Abstract
The effects of water-saving irrigation on emissions of greenhouse gases and soil prokaryotic communities were investigated in an experimental rice field. The water layer was kept at 1–2 cm in the water-saving (WS) irrigation treatment and at 6 cm in the continuous flooding (CF) irrigation treatment. WS irrigation decreased CH4 emissions by 78 % and increased N2O emissions by 533 %, resulting in 78 % reduction of global warming potential compared to the CF irrigation. WS irrigation did not affect the abundance or phylogenetic distribution of bacterial/archaeal 16S rRNA genes and the abundance of bacterial/archaeal 16S rRNAs. The transcript abundance of CH4 emission-related genes generally followed CH4 emission patterns, but the difference in abundance between mcrA transcripts and amoA/pmoA transcripts best described the differences in CH4 emissions between the two irrigation practices. WS irrigation increased the relative abundance of 16S rRNAs and functional gene transcripts associated with Anaeromyxobacter and Methylocystis spp., suggesting that their activities might be important in emissions of the greenhouse gases. The N2O emission patterns were not reflected in the abundance of N2O emission-related genes and transcripts. We showed that the alternative irrigation practice was effective for mitigating greenhouse gas emissions from rice fields and that it did not affect the overall size and structure of the soil prokaryotic community but did affect the activity of some groups.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Methane (CH4) and nitrous oxide (N2O) have been estimated to contribute approximately 14 and 8 %, respectively, of global anthropogenic emissions of greenhouse gases, and agriculture is suggested as a primary source of these gases [1]. Among the major cereal crops, rice has the highest global warming potential (GWP), which is largely driven by CH4 emissions from paddy soils [2]. Strategies for mitigating CH4 emissions from rice fields included alternative practices of irrigation, residue management, and fertilizer management [3–5].
Previous studies have shown that alternative irrigation practices in rice fields effectively reduced CH4 emissions without a significant loss in crop yield [3, 6–8]. Although many of the practices were found to increase N2O emissions, the resulting increase in GWP was estimated to be relatively minor compared to the effect of reduced CH4 emissions on GWP [5, 8, 9]. However, some investigators have observed that the inverse relationship between emissions of CH4 and N2O can be altered depending on soil properties, the application rate of nitrogen fertilizer, or the duration of the drainage period [10, 11].
The shift in the emission patterns of greenhouse gases resulting from alternative irrigation practices was explained by variations in oxidation-reduction (redox) potential and related microbiological activity in the rice paddy soils [9, 12, 13]. Drainage-induced aeration of soil results in regeneration of electron acceptors such as ferric iron and sulfate, and microorganisms that use these substrates can outcompete methanogens, leading to decreased CH4 emission [14, 15].
Molecular approaches, which can overcome limitations of culture-dependent studies, have also been adopted to describe variability in microbial community structure. Watanabe et al. [16] observed that an alternate wetting and drying practice (AWD) had no significant effect on abundance of archaeal 16S rRNA genes or methyl-coenzyme M reductase genes (mcrA genes) but moderately influenced microbial community structure in rice paddy soils. Ma and Lu [17] observed that soil drainage decreased the abundance of archaeal 16S rRNA genes and increased the abundance of particulate methane monooxygenase genes (pmoA genes) in rhizosphere soils and also increased the relative abundance of the type II methanotrophs Methylocystis/Methylosinus in the rhizosphere and surface soils. Ma et al. [18] demonstrated that intermittent drainage had a stronger effect on mcrA transcripts than on mcrA genes in rice paddy soils, suggesting that transcription of a functional gene is a more sensitive descriptor of activity than the abundance of the gene. To our knowledge, molecular approaches have not been used to explain variability in N2O emission patterns induced by alternative irrigation practices.
In the present study, we performed a field experiment for comprehensive examination of changes in prokaryotic communities that resulted from an alternative irrigation practice. We investigated the abundance and phylogenetic distribution of 16S rRNA genes and genes associated with greenhouse gas emissions at both the DNA and RNA levels. Our results demonstrated that the alternative irrigation affected the abundance of transcripts of CH4 but not of N2O emission-related genes. In addition, it was found that the activities of two bacterial genera, Anaeromyxobacter and Methylocystis, were increased under the alternative irrigation. Several new characteristics of bacterial and archaeal communities in relation to emissions of CH4 and N2O are described and discussed.
Materials and Methods
Experimental Site and Water Management
The field experiment was conducted at the experimental rice field of the National Institute of Crop Science, Suwon, South Korea (37°16′N, 126°59′E) in 2012. Two irrigation practices, continuous flooding (CF) and water-saving (WS), were conducted and each treatment consisted of three replicate plots (4 m × 4 m each) connected in series. The water level was maintained at 6 cm in CF plots and at 1–2 cm in WS plots using an automatic water level controller (Joeun Eng., Seoul, South Korea). Seedlings of Samgwang rice (Oryza sativa L. var. japonica) were transplanted on May 22 and harvested on October 10. Urea, fused phosphate, and potassium chloride fertilizers were applied at respective rates of 110 kg N ha−1, 30 kg P2O5 ha−1, and 30 kg K2O ha−1.
Gas Flux Measurements
CH4 and N2O fluxes were measured twice per week using the static chamber method. A rectangular polyacrylic chamber (0.6 m wide × 0.6 m long × 1 m high) was placed in each plot, and gas samples were taken with a syringe between 10:00 AM and 1:00 PM. The concentrations of CH4 and N2O were determined using a gas chromatograph (Varian 3800; Varian, Palo Alto, USA) with the working conditions presented in Table S1. Gas fluxes were calculated from the change in gas concentrations inside the chamber over 30 min, using the equation suggested by Hou et al. [12], and GWP was calculated by multiplying the cumulative emissions of CH4 and N2O by 21 and 310, respectively [19].
