Abstract
Diversity of lactic acid bacteria (LAB) species has been analyzed for three consecutive years (2006, 2007, and 2008) during alcoholic and malolactic fermentations of Tempranillo wine in a winery at La Rioja. The results showed differences in malolactic fermentation duration, and in both diversity of LAB species and diversity of Oenococcus oeni genotypes. O. oeni was shown to be the predominant species (73% of total isolates). Monitoring the different strains of O. oeni using pulsed-field gel electrophoresis of chromosomal DNA digested with SfiI and ApaI allowed detection of a total of 37 distinct genotypes, most of them comprised at least two isolates. Six appeared in more than one vintage, one of them being present in the three studied years. Moreover, four genotypes were indistinct of the strains isolated from the air of this same winery in 2007 vintage. The frequency of participation of each genotype varied from year to year, thus dominant genotypes at one year were minority or not present at another year. This suggests that distinct indigenous O. oeni strains are better adapted to the different winery conditions every year. Predominant genotypes that appeared in more than one vintage and lead to quality wines with low histamine contents could be considered as interesting for selecting of new malolactic starter cultures.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The microorganisms play an essential role in winemaking. Yeast is able to convert sugar from grapes into alcohol and many other changes that lead to wine [1]. Lactic acid bacteria (LAB) are responsible to maintain the result of the yeast metabolism and to increase the complexity and microbial stability of this wine. To control the fermentation process, it is necessary to know and to understand the complex microbiota involved in them [2]. First studies of microbial ecology of wine were focused on the study of yeast; and subsequently, the difficulties encountered in carrying out malolactic fermentation (MLF) increased interest in the study of bacteria in wine [3]. In recent years, much work has been done on the study of the bacteria responsible for the transformation of malic acid into lactic acid and CO2, and all of them have concluded that Oenococcus oeni is the main agent of this biological deacidification, and therefore one of the most interesting oenological species [4–6]. This species has been studied at genetic and phenotypic level with the aim of selecting those most appropriate for marketing and promoting a reliable and quick fermentation [7, 8]. The development of efficient malolactic starters has therefore become one of the main challenges for oenological research in recent years. Several reports have shown that the success of these starters depends on strain and is influenced by a variety of conditions, including adaptation to the conditions of production of each wine [9–11]. Moreover, recent studies of the LAB strain diversity during consecutive vintages have shown the presence of some genotypes in several wineries and suggest the existence of a population of cosmopolitan O. oeni strains in a determinate wine-growing region [12]. Therefore, it is necessary to study previously the representative and best-adapted microbiota to the type of wine and winemaking procedures in each elaboration area. Studies conducted so far have been made mainly at the MLF stage, with other grape varieties or only in 1 year [7, 12–14]. In the present study, we analyze both the LAB diversity and the different O. oeni strains throughout alcoholic fermentation (AF) and MLF in order to know how they evolve and which one predominates in three consecutive vintages. This study contributes to a first knowledge of bacterial ecology from year to year in a winery at La Rioja region and in wine elaborated from Tempranillo, the classic red grape variety of Spain and native of Rioja Appellation.
Methods
Wine Production and Wine Samples
Traditional red wines elaborated from c.v. Tempranillo local grapes of 2006, 2007, and 2008 vintages, at one winery at the Spanish northern region of Rioja, were used in this study. Winemaking practices were typical of this wine-producing area: AF was conducted in the presence of grape skins, seeds, and stalks after the addition of sulfur dioxide and until the residual reducing sugar content was under 2 g/l. At this endpoint of AF, wines were drawn off into cement tanks and were allowed to undergo spontaneous MLF with the endogenous microbiota (no starter inoculum was used). The sampled winery had never used commercial starters for MLF. One fermentation tank was sampled in each vintage. Wine samples were collected aseptically for chemical and microbiological analyses at different times: must (stage 1), tumultuous AF (density around 1,025; stage 2), at the end of AF (<2 g/l glucose + fructose; stage 3), consumption of 10% (stage 4), 60% (stage 5) of the initial malic acid, and l-malic acid concentration <0.5 g/l (stage 6).
