Abstract
The dominant factors controlling soil bacterial community variation within the tropics are poorly known. We sampled soils across a range of land use types—primary (unlogged) and logged forests and crop and pasture lands in Malaysia. PCR-amplified soil DNA for the bacterial 16S rRNA gene targeting the V1–V3 region was pyrosequenced using the 454 Roche machine. We found that land use in itself has a weak but significant effect on the bacterial community composition. However, bacterial community composition and diversity was strongly correlated with soil properties, especially soil pH, total carbon, and C/N ratio. Soil pH was the best predictor of bacterial community composition and diversity across the various land use types, with the highest diversity close to neutral pH values. In addition, variation in phylogenetic structure of dominant lineages (Alphaproteobacteria, Beta/Gammaproteobacteria, Acidobacteria, and Actinobacteria) is also significantly correlated with soil pH. Together, these results confirm the importance of soil pH in structuring soil bacterial communities in Southeast Asia. Our results also suggest that unlike the general diversity pattern found for larger organisms, primary tropical forest is no richer in operational taxonomic units of soil bacteria than logged forest, and agricultural land (crop and pasture) is actually richer than primary forest, partly due to selection of more fertile soils that have higher pH for agriculture and the effects of soil liming raising pH.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Southeast Asia is one of the major hot spots of biodiversity [1]. It has been reported that in this region the above-ground diversity has been severely affected by land use changes [2]. Deforestation and agricultural intensification are the most pervasive land use changes in Southeast Asia. In comparison to other tropical regions, Southeast Asia has the highest deforestation rate [3, 4], which has impacted its rich and unique biodiversity [3, 5]. On the other hand, conversion of land to agricultural use such as oil palm plantations has even more detrimental impacts [6, 7]. For larger organisms in the tropics, such as plants, insects, birds, or amphibians, there is clear differentiation in species composition and diversity between agricultural and nonagricultural forest environments [8, 9], although, there is evidence that a good proportion of forest species can survive in secondary forests, logged forests, and even exotic tree plantations [9, 10].
However, very little is known about below-ground diversity in the tropics of Southeast Asia, and the impact of land use upon it. Bacteria constitute a major portion of the biodiversity in soils [11, 12] and play an essential role in soil processes [13], which ultimately affect the functioning of terrestrial ecosystems. It is important to know the factors that influence the biodiversity of soil bacterial communities, to understand how these communities are structured, and also to predict ecosystem responses to a changing environment.
There have been some studies that have investigated the effects of land use change on the structure of microbial communities in the tropics. Borneman and Triplett [14] detected significant differences between soil microbial community structure in a mature forest soil and an adjacent pasture soil in eastern Amazonia. Nusslein and Tiedje [15] reported significant changes in soil bacterial community composition due to change in vegetation cover of a Hawaiian soil from forest to pasture. Bossio et al. [16] also found similar results in eastern Kenya. In addition, they found that the soil bacterial community at a regenerating secondary forest on one site was more similar to an indigenous forest at another site than it was to nearby agricultural sites. Jesus et al. [17] found that the bacterial community composition and structure in western Amazon soils were significantly more correlated to changes in soil attributes than land use. However, all these studies used traditional molecular methods such as denaturing gradient gel electrophoresis, terminal restriction fragment length polymorphism, cloning, and Sanger sequencing. These approaches are often limited to the analysis of a relatively small number of clones and a few different soil samples. Taking into account the large bacterial community size and the heterogeneity of soils, only a tiny fraction of the bacterial diversity was unraveled by these studies. With the recent development of high-throughput pyrosequencing of 16S rRNA gene [11], in-depth analysis of soil bacterial communities has now become possible.
The present study provides the most thorough research to date on large-scale variation in soil bacterial diversity across different land use types in Malaysia, one of the major hotspots of biodiversity in Southeast Asia. We used pyrosequencing to analyze bacterial community structure across four land use types. Our main objectives were to (1) identify whether and how the land use (forest vs. agriculture) influences the structure of bacterial communities and (2) identify environmental factors linked to differences in the structure and diversity of those communities.