Soil Sampling and Chemical Characterization
Soil samples for microbial characterization were collected on May 29, July 9, August 2, September 6, and October 8. Soil cores (1.8-cm diameter, 15-cm height) were collected from the center and four corners of each plot, mixed thoroughly, and stored at −70 °C until molecular analyses. When the field experiment had terminated, an additional set of soil samples was collected and the following analyses were conducted: pH and electrical conductivity (1:5 water extraction); organic matter [20]; total nitrogen (Kjeldahl method); available P2O5 (Lancaster method) [21]; and the contents of exchangeable Ca2+, Mg2+, and K+ (1 M NH4-acetate, pH 7.0, ICP-AES; GBC Scientific Equipment, Melbourne, Australia).
Nucleic Acid Extraction and cDNA Preparation
Soil RNA and DNA were extracted using the RNA PowerSoil Total RNA Isolation Kit and the RNA PowerSoil DNA Elution Accessory Kit (MO BIO Laboratories, Carlsbad, USA), respectively, according to the manufacturer’s instructions. Aliquots of RNA extracts were treated by RQ1 RNase-Free DNase (Promega, Madison, USA) and purified by using the RNeasy MinElute Cleanup kit (Qiagen, Hilden, Germany). cDNA was synthesized using the GoScript reverse transcription system and random hexamers (Promega, Madison, USA) according to the manufacturer’s instructions.
Quantification by Real-Time PCR
The following genes and their transcripts were quantified using the CFX96 system (Bio-Rad, Hercules, USA): bacterial and archaeal 16S rRNA genes, mcrA genes, ammonia monooxygenase and particulate methane monooxygenase genes (amoA/pmoA genes), quinol-oxidizing nitric oxide reductase genes (qnorB genes), cytochrome-oxidizing nitric oxide reductases genes (cnorB genes), and nitrous oxide reductase genes (nosZ genes). The mcrA genes and amoA/pmoA genes were selected for monitoring CH4 production and consumption, respectively, and the qnorB/cnorB genes and nosZ genes were selected for monitoring N2O production and consumption, respectively. The primers and PCR conditions are presented in Table S2. The primers to quantify the functional genes were designed based on reference sequences obtained from the KEGG database (http://www.genome.jp/kegg/). The reaction solution contained 10 μl of iQ SYBR Green Supermix (2x, Bio-Rad, Hercules, USA), 0.4 μM of each primer, 1 mg/ml of BSA, and 1 μl of DNA (10:1 dilution) or cDNA, in a total volume of 20 μl. Linearized plasmids containing the corresponding genes were used as standards for all assays. Efficiencies for PCR reactions were between 64 and 85 % and standard curves had r 2 > 0.98.
Pyrosequencing
The phylogenetic diversities of the following genes and their transcripts were examined using pyrosequencing analysis: bacterial and archaeal 16S rRNA gene, mcrA, amoA/pmoA, qnorB, and nosZ. Only soil samples collected on August 2 were used in this analysis, when the emissions of CH4 and N2O were at their maximum levels among the soil sampling dates (see text). For 16S rRNA sequences, soil samples from the three replicate plots were analyzed; for the other genes and their transcripts, a soil sample from the middle one of the three replicate plots was analyzed. The 50-μl reaction mixture contained 1× Phusion HF buffer, 0.2 mM of each of the deoxynucleoside triphosphates (dNTPs), 0.4 μM of each primer, 1 mg/ml of BSA, 2.5 μl DMSO, 1 U of Phusion High-Fidelity DNA polymerase (New England Biolabs, Herts, UK), and 1 μl of DNA (10:1 dilution) or cDNA. The primers and PCR conditions are presented in Table S3. The PCR products were gel-purified with the QIAquick Gel extraction kit (Qiagen, Hilden, Germany); pyrosequencing of the PCR products was performed by the National Instrumentation Center for Environmental Management (Seoul, South Korea) using a 454 GS FLX Titanium Sequencing System (Roche, Mannheim, Germany), according to the manufacturer’s instructions.
Processing of Pyrosequencing Data
All pyrosequencing data were processed using the Mothur software package (version 1.27.0) [22] if not mentioned otherwise. Briefly, prefiltered flowgrams of the pyrosequencing reads were clustered using the PyroNoise algorithm [23], and chimeric sequences were removed using UCHIME [24]. The number of pyrosequencing reads was normalized among the samples for each of the genes. For bacterial and archaeal 16S rRNA sequences, the qualified sequences were classified to the genus level using the Bayesian method based on the Ribosomal Database Project (RDP) taxonomy (version 9) [25] and the SILVA taxonomy (version 111) [26], respectively. The sequences were aligned using the SINA aligner (http://www.arb-silva.de/aligner) [27] and then clustered into operational taxonomic units (OTUs) at 97 % similarity cutoff using the average-neighbor algorithm. Richness estimators and diversity indices were calculated using the summary.seqs command in the Mothur package. Weighted Fast UniFrac analysis [28] was performed using a phylogenetic tree constructed with the FastTree program (http://meta.microbesonline.org/fasttree) [29]. For functional genes and their transcripts, the qualified sequences were subjected to the RDP FrameBot for frameshift corrections (http://fungene.cme.msu.edu). The frameshift-corrected sequences were translated to protein sequences and aligned using the Clustal Omega program [30] with the pre-aligned sequences provided by the FunGene database as the reference in the Mobyle Portal (http://mobyle.pasteur.fr). The aligned sequences were imported into the ARB software package (version 5.5) [31] and the distance matrixes were calculated using the ARB neighbor-joining method. The numbers of the unique sequences were calculated using the average-neighbor algorithm based on the distance matrix.
Phylogenetic Analysis
For 16S rRNA sequences, predominant OTUs and closely related sequences obtained from GenBank were aligned as mentioned above. The aligned sequences were imported into the ARB software package and almost full-length sequences (≥1,300 bp) were used for construction of the initial tree via the maximum-likelihood algorithm (Phylip). Shorter sequences (<1,300 bp) were added to this tree using the ARB parsimony tool, which allows the addition of short sequences to phylogenetic trees without changing the global tree topology [32]. The relative abundances of dominant OTUs in each sample were represented as a heatmap with the iTOL tool (http://itol.embl.de) [33]. For the other genes and their transcripts, the maximum-likelihood trees were constructed from the aligned protein sequences using the FastTree program.
