ocean through Joá channel, which is very close to the BT sampling site. GB is a brackish environment that, despite receiving several sources of wastewater discharges (i.e., domestic, rural, hospital, and industrial) also provides food and leisure to local inhabitants. CC is a highly impacted channel, within Guanabara Bay, that underwent eutrophication due to the massive amounts of organic matter released in its waters [57]. These six samples were used to assess the diversity of ampicillin-resistant bacteria.
Bacterial Liquid Cultures
One milliliter of water from each of the six samples was used to inoculate 50 mL of LB culturing medium, so to promote the growth of aerobic heterotrophic bacteria. Samples were inoculated in three distinct ampicillin concentrations: 0 mg/L (Control), 20 mg/L (clinical concentration), and 1 g/L (super-resistance concentration) [15]. Culturing was performed in aerobic conditions at 37 °C for 24 h. Super-resistance (1 g/L) cultures that showed growth after 24 h were used to inoculate fresh medium supplemented with an even higher concentration of ampicillin: 12 g/L, which were allowed to grow in the same conditions (Fig. 1b).
Construction of 16S rRNA Gene Libraries
Cultures were centrifuged, and pelleted material was recovered. Cell lysis was performed using sodium dodecyl sulfate and proteinase K followed by phenol-chlorophorm DNA extraction as previously described [58]. DNA integrity was checked by 1 % (w/v) agarose gel electrophoresis.
DNA extracted from cultures was used as template for the amplification of the 16S rRNA gene through PCR using universal bacterial primers 27BF (5′-AGAGTTTGATCCTGGCTCAG-3′) [35] and 907RAB (5′-TTTGAGTTTMCTTAACTGCC-3′) [61]. PCR, product purification, cloning, and gene library construction were carried out as previously described [52].
Sequencing Reactions
Sequencing reactions were carried out with forward primer 27BF [35]. ∼400 ng of plasmidial DNA was obtained from each clone and processed with Big Dye terminator v3.1 Cycle sequencing Kit (Applied Biosystems). Products were analyzed in an Applied Biosystems ABI Prism 3730 automated DNA sequencer. Electropherograms were converted to FASTA format through Phred software [23].
Sequence Analysis
Each culture from Ilha Grande (MS, PB, and PR) provided 64 clones for sequencing. For samples from Rio de Janeiro city: control cultures (no antibiotic: BT1, GB1, and CC1) and ampicillin cultures (BTAmp, GBAmp, and CCAmp) provided 96 clones each for sequencing. Half the clones from the antibiotic supplemented libraries were from clinical concentration (20 mg/L) cultures (BT2, GB2, and CC2), and half were originated from super-resistance cultures, 24 of them from 1 g/L cultures (BT3, GB3, and CC3) and 24 clones from 12 g/L cultures (BT4, GB4, and CC4).
Anomalous sequences detected by software Mallard [4] were removed from further analysis. A total of 568 high quality sequences were deposited at GenBank under accession numbers JQ480652–JQ480817 and KC208065–KC208466. Sequences were compared by BLASTn search software [2] against GenBank (http://www.ncbi.nlm.nih.gov/) and EzTaxon (http://www.eztaxon.org/) [11] databases, from which reference sequences were retrieved. Taxonomic affiliation of operational taxonomic units (OTUs) was performed using Ribosomal Database Project (RDP) classifier tool (http://rdp.cme.msu.edu/classifier) [12]. Sequences were aligned through MUSCLE software [22] and grouped as OTUs at 97 % stringency using MOTHUR [47]. Aligned sequences were used to construct a phylogenetic tree through neighbor-joining algorithm using distances calculated by the Kimura-2 method through MEGA 5 [55]. Bootstrap tests were conducted in 1,000 replicates.
UniFrac metric analysis [38] was used to generate unweighted principal coordinate analysis (PCA) based on sequence data retrieved from gene libraries through MOTHUR [47]. UniFrac statistical test was applied to determine if the structure of PCA groups was significantly different. MOTHUR was also used to generate rarefaction curves and to calculate the non-parametric richness estimators Chao1 and ACE (abundance based coverage estimator), and the Shannon–Weaver diversity index (H′).
Bacterial Isolates
To analyze the antibiotic resistance profiles of bacteria discharged in Barra da Tijuca beach, we performed antibiogram tests in isolates from the Jacarepaguá lagoon complex. This ecosystem, located within Rio de Janeiro city is composed of four major brackish lakes: Tijuca, Camorim, Jacarepaguá, and Marapendi (Fig. 1a), connected to the Atlantic Ocean through Joá channel, in Barra da Tijuca (BT) beach. Twenty-two isolates were obtained from water samples of the lagoons (indicated by stars in Fig. 1a) using blood agar medium (DIFCO) supplemented with 5 % sheep blood and four types of selective culture media. One milliliter of each sample was inoculated on 10 mL of brain heart infusion (BHI) liquid medium and incubated at 37 °C. After 24 h, cultures were streaked onto BHI, cetrimide agar, MacConkey Agar, and mannitol salt agar plates and then incubated at 37 °C for 24–48 h. This media were selected so to favor the growth of potentially pathogenic bacteria. Organisms were classified through sequencing of their 16S rRNA gene. Isolates were deposited at the CMRVS collection of microorganisms at FIOCRUZ, Rio de Janeiro, Brazil under accession numbers P4370-P4391.