Materials and Methods
Site Description
Samples were taken from forest and non-forest sites within the lowland equatorial tropical rainforest biome [18] at sites scattered across central and southern Malay Peninsula and Northern Borneo (Table 1). All sites sampled have an equatorial–wet climate, with precipitation equaling or exceeding potential evapotranspiration in all months of the year but two distinct peaks of rainfall in April–May and October–November in peninsular Malaysia [19], where as in May–June and October–January in northern Borneo[20]. The mean annual temperature is approximately 26.5°C throughout Malaysia with almost no variability in mean monthly temperature [19].
In late February 2009, 28 composite samples were collected from primary forest (no record of logging or tree planting in the last 100 years), logged forest (records of logging or planting with native species in the last 100 years), and crop and pasture sites (number of samples per land use type are shown in Table 1). Species cultivated at the crop sites were banana, lemongrass, oil palm, papaya, sugarcane, and tapioca.
Sampling distribution was nonrandom due to a combination of factors. Sampling was determined partly by the logistics of travel time available during fieldwork on other studies. However, samples were deliberately chosen to represent a range of terra firme forest types, soils, and land use types, in lowland Malaysia—while at the same time spaced to avoid spatial clustering. Agricultural sites were sampled during travel between forest sites, their sampling determined by availability of access roads, and ability to secure sampling permission from the farmer or landowner. Agricultural samples were also taken based on crop type (our intention being to sample a range of common Malaysian crops) and pH based on a preliminary pH sample taken at the field site before sampling. We deliberately chose samples to represent a range of soil pH levels. The localized, nonrandom distribution of areas of particular pH ranges also prevented random sampling from being a time-effective method. In agricultural sites, only fields with crops close to maturity were sampled, rather than bare fields or stubble. Since liming and fertilizer application tend to take place during earlier growth stages, this will avoid spurious effects of recent application of these chemicals. Fields in which freshly applied fertilizer pellets were visible at the ground surface were not sampled.
Soil Collection and DNA Extraction
Samples were taken at least 1 km apart. Each sampling point consisted of 1 ha and consisted of five pooled samples. This method, used by Fierer and Jackson [21], is intended to factor out very local and transient effects (e.g., a single newly fallen leaf releasing hydrogen ions) which might confuse a picture discernible on a larger scale. The intention here was to focus on a large scale rather than highly localized patterns, which would require a separate study. At each hectare sampling point, we took a scoop of approximately 200 g of soil from the top 5 cm of B horizon soil from each of the four corners of the hectare. An additional sample was also taken in the center of this hectare, and the five samples were then thoroughly homogenized in the same sterile bag. For sample collection, a sterilized trowel was used and cleaned thoroughly between successive samples. Soil samples were composited, and stored at 4°C for up to 12 h before the samples were sieved through 4-mm mesh and simultaneously stored at −80°C prior to DNA extraction.
DNA was extracted from each of the collected soil samples using the Power Soil DNA extraction kit (Mo Bio Laboratories, Carlsbad, CA, USA) according to the manufacturer’s protocol, with 0.25 g of soil (dry wt.). The purified DNA was resuspended in 50 μl of solution S6 (MoBio Laboratories) and stored at −80°C until PCR amplification.
Amplification of 16S rRNA Genes and Pyrosequencing
The extracted DNA was amplified using primers targeting the V1 to V3 hypervariable regions of the bacterial 16S rRNA gene [22]. The primers used for bacteria were V1-9 F: 5′-X-AC-GAGTTTGATCMTGGCTCAG-3′ and V3-541R: 5′-X-AC-WTTACCGCGGCTGCTGG-3′ (where X barcode is uniquely designed for each soil sample, followed by a common linker AC). Polymerase chain reactions were carried out under the following conditions: initial denaturation at 94°C for 5 min, followed by 10 cycles of denaturation at 94°C for 30 s, annealing at 60°C to 55°C with a touchdown program for 45 s, and elongation at 72°C for 90 s. This was followed by an additional 20 cycles of denaturation at 94°C for 30 s, annealing at 55°C for 45 s, and elongation at 72°C for 90 s. The amplified products were purified using the QIAquick PCR purification kit (Qiagen, CA, USA). Amplicon pyrosequencing was performed by Macrogen Incorporation (Seoul, Korea) using 454/Roche GS-FLX Titanium Instrument (Roche, NJ, USA).
