Abstract
Biodeterioration of archaeological sites and historic buildings is a major concern for conservators, archaeologists, and scientists involved in preservation of the world's cultural heritage. The Maya archaeological sites in southern Mexico, some of the most important cultural artifacts in the Western Hemisphere, are constructed of limestone. High temperature and humidity have resulted in substantial microbial growth on stone surfaces at many of the sites. Despite the porous natureof limestone and the common occurrence of endolithic microorganisms in many habitats, little is known about the microbial flora living inside the stone. We found a large endolithic bacterial community in limestone from the interior of the Maya archaeological site Ek' Balam. Analysis of 16S rDNA clones demonstrated disparate communities (endolithic: >80% Actinobacteria, Acidobacteria, and Low GC Firmicutes; epilithic: >50% Proteobacteria). The presence of differing epilithic and endolithic bacterial communities may be a significant factor for conservation of stone cultural heritage materials and quantitative prediction of carbonate weathering.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The interaction of microorganisms and building stone iscomplex and has been reviewed extensively [11, 13, 23, 45–47, 64]. Visible microbial growth and associated pigments may result in undesirable aesthetic changes to historically or culturally important stone buildings and objects [5], whereas the production of organic [10, 63] or inorganic [47] acids and extracellular polysaccharides (EPS) [36, 37] may lead to deterioration of the stone. On the other hand, accumulating evidence points to a protective role for microorganisms in some situations [10, 26, 42, 55, 60].
The Maya archaeological sites in southern Mexico are among the most important cultural artifacts in the Western Hemisphere. The ruins are constructed of limestone, and deterioration is the result of cyclic changes in temperature and humidity and microbial growth [16, 24]. A small number of studies have examined the microbiota of Mayan archaeological sites. These studies have demonstrated the presence of a diverse microbial community, including heterotrophic bacteria, Cyanobacteria, algae, fungi, and lichens, on the surface of stonefrom the ruins [8, 12, 27, 28, 34, 35, 61, 62]. Despite the porous nature of limestone, all of these studies have been limited to examination of growth of the surface microbiota.
The presence of endolithic microorganisms has been demonstrated in a number of habitats, such as both hot and cold deserts [9, 32], the deep subsurface [2, 33], and rock from an ocean trench [19]. There have been few studies on the endolithic microbiota of culturally important structures, and these have been limited to the examination of chasmoendolithic Cyanobacteria [44, 45] and penetration of stone by fungi [3, 17, 20, 53].
The purpose of this study was to compare the epilithic and endolithic bacterial communities of limestone from the interior of the Maya site Ek' Balam. Total community DNA was extracted from stone samples, and a clone library was constructed after polymerase chain reaction (PCR) amplification. Analysis of clone sequences indicated substantial differences between the epilithic and endolithic communities.
Methods
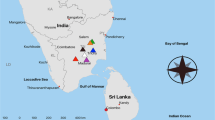
Samples were collected from Room 44 of the acropolis at Ek' Balam, Yucatan, Mexico (Fig. 1). Stone cores (n = 6) were removed from a location ca. 150 cm high on the interior of the northern wall. Electron microprobe and X-ray diffraction analyses indicated that the stone was primarily composed of calcite (>98%) and contained small amounts of magnesium, glauconite mica, K-feldspar, and Ti–Fe oxide. Samples were removed using a rock chisel sterilized with ethanol wipes and placed in sterile Whirlpak (Nasco, Fort Atkinson, WI, USA) bags on ice. In the laboratory stone samples were separated into epilithic (surface: 1 mm) and endolithic (remaining 1–2 cm) fractions in a laminar flow hood using a sterile rock chisel and hammer. Samples were then weighed in Whirlpak bags and pulverized by wrapping the bags in layers of autoclaved aluminum foil and crushing the stone with a hammer. Three epilithic and three endolithic samples were frozen in TE (100 mM Tris–HCl, 10 mM EDTA, pH 8.0) and an equal number were preserved with 1% formaldehyde.
Total numbers of bacteria in samples preserved with 1% formaldehyde were enumerated after staining with 4′,6-diamidino-2-phenylindole (DAPI) [39]. Samples were first sonicated (Branson model 2510; Danbury, CT, USA) for 5 min to detach cells from the stone. Bacteria were concentrated by filtration (15 kPa vacuum) onto 0.22-μm pore size black polycarbonate membranes (Poretics, Livermore, CA, USA), stained for 5 min with 1.0 mL of 1.0 μg/mL DAPI, and rinsed with 1.0 mL deionized water. Bacteria were enumerated using epifluorescence microscopy.