Resistance profiles of the isolates were determined by disc diffusion method in modified Mueller–Hinton agar. Antibiotic discs used were the following: piperacillin/tazobactam (100/10 μg), ticarcillin/clavulanic (75/10 μg), ceftazidime (30 μg), imipenem (10 μg), cefepime (30 μg), meropenem (10 μg), polymyxin B (300 IU), aztreonam (30 μg), gentamicin (10 μg), tobramycin (10 μg), ciprofloxacin (5 μg), and norfloxacin (10 μg). Discs were placed at a distance of 30 mm between each to other, on Mueller–Hinton agar plates previously inoculated with 0.5 McFarland bacterial suspensions. Measurements were taken after 18 h and after 5 days of incubation at 37 °C. Reference strains for antibiotic disc controls were Escherichia coli 00033 (ATCC 25922), Pseudomonas aeruginosa 00099 (ATCC 853), and Staphylococcus aureus INCQS 00015 (25923). Results were interpreted according to the CLSI criteria [44, 50].
Additionally, we obtained five isolates from Guanabara Bay water samples. These bacteria were tested for their ability to resist three antibiotics, i.e., ampicillin, tetracycline, and kanamycin, and then identified through sequencing of their 16S rRNA gene. Following these steps, we sought to obtain plasmids from these organisms to shed light on the molecular mechanisms of antibiotic resistance among these bacteria.
Results
Enrichment Cultures
Water samples were collected from rainforest (PR, PB, and MS) and urban (BT, GB, and CC) environments of Rio de Janeiro (Fig. 1a). These samples were used to inoculate bacterial cultures in four distinct ampicillin concentrations: control (0 mg/L), clinical level (20 mg/L), and super-resistance (1 and 12 g/L) (Fig. 1b). All cultures from urban environments yielded growth after 24 h of culturing at 37 °C, in all ampicillin concentrations. A distinct pattern was observed for the samples from Ilha Grande, for which growth was only observed in ampicillin-free culturing media.
Principal Coordinate Analysis and Beta-Diversity
We used 16S rRNA sequences obtained from the cultures to visualize the degree of similarity between these bacterial communities through UniFrac. Since no resistant bacteria were detected among Ilha Grande samples, this dataset was excluded from this analysis. PCA was carried grouping 16S rRNA gene sequences in two distinct patterns. First, sequences were separated in six groups, according to their original environment and absence (BT1, GB1, and CC1) or presence (BTAmp, GBAmp, and CCAmp) of ampicillin in culturing medium, regardless of concentration (Fig. 2a). Ampicillin-free community GB1 and its ampicillin-supplemented counterpart GBAmp were separated by PC1, which explained 30.1 % of variation. Meanwhile, BT1 and BTAmp groups were separated by PC2, which explained 24.2 % of variation. CC1 and CCAmp were separated by neither PC1 nor PC2. UniFrac test classified differences in the structure of bacterial communities with ampicillin (BTAmp, GBAmp, and CCAmp) or without it (BT1, GB1, and CC1) as significant (p < 0.001).
Principal coordinate analysis. Scatter plots of principal coordinate analysis generated through the UniFrac metrics. a Samples grouped according to source of sample and absence (BT1, GB1, and CC1) or presence (BTAmp, BGAmp, and CCAmp) of ampicillin in culturing medium. b Samples grouped according to source of sample and ampicillin concentration: 0 mg/L (BT1, GB1, and CC1); 20 mg/L (BT2, GB2, and CC2); 1 g/L (BT3, GB3 and CC3); and 12 g/L (BT4, GB4, and CC4)
Sequences were then separated in 12 groups according to original environment and the four distinct levels of ampicillin concentration in culturing medium (Fig. 2b). In this case, PC1 explained 24.0 % of variation while PC2 explained 18.9 %. Sequences from 1 and 12 g/L cultures originated from the same sampling site always clustered together. In addition, all Cunha Channel samples (CC1, CC2, CC3, and CC4) clustered near to each other. The same was not observed for libraries from Barra da Tijuca and Guanabara Bay, which were scattered through the plot.
Guanabara Bay libraries yielded the highest values of Shannon–Weaver diversity index and of the richness estimators ACE and Chao1 at 97 % stringency. Intermediate values were detected for Ilha Grande (IG1) and Cunha Channel samples (CC). The lowest values were detected for Barra da Tijuca libraries, but the diversity of BTAmp was much higher than that of BT1 (Table 1). These results were corroborated by rarefaction curves generated by MOTHUR (Data not show) that indicate GB libraries as the most diverse, IG and CC of intermediate diversity, and BT as the least diverse. In addition, rarefaction curves of BT1 and BTAmp libraries reached a plateau, thus suggesting that these were the only two gene libraries that fully covered species diversity within bacterial cultures.