Chemical Analysis of the Musts and Wines
Alcohol degree, total acidity, volatile acidity, pH, free and total SO2, and reducing sugars were measured according to the European Community Official Methods [15]. Color intensity was calculated as the sum of optical density (OD) 420, 520, and 620 nm and total phenols as OD 280 nm [15]. Histamine was analyzed by reverse-phase high-performance liquid chromatography using the method reported by López [16]. MLF was followed by measuring wine l-malic acid and l-lactic acid content using kits for Enzymatic BioAnalysis (Boehringer-Mannheim/R-Biopharm, Darmstadt, Germany).
Bacterial Enumeration and Isolation
Wine samples were diluted in sterile saline solution and plated on de Man, Rogosa, and Sharpe agar (Scharlau Chemie S.A., Barcelona, Spain) plates supplemented with tomato juice (10% v/v), fructose (6 g/l), cysteine-HCl (0.5 g/l), d,l-malic acid (5 g/l), and 50 mg of pymaricine/l (Acofarma, S. Coop., Terrassa, Spain). Samples were incubated at 30°C under strict anaerobic conditions (Gas Pak System, Oxoid Ltd., Basingstoke, England, UK) for at least 10 days, and viable counts were reported as the number of colony-forming units per milliliter (CFU/ml). Fifteen colonies from each wine sample were selected for reisolation and identification. Isolates were stored in 20% sterile skim milk (DIFCO) at −20°C.
Species Identification
Species identification was carried out by previously recommended methods, which included bacteria morphology, Gram staining, and catalase [17]. O. oeni, Lactobacillus plantarum, and Lactobacillus brevis species was confirmed by the species-specific polymerase chain reaction (PCR) method [18, 19]. In case of identification of other unknown species, PCR amplification of partial 16S rRNA genes was performed with WLAB1 and WLAB2 as previously described [20]. PCR products were sequenced by Macrogen Inc. (Seoul, South Korea) and sequences were used for comparison to the data in GenBank using the Basic Local Alignment Search Tool [21].
Strain Typing of O. oeni
Pulsed-field gel electrophoresis (PFGE) was carried out according to the method described by Birren and Lai [22] with some modifications [13] for agarose block preparation. Macrorestriction analysis were performed with two separate enzymes: SfiI endonuclease following the method reported by Lopez et al. [13], and ApaI following the method reported by Ruiz et al. [23] with some modifications for optimal separation of fragments: 1.2% (w/v) agarose gels were submitted to 24 h with a pulse ramping between 0.5 and 20 s at 14°C and 6 V/cm in a CHEF DRII apparatus (Bio-Rad).
Numerical Analysis of Gel Images
The FPQuest™ software version 5.1 (Bio-Rad, Hercules, CA, USA) was used for conversion, normalization, and further processing of images. Comparison of the obtained PFGE patterns was performed with Pearson's product–moment correlation coefficient and the unweighted pair group method using arithmetic averages.
Results and Discussion
Analytical composition of must and wines during three vintages are shown in Table 1. Data for chemical parameters were within the usual range for this vinification area [16, 24]. Alcohol content ranged between 13.0% and 14.0%, pH was between 3.64 and 3.32, and free SO2 level was between 4.24 and 18.1 mg/l, at the end of AF. During MLF, a decrease in total acidity and a subsequent increase in pH were observed. In addition, an increase in volatile acidity and a decrease in parameters related to wine color (total phenols and color intensity) were noted as expected (Table 1). Thus, the wine from 2008 vintage showed more restrictive parameters for microbial growth, except for SO2 (higher in 2007), whereas the wine from 2006 vintage presented higher pH and the lowest values of alcohol content, SO2, total phenols, and color intensity. Regarding histamine concentrations in wines, results showed low levels of this amine in all studied vintages in comparison with other values reported for spontaneous MLF [25].