Samples frozen in TE were thawed at room temperature and DNA was extracted using the UltraClean Soil DNA Kit (MoBio Laboratories, Carlsbad, CA, USA). A portion of the 16S rDNA gene was amplified using the bacterial primers 341f (5′-CCTACGGGAGGCAGCAG-3′) [30] and 907r (5′-CCCCGTCAATTCATTTGAGTTT-3′) [31] as described by Schabereiter-Gurtner et al. [48]. PCR reactions were conducted in 50-μL volumes and contained 25 pmol of each primer, 0.2 mM of each dNTP, 5.0 μL of 10× PCR buffer (200 mM Tris–HCl pH 8.4, 500 mM KCl), 2 mM MgCl2, 2 U of Taq DNA polymerase, 4 μL of template DNA from the extractions, and deionized water. PCR was performed in a Robocycler Gradient 96 thermocycler (Stratagene, La Jolla, CA, USA) using a touchdown PCR program [48].
PCR products were cloned into the pCR 2.2-TOPO vector and transformed into competent Escherichia coli as described in the manufacturer's instructions (TOPO TA Cloning Kit K4500-01; Invitrogen, Carlsbad, CA). Plasmid preparation and sequencing were performed at the Dana Farber/Harvard Cancer Center High-Throughput DNA Sequencing Facility (Cambridge, MA, USA) using the McPrep Plasmid Purification system (Agencourt Bioscience, Beverly, MA, USA) and a 3700 DNA Analyzer (Applied Biosystems, Foster City, CA, USA) as described in the manufacturer's instructions.
We tested for Chimeric sequences using the Chimera Check program on the Ribosomal Database Project II website [7]. Unaligned sequences were compared to the National Center for Biotechnology Information database using the BLAST search program to find closely related sequences [1]. Alignments were constructed using ClustalX [59]. Phylogenetic analysis was performed using Paup 4.0 beta 10 [57]. Trees were constructed using neighbor-joining distances with 1000 bootstrap replicates. Groupings that occurred in less than 50% of replicates were excluded.
Rarefaction curves were calculated to determine if a sufficient number of clones had been sequenced to compare the two communities, and to compare the number of groups (phyla or in the case of the Proteobacteria, subphyla)found in the two communities. Rarefaction curves [21, 52] for the communities were calculated using:
where
-
E(Ŝ n ) = the expected number of groups in a random sample of individuals
-
S = the total number of groups in the entire collection
-
N i = the number of clones of group i
-
N = the total number of clones in the collection ∑N i
-
n = value of sample size (number of clones) chosen for standardization (n ≤ N)
-
\({\left( {\begin{array}{*{20}c} {N} \\ {n} \\ \end{array} } \right)}\) = number of combinations of n clones that can be chosen from a set of N clones, or [N!/n! (N − n)!]
Composition of the communities was compared using two similarity measurements. The Renkonen index [21] was calculated as
where
-
P = similarity between community 1 and 2
-
p1i = proportion of group i in community 1
-
p2i = proportion of group i in community 2.
The simplified Morisita index of similarity [18, 21] was calculated as
where
-
CH = the simplified Morisita index of similarity
-
X ij , X ik = the proportion of group i in sample j and sample k.
Results
Stone samples were collected from the Maya site Ek' Balam to compare the epilithic and endolithic bacterial communities. Large bacterial communities were found in both epilithic (7.3 × 107 ± 6.8 × 107 bacteria/g) and endolithic (2.9 × 106 ± 2.3 × 106 bacteria/g) habitats (mean ± SE, n = 3). To compare the taxonomic composition of the epilithic and endolithic bacterial communities, we prepared 16S rDNA clone libraries using DNA extracted from stone cores. Sequences from 110 clones (63 epilithic and 47 endolithic) were compared to the GenBank database using BLAST to determine their phylogenetic affiliation at the phylum or subphylum level. Clones were placed into groups having either 98% sequence similarity or the closest BLAST match to the same sequence (Tables 1 and 2). Rarefaction curves (Fig. 2) indicated that a sufficient number of clones had been analyzed because few new groups were likely to be found. Additionally, the epilithic community was more diverse than the endolithic community.
Among the epilithic clones, the majority of sequences were related to Proteobacteria, with the α-Proteobacteria being the most abundant (Figs. 3A and B). There was a cluster of sequences within the α-Proteobacteria primarily consisting of clones from epilithic samples. Within this cluster, two of the sequences were closely related to sequences previously obtained from stone cultural heritage materials. The closest BLAST match to clone Endob04 was from Altamira Cave, Spain (Table 2), whereas the closest BLAST match to clone Epih08 was from the wall of Llonín Cave, Spain (Table 1, [50]). A large number of clones from the epilithic community were closely related to Actinobacteria (Fig. 3C), and a number of clones from the epilithic community were closely related to the photosynthetic groups Cyanobacteria and Chloroflexi (Fig. 3D). Small numbers of clones from the epilithic community were closely related to the Nitrospirae and Candidate Division OP10 (Fig. 3D) as well as the Acidobacteria, Cytophaga–Flavobacterium–Bacteroides (CFB) group, and Low GC Firmicutes (Fig.3E).