Taxonomic Assignment
16S rRNA gene sequences obtained from cultures were annotated using the RDP classifier tool. Taxonomic assignment showed that clones from Ilha Grande libraries were affiliated with only two classes: Gammaproteobacteria (80 %) and Bacilli (20 %), which belong to phyla Proteobacteria and Firmicutes, respectively (Fig. 3). Among Gammaproteobacteria, orders Vibrionales, Aeromonadales, and Pseudomonadales were the most representative. Most OTUs from Ilha Grande were assigned to genus Vibrio (Fig. 4). Enterobacteriales and Alteromonadales were also identified among cultures from this habitat, both comprising <4 % of all OTUs.
Taxonomic assignment of 16S rRNA gene sequences from gene libraries. Assignment was performed through the RDP classifier tool. Samples from IG were grouped and analyzed as a single dataset. Samples from the city of Rio de Janeiro were divided according to environment and presence or absence of ampicillin in culturing medium, regardless of concentration
Neighbor-joining phylogenetic tree based on the alignment of sequences from 16S rRNA gene libraries. Distances were calculated by the Kimura-2 method. Reference sequences and their respective accession numbers are showcased in bold. Bootstrap analysis was conducted in 1,000 replicates. Bootstrap values below 50 % are not shown
Barra da Tijuca (BT1 and BTAmp) libraries were dominated by order Vibrionales (73 %) followed by Bacteroidetes (17 %) (Figs. 3 and 4). The majority of OTUs from library BT1 was affiliated with genus Vibrio; a single OTU from this sample was assigned to genus Stenotrophomonas. On the other hand, none of the OTUs, from the culture supplemented with clinical concentrations of ampicillin (BT2), were assigned to order Vibrionales. Instead, clones from library BT2 were mostly represented by order Flavobacteriales, specifically by Chryseobacterium. BT2 was also represented by orders Burkholderiales, Pseudomonadales and Enterobacteriales. All clones from libraries BT3 and BT4 (super-resistance cultures) were affiliated to genus Vibrio.
The most representative classes from GB libraries were Gammaproteobacteria (51 %), Clostridia (34 %), and Flavobacteria (13 %) (Fig. 3). Clones from library GB1 were mostly affiliated to Clostridiales and Vibrionales. The diversity of GB3 was divided among orders Vibrionales, Aeromonadales, Enterobacteriales, Flavobacteriales, and Clostridiales. Clones from libraries GB3 and GB4 (super-resistance concentrations) were exclusively associated to orders Aeromonadales, Xanthomonadales, Pseudomonadales, and Flavobacteriales.
The most representative classes among CC samples were Gammaproteobacteria (73 %), Bacteroidia (15 %), and Clostrida (8 %) (Figs. 3 and 4). Most of the diversity of CC2 was distributed among orders Clostridia and Bacteroidia. Meanwhile, clones from samples CC3 and CC4 were distributed in several groups, but Enterobacteriales was the most representative in both of them. Unlike the other 16S libraries, most orders present in CC samples were retrieved from all ampicillin concentrations, with the exception Pseudomonadales, Campylobacteriales, and Burkholderiales, which were exclusive from ampicillin-supplemented cultures. Nevertheless, those were underrepresented groups, all harboring <3 OTUs each.
Several OTUs from the ampicillin-supplemented cultures were closely related to sequences of potentially pathogenic organisms (e.g., Vibrio cholerae and Klebsiella pneumoniae) or to genera which include pathogenic species (e.g., Bacillus, Pseudomonas, and Clostridium). The impacted habitats GB and CC were particularly rich in such taxa and also presented several OTUs closely related to groups, which are characteristic of mammalian gut (e.g., Bacteroidales and Enterobacteriales).
Antibiotic Resistance Profiles of Isolates from Jacarepaguá Lagoons and Guanabara Bay
We obtained 22 isolates from the Jacarepaguá lagoon system and determined their antibiotic resistance profiles through the disc diffusion method (Table 2). Isolates identified as Enterococcus gallinarum, P. aeruginosa, and Vibrio fluvialis were classified as multiresistant (i.e., presented resistance to more than three different classes of antibiotics). Twelve of the 22 isolates were resistant to at least one of the antibiotics tested. Among these antibiotic-resistant bacteria, some are known pathogens, like P. aeruginosa, Shigella sp., and V. cholerae, which can be responsible for severe infections in humans.
In addition, we obtained five isolates from Guanabara Bay (GB). Two were identified as K. pneumoniae resistant to ampicillin, tetracycline, and kanamycin. The three other isolates were resistant to ampicillin only. Two of them were classified as Aeromonas sp. and the remaining one as Acinetobacter calcoaceticus. We obtained a high molecular weight plasmid from K. penumoniae, which granted ampicillin resistance upon elctrotransformation into competent DH10B E. coli cells.
