Abstract
The community structure of methanogenic Archaea on anoxically incubated rice roots was investigated by amplification, sequencing, and phylogenetic analysis of 16S rRNA and methyl-coenzyme M reductase (mcrA) genes. Both genes demonstrated the presence of Methanomicrobiaceae, Methanobacteriaceae, Methanosarcinaceae, Methanosaetaceae, and Rice cluster I, an uncultured methanogenic lineage. The pathway of CH4 formation was determined from the 13C-isotopic signatures of the produced CH4, CO2 and acetate. Conditions and duration of incubation clearly affected the methanogenic community structure and the pathway of CH4 formation. Methane was initially produced from reduction of CO2 exclusively, resulting in accumulation of millimolar concentrations of acetate. Simultaneously, the relative abundance of the acetoclastic methanogens (Methanosarcinaceae, Methanosaetaceae), as determined by T-RFLP analysis of 16S rRNA genes, was low during the initial phase of CH4 production. Later on, however, acetate was converted to CH4 so that about 40% of the produced CH4 originated from acetate. Most striking was the observed relative increase of a population of Methanosarcina spp. (but not of Methanosaeta spp.) briefly before acetate concentrations started to decrease. Both acetoclastic methanogenesis and Methanosarcina populations were suppressed by high phosphate concentrations, as observed under application of different buffer systems. Our results demonstrate the parallel change of microbial community structure and function in a complex environment, i.e., the increase of acetoclastic Methanosarcina spp. when high acetate concentrations become available.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Flooded rice fields are an important source of the greenhouse gas CH4, contributing about 13% to the total CH4 budget [24]. The microbiota on rice roots may contribute a significant portion, since up to 50% of the emitted CH4 originates from plant photosynthates that are excreted from the roots [40], and about 3–6% of the photosynthetically fixed CO2 is converted to CH4 and emitted [9]. The root-associated methanogenesis possibly affects not only the amount but also the isotopic signature of the emitted CH4 [8, 21].
Rice field soil and rice roots have also become a model system for the microbial ecology of wetlands [4]. The methanogenic archaeal biota on rice roots seems to be different from that in the anoxic soil [17, 18]. Likewise, the pathway of CH4 production and its temporal dynamics was found to be different in root and soil [5, 8, 23]. Acetoclastic methanogenic activity, in particular, was found to increase with time in root incubations and to be subject to inhibition by phosphate [6, 7]. However, data on the methanogenic archaeal community structure on rice roots are scarce. In particular, the contribution of the different major methanogenic groups to CH4 formation and their population dynamics upon onset of methanogenesis are unknown. Furthermore, analysis of the methanogenic community structure on rice roots has been restricted to analysis of 16S rRNA genes. Functional marker genes, such as the mcrA gene encoding the methyl-coenzyme M reductase, the key enzyme of methanogenesis [12], have not been studied. This specific functional marker has been applied for the selective investigation of methanogenic communities in different anoxic habitats [2, 11, 16, 19, 25, 26, 30, 31, 33].
In this study, clone libraries targeting both 16S rRNA and mcrA genes of methanogens on anoxically incubated excised rice roots were constructed to compare the populations to those detected previously in anoxic bulk soil [3, 10, 18]. Furthermore, we investigated temporal changes in the methanogenic archaeal community in correlation to the predominant pathways of CH4 formation. Such analyses are necessary to improve our current knowledge of factors controlling the presence and activity of methanogenic populations on the surface of rice roots.
Materials and Methods
Incubation Experiments and Chemical Analyses
The growth of rice plants and the preparation of rice roots have already been described in detail [5]. Briefly, rice plants (Oryza sativa, var. Roma, type japonica) were grown in the greenhouse. After 12–14 weeks, plants were removed from the soil, and the roots were washed first with tap water, then with sterile demineralized water that was bubbled with N2. The roots were then cut off with a razor blade, and aliquots (10 g) were incubated in moist state at room temperature (about 25°C) under a N2 atmosphere, using stoppered glass bottles (150 mL volume) [5]. Alternatively, the roots were incubated in 50 mL anoxic sterile carbonate or phosphate buffer. The carbonate buffer consisted of demineralized water plus 5 g marble granules (0.5–2.0 mm size, consisting of CaCO3; Merck, Darmstadt, Germany), the phosphate buffer of 50 mM KH2PO4, 17 mM NaCl and 0.2 mM MgCl2 (pH 7.0) [6].