Environmental Variables
To measure environmental variables, the remainder of each soil samples after DNA extraction was analyzed. Soil samples were oven dried at 60°C until constant weight. Soil pH was measured in water at the soil to solution ratio of 1:2 using a pH meter. Total nitrogen was determined by sulfuric acid digestion using Se, CuSO4, and K2SO4 as catalysts, with 1 g of soil. Total N in the digest was determined by the regular Kjeldahl distillation method [23]. Total carbon was determined by the Carbon Analyzer Leco CR-412 (Leco Corporation, St. Joseph, MI, USA), with 1 g of soil. Exchangeable potassium was estimated using 1 M ammonium acetate buffered at pH 7 [24] and determined by using atomic absorption spectroscopy using 3 g of soil. Available phosphorus was determined by the method of Bray and Kurtz [25] by autoanalyzer with 3 g of soil.
Processing of Pyrosequencing Data and Taxonomic Analysis
All the sequences were processed and analyzed following the procedures described previously [22]. The total sequencing reads were divided and assigned to each sample by recognition of the unique barcode, followed by trimming sequences by removing barcode, linker, and primer sequences at both sides. The resultant sequences were subjected to a filtering process where only reads containing 0-1 ambiguous base calls (Ns) and 300 or more base pairs were selected for the further analysis. Nonspecific PCR amplicons that showed no match with the 16S rRNA gene database upon BLASTn search (expectation value of >10−5) were also removed from the subsequent analyses. Putative chimeric sequences were detected and screened using a similarity-based approach, which splits each query sequence into two even-length fragments and then assigns each fragment to a taxon using BLAST search against EzTaxon-extended database (http://eztaxon-e.ezbiocloud.net/), followed by removal of the sequences when two fragments differ at the order level or percent identities are greater than 95% for both fragments despite assigned to different taxonomies. All sequences were classified using EzTaxon-extended database.
Phylogenetic Analysis
We used the Mothur platform (http://www.mothur.org) to compare the community-level bacterial diversity across all 28 soils [26]. The number of phylotypes (richness) was calculated with a 97% sequence similarity cutoff based on sequence alignment against EzTaxon-aligned bacterial reference sequences. We also estimated diversity using Faith’s index of phylogenetic diversity (Faith’s PD) [27], to avoid the single level of taxonomic resolution. We standardize the number of sequences across samples to avoid incomparability of measurements resulting from different-sized samples. To do this, we picked the minimum number of sequences present in a sample that was 555 sequences and randomly select this number of sequences from each of samples. We calculated both richness and Faith’s PD values from this subset of 555 sequences per sample.
The richness and Faith’s PD value was also obtained for specific lineages of bacteria (Alphaproteobacteria, Beta/Gammaproteobacteria, Acidobacteria, and Actinobacteria). For this lineage-specific richness and Faith’s PD estimation analyses, we limited the number of sequences to 150, 90, 50, and 50 randomly selected sequences per soil for Acidobacteria, Alphaproteobacteria, Beta/Gammaproteobacteria, and Actinobacteria, respectively.
Statistical Analysis
All statistical analyses were performed on a random subset of 555 sequences per soil sample to avoid effects on diversity metrics due to different number of reads among samples. The difference in overall community composition between each pair of samples was determined using the Bray–Curtis distance [28], which estimates the community distance between two samples.
Relationships between Bray–Curtis distance of bacterial community, land use (forest vs. agriculture), and soil properties were assessed using PRIMER v6 [29]. To look at the effect of land use on bacterial community, we performed an analysis of similarity (ANOSIM) with pairwise Bray–Curtis distance as response variable and land use (forest vs. agriculture) as factor. The RELATE function (a Mantel-type test) was used to determine if community-level phylogenetic distances were significantly correlated to soil properties. ANOSIM and RELATE analyses here involve distance matrices (i.e., pairwise distances between samples); pairwise distances are non-independent and therefore statistical significance needs to be evaluated by random permutation. The samples were permuted 999 times and for each time the statistic was calculated; significance is evaluated by comparing the statistic obtained from the real data against the distribution of the statistic obtained by the 999 random permutations. Statistical significance was considered at P < 0.05. Nonmetric multidimensional scaling (NMDS) was used to visualize the Bray–Curtis distance of bacterial community between each pair of samples.