Viable LAB counts and l-malic acid concentration in wines during the total fermentation process are shown in Fig. 1. AF completion lasted for 6, 16, and 11 days in 2006, 2007, and 2008 vintages, respectively. Counts during AF were in the range of 102–103 CFU/mL, increasing to 107–108 CFU/mL during MLF, similar to spontaneous MLF results reported by other authors [12, 14]. The development of the MLF was related to the viable population of LAB and there was a relation between bacterial population and decrease in l-malic acid. Important differences in MLF duration were observed between vintages and MLF completion lasted for 21, 239, and 136 days (calculated as total fermentation days minus days of AF completion) in 2006, 2007, and 2008 vintages, respectively. The lack of temperature control in the winery (wine temperature below 12°C after AF in 2007) was the determinant factor in these differences, but factors such as pH, composition of the wine, and the interaction with other microorganisms implicated in the fermentation could also influence, as has been reported by other authors [2, 7, 26].
Table 2 shows the number of isolates and the percentage of the LAB species identified at each stage and year of vinification. A total of 251 LAB isolates were recovered. The greatest diversity of LAB species was detected during the AF (stages 1–3). O. oeni was present in all studied stages of the fermentation process. It was isolated in must and tumultuous AF in 2006 and 2008 vintages, and was the only species isolated at MLF in the 3 years, being therefore the predominant species, followed by L. plantarum and L. mali. The other non-O. oeni species appeared at stages 1–3 in variable rates in the three vintages. A similar distribution of species has been also reported by other authors [3, 11] and they also concluded that O. oeni is the main responsible species for MLF. The diversity of species found at each year was different, the number of species isolated in 2007 being almost double that in 2006 and 2008 and missing O. oeni until the end of the 2007 AF, a fact that could also influence the MLF duration as indicated above.
Identification of the O. oeni strains in this study was successfully achieved by PFGE of DNA digested with SfiI and each strain presented a characteristic PFGE pattern. Digestions with ApaI enzyme were not more discriminating than SfiI restriction (data not shown). Cluster analysis and visual inspection of the PFGE patterns from the 187 O. oeni isolates revealed 24, 5, and 15 unrelated patterns in 2006, 2007, and 2008 vintages, respectively (Fig. 2, Table 3). The index of diversity was also higher in 2006 than in the other years.
The comparison of PFGE patterns between the 3 years gave a total number of 37 distinct genotypes. Table 4 shows the number of different O. oeni genotypes taking part in each vintage, the stages in which each genotype was isolated and the frequency of their appearance at each vintage. Twenty-one PFGE patterns comprised several isolates (between 2 and 33), 16 strains showed unique PFGE patterns (results not shown) with percentages of appearance ranged between 1 and 2 at each year (Table 4).
This is the first study, to our knowledge, that analyzes O. oeni strain variability along the whole fermentation process in three consecutive years. Curiously, only 14 genotypes appeared at AF (stages 1–3), six were present at all MLF stages [1, 9, 24], and only three of them were also detected at stage 3. Therefore, no genotype was isolated in all fermentation stages (Table 4).
Seven genotypes (9, 13, 18, 20, 25, 26, and 34) could be described as predominant because they were detected most frequently (>10%). Genotypes 3, 13, 17, 18, 20, and 25 appeared in more than one vintage and only one genotype [25] was detected in the three studied years. There was also four genotypes (18, 20, 25, and 34) indistinct of the strains isolated by Garijo et al. [27] from the air of this winery in 2007 vintage. The frequency of participation of each genotype varied from year to year, thus dominant genotypes at 1 year were minority or not present at another year.
Comparing coincident genotypes for the three vintages, it was observed that 2006 and 2008 shared the same number [1] of patterns and only one was different from year to year but their frequency of appearance was very different, which suggests the adaptation of O. oeni strains that grow spontaneously in wine to the winery conditions every year. Similar results were reported by other authors in studies of population of bacteria and yeast [7, 14, 28].