Statistical Analysis
The differences in the gas emission, richness estimator, diversity index, and the gene/transcript abundance between CF and WS plots were analyzed with Student’s t tests. Differences in the abundance of taxa were determined with one-way analysis of variance (ANOVA) followed by Tukey’s pairwise comparison. p values < 0.05 were considered significant. All analyses were performed using OriginPro 8 SR4 software (OriginLab Corp., Northampton, USA).
DNA Sequence Data
The raw pyrosequencing data are available in the NCBI Sequence Read Archive under the accession numbers SRX318152-SRX318157 and SRX326402-SRX326407.
Results
Soil Characteristics and Crop Yield
The chemical properties of the rice paddy soils are presented in Table 1. No noticeable differences were observed between the two irrigation practices with the exception of available P2O5, which was about two times higher in CF than in WS plots. Grain yield of rice was 5.83 ton ha−1 in CF plots and 6.25 ton ha−1 in WS plots.
CH4 and N2O Emissions
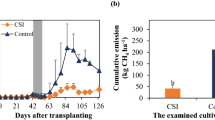
The two irrigation practices produced dramatic differences in the patterns of greenhouse gas emissions during the rice-growing season (Fig. 1). In CF plots, CH4 emissions increased rapidly from July 16 and peaked on August 6, while in WS plots, CH4 emissions remained relatively low during the entire experimental period (Fig. 1a). CH4 emissions were significantly higher in CF plots than in WS plots on most of the measurement dates. In contrast, N2O emissions initially decreased in both treatments and increased again only in WS plots, peaking on August 16 (Fig. 1b). In CF plots, N2O emissions were almost nondetectable after June 18. Cumulative emissions were 286 kg CH4 ha−1 and 0.003 kg N2O ha−1 in CF plots and 62 kg CH4 ha−1 and 0.019 kg N2O ha−1 in WS plots, resulting in a reduction in GWP from 6.02 ton CO2 ha−1 in CF plots to 1.31 ton CO2 ha−1 in WS plots.
Seasonal variability of CH4 (a) and N2O (b) emissions from rice paddy soils under continuous flooding (CF) and water-saving (WS) irrigation practices. Arrows denote the dates on which soils were sampled for microbiological characterization. Asterisks indicate significant differences (p < 0.05) between CF and WS plots on the corresponding date. Values are means ± standard errors, n = 3
Quantification by Real-Time PCR
Bacterial and Archaeal 16S rRNAs
The variability in abundance of bacterial and archaeal 16S rRNA genes and 16S rRNAs during the rice-growing season are illustrated in Fig. 2. No significant difference was observed between the two irrigation practices, indicating that WS irrigation did not have a substantial effect on the total abundance or activity of soil prokaryotes. Interestingly, bacterial abundance and activity peaked in August (Fig. 2a), while those of archaea tended to increase until the end of the experiment (Fig. 2b). The extent of variability was much higher in the 16S rRNAs (14.4- to 28.8-fold increase from the minimum to the maximum) than in the 16S rRNA genes (2.2- to 3.1-fold increase).
Seasonal variability in abundance of bacterial 16S rRNA genes/16S rRNAs (a) and archaeal 16S rRNA genes/16S rRNAs (b) in rice paddy soils under continuous flooding (CF) and water-saving (WS) irrigation practices. Values are means ± standard errors, n = 3. “DNA” and “RNA” in the legend denote 16S rRNA genes and 16S rRNAs, respectively
mcrA and amoA/pmoA
The variability in abundance of CH4 emission-related genes and transcripts is illustrated in Fig. 3. The abundance of the mcrA transcripts was more reflective of CH4 emissions than was the abundance of the mcrA genes (Fig. 3a): mcrA transcripts peaked on August 2 and July 9 in CF plots and WS plots, respectively, and decreased thereafter in agreement with CH4 emissions in the respective plots (Fig. 1a, when confined to emissions on the soil sampling dates). On the other hand, the abundance of mcrA genes also peaked on August 2 and July 9 in CF and WS plots, respectively, then decreased, and increased again on October 8. In addition, the extent of variability was higher in mcrA transcripts (4.2- to 4.8-fold increase from the minimum to the maximum) than in mcrA genes (1.6- to 2.5-fold increase). The abundance of mcrA transcripts was higher in CF plots than in WS plots on August 2 and September 6 (Fig. 3a), when CH4 emissions were significantly higher in CF plots; however, these differences were not statistically significant.
Seasonal variability in abundance of mcrA genes and transcripts (a), amoA/pmoA genes and transcripts (b), and mcrA genes − amoA/pmoA genes and mcrA transcripts − amoA/pmoA transcripts (c) in rice paddy soils under continuous flooding (CF) and water-saving (WS) irrigation practices. Values are means ± standard errors, n = 3. Asterisks indicate significant differences (p < 0.05) between CF and WS irrigation on the corresponding dates. “DNA” and “RNA” in the legend denote genes and transcripts, respectively
The abundance of amoA/pmoA genes fluctuated during the rice-growing season (Fig. 3b) and showed no relationship to CH4 emissions, while the abundance of amoA/pmoA transcripts peaked on August 2 and decreased thereafter, indicating that methanotrophic activity was the highest on August 2. The abundance of amoA/pmoA genes and transcripts did not differ significantly between the two irrigation practices with the exception of amoA/pmoA transcripts on October 8, when CH4 emissions were not different between CF and WS (Fig. 1a). When the difference in abundance between mcrA transcripts and amoA/pmoA transcripts was plotted against time, it followed the patterns of CH4 emissions in the respective plots and was higher in CF plots on August 2, September 6, and October 8, although a significant difference was observed only on August 2 (Fig. 3c). The difference in abundance between mcrA genes and amoA/pmoA genes showed significant differences between the two irrigation practices on September 6, but again did not follow the patterns of CH4 emissions.
qnorB, cnorB, and nosZ
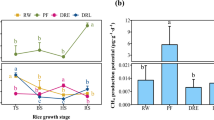
The abundance of N2O emission-related genes and transcripts generally fluctuated and showed large variability among the replicate plots (Fig. 4a–c). Significant differences between the irrigation practices were not observed during most of the soil sampling dates. Differences in abundance between genes/transcripts related to N2O production (qnorB and cnorB) and those related to N2O consumption (nosZ) also did not differ significantly between the two irrigation practices (Fig. 4d). The abundance of qnorB genes was 6.4- to 12.8-fold higher than that of cnorB genes and 55.8- to 82.8-fold higher than that of nosZ genes (Fig. S1a), and the abundance of qnorB transcripts was 2.1- to 13.4-fold higher than that of cnorB transcripts and 1.7- to 8.7-fold higher than that of nosZ transcripts during the rice-growing season (Fig. S1b).