Difference on soil properties between forest and agricultural land was determined by using Student’s t test for soil pH, total C, total N, C/N ratio, exchangeable K, and available P, which were normally distributed, and Wilcoxon test for K because its distribution was non-normal. Correlations between soil properties were analyzed by using VARCLUS procedure in the Hmisc R package. A significant correlation was found only between total C and total N (Fig. S1); therefore we removed total N from the analysis. To test whether soil properties that are significantly correlated to Bray–Curtis distance of bacterial community are also correlated to richness and Faith’s PD values, we performed regression analysis using linear and polynomial functions (quadratic and cubic), and the one with better fit was chosen. Regression lines were drawn by using loess function implemented in R software. A Wilcoxon test was performed to evaluate if phylotype richness and phylogenetic diversity differ between forest and agricultural land. Student’s t test, Wilcoxon test, VARCLUS procedure, regression analysis, rarefaction curves, and heat map were performed using R software package 2.13.1 (http://www.R-project.org).
Results
General analysis of the Pyrosequencing-Derived Dataset
Across all 28 soil samples, we obtained 74,802 quality sequences in total, which were classified into 27,318 operational taxonomic units (OTUs) at 97% similarity level. On average, each individual sample was represented by 2,671 classifiable sequences, with a range of 555 to 8,184 sequences per sample. The dominant taxa across all soil samples were Acidobacteria (35.8%), Alphaproteobacteria (18.1%), Beta/Gammaproteobacteria (9.4%), Chloroflexi (6.8%), Deltaproteobacteria (6.0%), and Actinobacteria (6.3%) respectively (Fig. S2 and Table S1). Most samples showed no sign of reaching an asymptote in OTU richness among the total number of reads (often over 1,000) available in the rarefaction analysis (Fig. S3).
Land Use and Bacterial Community
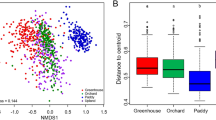
The effect of land use was tested on pairwise Bray–Curtis distances of total bacterial community as well as of four major taxa (Acidobacteria, Alphaproteobacteria, Beta/Gammaproteobacteria, and Actinobacteria). Significant differences between bacterial community composition for the different land use types were observed for total bacterial community (r = 0.14), Acidobacteria (r = 0.11), Alphaproteobacteria (r = 0.11), and Beta/Gammaproteobacteria (r = 0.08) (P < 0.05 in all cases), but not for Actinobacteria (r = 0.03, P = 0.1). NMDS also indicated some differentiation according to the different land use categories (Fig. 1a).
Soil Properties and Bacterial Community
Total C, exchangeable K, and available P differed significantly between land use types (P < 0.5 in all cases), while soil pH, total N, and C/N ratio did not vary significantly across the landscape (Table 2). The RELATE test indicated that soil pH, total C, and C/N ratio were significantly correlated with pairwise Bray–Curtis distances of total bacterial community as well as with Bray–Curtis distances of the four major taxa (Table 3). Among the soil parameters measured, soil pH was the most strongly correlated to the pairwise Bray–Curtis distances. Regression between the scores on the first NMDS axis and soil pH also indicated a strong relationship between the bacterial community composition and soil pH (Fig. 1b). However, particular agricultural land use categories clustered strongly together, presumably partly as a result of similar pH values. For example, pasture lands form one cluster (except for one sample), and oil palm plantations form another, despite the fact that they are geographically scattered (Fig. 1a).
Regression analysis results showed that soil pH, total C, and C/N ratio were significantly correlated to both phylotype richness and phylogenetic diversity of the total bacterial community, as well as of the four major taxa (Table 4). Of these, soil pH gave the most significant correlation (P < 0.0001) with both phylotype richness and phylogenetic diversity, when samples from distinct land use types are combined to encompass a wide pH range (Fig. 2). For all bacterial groups combined, there is a peak of diversity close to neutral values (pH 7.0) (Fig. 2). Soil pH was also significantly correlated (P < 0.05) to relative abundances of the dominant bacterial taxa (Fig. 3). These results indicate that pH is the best predictor of overall bacterial diversity across many soils. It was also found that the richness (W = 179, P < 0.0001) and diversity (W = 177, P < 0.0001) of agricultural sites were significantly higher than forest sites (Fig. 4).