Discussion
Antibiotic therapy leads to drastic shifts in the species composition of the human-associated microbiome, which includes decreases in bacterial richness and diversity [19]. During this process, resistant bacteria are positively selected, showing higher abundance following antibiotic treatment [31]. As the use of antibiotics increases, so does the selective pressure imposed upon bacterial communities. In the environment, bacteria may exchange genes through horizontal gene transfer, one of the main mechanisms by which resistance traits can spread [69]. Aquatic habitats that receive wastewater discharges are hotspots for horizontal gene transfer, in such sites distinct bacterial communities from several sources come together creating an ideal environment for gene exchange [27, 39, 45].
Previous studies that analyzed our sampling sites revealed remarkable differences in the physical and chemical properties of these habitats. The levels of variables such as total phosphorus and inorganic nitrogen reported for Guanabara Bay are much higher than those measured at Ilha Grande sites [52, 57]. Meanwhile, Cunha Channel showed even higher concentrations of these compounds and was also characterized by very low levels of dissolved oxygen [57]. These studies also reported on the diversity of microorganisms that dwells in these habitats through culturing independent approaches.
Few species were found in common between the libraries presented here and those previously reported for these habitats. This result was expected since the culture independent analysis will cover a distinct portion of the diversity than that assessed by our culturing approach. In agreement with our findings, the diversity of microorganisms in these sites was shown to be influenced by the degree of pollution of these habitats [52, 57, 70]. Taxa of Cyanobacteria, Actinobacteria, and Alphaproteobacteria were absent from our samples, even though they were common in the 16S gene libraries reported for the previous studies, a possible consequence of the culturing method employed here. Meanwhile, many of the potentially pathogenic organisms reported in the present study were not detected by previous analyzes nor was their ability to resist antibiotics assessed.
Here, we show that ampicillin-resistant bacteria are widespread in impacted aquatic environments of Rio de Janeiro. Resistant bacteria were not detected among environments from Ilha Grande. On the other hand, these organisms were successfully cultured in all ampicillin concentrations from all impacted environments within the city of Rio de Janeiro. These results show that these urban environments are an important source of antibiotic-resistant bacteria. It is likely that resistant organisms are present in the environments of the island but could not be detected by our culturing methodology and class of antibiotic tested.
As expected, some of the organisms that grew in the antibiotic-supplemented cultures are intrinsically resistant to ampicillin and to other beta-lactam antibiotics (e.g., Pseudomonas sp., Acinetobacter sp., and Klebsiella sp. [78]). Some of them were retrieved from the control cultures (no antibiotic) but not from those supplemented with ampicillin. This can occur if, by chance, these organisms were absent from the 1-mL aliquots used to inoculate the cultures, or if they were present in very low concentrations and were incapable of growing to detectable levels during the 24 hours culturing. In addition, these organisms may not have had time to adapt to the antibiotic supplemented cultures. Since there is a trade-off between bacterial fitness and antibiotic resistance, these organisms fine tune the expression of their resistance mechanisms [79]. Unless these organisms could adapt fast enough to the culturing medium supplemented with antibiotics ranging from clinical concentrations up to doses 600 times higher, they would be incapable of growing, despite being intrinsically resistant to these drugs.
Among our urban sampling sites, Barra da Tijuca beach is the least impacted. PCA (Fig. 2) and phylogenetic tree (Fig. 3) revealed that the species composition of cultures with ampicillin (BTAmp) and without it (BT1) is clearly distinct. In addition, rarefaction curves and diversity indexes indicate that the species diversity of BTAmp is much higher than that of its antibiotic free counterpart BT1. Together, these results show that the presence of ampicillin in the culturing medium caused a clear shift in the community composition of cultures from this habitat.
In opposition to what was observed for BT samples, the extremely polluted Cunha Channel showed very similar communities in CC1 and CCAmp libraries (Figs. 2 and 3). Antibiotic resistance is probably extensively disseminated in this highly impacted environment, as most taxa that grew in antibiotic-free medium were also detected in ampicillin-supplemented cultures. This may be consequence of high amounts of untreated sewage carrying resistant strains released in this site. Hence, the presence of ampicillin produces little impact in the species composition of cultures of organisms retrieved from CC.
Diversity indexes indicated that cultures inoculated with Guanabara Bay water samples are the most diverse. This is probably a consequence of the intense mixing of water masses that occurs within the bay [29], which favors a more diverse community. This brackish environment receives seawater from the Atlantic Ocean and freshwater from several rivers of its surroundings. Additionally, wastewaters from urban, rural, and industrial sources are constantly released into the Bay, often without any treatment prior to release. These characteristics contribute to the increased bacterial diversity of this site [57]. Such diversity may contribute for the widely disseminated antibiotic resistance detected in this habitat. A metagenomic study revealed that several drug resistance genes can be found at Guanabara Bay, including those that encode tolerance to beta-lactams, erythromycin, methicillin, vancomycin, and also genes encoding multidrug resistance efflux pumps [29]. Together with our results, this information indicates that GB is an important source of resistance genes, plasmids encoding antibiotic resistance, and also of multi- and super-resistant bacteria, including pathogenic species.