Phylogenetic relationships based on partial 16S rDNA sequences of epilithic and endolithic clones isolated from Ek' Balam with sequences from members of the (A) α-Proteobacteria; (B) β-, γ-, and δ-Proteobacteria; (C) Actinobacteria; (D) Nitrospirae, Chloroflexi, Candidate Division OP10, and Cyanobacteria; and (E) Acidobacteria, Gemmatimonadetes, Cytophaga–Flavobacterium–Bacteroides, and Low GC Firmicutes. Neighbor joining trees; bootstrap values based on 1000 replicates are indicated for branches supported by >50% of trees. Scale bar represents 0.1 nucleotide changes per position.
The largest number of endolithic clones were closely related to the Actinobacteria (Table 2 and Fig. 3C). A cluster of sequences from primarily endolithic clones (Endog09 and related) within the Actinobacteria were closely related to organisms from peat soils [29]. In addition, a large percentage of the clones contained sequences similar to the Acidobacteria and the Low GC Firmicutes (Fig. 3E). A cluster of sequences (Endog10 and related) within the Acidobacteria contained clones primarily related to soil microorganisms (Fig. 3E, Table1), whereas clones Endoe11 and Epig02 were closely related to Bacillus barbaricus isolated from an experimental wall painting (Fig. 3E, Tables 1 and 2; [58]). A much smaller number of clones were closely related to the Proteobacteria (Figs. 3A and B) than had been found in the epilithic community, and two of the clones were most similar to sequences of organisms from the Gemmatimonadetes group (Fig. 3E). Many of the groups found in the epilithic community were not detected in endolithic samples (i.e., Cyanobacteria, Chlorflexi, CFB, Nitrospirae, and Candidate Division OP10).
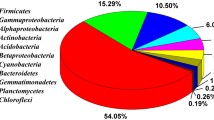
Taxonomic composition of the epilithic and endolithic communities appeared quite different (Fig. 4). Over 50% of clones from the epilithic samples were related to the Proteobacteria. Large percentages of the epilithic clones were related to the Acidobacteria, Actinobacteria, and Low GC Firmicutes. Similarity of the communities determined using the Renkonen index (RI) was 0.5, while the simplified Morisita index (SMI) returned a similarity of 0.6.
Discussion
We compared the epilithic and endolithic bacterial communities from limestone used in construction of the Acropolis at the Maya archaeological site Ek' Balam. The epilithic community was dominated by Proteobacteria with substantial numbers of Actinobacteria and the presence of photosynthetic microorganisms, whereas the endolithic community was dominated by Actinobacteria and contained large numbers of Acidobacteria and Low GC Firmicutes. Statistical indices of similarity demonstrated that the two communities differed significantly.
The Renkonen index (RI) and the simplified Morisita index (SMI) yielded different measures of similarity (0.5 and 0.6, respectively). Both indices have theoretical minima and maxima of 0.0 and 1.0, but the expected maximum similarity may be lower, depending on sample size and community diversity. Extrapolating from calculations of expected maxima for different sample sizes and diversities [66], the maximum value of the RI should be in the range of 0.6–0.8, whereas the expected maximum for the SMI, which is less affected by sample size, should fall between 0.8 and 1.0. Extrapolation to small sample sizes is valid when community diversity is low (we identified 13 bacterial groups in our samples, 25% lower than the smallest diversity level examined by Wolda [66]). Because the expected maximum of the RI is substantially lower than that of the SMI, the two indices yield comparable estimates of similarity. The RI score of 0.5 and the SMI score of 0.6 each represents 60–80% of the expected maxima. Both similarity measurements clearly demonstrated that there were dissimilar epilithic and endolithic bacterial communities present on the Maya limestone.
Previous studies of microorganisms on Maya stone have been exclusively based on culture methods and microscopy. Consequently, Cyanobacteria and a few groups of readily culturable heterotrophic bacteria (e.g., Pseudomonas sp. and Bacillus sp.) were identified as the most common organisms [12, 61]. The dominance of Pseudomonas and Bacillus is not surprising, given that these genera are readily culturable. We found that a large percentage of the clones in our study were closely related to the Proteobacteria and Low GC Firmicutes, perhaps indicating that the dominance of Pseudomonas and Bacillus in culturing studies is, in some respects, an accurate description of the bacterial community on Maya stone.