Seasonal variability in abundance of qnorB genes and transcripts (a), cnorB genes and transcripts (b), nosZ genes and transcripts (c), and qnorB genes + cnorB genes − nosZ genes and qnorB transcripts + cnorB transcripts − nosZ transcripts (d) in rice paddy soils under continuous flooding (CF) and water-saving (WS) irrigation practices. Values are means ± standard errors, n = 3. Asterisks indicate significant differences (P < 0.05) between CF and WS irrigation on the corresponding dates. “DNA” and “RNA” in the legend denote genes and transcripts, respectively
Phylogenetic Analysis
Bacterial and Archaeal 16S rRNA
A summary of the pyrosequencing data is presented in Tables 2 and S4. No significant differences in the number of OTUs, richness estimators, or diversity indices were observed between the two irrigation practices. When weighted Fast UniFrac analysis was performed to compare prokaryotic community structure among the soil samples, the bacterial and archaeal communities obtained using 16S rRNA genes were separated from those obtained using 16S rRNAs by principal coordinate 1 (Fig. S2). It appeared that irrigation practices had no effect on the global community structure of the soil prokaryotes. The composition of bacterial phyla did not differ significantly between the two irrigation practices (Fig. 5a). The abundance of pyrosequencing reads affiliated with Proteobacteria was significantly higher in the 16S rRNAs than in the 16S rRNA genes, while the situation was reversed for Chloroflexi, suggesting that the metabolic activity of Proteobacteria was higher than that of Chloroflexi. On the other hand, the abundance of 16S rRNAs affiliated with Anaeromyxobacter and Methylocystis was significantly higher in WS plots (8.2 and 1.3 %, respectively) than in CF plots (4.2 and 0.5 %, respectively) (Fig. 5b); these differences were also observed at the species level (Fig. S3a). The abundance of 16S rRNAs belonging to the OTUs 4478, 4946, 5036, 5042, and 5061, which were clustered with the 16S rRNA gene of Anaeromyxobacter dehalogenans 2CP-1T (AF382396), were more abundant in WS than in CF plots, although these differences were not significant. OTU 4604, which showed 98.6 % of 16S rRNA gene similarity with Methylocystis echinoides IMET 10491T (AJ458473), occupied 0.33 % of bacterial 16S rRNAs in CF plots and 1.17 % in WS plots, and this difference was significant.
Relative abundance of bacterial phyla (a), predominant bacterial genera (average abundance > 0.55 %) (b), archaeal phyla (c), and Euryarchaeotal orders (d) in rice paddy soils under continuous flooding (CF) and water-saving (WS) irrigation practices. Values are means ± standard errors, n = 3. Letters indicate significant differences (p < 0.05) among samples for a given taxonomic group. “DNA” and “RNA” in the legend denote 16S rRNA genes and 16S rRNAs, respectively
The distribution of archaeal phyla in paddy soils did not differ significantly according to irrigation practice (Fig. 5c). The relative abundance of six orders of phylum Euryarchaeota is presented in Fig. 5d. Methanosarcinales was the most abundant Euryarchaeotal order (28 and 24 % of archaeal 16S rRNA genes and 38 and 39 % of archaeal 16S rRNAs in CF and WS plots, respectively). At the species level, OTUs affiliated with GOM Arc I and Methanosaetaceae, which belong to the order Methanosarcinales, dominated the Euryarchaeotal 16S rRNA genes and 16S rRNAs (Fig. S3b).
mcrA and amoA/pmoA
Although the mcrA genes and transcripts were distributed among the orders Methanosarcinales, Methanomicrobiales, and Methanobacteriales, group 1 (belonging to Methanosarcinales) occupied most of the sequences (89–99 %, Fig. 6a). Sequences affiliated with group 1 have frequently been found in rice paddy soils (Fig. S4a) but only as minor members [34–36].
Phylogenetic trees of genes and transcripts related to emissions of the greenhouse gases from rice paddy soils under continuous flooding (CF) and water-saving (WS) irrigation (in bold): mcrA (a), amoA/pmoA (b), qnorB (c), and nosZ (d). Reference sequences obtained from the FunGene and GenBank databases were also included. The phylogenetic trees were constructed based on the homology of deduced amino acid sequences with average lengths of 133 (mcrA), 100 (amoA/pmoA), 113 (qnorB), and 153 (nosZ) amino acids. Local support values (≥70) calculated using the Shimodaira-Hasegawa (SH) test are indicated at each node. The sequences were grouped arbitrarily based on the local support values and the distances among groups. A representative reference sequence is presented for each group and the isolation sources are indicated where known. Relative abundances of the groups or the sequences in each sample are represented by color gradients (heatmaps). “DNA” and “RNA” in the legend denote genes and transcripts, respectively
The phylogenetic distribution of the amoA/pmoA genes and transcripts differed dramatically (Fig. 6b). About 98 % of the amoA/pmoA genes were affiliated with amoA genes from ammonia-oxidizing bacteria in both treatments, while 73 (CF) and 89 % (WS) of amoA/pmoA transcripts were affiliated with pmoA genes from methanotrophic bacteria. Although pmoA transcripts were associated with both type I and type II methanotrophs, many (49 and 68 % in CF and WS treatments, respectively) were grouped with pmoA genes from type II methanotrophs such as Methylocystis and Methylosinus spp. (group 7 in Fig. 6b). The higher abundance of pmoA transcripts belonging to type II methanotrophs in WS treatment compared to CF treatment was in agreement with the above result (Fig. 5b), where 16S rRNAs affiliated with Methylocystis were significantly more abundant in WS plots than in CF plots.