Gas samples (0.25–1.0 mL) and liquid samples (0.5 mL) were analyzed by gas chromatography and HPLC as described [5]. Stable isotope analysis of 13C/12C in gas and liquid samples was performed using a gas chromatograph combustion isotope ratio mass spectrometry (GCC-IRMS) system purchased from Finnigan (Thermoquest, Bremen, Germany) [8]. The fractions of CH4 produced from reduction of CO2 and acetate cleavage were calculated from the 13C data of CH4, CO2 and acetate using the mass balance equations described by Conrad et al. [8], and assuming fractionation for the conversion of acetate to CH4 and of CO2 to CH4 by −21‰ and −70‰, respectively.
Molecular Analyses
Root materials were taken before incubation as control and after incubation in presence of buffer or without additional buffer. The roots were cut in 1-mm pieces, and 0.5 g of roots was transferred to the 2-mL tubes containing DNA extraction buffer. The tubes were shaken vigorously to detach the microorganisms from rice roots before DNA extraction. DNA extraction from root material, PCR amplification of partial 16S rRNA and mcrA genes, cloning of amplicons, and sequence analysis were performed as published before [3, 26]. Phylogenetic trees were calculated from aligned 16S rRNA and deduced McrA amino acid sequences with the ARB software package using neighbor-joining, FITCH, maximum-parsimony, and maximum-likelihood methods as described earlier [15, 26]. Tree topologies reconstructed with different algorithms were compared and, in case of inconsistent branching orders, adjusted to a consensus tree by introducing multifurcations. T-RFLP analyses was done with 5′,6-carboxyfluorescein (FAM)-labeled 16S rDNA [3] and mcrA amplicons [26] on an ABI 373 DNA sequencer in GeneScan mode, and relative signal intensities of T-RFs were inferred by peak area integration.
Accession Numbers
The sequences generated in this study were deposited in the GenBank database under the following accession numbers: mcrA clones, MRR-mcr1 to MRR-mcr75 under AY125601 to AY125675; 16S rDNA clones, MRR1 to MRR49 under AY125676 to AY125724.
Results
Methane Production in Rice Root Incubations
The production of CH4 was highest in rice roots suspended in carbonate buffer, followed by rice roots incubated in moist state (without additional buffer), and was lowest in rice roots suspended in phosphate buffer (Fig. 1A). Acetate accumulated in the rice root incubations (only measured in the buffered suspensions) to millimolar concentrations (Fig. 1B). Eventually, acetate decreased again, but not in the phosphate-buffered root incubations (Fig. 1B) or only after a longer delay (data not shown). These data confirm earlier observations [5, 6, 7, 23].
The relative contribution of H2/CO2-dependent and acetate-dependent methanogenesis to CH4 production was determined by following the δ13C in CO2, CH4 and acetate. The experimental procedure and a detailed data set obtained from anoxic root incubations (carbonate buffer) has already been published [8]. Similar data sets were obtained in the present study, both for incubations in carbonate and phosphate buffer. These experiments showed that the contribution of acetate-dependent methanogenesis to CH4 production was ~40% and was reduced to ~20% when the roots were incubated in phosphate buffer (Table 1).
Archaeal 16S rRNA and Methyl-Coenzyme M Reductase Genes Retrieved from Rice Roots
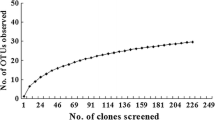
DNA was extracted from four different rice root incubations, and each extract was used to generate clone libraries of amplified partial 16S rRNA and mcrA genes. The rice roots were from (i) 1 day incubation without buffer, (ii) 1 day incubation in phosphate buffer, (iii) 20 days incubation in carbonate buffer, and (iv) 20 days incubation in phosphate buffer. The libraries were screened randomly by T-RFLP analysis to recover clones representing different T-RFs. A total of 70 and 80 clones were screened from the 16S rRNA and mcrA gene clone libraries, respectively. Gene fragments from 49 and 75 of these clones, respectively, were sequenced and phylogenetic trees were calculated, both for the 16S rRNA genes (Fig. 2) and deduced McrA amino acid sequences (Fig. 3). The remaining 21 and 5 clones of the 16S rRNA and mcrA gene clone libraries represented highly redundant T-RF and were not sequenced, since these T-RF were already sufficiently covered by other clones.