Studies using molecular techniques to examine epilithic bacteria on stone cultural heritage items have found a diverse community, including Proteobacteria, Actinobacteria, the Cytophaga–Flavobacterium–Bacteroides (CFB) group, and Acidobacteria [15, 43, 48]. Similarly, we found that large percentages of clones from Ek' Balam were related to the Proteobacteria, Actinobacteria, and Acidobacteria groups. Additionally, a large percentage of the endolithic clones from Ek' Balam were closely related to the Low GC Firmicutes group. Other molecular studies of bacteria on stone buildings and cultural heritage objects may have failed to detect these organisms as an important component of the stone microbiota because they have only sampled the stone surface.
The bacterial community from Ek' Balam is also similar in some respects to bacterial communities found in caves. Culture-based studies have reported large numbers of Actinobacteria in caves [14, 25]. Molecular studies of epilithic bacteria and soil from caves have shown that close to 50% of the community is related to Proteobacteria and a large percentage of clones are closely related to Acidobacteria [49, 50]. These studies differed in the percentage of the community related to Actinobacteria (20% and 5%), and a maximum of 11% of clones were closely related to the Firmicutes. Similarities between the epilithic bacteria at Ek' Balam and the epilithic and soil bacteria in caves might be expected because the environments are comparable (i.e., rock surfaces, oligotrophic, low light levels).
Cyanobacteria have frequently been found in endolithic habitats. In extreme environments, endolithic growth provides protection from low temperature, UV radiation, and desiccation [32]. Reports of endolithic microorganisms in stone buildings and statues have detailed the growth of chasmoendolithic Cyanobacteria inhabiting cracks and fissures in the stone [44, 45]. However, we did not find any clones closely related to Cyanobacteria in our endolithic samples. On external stone and building surfaces, high light levels may favor endolithic growth by Cyanobacteria. Our samples were collected from an interior room where no direct light reached the stone surface, and light intensity may have limited the growth of endolithic Cyanobacteria.
Endolithic Cyanobacteria may cause the deterioration of culturally important stone objects. Water absorption by the biofilm matrix results in shrinking and swelling of the EPS, causing mechanical stress that opens cracks and fissures in the stone [64]. In the case of endolithic Cyanobacteria, this process may lead to the exfoliation of surface layers and crusts [22]. Conceivably, the biofilm matrix produced by the endolithic bacteria described in this study may function in a similar manner. Furthermore, much of the EPS in these previous studies may have originated from the largely ignored heterotrophic bacterial community associated with the Cyanobacteria.
The observation of a phylogenetically distinct population of microorganisms in endolithic limestone environments also affects our understanding of weathering of carbonate minerals. These minerals comprise 4% of the earth's crust and represent the most reactive mineral species [40]. Their cycling influences the chemistry of oceans [38, 51] and partially regulates local alkalinity in terrestrial environments [56], which can affect the transport of anthropogenic pollutants, especially heavy metals [41, 54]. The role of microorganisms in dissolution and precipitation of carbonate minerals is the focus of current research efforts and remains unclear. Microorganisms have been observed to accelerate the mineral dissolution rate through production of simple acids [6]. Additionally, extracellular polysaccharides in microbial biofilms can accelerate [37] or retard dissolution rates [4, 65]. These polymers may also act to stabilize secondary mineral precipitation with contaminant metal species. The observation of a distinct endolithic microbial population affects our current understanding of limestone weathering and leads to increased complexity in geochemical modeling.
In conclusion, we have demonstrated the presence of an endolithic bacterial community in limestone from the Maya site Ek' Balam that is distinctly different from the community on the stone surface. The presence of a previously undescribed endolithic microbial community has important implications for the protection of the world's archaeological sites and historic buildings, and for our understanding of biogeochemical processes controlling dissolution of carbonate minerals.