qnorB and nosZ
The qnorB genes and transcripts showed homologies with those from Acidobacteria, Proteobacteria, Firmicutes, and Cyanobacteria at the phylum level (Fig. 6c). The qnorB sequences assigned to group 10, which was related to the qnorB gene from A. dehalogenans, were most abundant in qnorB transcripts (16 and 24 % in CF and WS plots, respectively) but were not detected in qnorB genes. The higher abundance of qnorB transcripts belonging to group 10 in WS than in CF was in agreement with the above result (Fig. 5b), where the 16S rRNAs affiliated with Anaeromyxobacter spp. were significantly more abundant in WS plots than in CF plots.
Unlike the qnorB sequences, the nosZ sequences obtained from the rice paddy soils were distributed among nosZ genes only from Proteobacteria (Fig. 6d). Most nosZ genes and transcripts (70–79 %) were assigned to group 1, which included sequences from Betaproteobacteria such as Aromatoleum, Azoarcus, Rhodoferax, and Thiobacillus (Fig. S4b). Noticeable differences in abundance of nosZ sequences between the two irrigation practices were not observed.
Discussion
The water-saving (WS) irrigation reduced CH4 emissions by 78 % and increased N2O emissions by 533 % compared to continuous flooding (CF) irrigation, without loss of crop yield. This resulted in a 78 % reduction of GWP in the rice field, which exceeded reductions obtained by other alternative irrigation practices such as intermittent drainage (27–61 %) [6–8], indicating that WS irrigation is effective for mitigating emissions of the greenhouse gases from rice fields.
The significant difference in CH4 emissions between the irrigation practices was best described by the difference in abundance between mcrA transcripts and amoA/pmoA transcripts, indicating that WS irrigation simultaneously affected methanogenic and methanotrophic activities. Zhang et al. [37] also showed that intermittent drainage reduced CH4 production potential and increased CH4 oxidation potential, thus decreasing CH4 emissions from rice fields.
Unlike the CH4 emission-related genes, the transcript abundance of N2O emission-related genes could not explain either the N2O emission patterns or the differences in N2O emissions between the two irrigation practices (Fig. 4). Two recent studies demonstrated the existence of novel nosZ genes that are difficult to detect with currently available primers, and these genes are abundant in various environments [38, 39]. Jones et al. [38] showed that nosZ sequences obtained using newly designed primers were grouped with the nosZ genes from Bacteroidetes, Gemmatimonadetes, Chloroflexi, and Proteobacteria; in contrast, the nosZ sequences obtained in the present study were associated only with genes from Proteobacteria (Fig. 6d). The discrepancy between the N2O emission patterns and the molecular results may have arisen from microbial groups with undetected nosZ genes.
There are two possible explanations for the observed differences in N2O emissions between WS and CF irrigations. The thicker layer of water in CF plots might facilitate microbial conversion of N2O to N2 by increasing the residence time of N2O [40]. On the other hand, reduction of N2O may have been inhibited in WS plots because it is known that nitrous oxide reductase can be irreversibly inactivated by O2 exposure [41, 42].
Analysis of community structure based on 16S rRNAs showed that the relative abundance of Anaeromyxobacter and Methylocystis spp. increased in WS plots compared to CF plots (Fig. 5b). It was shown that bacteria affiliated with Anaeromyxobacter were capable of utilizing acetate under methanogenic conditions [43] and A. dehalogenans, the sole species of the genus, can utilize various electron acceptors, such as chlorinated phenols, nitrate, oxygen, and ferric iron [44, 45]. We presume that the relatively oxic conditions induced by WS irrigation increased the availability of electron acceptors for Anaeromyxobacter spp., leading to their enrichment in WS plots. However, it is unclear whether Anaeromyxobacter spp. affected methanogenesis by competing with methanogens for acetate. On the other hand, genome sequencing analysis revealed that A. dehalogenans possesses norB and nosZ genes [46], and indeed, norB transcripts related to this species were abundant in the rice paddy soils, particularly in WS plots (Fig. 6c).
Ma and Lu [17] observed that intermittent drainage significantly increased the abundance of Methylocystis/Methylosinus in rhizosphere and surface soils, but not in bulk soils, suggesting that the Methylocystis/Methylosinus group responds to oxic conditions induced by intermittent drainage more rapidly than other methanotrophs. Our results showed that this group of methanotrophs can be enriched in bulk soils by the alternative irrigation practices. On the other hand, methanotrophs are known to reduce NO2 − to N2O, similar to ammonia oxidizers [47]. Mandernack et al. [48] showed that type II methanotrophs were important in NH3 oxidation and concomitant production and emission of N2O from landfill soils. Further studies will be needed to clarify the roles of Anaeromyxobacter and Methylocystis in the emissions of greenhouse gases.
GOM Arc I was previously known as ANME-2d because this group was phylogenetically related to the anaerobic methanotrophs ANME-2 [49, 50], but its role in methane biogeochemistry remains unclear [51, 52]. Because ANME clusters, including GOM Arc I, have not previously been detected in rice field soils [53, 54], identification of the exact role of this group of archaea in CH4 emissions in rice fields will be an interesting subject.
In this study, primers amplifying both amoA and pmoA genes were used for monitoring CH4 oxidation because ammonia monooxygenase of NH3-oxidizing bacteria has also the capacity to oxidize CH4 to methanol [55]. In DNA-based analysis, amoA sequences were more abundant than pmoA sequences, but in RNA-based one, the situation was reversed (Fig. 6b). This result suggests that the abundance of NH3-oxidizing bacteria was higher than that of CH4-oxidizing bacteria, but that CH4 oxidation was mediated mainly by CH4-oxidizing bacteria. This is consistent with previous reports showing that the contribution of nitrifying bacteria to CH4 oxidation was insignificant in soils [56, 57].