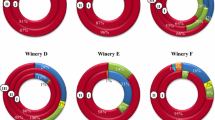
Figure 3 shows the population dynamics of O. oeni genotypes in this study. Interestingly, most fermentation stages showed mixed O. oeni strains populations, which confirmed that several strains of O. oeni occur in a single spontaneous MLF [13, 14, 29]. The number of different genotypes identified ranged from 0 to 5 and from 3 to 9 at stages 1–3 and 4–6, respectively. The 2006 vintage showed equal or higher number of genotypes than the other vintages at all stages, except for stage 2; and 2008 was the only year in which the same genotype persisted as majority at all three MLF moments studied, although different genotypes coexisted at the same moment.
Summarizing, this study contributes to a better understanding of microbial diversity of LAB populations from year to year in red wine. These results all together indicated the high diversity of indigenous O. oeni strains responsible for MLF of the wines in this study and the complexity of the ecology involved in a fermenting wine. Several genotypes [13, 17, 25] can be considered as interesting O. oeni strains because, in addition to being isolated at more than 1 year in wines with no significant production of histamine, they were dominant in most of the MLF stages at each vintage. This could be the result of successful adaptation to winemaking conditions and suggests their potential utility for the selection of new malolactic starter cultures as individual or mixed strains. Further investigation is required in more wineries at several vintages to determine coincident genotypes between wineries and distinctive genotypes at each winery that could contribute to the peculiarity of the final product. Moreover, genetic and technological characterization of the predominant strains will be necessary to carry out a starter culture selection as has been performed in other winemaking regions.
References
Carrascosa AV, Muñoz R, González R (2005) Microbiología del vino. AMV Ediciones, Madrid
Lonvaud-Funel A (1999) Lactic acid bacteria in the quality improvement and depreciation of wine. Antonie Van Leeuwenhoek, Int J Gen Mol Microbiol 76:317–331
Fugelsang KC, Edwards CG (2007) Microbial ecology during vinification. In: Fugelsang KC, Edwards CG (eds) Wine microbiology. Practical applications and procedures, second edition. Springer, New York, pp 88–91
Guerrini S, Bastianini A, Blaiotta G, Granchi L, Moschetti G, Coppola S, Romano P, Vincenzini M (2003) Phenotypic and genotypic characterization of Oenococcus oeni strains isolated from Italian wines. Int J Food Microbiol 83:1–14
Larisika M, Claus H, König H (2008) Pulsed-field gel electrophoresis for the discrimination of Oenococcus oeni isolates from different wine-growing regions in Germany. Int J Food Microbiol 123:171–176
Solieri L, Genova F, De Paola M, Giudici P (2010) Characterization and technological properties of Oenococcus oeni strains from wine spontaneous malolactic fermentations: a framework for selection of new starter cultures. J Appl Microbiol 108:285–298
Reguant C, Carreté R, Constantí M, Bordons A (2005) Population dynamics of Oenococcus oeni strains in a new winery and the effect of SO2 and yeast strain. FEMS Microbiol Lett 246:111–117
Renouf V, Delaherche A, Claisse O, Lonvaud-Funel A (2008) Correlation between indigenous Oenococcus oeni strain resistance and the presence of genetic markers. J Ind Microbiol Biotechnol 35:27–33
Coucheney F, Desroche N, Bou M, Tourdot-Maréchal R, Dulau L, Guzzo J (2005) A new approach for selection of Oenococcus oeni strains in order to produce malolactic starters. Int J Food Microbiol 105:463–470
Izquierdo PM, Garcia E, Martinez J, Chacon JL (2004) Selection of lactic bacteria to induce malolactic fermentation in red wine of cv Cencibel. Vitis 43:149–153
Ruiz P, Izquierdo PM, Sesena S, Palop ML (2010) Selection of autochthonous Oenococcus oeni strains according to their oenological properties and vinification results. Int J Food Microbiol 137:230–235