References
SF Altschul TL Madden AA Schäffer J Zhang Z Zhang W Miller DJ Lipman (1997) ArticleTitleGapped BLAST and PSI-BLAST: a new generation of protein database search programs Nucleic Acids Res 25 3389–3402 Occurrence Handle10.1093/nar/25.17.3389 Occurrence Handle1:CAS:528:DyaK2sXlvFyhu7w%3D Occurrence Handle9254694
PS Amy DL Haldeman D Ringelberg DH Hall C Russell (1992) ArticleTitleComparison of identification systems for classification of bacteria isolated from water and endolithic habitats within the deep subsurface Appl Environ Microbiol 58 3367–3373 Occurrence Handle16348791
C Ascaso J Wierzchos R Castello (1998) ArticleTitleStudy of the biogenic weathering of calcareous limestones caused by lichen and endolithic microorganisms Int Biodeterior Biodegrad 42 29–38 Occurrence Handle1:CAS:528:DyaK1cXnslGlsr4%3D Occurrence Handle10.1016/S0964-8305(98)00043-2
JF Banfield WW Barker SA Welch A Taunton (1999) ArticleTitleBiological impact on mineral dissolution: application of the lichen model to understanding mineral weathering in the rhizosphere Proc Natl Acad Sci USA 96 3404–3411 Occurrence Handle10.1073/pnas.96.7.3404 Occurrence Handle1:CAS:528:DyaK1MXjslCisbc%3D Occurrence Handle10097050
M Bassi A Ferrari M Realini C Sorlini (1986) ArticleTitleRed stains on the Certosa of Pavia: a case of biodeterioration Int Biodeterior 3 201–205
BE Christensen WG Characklis (1990) Physical and chemical properties of biofilms WG Characklis KC Marshall (Eds) Biofilms Wiley New York
JR Cole B Chai TL Marsh RJ Farris Q Wang SA Kulam S Chandra DM McGarrell TM Schmidt GM Garrity JM Tiedje (2003) ArticleTitleThe Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy Nucleic Acids Res 31 442–443 Occurrence Handle10.1093/nar/gkg039 Occurrence Handle1:CAS:528:DC%2BD3sXhvFSls7o%3D Occurrence Handle12520046
CA Crispim PM Gaylarde CC Gaylarde (2003) ArticleTitleAlgal and cyanobacterial biofilms on calcareous historic buildings Curr Microbiol 46 79–82 Occurrence Handle10.1007/s00284-002-3815-5 Occurrence Handle1:CAS:528:DC%2BD3sXhtlOhsrc%3D Occurrence Handle12520359
JR Torre ParticleDe la BM Goebel EI Friedmann NR Pace (2003) ArticleTitleMicrobial diversity of cryptoendolithic communities from the McMurdo Dry Valleys, Antarctica Appl Environ Microbiol 69 3858–3867 Occurrence Handle12839754 Occurrence Handle10.1128/AEM.69.7.3858-3867.2003 Occurrence Handle1:CAS:528:DC%2BD3sXlsFagt74%3D
MP Bonaventura ParticleDi M Gallo Particledel P Cacchio C Ercole A Lepidi (1999) ArticleTitleMicrobial formation of oxalate films on monument surfaces: bioprotection or biodeterioration? Geomicrobiol J 16 55–64 Occurrence Handle10.1080/014904599270749
CC Gaylarde LHG Morton (1999) ArticleTitleDeteriogenic biofilms on buildings and their control: a review Biofouling 14 59–74
PM Gaylarde CC Gaylarde PS Guiamet SGG Saravia Particlede HA Videla (2001) ArticleTitleBiodeterioration of Mayan buildings at Uxmal and Tulum, Mexico Biofouling 17 41–45
PS Griffin N Indictor RJ Koestler (1991) ArticleTitleThe biodeterioration of stone: a review of deterioration mechanisms, conservation case histories, and treatment Int Biodeterior 28 187–207 Occurrence Handle10.1016/0265-3036(91)90042-P
I Groth P Schumann L Laiz S Sanchez-Moral JC Canaveras C Saiz-Jimenez (2001) ArticleTitleGeomicrobiological study of the Grotta dei Cervi, Porto Badisco, Italy Geomicrobiol J 18 241–258 Occurrence Handle1:CAS:528:DC%2BD3MXmvFKkurg%3D Occurrence Handle10.1080/01490450152467778
C Gurtner J Heyrman G Pinar W Lubitz J Swings S Rolleke (2000) ArticleTitleComparative analysis of the bacterial diversity on two different biodeteriorated wall paintings by DGGE and 16S rDNA sequence analysis Int Biodeterior Biodegrad 46 229–239 Occurrence Handle1:CAS:528:DC%2BD3MXltVSnug%3D%3D Occurrence Handle10.1016/S0964-8305(00)00079-2
ME Hale SuffixJr (1983) The Biology of Lichens EditionNumber3 Edward Arnold London 137–138
P Hirsch FEW Eckhardt RJ Palmer SuffixJr (1995) ArticleTitleFungi active in weathering of rock and stone monuments Can J Bot 73 S1384–S1390 Occurrence Handle10.1139/b95-401