Orders Alteromonadales and Bacillales were detected only in ampicillin-free samples, which could mean that these groups have fewer members capable of resisting this antibiotic in the habitats sampled. Meanwhile, orders Burkholderiales, Flavobacteirales, and Campylobacterales were detected exclusively in the ampicllin-supplemented libraries (Fig. 3). These organisms probably were present in the control cultures, but in lower abundances, therefore could not be detected due to our limited number of sequences. Vibrionales was the only order to be detected in cultures from all the environments in all ampicillin concentrations, suggesting that members of this group are an important reservoir of antibiotic resistance mechanisms in the environments studied.
Bacteroidetes, Firmicutes, and Enterobacteria are indicators of fecal contamination [19, 31]. These groups were scarce among Barra da Tijuca and Ilha Grande libraries. On the other hand, such taxa were well represented in Guanabara Bay and Cunha Channel libraries, in which several clones were affiliated with bacteria from mammalian gut and in some cases human gut specifically (Fig. 4). Human associated bacteria were shown to be an important source of antibiotic resistance features [53]. OTUs from libraries GB and CC were frequently classified as pathogenic organisms. In addition, some of the bacteria isolated from Jacarepaguá lagoon system and Guanabara bay were classified as multiresistant opportunistic pathogens. Free-living bacteria are constantly inoculated into the human organism, through direct contact with these sites or indirectly through food, air, and water [69]. As the urban environments BT, GB, and CC and the Jacarepaguá lagoon system are sources of food and leisure for the local inhabitants, the presence of antibiotic-resistant bacteria in these sites poses a serious threat to the well-being of the local population.
Several OTUs were shared between samples from Ilha Grande (PB, PR, and MS) and the sewage contaminated sites from the city of Rio de Janeiro (BT, GB, and CC) sites, even though the bacteria retrieved from Ilha Grande were not capable of growing in the presence of ampicillin (Fig. 4). In addition, strains isolated from the Jacarepaguá lagoon system that belong to the same species presented distinct antibiotic susceptibility profiles (e.g., P.aeruginosa and V. cholerae, Table 2). Organisms that belong to the same species share the same core genome; therefore, their differences in tolerance to antibiotics is probably associated with the presence of mobile genetic elements, which are considered important spreaders of antibiotic resistance genes in the environment [7]. The plasmid obtained from K. pneumoniae is an example of these elements, and the occurrence of it is evidence that plasmids play a central role in disseminating antibiotic resistance in the sampled habitats.
The isolates identified as V. cholerae, obtained from the Jacarepaguá Lagoon systems, showed remarkable antibiotic resistance profiles. This waterborne pathogen can become resistant through several genetic mechanisms, such as integrons, conjugative plasmids, and spontaneous mutations [72]. The genomic flexibility of this organism is reflected in the resistance profiles of our isolates that, despite being classified as the same species, responded differently to seven antibiotics. In addition, the resistance profiles of our V. cholerae isolates are distinct from the ones previously reported for several clinical and environmental strains of V. cholerae from throughout the world [72]. The resistance profile of our V. fluvialis isolate was different from that which was previously reported for other isolates of this species [73]. Isolates identified as P. aeruginosa were sensible to all antibiotics tested. The susceptibility profiles of this environmental isolates is remarkable as many clinical strains of P. aeruginosa are capable of resisting several classes of antibiotics [74]. Meanwhile, all three isolates of P. pseudoalcaligenes were resistant to Aztreonam, one of them was also resistant to cefepime and another to ticarcillin/clavulanic acid. Some strains of this species, which is rarely pathogenic, are resistant to several antibiotics [75–77], but to our knowledge, there are no reports of it being resistant to aztreonam. Variation in the antibiotic susceptibility profiles was also evident for isolates identified as Exiguobacterium, one isolate was susceptible to all drugs while another was resistant to four different antibiotics. Information on the susceptibility profile of this genus is currently scarce, thus our results help to elucidate the potential role of these (and also of the other) organisms as reservoirs of resistance mechanisms in urban aquatic environments.