Relationship between soil pH and the number of phylotypes (left) and phylogenetic diversity (right) of total bacterial community and four dominant bacterial taxa. Adjusted R 2 values with the associated P values are shown for each taxonomic group. Lines represent the best-fit model to the data (single asterisk linear; double asterisks quadratic; triple asterisks cubic)
In terms of both phylum and species level, comparison of relative abundances also revealed many apparent relationships to pH (Fig. 5). The heat map of 30 most abundant OTUs shows that among the dominant OTUs in these samples, no single OTU is abundant at all pH levels, although each shows its own pattern peaking at low, intermediate or high pH levels (Fig. 5). It thus seems that individual bacterial “species” are specialized in their niche to certain pH levels.
Discussion
Our results revealed that bacterial community structure was significantly correlated to both land use and soil parameters (especially soil pH). It is difficult to separate the influence of land use alone, as opposed to soil parameters, as there are significant differences in nutrient concentrations (total C, exchangeable K, and available P) between land use types. Nevertheless, the relationship to soil pH in particular is so striking that it seems that this—rather than land use—is dominant in bacterial ecology in tropical Southeast Asia.
Our results are in agreement with other studies in the tropics [17] and elsewhere [30, 31], where both land use and soil parameters were found to be key factors in determining the diversity and structure of bacterial communities. In the present study, soil pH was found to be the best predictor of bacterial community composition and diversity across all land use types: primary forest, logged forest, and crop and pasture land. Whatever the ecological and evolutionary factors behind these relationships, they appear to be as important in equatorial tropics of Southeast Asia as they are in any other biome or region of the world [17, 21, 32–35].
It is also clear from our data that all four dominant taxa in these samples (Alphaproteobacteria, Beta/Gammaproteobacteria, Acidobacteria, and Actinobacteria) show a similar trend with pH to that seen for the total bacterial community, in general. It is interesting that although the abundance of Acidobacteria shows the “expected” relationship seen in many other parts of the world [34], peaking at low pH values, their OTU richness as well as their diversity shows the same pattern as the bacterial community as a whole—peaking around neutral pH. In fact, none of the taxa we studied in greater detail shows a pattern of greater diversity at low pH: where they show any significant trend, they each peak in diversity around neutral.
In terms of land use, it is clear that the predictors of diversity in larger organisms such as trees or birds [8, 9, 36] do not hold true for soil bacteria. Generally, from this study, it seems that tropical agricultural environments, which are depauperate in diversity of larger organisms, are actually more diverse than primary tropical forests when it comes to soil bacteria. However it is important to note that the rarefaction curves for OTU richness (Fig. S3) did not approach an asymptote, indicating that many more sequences need to be retrieved to census the entire microbial communities.
The pattern we see is despite the supposition that greater diversity of plant species might be able to give more diverse bacteria specialized to the roots zone of particular plant species or to the decomposing litter beneath each species of plant. It appears likely that the greater bacterial diversity of the agricultural environments is largely due to their higher soil pH, which partly relates to liming by farmers, and partly to deliberate choice of soils which have naturally higher pH due to their better crop yields. It is interesting that limestone forest, with its high pH, has among the highest soil diversity values for the forest sites, despite its generally low tree species diversity [37].
Attention should now focus on why the striking diversity relationship with pH occurs, and why it is so general, both within and outside the tropics. Lauber et al. [34] suggested that near-neutral pH might be regarded as a proxy for physiological availability of a variety of nutrients, such as phosphorus, whose chemistry varies with pH value. Though there is no correlation between bacterial diversity and available P, in our samples, the trends observed could well be the result of precise and distinct chemical forms of these elements at different pH values. Additionally, Lauber et al. [34] note that the internal pH of bacterial cells is normally close to neutral. An external pH environment similar to this intracellular value may mean less energy expenditure on maintaining internal pH and fewer specialized adaptations. A low pH environment in the tropics could be regarded as a “stressful” environment, sensu Grime [38], demanding specialized adaptations that relatively few taxa have been able to acquire. However, part of the usual explanation among ecologists for why stressful/extreme environments are often poor in species is that these environments are rare and ephemeral in time and space, preventing evolutionary adaptation. This does not seem a convincing explanation in the tropics, where low pH soils with low bacterial diversity predominate. By offering a stable, widespread environment, these acidic soils would be expected to have accumulated the greatest diversity of bacteria.