HS Horn (1966) ArticleTitleMeasurement of “overlap” in comparative ecological studies Am Nat 100 419–424 Occurrence Handle10.1086/282436
F Inagaki K Takai K Tetsushi Y Sakihama A Inoue K Horikoshi (2002) ArticleTitleProfile of microbial community structure and presence of endolithic microorganisms inside a deep-sea rock Geomicrobiol J 19 535–552 Occurrence Handle1:CAS:528:DC%2BD3sXntl2nuw%3D%3D Occurrence Handle10.1080/01490450290098577
RJ Koestler AE Charola M Wypyski JJ Lee (1985) Microbiologically induced deterioration of dolomitic and calcitic stone as viewed by scanning electron microscopy G Felix (Eds) Vth International Congress on Deterioration and Conservation of Stone Presses Polytechniques Romandes Lausanne, Switzerland
CJ Krebs (1999) Ecological Methodology EditionNumber2 Benjamin-Cummings Menlo Park, CA
WE Krumbein (1988) Microbial interactions with mineral materials DR Houghton RN Smith HOW Eggins (Eds) Biodeterioration 7 Elsevier London 78–100
R Kumar AV Kumar (1999) Biodeterioration of stone in tropical environments The Getty Conservation Institute Los Angeles, CA
R Kumar WS Ginell (1995) Evaluation of consolidants for protection of weak Maya limestone Methods of Evaluating Products for the Conservation of Porous Building Materials in Monuments, Preprints of the International Colloquium ICCROM Rome
L Laiz M Gonzalez-Delvalle B Hermosin A Ortiz-Martinez C Saiz-Jimenez (2003) ArticleTitleIsolation of cave bacteria and substrate utilization at different temperatures Geomicrobiol J 20 479–489 Occurrence Handle1:CAS:528:DC%2BD3sXnvVOlu7c%3D Occurrence Handle10.1080/713851125
A Lüttge PG Conrad (2004) ArticleTitleDirect observation of microbial inhibition of calcite dissolution Appl Environ Microbiol 70 1627–1632 Occurrence Handle15006787 Occurrence Handle10.1128/AEM.70.3.1627-1632.2004 Occurrence Handle1:CAS:528:DC%2BD2cXisVKis7w%3D
CJ McNamara TD Perry M Zinn M Breuker R Mitchell (2002) Biodeterioration of concrete and stone B Little (Eds) Microbiologically Influenced Corrosion NACE International Houston, TX
CJ McNamara TD Perry M Zinn M Breuker G Hernandez-Duque R Mitchell (2003) Microbial processes in the deterioration of Mayan archaeological buildings in southern Mexico RJ Koestler VH Koestler AE Charola FE Nieto-Fernandez (Eds) Art, Biology, and Conservation: Biodeterioration of Works of Art The Metropolitan Museum of Art New York
SA Morris S Radajewski TW Willison JC Murrell (2002) ArticleTitleIdentification of the functionally active methanotroph population in a peat soil microcosm by stable-isotope probing Appl Environ Microbiol 68 1446–1453 Occurrence Handle10.1128/AEM.68.3.1446-1453.2002 Occurrence Handle1:CAS:528:DC%2BD38XitFSjsbo%3D Occurrence Handle11872500
G Muyzer EC Waal ParticleDe AG Uitterlinden (1993) ArticleTitleProfiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA Appl Environ Microbiol 59 695–700 Occurrence Handle1:CAS:528:DyaK3sXit1Kktrk%3D Occurrence Handle7683183
G Muyzer A Teske CO Wirsen HW Jannasch (1995) ArticleTitlePhylogenetic relationships of Thiomicrospira species and their identification in deep-sea hydrothermal vent samples by denaturing gradient gel electrophoresis of 16S rDNA fragments Arch Microbiol 164 165–172 Occurrence Handle10.1007/BF02529967 Occurrence Handle1:CAS:528:DyaK2MXovFyjsrk%3D Occurrence Handle7545384
JA Nienow EI Friedmann (1993) Terrestrial lithophytic (rock) communities EI Friedmann (Eds) Antarctic Microbiology Wiley-Liss Inc. New York
CJ Newberry G Webster BA Cragg RJ Parkes AJ Weightman JC Fry (2004) ArticleTitleDiversity of prokaryotes and methanogenesis in deep subsurface sediments from the Nankai Trough, Ocean Drilling Program Leg 190 Environ Microbiol 6 274–287 Occurrence Handle10.1111/j.1462-2920.2004.00568.x Occurrence Handle14871211
O Ortega-Morales G Hernández-Duque L Borges-Gómez J Guezennec (1999) ArticleTitleCharacterization of epilithic microbial communities associated with Mayan stone monuments in Yucatan, Mexico Geomicrobiol J 16 221–232 Occurrence Handle1:CAS:528:DyaK1MXmtVKis7s%3D Occurrence Handle10.1080/014904599270604