Phylogenetic dendrogram showing the relation of selected archaeal 16S rDNA sequences retrieved from anoxically incubated excised rice roots in this study (MRR clones, in bold) to those of cultivated Archaea and to other environmental clone sequences. The tree was reconstructed from partial (~800 bp) 16S rRNA genes using the maximum likelihood algorithm; the bar represents 10% sequence divergence. Nomenclature of uncultured archaeal lineages according to [3, 10, 18].
Phylogenetic dendrogram showing the relation of selected methanogen McrA and MrtA sequences retrieved from anoxically incubated excised rice roots in this study (MRR-mcr clones, in bold) to those of cultivated methanogens and to other environmental clone sequences. The tree was reconstructed from deduced amino acid (~150 positions) sequences using the FITCH algorithm. The full circle indicates a manually introduced multifurcation where inconsistent branching topologies for different sequence clusters were observed with different treeing algorithms; the bar represents 10% sequence divergence. Nomenclature of uncultured methanogenic lineages according to [16, 26].
The following methanogenic groups were detected in the clone libraries of both genes, i.e., methanogens within the Methanosarcinaceae, the Methanosaetaceae, the Methanomicrobiaceae, and the Methanobacteriaceae. The detection of methanomicrobial mcrA genes on the rice roots was a surprise, since we had failed to detect them in anoxic bulk soil earlier [26]. Furthermore, Rice cluster I [18] was detected by both 16S rRNA and mcrA genes, showing also the presence of this uncultured methanogenic lineage [26], which is assumed to produce CH4 from H2/CO2 [13]. The as yet unidentified cluster of rice field soil mcrA genes (e.g., MRR-mcr24) related to the Methanosarcinales retrieved with low frequency from anoxic bulk soil [26] was also detected on rice roots in this study, which might prove valuable in future attempts to clarify the systematic affiliation of these sequences.
In addition to these different methanogenic lineages, the rice roots also contained 16S rRNA gene sequences related to uncultivated archaeal clusters that are probably not methanogenic (Fig. 2). These were the Thermoplasmales-related Rice cluster III (Group E3 Euryarchaeota [10]), Rice cluster V [18], and the Rice clusters IV and VI (Group C2 freshwater Crenarchaeota and Group C1b terrestrial Crenarchaeota, respectively [10]).
Archaeal Population Dynamics in Rice Root Incubations
Time-dependent shifts within the methanogen populations were followed by 16S rDNA-based T-RFLP analysis, a tool shown to give a reliable and quantitative measure of 16S rRNA gene abundance in archaeal communities [29]. The assignment of different characteristic 16S rDNA and mcrA-based T-RF to distinct lineages of Archaea and methanogens is shown in Tables 2 and 3. All peaks detected in T-RFLP analysis of 16S rRNA genes from rice roots incubated in moist state without additional buffer for 1 day (Fig. 4) were assignable to defined archaeal lineages (Table 2), notably those at 83 bp (Methanomicrobiaceae and Rice cluster IV), 91 bp (mainly Methanobacteriaceae), 185 bp (mainly Methanosarcinaceae), 284 bp (Methanosaetaceae), and 392 bp (Methanomicrobiaceae and Rice cluster I). The affiliation of T-RF on roots is thus similar to that in soil [3, 27, 34], with the exception that the 392-bp T-RF on roots, besides Rice cluster I, also represents members of the Methanomicrobiaceae. We did not find any 16S rDNA clones with terminal restriction sites that were not detected in direct T-RFLP analyses of root DNA.
T-RFLP analysis of the methanogenic archaeal community on anoxically incubated excised rice roots, which were incubated in moist state for one day (without additional buffer). Electropherograms were generated targeting archaeal 16S rRNA (A) and methanogen mcrA/mrtA (B) genes. Characteristic T-RFs of defined lineages are indicated according to their length (bp). Mb, Methanobacteriaceae; Msr, Methanosarcinaceae.
Also the T-RF of mcrA genes retrieved from rice roots (Fig. 4) could all be assigned to distinct sequence clusters in the mcrA phylogeny (Table 3), notably those at 238 bp (Rice cluster I), 394 bp (Methanosarcinaceae), and 403, 470, and 503 bp (Methanobacteriaceae). Also, the 427 bp T-RF represented members of the Methanosarcinaceae. Although it was not observed for any mcrA clone in this study, it has previously been described for anoxic bulk soil [26]. The T-RFLP analysis of rice root DNA did not show any T-RF characteristic for the mcrA genes of the Methanomicrobiaceae (262 or 275 bp) and Methanosaetaceae (147 bp). Also, other mcrA T-RF predicted from the clone library were not detected in T-RFLP analysis of rice root DNA (e.g., 241, 347, 406, 409 bp, Table 3).