Why then might neutral environments in the tropics have accumulated more species of bacteria? A possible explanation is that the specialized adaptations to maintain the intracellular pH level around neutral are much more easily lost than gained. A lineage that speciates into a new niche can easily do so by losing specialized low or high pH adaptations but is much less likely to regain such adaptations. Hence, lineages may be able to “fall into” relatively neutral pH environments but be less likely to escape from them in evolutionary time. The result is a net build up of bacterial taxa in niches closer to neutral soil pH. Evidence for this pattern might ultimately be found in tracing phylogenetic clades of bacteria within our data. Hypothetically, species differentiation towards the neutral middle of the pH gradient would be more frequent than movement outwards towards its extremes, with “neutral pH” clades remaining more conservative in terms of producing branches which move along the pH gradient.
As Lauber et al. [34] have suggested, there is a need for experimental study of the relationships between soil bacterial diversity and pH. Future studies should manipulate individual factors that are known to vary with pH, to better understand their role. Such studies should consider tropical soils, as much as soils from any other part of the world.
References
Myers N, Mittermeier RA, Mittermeier CG, da Fonseca GAB, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858
Hoffmann M et al (2010) The impact of conservation on the status of the world's vertebrates. Science 330:1503–1509
Sodhi NS, Koh LP, Brook BW, Ng PKL (2004) Southeast Asian biodiversity: an impending disaster. Trends Evol Ecol 19:654–660
Hansen MC, Stehman SV, Potapov PV, Loveland TR, Townshend JRG, DeFries RS, Pittman KW, Arunarwati B, Stolle F, Steininger MK, Carroll M, DiMiceli C (2008) Humid tropical forest clearing from 2000 to 2005 quantified by using multitemporal and multiresolution remotely sensed data. P Natl Acad Sci USA 105:9439–9444
Brook BW, Sodhi NS, Ng PKL (2003) Catastrophic extinctions follow deforestation in Singapore. Nature 424:420–423
Koh LP, Wilcove DS (2008) Is oil palm agriculture really destroying tropical biodiversity? Conserv Lett 1:60–64
Edwards DP, Hodgson JA, Hamer KC, Mitchell SL, Ahmad AH, Cornell SJ, Wilcove DS (2010) Wildlife-friendly oil palm plantations fail to protect biodiversity effectively. Conserv Lett 3:236–242
Fitzherbert EB, Struebig MJ, Morel A, Danielsen F, Bruhl CA, Donald PF, Phalan B (2008) How will oil palm expansion affect biodiversity? Trends Evol Ecol 23:538–545
Edwards DP, Larsen TH, Docherty TDS, Ansell FA, Hsu WW, Derhe MA, Hamer KC, Wilcove DS (2011) Degraded lands worth protecting: the biological importance of Southeast Asia's repeatedly logged forests. P Roy Soc B-Biol Sci 278:82–90
Sheldon FH, Styring A, Hosner PA (2010) Bird species richness in a Bornean exotic tree plantation: a long-term perspective. Biol Conserv 143:399–407
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AKM, Kent AD, Daroub SH, Camargo FAO, Farmerie WG, Triplett EW (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1:283–290
Fulthorpe RR, Roesch LFW, Riva A, Triplett EW (2008) Distantly sampled soils carry few species in common. ISME J 2:901–910
Bardgett RD, Freeman C, Ostle NJ (2008) Microbial contributions to climate change through carbon cycle feedbacks. ISME J 2:805–814
Borneman J, Triplett EW (1997) Molecular microbial diversity in soils from eastern Amazonia: evidence for unusual microorganisms and microbial population shifts associated with deforestation. Appl Environ Microbiol 63:2647–2653
Nusslein K, Tiedje JM (1999) Soil bacterial community shift correlated with change from forest to pasture vegetation in a tropical soil. Appl Environ Microbiol 65:3622–3626