O Ortega-Morales J Guezennec G Hernández-Duque CC Gaylarde PM Gaylarde (2000) ArticleTitlePhototrophic biofilms on ancient Mayan buildings in Yucatan, Mexico Curr Microbiol 40 81–85 Occurrence Handle10.1007/s002849910015 Occurrence Handle1:CAS:528:DC%2BD3cXotFSrtg%3D%3D Occurrence Handle10594218
TD Perry CJ McNamara R Mitchell G Hernandez-Duque (2003) An investigation of bacterial dissolution of Maya limestone: biodiversity and functional analysis C Saiz-Jimenez (Eds) Molecular Biology and Cultural Heritage Swets and Zeitlinger Lisse
TD Perry OW Duckworth CJ McNamara ST Martin R Mitchell (2004) ArticleTitleEffects of the biologically produced polymer alginic acid on macroscopic and microscopic calcite dissolution rates Environ Sci Technol 38 3040–3046 Occurrence Handle10.1021/es035299a Occurrence Handle1:CAS:528:DC%2BD2cXjs1ensLY%3D Occurrence Handle15224733
M Pilson (1998) An Introduction to the Chemistry of the Sea Prentice-Hall Upper Saddle River, NJ
KG Porter YS Feig (1980) ArticleTitleThe use of DAPI for identifying and counting aquatic microflora Limnol Oceanogr 25 943–948 Occurrence Handle10.4319/lo.1980.25.5.0943
L Raymond (1995) Petrology: The Study of Igneous, Sedimentary, and Metamorphic Rocks Brown Publishers Dubuque, IA
RJ Reeder M Nugent C Tait D Morris S Heald K Beck W Hess A Lazirotti (2001) ArticleTitleCoprecipitation of uranium (VI) with calcite: XAFS, micro-XAS, and luminescence characterization Geochim Cosmochim Acta 65 3491–3503 Occurrence Handle10.1016/S0016-7037(01)00647-0 Occurrence Handle1:CAS:528:DC%2BD3MXotVOrurY%3D
C Rodriguez-Navarro M Rodriguez-Gallego KB Chekroun MT Gonzalez-Muñoz (2003) ArticleTitleConservation of ornamental stone by Myxococcus xanthus-induced carbonate biomineralization Appl Environ Microbiol 69 2182–2193 Occurrence Handle10.1128/AEM.69.4.2182-2193.2003 Occurrence Handle1:CAS:528:DC%2BD3sXivFKqsL4%3D Occurrence Handle12676699
S Rolleke G Muyzer C Wawer G Wanner W Lubitz (1996) ArticleTitleIdentification of bacteria in a biodegraded wall painting by denaturing gradient gel electrophoresis of PCR-amplified gene fragments coding for 16S rRNA Appl Environ Microbiol 62 2059–2065 Occurrence Handle1:CAS:528:DyaK28XjtlGgsL4%3D Occurrence Handle8787403
C Saiz-Jimenez J Garcia-Rowe MA Garcia del Cura JJ Ortega-Calvo E Roekens R Grieken ParticleVan (1990) ArticleTitleEndolithic cyanobacteria in Maastricht limestone Sci Total Environ 94 209–220 Occurrence Handle10.1016/0048-9697(90)90171-P Occurrence Handle1:CAS:528:DyaK3cXksVGgtb4%3D
C Saiz-Jimenez (1999) ArticleTitleBiogeochemistry of weathering processes in monuments Geomicrobiol J 16 27–37 Occurrence Handle1:CAS:528:DyaK1MXhsl2itLo%3D Occurrence Handle10.1080/014904599270721
C Saiz-Jimenez (2001) The biodeterioration of building materials JC Stoecker (Eds) Microbiologically Influenced Corrosion, vol. 2 NACE International Houston
W Sand E Bock (1991) ArticleTitleBiodeterioration of mineral materials by microorganisms—biogenic sulfuric and nitric acid corrosion of concrete and natural stone Geomicrobiol J 9 129–138 Occurrence Handle1:CAS:528:DyaK38Xmtlykt74%3D
C Schabereiter-Gurtner G Pinar W Lubitz S Rolleke (2001) ArticleTitleAn advanced molecular strategy to identify bacterial communities on art objects J Microbiol Methods 45 77–87 Occurrence Handle10.1016/S0167-7012(01)00227-5 Occurrence Handle1:CAS:528:DC%2BD3MXislGktrg%3D Occurrence Handle11311392
C Schabereiter-Gurtner C Saiz-Jimenez G Pinar W Lubitz S Rolleke (2002) ArticleTitleAltamira cave Paleolithic paintings harbor partly unknown bacterial communities FEMS Microbiol Lett 211 7–11 Occurrence Handle1:CAS:528:DC%2BD38XktFKgurY%3D Occurrence Handle12052543 Occurrence Handle10.1111/j.1574-6968.2002.tb11195.x
C Schabereiter-Gurtner C Saiz-Jimenex G Pinar W Lubitz S Rolleke (2004) ArticleTitlePhylogenetic diversity if bacteria associated with Paleolithic paintings and surrounding rock walls in two Spanish caves (Llonín and La Garma) FEMS Microbiol Ecol 47 235–247 Occurrence Handle1:CAS:528:DC%2BD2cXhtFWgurs%3D Occurrence Handle10.1016/S0168-6496(03)00280-0 Occurrence Handle19712338