In this work, we assess the influence of anthropogenic impacts in the diversity of antibiotic-resistant bacteria from aquatic habitats. We describe bacterial communities capable of tolerating extremely high doses (×600 the clinical concentrations) of ampicillin and also multiresistant bacteria widespread in urban environments, which spam a diverse array of bacterial phyla. Further analyses are necessary to characterize the influence of anthropogenic impacts in the diversity of antibiotic-resistant bacteria; nevertheless, our data provides evidence that there are remarkable differences in the diversity of these organisms among our sampling sites and that resistance and super-resistance to antibiotics is widespread in the urban aquatic environments studied. Public policies aimed at mitigating damages caused by resistant bacteria rely on the necessity of determining which environments are potential sources of resistance traits that pose a threat to human populations [40–42]. This information can help to develop strategies to mitigate the spread of antibiotic resistance, consequently reducing damages to human welfare caused by resistant and multiresistant pathogenic species [43]. Our work, suggests that additionally to making conscious use of antibiotics [10, 30, 62], managing pollution impacts to aquatic environments may be a relevant strategy to achieve that goal, so that the effectiveness of these drugs is preserved, ensuring the benefits they bring to humanity [63].
References
Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J (2010) Call of the wild: antibiotic resistance genes in natural environments. Nat Rev Microbiol 8:251–259
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 5:403–410
Andersson DI, Levin BR (1999) The biological cost of antibiotic resistance. Curr Opin Microbiol 2:489–493
Ashelford KE, Chuzhanova NA, Fry JC, Jones AJ, Weightman AJ (2006) New screening software shows that most recent large 16S rRNA gene clone libraries contain chimeras. Appl Environ Microbiol 72:5734–5741
Ball M, Gómez W, Magallanes X, Rosales R, Melfo A, Yarzábal L.A (2013) Bacteria recovered from high-altitude, tropical glacier in Venezuelan Andes. World J Microbol Biotechnol
Baquero F, Martínez JL, Cantón R (2008) Antibiotic and antibiotic resistance in water environments. Curr Opin Biotechnol 19:260–265
Bennett PM (2008) Plasmid encoded antibiotic resistance: acquisition and transfer of antibiotic resistance genes in bacteria. Br J Pharmacol 153(Suppl 1):S347–S357
Bhullar K, Waglechner N, Pawlowski A, Koteva K, Banks E.D, Johnston M.D. et al. (2012) Antibiotic resistance is prevalent in an isolated cave microbiome. PLoS ONE
Cantón R (2009) Antibiotic resistance genes from the environment: a perspective through newly identified antibiotic resistance mechanisms in the clinical setting. Clin Microbiol Infect 15(Suppl 1):20–25
Carlet J, Jarlier V, Harbarth S, Voss A, Gossens H, Pittet D (2012) Ready for a world without antibiotics? The Pensières antibiotic resistance call to action. Antimicrob Restist Infect Control 14:11–23
Chun J, Lee JH, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261
Cole JR, Wang Q, Crdenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarell DM, Marsh T, Garrity GM, Tiedje JM (2009) The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37:141–145
Czekalski N, Berthold T, Caucci S, Egli A, Bürgmann H (2012) Increased levels of multiresistant bacteria and resistance genes after wastewater treatment and their dissemination into Lake Geneva, Switzerland. Front Microbiol 3:106
D'Costa VM, Griffiths E, Wright GD (2007) Expanding the soil antibiotic resistome: exploring environmental diversity. Curr Opin Microbiol 10:481–489
Dantas G, Sommer MOA, Oluwasegun RD, Church GM (2008) Bacteria subsisting on antibiotics. Science 320:100–103
Davies J (2006) Are antibiotics naturally antibiotics? J Indust Microbiol Biotechnol 33:496–499
De Souza MJ, Nair S, Loka Bharathi PA, Chandramohan D (2006) Metal and antibiotic-resistance in psychrotrophic bacteria from Antarctic Marine waters. Ecotoxicology 15:379–384
Deris JB, Kim M, Zhang Z, Okano H, Hersmen R, Groisman A, Hwa T (2013) The innate growth bistability and fitness landscapes of antibiotic-resistant bacteria. Science 342:12374351–12374358
Dethlefsen, L., Huse, S, Sogin, M.L., Relman, D.A. (2008) The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS Biology.
D’Costa VM, King CE, Kalan L, Morar M, Sung WWL, Schwarz C et al (2011) Antibiotic resistance is ancient. Nature 477:457–461
D’Costa VM, McGrann KM, Hughes DW, Wright GD (2006) Sampling the antibiotic resistome. Science 311:374–377
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Edwing B, Hillier L, Wendl M, Green P (1998) Basecalling of automated sequencer traces using phred. I. Accuracy assessment. Gen Res 8:175–185
Forbserg KJ, Reyes A, Wang B, Selleck EM, Sommer MOA, Dantas G (2012) The shared antibiotic resistome of soil bacteria and human pathogens. Science 337:1107–1111
Gaze WH, Krone SM, Joakim-Larsson DG, Li XZ, Robinson JA, Simonet P et al (2013) Influence of humans on the evolution and mobilization of environmental antibiotic resistome. Emerg Infect Dis 19:1–7
Gifford DR, MacLean RC (2013) Evolutionary reversals of antibiotic resistance in experimental populations of Pseudomonas aeruginosa. Evolution 67:2973–2981
Gillings MR, Stokes HW (2012) Are humans increasing bacterial evolvability? Trends Ecol Evol 27:346–352
Goh E, Yim G, Tsui W, McClure J, Surette MG, Davies J (2002) Transcriptional modulation of bacterial gene expression by subinhibitory concentrations of antibiotics. PNAS 99:17025–17030
Gregoracci G.B, Nascimento J.R, Cabral A.S, Paranhos R, Valentin J.L, Thomspon, C.C, Thompson F.L (2012) Structuring of bacterioplankton diversity in a large tropical bay. PLoS ONE
Höjgård S. Antibiotic resistance—why is the problem so difficult to solve? (2012) Infect Ecol Epidemol.