In previous studies, nitrite reductase genes (nirK/nirS) and nosZ genes have been the primary marker genes for denitrification [58] and norB genes have rarely been used, probably because many nondenitrifiers such as pathogens [59], nitrifiers [60], and methanotrophs [61] also possess norB genes. It has also been shown that the ratio of the abundance of nir and nosZ genes might influence potential N2O emissions [62, 63]. However, our results suggest that norB genes may be better descriptors of N2O emissions, at least in rice paddy soils. The most active qnorB group was closely associated with the qnorB gene from A. dehalogenans (group 10 in Fig. 6c), which does not possess nir genes [39]. In addition, groups 1 and 2 (other active qnorB groups) were clustered with qnorB genes from “Candidatus Koribacter versatilis” and “Candidatus Solibacter usitatus,” respectively, which also lack denitrification genes [64]. In addition, the gene and transcript abundance of qnorB genes was much higher than that of cnorB and nosZ genes (Fig. S1), suggesting that N2O emissions in rice fields might be mediated primarily by the expression of qnorB genes. The importance of nondenitrifying bacteria and their qnorB genes to N2O emissions from rice fields requires further study.
The bacterial and archaeal 16S rRNA numbers were generally higher than the 16S rRNA gene numbers, while the transcript numbers of all functional genes were lower than the gene numbers. It is believed that mRNAs were more degraded than were rRNAs during our experimental procedure because rRNA is more stable than mRNA due to its complex secondary structure [65–67]. Therefore, it is thought that the copy numbers obtained in our study must not be considered as absolute values, but considered as relative values that are meaningful only for comparison between different treatments.
It was shown that the bacterial activity peaked in August and decreased thereafter (Fig. 2a), whereas the archaeal one continuously increased during the rice-growing season (Fig. 2b). Because methane emissions (Fig. 1a) and the abundance of mcrA transcripts (Fig. 3a) peaked in August and decreased thereafter, it is thought that the increased activity of archaea after August was due to the other archaeal groups except methanogens. Many predominant archaeal sequences were clustered with ammonia-oxidizing archaea and environmental sequences, but the ecological functions of which are unknown (Fig. S3b). Investigation on the functions and activities of these archaeal groups will help to explain the seasonal variation of total archaeal activity in rice paddy soils.
In this study, only bulk soils were investigated to identify the effects of water-saving irrigation on soil prokaryotic communities. However, the rice rhizosphere was reported to be the main site for CH4 production and oxidation [56, 68, 69]. Ma et al. showed that intermittent drainage resulted in distinct effects on methanotrophic communities in bulk, rhizosphere, and surface soils [70]. The rhizosphere was also found to be the major site for coupled nitrification-denitrification in rice field soils [71, 72]. Based on these findings, investigation of the prokaryotic communities in rhizosphere soils will need to be included in future work to fully explain the effects of alternative irrigation practices on emissions of CH4 and N2O.
In conclusion, our results showed that WS irrigation could effectively reduce the GWP of rice paddy soils without a reduction in crop yield. The WS irrigation did not affect the size or structure of soil prokaryotic communities but did affect the activity of some groups. This indicates that an RNA-based approach can provide a more realistic picture of the microbial ecology of rice paddy soils than a DNA-based approach, as revealed by previous studies [18, 73, 74]. Our results provide several new findings about the effects of alternative irrigation practices on soil prokaryotic communities and are expected to contribute to the development of more effective methods for mitigating greenhouse gas emissions from rice fields.
References
IPCC (2007) IPCC fourth assessment report: climate change 2007 synthesis report. IPCC, Geneva
Linquist B, van Groenigen KJ, Adviento-Borbe MA, Pittelkow C, van Kessel C (2012) An agronomic assessment of greenhouse gas emissions from major cereal crops. Glob Chang Biol 18:194–209
Yan X, Yagi K, Akiyama H, Akimoto H (2005) Statistical analysis of the major variables controlling methane emission from rice fields. Glob Chang Biol 11:1131–1141
Linquist BA, Adviento-Borbe MA, Pittelkow CM, van Kessel C, van Groenigen KJ (2012) Fertilizer management practices and greenhouse gas emissions from rice systems: a quantitative review and analysis. Field Crop Res 135:10–21
Yan X, Akiyama H, Yagi K, Akimoto H (2009) Global estimations of the inventory and mitigation potential of methane emissions from rice cultivation conducted using the 2006 Intergovernmental Panel on Climate Change Guidelines. Global Biogeochem Cy 23:GB2002
Yang S, Peng S, Xu J, Luo Y, Li D (2012) Methane and nitrous oxide emissions from paddy field as affected by water-saving irrigation. Phys Chem Earth Pt A/B/C 53–54:30–37
Suryavanshi P, Singh YV, Prasanna R, Bhatia A, Shivay YS (2013) Pattern of methane emission and water productivity under different methods of rice crop establishment. Paddy Water Environ 11:321–329
Hou H, Peng S, Xu J, Yang S, Mao Z (2012) Seasonal variations of CH4 and N2O emissions in response to water management of paddy fields located in Southeast China. Chemosphere 89:884–892
Jiao Z, Hou A, Shi Y, Huang G, Wang Y et al (2006) Water management influencing methane and nitrous oxide emissions from rice field in relation to soil redox and microbial community. Commun Soil Sci Plant 37:1889–1903