Root DNA was extracted from successive time points in parallel to the CH4 production measurements shown in Fig. 1 and analyzed by 16S rDNA- and mcrA-targeted T-RFLP fingerprinting. Because of the highly degenerate mcrA-targeted primer set, this assay could not be utilized for quantitative community analyses [29]. The relative frequency of characteristic 16S rDNA T-RF of different groups of Archaea changed significantly with time and condition of the root incubations (Fig. 5). Initially, all incubations exhibited only a low frequency (up to 11%) of the T-RF characteristic for acetoclastic methanogens (Methanosarcinaceae and Methanosaetaceae), whereas the frequency of T-RF characteristic for H2/CO2-dependent methanogens (Methanomicrobiaceae and Rice cluster I, in particular) was high (up to 42 and 35%, respectively). With progress of incubation, the acetoclastic Methanosarcinaceae (but not Methanosaetaceae) became more frequent (up to 41% on day 6 in carbonate buffer). This increase was strongly delayed in phosphate-buffered incubations (Fig. 5); here the abundance of the methanosarcinal T-RF only increased strongly after 20 days. The relative increase in abundance of Methanosarcinaceae T-RF consistently occurred just before the acetate concentrations in the incubations started to decrease (Fig. 1), indicating increased acetate consumption by a growing population of acetoclastic methanogens. After 2 days, the T-RF of 392 bp (representing Rice cluster I and Methanomicrobiaceae) exhibited a relatively higher frequency in the carbonate-buffered than in the other root incubations, which in turn exhibited a relatively higher frequency of the T-RF at 83 bp (characteristic for Methanomicrobiaceae and crenarchaeotal Rice cluster IV). The functional consequences of these population shifts, however, remain to be elucidated.
Time course of archaeal population dynamics on anoxically incubated excised rice roots as determined by 16S rDNA targeted T-RFLP analysis. Shown is the contribution percentage of individual T-RF to the total integrated fluorescence; percentages above 8% are given. Minor T-RF have been combined as “diverse”. Mm, Methanomicrobiaceae; Msa, Methanosaetaceae; Msr, Methanosarcinaceae; Mb, Methanobacteriaceae; RC, Rice cluster.
Discussion
Methanogenesis becomes the dominant pathway for anaerobic degradation of organic matter when environmental concentrations of inorganic electron acceptors such as nitrate, sulfate, or ferric iron are low. Otherwise, nitrate reducers, sulfate reducers, or iron reducers compete successfully for the methanogenic substrates H2 and acetate and thus suppress methanogenesis. The dynamic interplay between inorganic electron acceptors and methanogenic pathways has been shown in several vegetated methanogenic environments, e.g., rice fields [22, 37] or peatlands [1, 36]. Recently, it was shown that nitrate and sulfate affect not only the activity but also the community structure of methanogenic archaea using rice roots as model system [35]. However, little is known about the factors that influence the structure within the methanogenic community, i.e., the hydrogenotrophic and acetoclastic methanogenic archaea.
Our results establish a link between functionality and structure of a methanogenic population, i.e, the accumulation of acetate in rice root incubations leading to an increased relative abundance of the acetoclastic Methanosarcinaceae and a subsequently enhanced consumption of acetate. The pathway of CH4 formation on the roots consequently shifted to a higher contribution of acetate as methanogenic precursor. The establishment of an abundant Methanosarcina population was obviously crucial for this pathway of CH4 formation. High phosphate concentrations, which delayed the development of methanosarcinal populations, also delayed acetoclastic methanogenesis. Inhibition of acetoclastic methanogenesis by high phosphate has been observed before [6, 7], but it was unknown whether the effect was only due to inhibition of enzyme activity or also to inhibition of growth. Obviously, proliferation of the acetoclastic Methanosarcina root population was negatively affected by high phosphate levels.
There are only few studies in which changes in methanogenic community function were successfully correlated to changes in community structure, and rarely were the observed community changes as pronounced as in the present study. In rice field soil incubations, for example, the relative abundance of the Methanosarcinaceae change only to small extent during the course of anoxic incubation [27], and an increased acetoclastic methanogenesis is mainly accompanied by increasing rRNA expression levels of existing Methanosarcina and Methanosaeta populations [28]. Brief exposure of methanogenic rice field soil to 50°C results in a pronounced inhibition of acetoclastic methanogenesis but only in a relatively small change in the acetoclastic methanogenic populations [41]. Prolonged incubation at 50°C, on the other hand, results in a drastic change in the methanogenic microbial community, resulting in the dominance of Rice cluster I and the production of CH4 from H2/CO2 exclusively [13].