Jernberg C, Löfmark S, Edlund C, Jansson JK (2007) Long-term ecological impacts of antibiotic administration on the human intestinal microbiota. ISME J 1:56–66
Juhas M (2013) Horizontal gene transfer in human pathogens. Crit Rev in Microbiol
Kristiansson E, Fick J, Janzon A, Grabic R, Rutgersson C, Weijdegår B, Söderström H, Larsson D.G (2011) Pyrosequencing of antibiotic-contaminated river sediments reveals high levels of resistance and gene transfer elements. PLoS ONE
Kümmerer K (2009) Antibiotics in the aquatic environment—a review. Part II. Chemosphere 75:435–441
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematic. Wiley, New York, pp 115–175
Levy SB, Marshall B (2004) Antibacterial resistance worldwide: causes, challenges and responses. Nat Med 10(12 Suppl):122–129
Linares JF, Gustafsson I, Baquero F, Martínez JL (2006) Antibiotics as intermicrobial signaling molecules instead of weapons. PNAS 103:19484–19489
Lozupone C, Hamady M, Knight R (2006) UniFrac - an online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinforma 7:371–384
Lupo A, Coyne S, Berendonk TU (2012) Origin and evolution of antibiotic resistance: the commom mechanisms of emergence and spread in water bodies. Front Microbiol 3:18
Martínez JL (2008) Antibiotics and antibiotic resistance genes in natural environments. Science 321:365–367
Martínez JL (2009) Environmental pollution by antibiotics and antibiotic resistance determinants. Environ Pollut 157:2893–2902
Martínez JL (2012) Natural antibiotic resistance and contamination by antibiotic resistance determinants: the two ages in the evolution of resistance to antimicrobials. Front Microbiol 3:1
Martínez JL, Baquero F, Andersson DI (2007) Predicting antibiotic resistance. Nat Rev Microbiol 5:958–965
Pai V, Rao VI, Rao SP (2010) Prevalence and antimicrobial susceptibility pattern of methicillin-resistant Staphylococcus aureus [MRSA] isolates at a tertiary care hospital in Mangalore, South India. J Lab Phys 2:82–84
Pruden A, Pei R, Storteboom H, Carlson KH (2006) Antibiotic resistance genes as emerging contaminants: studies in Northern Colorado. Environ Sci Technol 40:7745–7750
Santoro DO, Romão CMCA, Clementino MM (2012) Decreased aztreonam susceptibility among Pseudomonas aeruginosa isolates from hospital effluent treatment system and clinical samples. Int J Environ Health Res 22:560–570
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Schwartz T, Kohnen W, Jansen B, Obst U (2003) Detection of antibiotic-resistant bacteria and their resistance genes in wastewater, surface water and drinking water biofilms. FEMS Microbiol Ecol 43:325–335
Segawa T, Takeuchi N, Rivera A, Yamada A, Yoshimura Y, Barcaza G et al (2013) Distribution of antibiotic resistance in glacier environments. Environ Microbiol Rep 5:127–134
Sejas LM, Silbert S, Reis AO, Sader HS (2003) Avaliação da qualidade dos discos com antimicrobianos para testes de disco-difusão disponíveis comercialmente no Brasil. Jornal Brasileiro de Patologia e Medicina Laboratorial 39:27–35
Sengupta S, Chattopadhyay MK, Grossart HP (2013) The multifaceted roles of antibiotics and antibiotic resistance in nature. Front Microbiol 4:47
Silveira CB, Vieira RP, Cardoso AM, Paranhos R, Albano RM, Martins OB (2011) Influence of salinity on bacterioplankton communities from the Brazilian rain forest to the coastal Atlantic Ocean. PloS One
Sommer MO, Dantas G, Church GM (2009) Functional characterizathion of the antibiotic resitance reservoir in the human microflora. Science 325:1128–1131
Tacão M, Correia A, Henriques I (2012) Resistance to broad-spectrum antibiotics in aquatic systems: anthropogenic activities modulate the dissemination of blaCTX-M-like genes. Appl Environ Microbiol 78:4134–4140
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular Evolutionary Genetics Analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thevenon F, Adatte T, Wildi W, Poté J (2012) Antibiotic resistant bacteria/genes dissemination in lacustrine sediments highly increased following cultural eutrophication of Lake Geneva (Switzerland). Chemosphere 86:468–476