Izquierdo PM, Ruiz P, Seseña S, Palop ML (2009) Ecological study of lactic acid microbiota isolated from Tempranillo wines of Castilla–La Mancha. J Biosc Bioeng 108:220–224
López I, Tenorio C, Zarazaga M, Dizy M, Torres C, Ruiz-Larrea F (2007) Evidence of mixed wild populations of Oenococcus oeni strains during wine spontaneous malolactic fermentations. Eur Food Res Technol 226:215–223
Ruiz P, Izquierdo PM, Sesena S, Palop ML (2010) Analysis of lactic acid bacteria populations during spontaneous malolactic fermentation of Tempranillo wines at five wineries during two consecutive vintages. Food Control 21:70–75
European Community (1990) Community methods for the analysis of wine. Commission Regulation (ECC) no. 26/76/90 of 17 September 1990. Off J Eur Commun 33(L272):1–191
López R (2009) Control de la fermentación maloláctica en vinos tintos de Rioja. Influencia en su calidad higiénica, físico-química y sensorial. Thesis, University of La Rioja, Spain
Holt JG, Krieg NR, Sneath PHA, Staley JT, Williams ST (1994) In: Hensyl WR (ed) Bergey's manual of determinative bacteriology, 9th edn. Baltimore, Williams and Wilkins
Beneduce L, Spano G, Vernile A, Tarantino D, Massa S (2004) Molecular characterization of lactic acid populations associated with wine spoilage. J Basic Microbiol 44:10–16
Zapparoli G, Torriani S, Pesente P, Dellaglio F (1998) Design and evaluation of malolactic enzyme gene targeted primers for rapid identification and detection of Oenococcus oeni in wine. Lett Appl Microbiol 27:243–246
López I, Ruiz-Larrea F, Cocolin L, Orr E, Phister T, Marshall M, VanderGheynst J, Mills DA (2003) Design and evaluation of PCR primers for analysis of bacterial populations in wine by denaturing gradient gel electrophoresis. Appl Environ Microbiol 69:6801–6807
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Birren B, Lai E (1993) Preparation of DNA for pulsed field analysis. In: Pulsed field gel electrophoresis. A practical guide. Academic Press Inc, San Diego
Ruiz P, Izquierdo PM, Sesena S, Palop ML (2008) Intraspecific genetic diversity of lactic acid bacteria from malolactic fermentation of Cencibel wines as derived from combined analysis of RAPD-PCR and PFGE patterns. Food Microbiol 25:942–948
Boletin de maduración del Consejo Regulador de la D.O.Ca. Rioja. http://www.riojawine.com/es/pdfs/2007_boletin7.pdf Accessed 20 June 2011
López I, López R, Santamaría P, Torres C, Ruíz-Larrea F (2008) Performance of malolactic fermentation by inoculation of selected Lactobacillus plantarum and Oenococcus oeni strains isolated from Rioja red wines. Vitis 47:123–129
Du Plessis HW, Dicks LMT, Pretorius IS, Lambrechts MG, Du Toit M (2004) Identification of lactic acid bacteria isolated from South African brandy base wines. Int J Food Microbiol 91:19–29
Garijo P, Lopez R, Santamaria P, Ocon E, Olarte C, Sanz S, Gutierrez AR (2009) Presence of lactic bacteria in the air of a winery during the vinification period. Int J Food Microbiol 136:142–146
Gutiérrez AR, Santamaría P, Epifanio S, Garijo P, López R (1999) Ecology of spontaneous fermentation in one winery during 5 consecutive years. Lett Appl Microbiol 29:411–415
Renouf V, Vayssieres LC, Claisse O, Lonvaud-Funel A (2009) Genetic and phenotypic evidence for two groups of Oenococcus oeni strains and their prevalence during winemaking. Appl Microbiol Biotechnol 83:85–97
Acknowledgments
This work was supported by funding and predoctoral grant (B.O.R., 6 March, 2009) of the Government of La Rioja, the I.N.I.A. project RTA2007-00104-00-00, and FEDER of the European Community and was made possible by the collaborating winery.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
González-Arenzana, L., López, R., Santamaría, P. et al. Dynamics of Indigenous Lactic Acid Bacteria Populations in Wine Fermentations from La Rioja (Spain) During Three Vintages. Microb Ecol 63, 12–19 (2012). https://doi.org/10.1007/s00248-011-9911-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-011-9911-y