Cai Z-C, Xing G-X, Shen G-Y, Xu H, Yan X-Y et al (1999) Measurements of CH4 and N2O emissions from rice paddies in Fengqiu, China. Soil Sci Plant Nutr 45:1–13
Berger S, Jang I, Seo J, Kang H, Gebauer G (2013) A record of N2O and CH4 emissions and underlying soil processes of Korean rice paddies as affected by different water management practices. Biogeochemistry 1–16
Hou AX, Chen GX, Wang ZP, Cleemput OV, Patrick WH Jr (2000) Methane and nitrous oxide emissions from a rice field in relation to soil redox and microbiological processes. Soil Sci Soc Am J 64:2180–2186
Krüger M, Frenzel P, Conrad R (2001) Microbial processes influencing methane emission from rice fields. Glob Chang Biol 7:49–63
Achtnich C, Bak F, Conrad R (1995) Competition for electron donors among nitrate reducers, ferric iron reducers, sulfate reducers, and methanogens in anoxic paddy soil. Biol Fert Soils 19:65–72
Ratering S, Conrad R (1998) Effects of short-term drainage and aeration on the production of methane in submerged rice soil. Glob Chang Biol 4:397–407
Watanabe T, Hosen Y, Agbisit R, Llorca L, Katayanagi N et al (2013) Changes in community structure of methanogenic archaea brought about by water-saving practice in paddy field soil. Soil Biol Biochem 58:235–243
Ma K, Lu Y (2011) Regulation of microbial methane production and oxidation by intermittent drainage in rice field soil. FEMS Microbiol Ecol 75:446–456
Ma K, Conrad R, Lu Y (2012) Responses of methanogen mcrA genes and their transcripts to an alternate dry/wet cycle of paddy field soil. Appl Environ Microbiol 78:445–454
IPCC (1997) Greenhouse gas emissions from agricultural soils. In: Houghton JT et al. JT (eds) Greenhouse gas inventory reference manual revised 1996 IPCC guidelines for national greenhouse gas inventories. IPCC/OECD/IES. UK Meteorological Office, Bracknell
Tuyrin IV (1951) To a method of the analysis for comparative structure studying soil organic matter or humus//Proc. of soil science institute by V. V. Dokuchaev (Trudi Pochvennogo Instituta by V. V. Dokuchaev) 38a: 5–21
NIAST (1988) Methods of soil chemical analysis. National Institute of Agricultural Science and Technology (NIAST), Rural Development Administration (RDA), Suwon, Korea
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Quince C, Lanzen A, Davenport R, Turnbaugh P (2011) Removing noise from pyrosequenced amplicons. BMC Bioinforma 12:38
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Cole JR, Wang Q, Cardenas E, Fish J, Chai B et al (2009) The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37:D141–D145
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W et al (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196
Pruesse E, Peplies J, Glöckner FO (2012) SINA: accurate high throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28:1823–1829
Hamady M, Lozupone C, Knight R (2009) Fast UniFrac: facilitating high-throughput phylogenetic analyses of microbial communities including analysis of pyrosequencing and PhyloChip data. ISME J 4:17–27
Price MN, Dehal PS, Arkin AP (2009) FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26:1641–1650
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K et al (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Ludwig W, Strunk O, Westram R, Richter L, Meier H et al (2004) ARB: a software environment for sequence data. Nucleic Acids Res 32:1363–1371
Ludwig W, Strunk O, Klugbauer S, Klugbauer N, Weizenegger M et al (1998) Bacterial phylogeny based on comparative sequence analysis (review). Electrophoresis 19:554–568
Letunic I, Bork P (2011) Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39:W475–W478
Xu Y, Ma K, Huang S, Liu L, Lu Y (2012) Diel cycle of methanogen mcrA transcripts in rice rhizosphere. Environ Microbiol Rep 4:655–663
Lueders T, Chin K-J, Conrad R, Friedrich M (2001) Molecular analyses of methyl-coenzyme M reductase α-subunit (mcrA) genes in rice field soil and enrichment cultures reveal the methanogenic phenotype of a novel archaeal lineage. Environ Microbiol 3:194–204
Conrad R, Klose M, Noll M, Kemnitz D, Bodelier PLE (2008) Soil type links microbial colonization of rice roots to methane emission. Glob Chang Biol 14:657–669
Zhang G, Ji Y, Ma J, Xu H, Cai Z et al (2012) Intermittent irrigation changes production, oxidation, and emission of CH4 in paddy fields determined with stable carbon isotope technique. Soil Biol Biochem 52:108–116
Jones CM, Graf DRH, Bru D, Philippot L, Hallin S (2013) The unaccounted yet abundant nitrous oxide-reducing microbial community: a potential nitrous oxide sink. ISME J 7:417–426
Sanford RA, Wagner DD, Wu Q, Chee-Sanford JC, Thomas SH et al (2012) Unexpected nondenitrifier nitrous oxide reductase gene diversity and abundance in soils. Proc Natl Acad Sci U S A 109:19709–19714
Drury CF, Findlay WI, McKenney DJ (1992) Nitric oxide and nitrous oxide production from soil: water and oxygen effects. Soil Sci Soc Am J 56:766–770
Richardson D, Felgate H, Watmough N, Thomson A, Baggs E (2009) Mitigating release of the potent greenhouse gas N2O from the nitrogen cycle—could enzymic regulation hold the key? Trends Biotechnol 27:388–397
Morley N, Baggs EM, Dörsch P, Bakken L (2008) Production of NO, N2O and N2 by extracted soil bacteria, regulation by NO2 − and O2 concentrations. FEMS Microbiol Ecol 65:102–112
Hori T, Noll M, Igarashi Y, Friedrich MW, Conrad R (2007) Identification of acetate-assimilating microorganisms under methanogenic conditions in anoxic rice field soil by comparative stable isotope probing of RNA. Appl Environ Microbiol 73:101–109
Sanford RA, Cole JR, Tiedje JM (2002) Characterization and description of Anaeromyxobacter dehalogenans gen. nov., sp. nov., an aryl-halorespiring facultative anaerobic Myxobacterium. Appl Environ Microbiol 68:893–900