Interestingly, we observed a population shift for the acetoclastic Methanosarcinaceae, but not the Methanosaetaceae. In fact, Methanosaetaceae were hardly detectable at all on the rice roots, in accordance with earlier studies [18]. We assume that Methanosarcinaceae predominated, since they can grow faster than Methanosaetaceae as long as acetate is available at relatively high concentrations. Methanosarcina spp. have a higher threshold and K m value for acetate compared to Methanosaeta spp., due to the different mechanisms of acetate activation, i.e., by acetate kinase/phosphotransacetylase versus acetyl-CoA synthetase, respectively [20]. Hence, it is plausible to find Methanosaeta spp. mainly in anoxic bulk soil, especially at low acetate concentrations [14], but not on rice roots with high acetate concentrations. Acetoclastic methanogens on decomposing rice straw, which also shows relatively high acetate concentrations, also consist mainly of Methanosarcina spp. [40].
In addition to the acetoclastic methanogens, there may also have been a change in the hydrogenotrophic methanogenic populations. In particular, we observed a relatively higher frequency of the 392 bp T-RF (characteristic for Rice cluster I and Methanomicrobiaceae) in the carbonate-buffered root incubations, whereas the other root incubations exhibited a relatively higher frequency of the 83 bp T-RF (characteristic for Methanomicrobiaceae and the crenarchaeotal Rice cluster IV). Since both T-RF were characteristic for hydrogenotrophic methanogens, it is unclear what the functional meaning of these different community structures is.
The 16S rDNA sequences retrieved from rice roots were similar to those from rice field soil sampled in Vercelli (Italy) [3, 27], Gapan (The Philippines), and Shenyang (China) [34]. Root and soil populations also showed similar characteristic T-RF for distinct phylogenetic groups. The only exception was the 392 bp T-RF, which in soil almost exclusively represents Rice cluster I methanogens, but on roots equally represents members of the Methanomicrobiaceae. This is likely to be explained simply by a much higher frequency of these methanomicrobial methanogens in rice root incubations (~10% of both 16S rDNA and mcrA clones in this study) compared to anoxic bulk soil (between 1 and 5% of 16S rDNA clones [3, 27]). The same assignment of the 392 bp T-RF to members of the Methanomicrobiaceae has previously been described in the methanogenic sediments of Lake Kinneret, where rice cluster I methanogens apparently do not exist [32]. Hence, it is mandatory to create a 16S rDNA sequence library for each environment, in which methanogenic community structure is to be characterized by T-RFLP analysis, in order to correctly assign T-RF to phylogenetic groups.
Most of the mcrA sequences retrieved from rice roots were similar to those from rice field soil [26]. In contrast to soil, however, we also detected members of the Methanomicrobiaceae on the rice roots. Recently, mcrA genes related to the Methanomicrobiales were retrieved from a Finnish fen [16] and a landfill [30] with different primer systems. Because of the degenerate primer system, T-RFLP analysis of mcrA gene does not result in a quantitative recovery of the genes of different phylogenetic lineages [29]. This may explain why we detected methanomicrobial mcrA genes in the clone library in the present study, but no corresponding T-RFs in the mcrA-targeted T-RFLP analysis. The detection of methanomicrobial mcrA sequences in our present study proves that the primer system utilized here [38] is capable of detecting these microorganisms. The fact that we did not detect such genes in anoxic bulk soil earlier [26] therefore was probably due to a low abundance of methanomicrobial methanogens rather than to an insufficient utility of the mcrA primers designed by Springer et al. [38] to detect all orders of methanogens, as suggested by Luton and co-workers [30].
In conclusion, our data clearly show that the structure of the methanogenic community on rice roots changed with incubation time and conditions, which corresponded to a functional change in the dominance of acetoclastic versus hydrogenotrophic methanogenesis. The predominance of Methanosarcina spp. versus Methanosaeta spp. was consistent with the prevailing acetate concentrations. The community structures on the basis of 16S rRNA and mcrA genes were mostly consistent. In future, more detailed analyses of the Methanomicrobiaceae, Methanobacteriaceae, and Rice cluster I may provide valuable clues for structural and functional relationships among the hydrogenotrophic methanogenic community.