WH Schlesinger (1997) Biogeochemistry: An Analysis of Global Change Academic Press San Diego, CA
D Simberloff (1978) Use of rarefaction and related methods in ecology KL Dickson J Cairns SuffixJr RJ Livingston (Eds) Biological Data in Water Pollution Assessment: Quantitative and Statistical Analysis, ASTM STP 652 American Society for Testing and Materials Philadelphia, PA
K Sterflinger WE Krumbein (1997) ArticleTitleDematiaceous fungi as a major agent for biopitting on Mediterranean marbles and limestones Geomicrobiol J 14 219–230
SL Stipp J Konnerup-Madsen K Franzreb A Kulik H Mathieu (1998) ArticleTitleSpontaneous movement of ions through calcite at standard temperature and pressure Nature 396 356–359 Occurrence Handle10.1038/24597 Occurrence Handle1:CAS:528:DyaK1cXnvVars7s%3D
S Stocks-Fischer JK Galinat SS Bang (1999) ArticleTitleMicrobiological precipitation of CaCO3 Soil Biol Biochem 31 1563–1571 Occurrence Handle10.1016/S0038-0717(99)00082-6 Occurrence Handle1:CAS:528:DyaK1MXls12nurg%3D
W Stumm JJ Morgan (1996) Aquatic Chemistry Wiley New York
DL Swofford (2003) PAUP*. Phylogenic Analysis Using Parsimony (*and other Methods), Version 4 Sinauer Associates Sunderland, MA
M Taubel P Kampfer S Buczolits W Lubitz HJ Busse (2003) ArticleTitleBacillus barbaricus sp. nov., isolated from an experimental wall painting Int J Syst Evol Microbiol 53 725–730 Occurrence Handle1:CAS:528:DC%2BD3sXkvFOju70%3D Occurrence Handle12807193 Occurrence Handle10.1099/ijs.0.02304-0
JD Thompson TJ Gibson F Plewniak F Jeanmougin DG Higgins (1997) ArticleTitleThe ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools Nucleic Acids Res 24 4876–4882 Occurrence Handle10.1093/nar/25.24.4876
P Tiano L Biagiotti G Mastromei (1999) ArticleTitleBacterial bio-mediated calcite precipitation for monumental stones conservation: methods of evaluation J Microbiol Methods 36 139–145 Occurrence Handle10.1016/S0167-7012(99)00019-6 Occurrence Handle1:CAS:528:DyaK1MXjs1Kntbw%3D Occurrence Handle10353808
HA Videla PS Guiamet SG Saravia Particlede (2000) ArticleTitleBiodeterioration of Mayan archaeological sites in the Yucatan Peninsula, Mexico Int Biodeterior Biodegrad 46 335–341 Occurrence Handle1:CAS:528:DC%2BD3MXitlKntL8%3D Occurrence Handle10.1016/S0964-8305(00)00106-2
Videla, HA, Guiamet, PS, de Saravia, SG, Maldonaldo, L (2001) Mechanisms of Microbial Biodeterioration of Limestone in Mayan Buildings. Paper No. 01250, Corrosion2001, NACE International, Houston
T Warscheid M Oelting WE Krumbein (1991) ArticleTitlePhysico-chemical aspects of biodeterioration processes in rocks with special regard to organic pollutants Int Biodeterior 28 37–48 Occurrence Handle1:CAS:528:DyaK38XhsFSkurg%3D Occurrence Handle10.1016/0265-3036(91)90032-M
T Warscheid J Braams (2000) ArticleTitleBiodeterioration of stone: a review Int Biodeterior Biodegrad 46 343–368 Occurrence Handle1:CAS:528:DC%2BD3MXitlKntLw%3D Occurrence Handle10.1016/S0964-8305(00)00109-8
SA Welch P Vandevivere (1994) ArticleTitleEffect of microbial and other naturally occurring polymers on mineral dissolution Geomicrobiol J 12 227–238 Occurrence Handle1:CAS:528:DyaK2MXnt1Cisrg%3D
H Wolda (1981) ArticleTitleSimilarity indices, sample size and diversity Oecologia 50 296–302 Occurrence Handle10.1007/BF00344966
Acknowledgments
This research was funded in part by a grant from the National Science Foundation (BES-9906337) to Harvard University and from the Consejo Nacional de Ciencia y Tecnologia to G. Hernandez-Duque. Additional support was provided by a grant from the Samuel H. Kress Foundation. The authors wish to thank the Instituto Nacional de Antropologia e Historia, Leticia Vargas de la Peña, and Alejandra Alonso Olvera for permission and help in collecting samples.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
McNamara, C.J., Perry, T.D., Bearce, K.A. et al. Epilithic and Endolithic Bacterial Communities in Limestone from a Maya Archaeological Site. Microb Ecol 51, 51–64 (2006). https://doi.org/10.1007/s00248-005-0200-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-005-0200-5