Vieira RP, Gonzalez AM, Cardoso AM, Oliveira DN, Albano RM, Clementino MM, Martins OB, Paranhos R (2008) Relationships between bacterial diversity and environmental variables in a tropical marine environment, Rio de Janeiro. Environ Microbiol 10:189–199
Vieira RP, Clementino MM, Cardoso AM, Oliveira DN, Albano RM, Gonzalez AM et al (2007) Archaeal Communities in a Tropical Estuarine Ecosystem: Guanabara Bay, Brazil. Microb Ecol 54:460–462
Vignesh S, Muthukumar K, James RA (2012) (2012) Antibiotic resistant pathogens versus human impacts: a study from three eco-regions of the Chennai coast, southern India. Mar Pollut Bull 64:790–800
Walsh F (2013) Investigating antibiotic resistance in non-clinical environments. Front Microbiol 4:19
Weissburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Wise R (2002) Antimicrobial resistance: priorities for action. J Antimicrob Chemother 49:585–586
World Health Organization (WHO) (2000) WHO Annual report on infectious disease: overcoming antimicrobial resistance. World Health Organization: Geneva, Switzerland, 2000. http://www.who.int/infectious-disease-report/2000/
World Health Organization (WHO) (2012) Fact sheet N°194: antimicrobial resistance. http://www.who.int/mediacentre/factsheets/fs194/en/index.html
Wright GD (2007) The antibiotic resistome: the nexus of chemical genetic diversity. Nat Rev Microbiol 5:175–186
Wright GD (2010) Antibiotic resistance in the environment: a link to the clinic? Curr Opin Microbiol 13:589–594
Yim G, Wang H, Davies J (2007) Antibiotics as signaling molecules. Philos Trans R Soc Lond B Biol Sci 362:1195–1200
Zhang Q, Lambert G, Liao D, Kim H, Robin K, Tung CK et al (2011) Acceleration of emergence of bacterial antibiotic resistance in connected microenvironments. Science 333:1764–1767
Zhang XX, Zhang T, Fang HH (2009) Antibiotic resistance genes in water environment. Appl Microbiol Biotechnol 82:397–414
Salloto GRB, Cardoso AM, Coutinho FH, Pinto LH, Vieira RP, Chaia C, Lima JL et al (2012) Pollution impacts on bacterioplankton diversity in a tropical urban coastal lagoon system. PLoS ONE 7(11):e51175
Singh GB (2004) β-Lactams in the new millennium. Part I: monobactams and carbapenems. Mini Rev Med Chem 4:69–92
Rajpara N, Patel A, Tiwari N, Bahuguna J, Antony A, Choudhury I et al (2009) Int J Antimicrob Agents 34(3):220–225
Ahmed AM, Nakagawa T, Arakawa E, Ramamurthy T, Shinoda S, Shimamoto T (2004) New aminoglycoside acetyltransferase gene, aac(3)-Id, in a class 1 integron from a multiresistant strain of Vibrio fluvialis isolated from an infant aged 6 months. J Antimicrob Chemother 53(6):947–951
Bonomo RA, Szabo D (2006) Mechanisms of multidrug resistance in Acinetobacter species and Pseudomonas aeruginosa. Clin Infect Dis 43:S49–S56
Quinteira S, Ferreira H, Peixe L (2005) First Isolation of blaVIM-2 in an environmental isolate of Pseudomonas pseudoalcaligenes. Antimicrob Agents Chemother 49(5):2140–2141
Tremaroli V, Fedi S, Turner RJ, Ceri H, Zannoni D (2008) Pseudomonas pseudoalcaligenes KF707 upon bioWlm formation on a polystyrene surface acquire a strong antibiotic resistance with minor changes in their tolerance to metal cations and metalloid oxyanions. Arch Microbiol 190(1):29–39
Nazl SA, Tariq P (2005) Prevalence and antibiogram pattern of Pseudomonas species causing secondary infectious among patients of pulomnary tuberculosis. Int Chem Pharm Med J 2(2):231–237
Rice LB (2009) The clinical consequences of antimicrobial resistance. Curr Opin Microbiol 12(5):476–481
Depardieu F, Podglajen I, Leclercq R, Collatz E, Courvalin P (2007) Modes and modulation of antibiotic resistance gene expression. Clin Microbiol Rev 20(1):79–114
Acknowledgments
This study was financed by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Fundação Carlos Chagas Filho de Amparo a Pesquisa do Estado do Rio de Janeiro (FAPERJ). We acknowledge genome studies facilities Johanna Döbereiner. Special thanks to Joyce Lemos Lima for preparing sequencing reactions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Coutinho, F.H., Silveira, C.B., Pinto, L.H. et al. Antibiotic Resistance is Widespread in Urban Aquatic Environments of Rio de Janeiro, Brazil. Microb Ecol 68, 441–452 (2014). https://doi.org/10.1007/s00248-014-0422-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-014-0422-5