Bossio DA, Girvan MS, Verchot L, Bullimore J, Borelli T, Albrecht A, Scow KM, Ball AS, Pretty JN, Osborn AM (2005) Soil microbial community response to land use change in an agricultural landscape of Western Kenya. Microb Ecol 49:50–62
Jesus ED, Marsh TL, Tiedje JM, Moreira FMD (2009) Changes in land use alter the structure of bacterial communities in Western Amazon soils. ISME J 3:1004–1011
Adams JM (2009) Species richness: patterns in the diversity of life. Springer-Praxis, Chichester
McGregor GR, Nieuwolt S (1998) Tropical climatology: an introduction to the climates of the low latitudes, 2nd edn. Wiley, New York
Walsh RPD, Newbery DM (1999) The ecoclimatology of Danum, Sabah, in the context of the world's rainforest regions, with particular reference to dry periods and their impact. Philos Trans R Soc B-Biol Sci 354:1869–1883
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103:626–631
Unno T, Jang J, Han D, Kim JH, Sadowsky MJ, Kim OS, Chun J, Hur HG (2010) Use of barcoded pyrosequencing and shared OTUs to determine sources of fecal bacteria in watersheds. Environ Sci Technol 44:7777–7782
Bremner JM, Mulvaney CS (1982) Nitrogen-total. In: Page AL, Miller RH, Kenny DR (eds) Method of soil analysis Part-2. American Society of Agronomy, Madison, pp 595–624
Thomas GW (1982) Exchangeable cations. In: Page AL, Miller RH, Kenny DR (eds) Method of soil analysis Part-2. American Society of Agronomy, Madison, pp 159–165
Bray RH, Kurtz LT (1945) Determination of total, organic, and available forms of phosphorus in soils. Soil Sci 59:39–45
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Faith DP (1992) Conservation evaluation and phylogenetic diversity. Biol Conserv 61:1–10
Magurran AE (2004) Measuring biological diversity. Blackwell, Oxford
Clarke KR, Gorley RN (2006) PRIMER v6: user manual/tutorial. Primer-E Ltd., Plymouth
Hartman WH, Richardson CJ, Vilgalys R, Bruland GL (2008) Environmental and anthropogenic controls over bacterial communities in wetland soils. Proc Natl Acad Sci USA 105:17842–17847
Wakelin SA, Macdonald LM, Rogers SL, Gregg AL, Bolger TP, Baldock JA (2008) Habitat selective factors influencing the structural composition and functional capacity of microbial communities in agricultural soils. Soil Biol Biochem 40:803–813
Chu H, Fierer N, Lauber CL, Caporaso JG, Knight R, Grogan P (2010) Soil bacterial diversity in the Arctic is not fundamentally different from that found in other biomes. Environ Microbiol 12:2998–3006
Griffiths RI, Thomson BC, James P, Bell T, Bailey M, Whiteley AS (2011) The bacterial biogeography of British soils. Environ Microbiol 13:1642–1654
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75:5111–5120
Lauber CL, Strickland MS, Bradford MA, Fierer N (2008) The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem 40:2407–2415
Berry NJ, Phillips OL, Ong RC, Hamer KC (2008) Impacts of selective logging on tree diversity across a rainforest landscape: the importance of spatial scale. Landscape Ecol 23:915–929
Whitmore TC, Flenley JR, Harris DR (1982) The tropics as the norm in biogeography. Geogr J 148:8–21
Grime JP (1987) Dominant and subordinate components of plant communities: implications for succession. In: Gray AJ, Crawley MJ, Edwards PJ (eds) Colonization, succession and stability. Blackwell, Oxford, pp 413–429
Acknowledgments
This work was supported by a Brain Gain grant from the Government of Malaysia and by SNU new faculty research grant to JM Adams. BM Tripathi was supported by the Graduate Scholarship for Excellent Foreign Students Program, SNU, South Korea.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 388 kb)
Rights and permissions
About this article
Cite this article
Tripathi, B.M., Kim, M., Singh, D. et al. Tropical Soil Bacterial Communities in Malaysia: pH Dominates in the Equatorial Tropics Too. Microb Ecol 64, 474–484 (2012). https://doi.org/10.1007/s00248-012-0028-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-012-0028-8