He Q, Sanford RA (2003) Characterization of Fe(III) reduction by chlororespiring Anaeromxyobacter dehalogenans. Appl Environ Microbiol 69:2712–2718
Thomas SH, Wagner RD, Arakaki AK, Skolnick J, Kirby JR et al (2008) The mosaic genome of Anaeromyxobacter dehalogenans strain 2CP-C suggests an aerobic common ancestor to the Delta-Proteobacteria. PLoS ONE 3:e2103
Stein LY, Yung YL (2003) Production, isotopic composition, and atmospheric fate of biologically produced nitrous oxide. Annu Rev Earth Pl Sc 31:329–356
Mandernack KW, Kinney CA, Coleman D, Huang Y-S, Freeman KH et al (2000) The biogeochemical controls of N2O production and emission in landfill cover soils: the role of methanotrophs in the nitrogen cycle. Environ Microbiol 2:298–309
Mills HJ, Martinez RJ, Story S, Sobecky PA (2005) Characterization of microbial community structure in Gulf of Mexico gas hydrates: comparative analysis of DNA- and RNA-derived clone libraries. Appl Environ Microbiol 71:3235–3247
Martinez RJ, Mills HJ, Story S, Sobecky PA (2006) Prokaryotic diversity and metabolically active microbial populations in sediments from an active mud volcano in the Gulf of Mexico. Environ Microbiol 8:1783–1796
Lloyd KG, Lapham L, Teske A (2006) An anaerobic methane-oxidizing community of ANME-1b archaea in hypersaline Gulf of Mexico sediments. Appl Environ Microbiol 72:7218–7230
Knittel K, Boetius A (2009) Anaerobic oxidation of methane: progress with an unknown process. Annu Rev Microbiol 63:311–334
Conrad R (2007) Microbial ecology of methanogens and methanotrophs. Adv Agron 96:1–63
Miyashita A, Mochimaru H, Kazama H, Ohashi A, Yamaguchi T et al (2009) Development of 16S rRNA gene-targeted primers for detection of archaeal anaerobic methanotrophs (ANMEs). FEMS Microbiol Lett 297:31–37
Bédard C, Knowles R (1989) Physiology, biochemistry, and specific inhibitors of CH4, NH4 +, and CO oxidation by methanotrophs and nitrifiers. Microbiol Rev 53:68–84
Bodelier PLE, Frenzel P (1999) Contribution of methanotrophic and nitrifying bacteria to CH4 and NH4 + oxidation in the rhizosphere of rice plants as determined by new methods of discrimination. Appl Environ Microbiol 65:1826–1833
Klemedtsson L, Jiang Q, Kasimir Klemedtsson Å, Bakken L (1999) Autotrophic ammonium-oxidising bacteria in Swedish mor humus. Soil Biol Biochem 31:839–847
Braker G, Conrad R (2011) Chapter 2—diversity, structure, and size of N2O-producing microbial communities in soils—what matters for their functioning? Adv Appl Microbiol 75:33–70
Hendriks J, Oubrie A, Castresana J, Urbani A, Gemeinhardt S et al (2000) Nitric oxide reductases in bacteria. BBA-Bioenergetics 1459:266–273
Casciotti KL, Ward BB (2005) Phylogenetic analysis of nitric oxide reductase gene homologues from aerobic ammonia-oxidizing bacteria. FEMS Microbiol Ecol 52:197–205
Stein LY, Klotz MG (2011) Nitrifying and denitrifying pathways of methanotrophic bacteria. Biochem Soc T 39:1826–1831
García-Lledó A, Vilar-Sanz A, Trias R, Hallin S, Bañeras L (2011) Genetic potential for N2O emissions from the sediment of a free water surface constructed wetland. Water Res 45:5621–5632
Philippot L, Andert J, Jones CM, Bru D, Hallin S (2011) Importance of denitrifiers lacking the genes encoding the nitrous oxide reductase for N2O emissions from soil. Glob Chang Biol 17:1497–1504
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM et al (2009) Three genomes from the phylum Acidobacteria provide insight into the life styles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056
Belasco JG, Biggins CF (1988) Mechanisms of mRNA decay in bacteria: a perspective. Gene 72:15–23
Higgins CF (1991) Stability and degradation of mRNA. Curr Opin Cell Biol 3:1013–1018
Hierro N, Esteve-Zarzoso B, González Á, Mas A, Guillamón JM (2006) Real-time quantitative PCR (QPCR) and reverse transcription-QPCR for detection and enumeration of total yeasts in wine. Appl Environ Microbiol 72:7148–7155
Gilbert B, Frenzel P (1998) Rice roots and CH4 oxidation: the activity of bacteria, their distribution and the microenvironment. Soil Biol Biochem 30:1903–1916
Philippot L, Hallin S, Börjesson G, Baggs EM (2009) Biochemical cycling in the rhizosphere having an impact on global change. Plant Soil 321:61–81
Ma K, Conrad R, Lu Y (2013) Dry/wet cycles change the activity and population dynamics of methanotrophs in rice field soil. Appl Environ Microbiol 79:4932–4939
Arth I, Frenzel P, Conrad R (1998) Denitrification coupled to nitrification in the rhizosphere of rice. Soil Biol Biochem 30:509–515
Arth I, Frenzel P (2000) Nitrification and denitrification in the rhizosphere of rice: the detection of processes by a new multi-channel electrode. Biol Fertil Soils 31:427–435
Wertz S, Dandie CE, Goyer C, Trevors JT, Patten CL (2009) Diversity of nirK denitrifying genes and transcripts in an agricultural soil. Appl Environ Microbiol 75:7365–7377
Yoshida M, Ishii S, Fujii D, Otsuka S, Senoo K (2012) Identification of active denitrifiers in rice paddy soil by DNA- and RNA-based analyses. Microbes Environ 27:456–461
Acknowledgement
This study was carried out with the support of “Research Program for Agricultural Science and Technology Development (Project No. PJ010036)”, National Academy of Agricultural Science, Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding authors
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1786 kb)
Rights and permissions
About this article
Cite this article
Ahn, JH., Choi, MY., Kim, BY. et al. Effects of Water-Saving Irrigation on Emissions of Greenhouse Gases and Prokaryotic Communities in Rice Paddy Soil. Microb Ecol 68, 271–283 (2014). https://doi.org/10.1007/s00248-014-0371-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-014-0371-z