References
GB Avery RD Shannon JR White CS Martens MJ Alperin (1999) ArticleTitleEffect of seasonal changes in the pathways of methanogenesis on the δ13C values of pore water methane in a Michigan peatland. Global Biogeochem Cycles 13 475–484 Occurrence Handle1:CAS:528:DyaK1MXkslOrurs%3D
KA Bidle M Kastner DH Bartlett (1999) ArticleTitleA phylogenetic analysis of microbial communities associated with methane hydrate containing marine fluids and sediments in the Cascadia Margin (ODP site 892b). FEMS Microbiol Lett 177 101–108 Occurrence Handle1:CAS:528:DyaK1MXls1SmsbY%3D Occurrence Handle10436927
KJ Chin T Lukow R Conrad (1999) ArticleTitleEffect of temperature on structure and function of the methanogenic archaeal community in an anoxic rice field soil. Appl Environ Microbiol 65 2341–2349
R Conrad P Frenzel (2002) Flooded soils. G Bitton (Eds) Encyclopedia of Environmental Microbiology. John Wiley & Sons New York 1316–1333
R Conrad M Klose (1999) ArticleTitleAnaerobic conversion of carbon dioxide to methane, acetate and propionate on washed rice roots. FEMS Microbiol Ecol 30 147–155 Occurrence Handle10.1016/S0168-6496(99)00048-3 Occurrence Handle1:CAS:528:DyaK1MXmt1emurs%3D Occurrence Handle10508939
R Conrad M Klose (2000) ArticleTitleSelective inhibition of reactions involved in methanogenesis and fatty acid production on rice roots. FEMS Microbiol Ecol 34 27–34 Occurrence Handle10.1016/S0168-6496(00)00071-4 Occurrence Handle1:CAS:528:DC%2BD3cXosVyrt7c%3D Occurrence Handle11053733
R Conrad M Klose P Claus (2000) ArticleTitlePhosphate inhibits acetotrophic methanogenesis on rice roots. Appl Environ Microbiol 66 828–831
R Conrad M Klose P Claus (2002) ArticleTitlePathway of CH4 formation in anoxic rice field soil and rice roots determined by 13C-stable isotope fractionation. Chemosphere 47 797–806 Occurrence Handle10.1016/S0045-6535(02)00120-0 Occurrence Handle1:CAS:528:DC%2BD38XjsVSqsrg%3D Occurrence Handle12079075
S Dannenberg R Conrad (1999) ArticleTitleEffect of rice plants on methane production and rhizospheric metabolism in paddy soil. Biogeochemistry 45 53–71 Occurrence Handle10.1023/A:1006085605184
EF DeLong NR Pace (2001) ArticleTitleEnvironmental diversity of Bacteria and Archaea. Syst Biol 50 470–478 Occurrence Handle10.1080/106351501750435040 Occurrence Handle1:STN:280:DC%2BD38zntVOmsw%3D%3D Occurrence Handle12116647
C Edwards et al. (1998) ArticleTitleMicrobiological processes in the terrestrial carbon cycle—methane cycling in peat. Atmos Environ 32 3247–3255 Occurrence Handle10.1016/S1352-2310(98)00107-1 Occurrence Handle1:CAS:528:DyaK1cXltFOhtbo%3D
U Ermler W Grabarse S Shima M Goubeaud RK Thauer (1997) ArticleTitleCrystal structure of methyl coenzyme M reductase—the key enzyme of biological methane formation. Science 278 1457–1462 Occurrence Handle9367957
A Fey KJ Chin R Conrad (2001) ArticleTitleThermophilic methanogens in rice field soil. Environ Microbiol 3 295–303 Occurrence Handle10.1046/j.1462-2920.2001.00195.x Occurrence Handle1:CAS:528:DC%2BD3MXlt1eht70%3D Occurrence Handle11422316
A Fey R Conrad (2000) ArticleTitleEffect of temperature on carbon and electron flow and on the archaeal community in methanogenic rice field soil. Appl Environ Microbiol 66 4790–4797 Occurrence Handle11055925
MW Friedrich (2002) ArticleTitlePhylogenetic analysis reveals multiple lateral transfers of adenosine-5′-phosphosulfate reductase genes among sulfate-reducing microorganisms. J Bacteriol 184 278–289 Occurrence Handle1:CAS:528:DC%2BD3MXpt1Cmtbk%3D Occurrence Handle11741869
PE Galand S Saarnio H Fritze K Yrjälä (2002) ArticleTitleDepth related diversity of methanogen Archaea in Finnish oligotrophic fen. FEMS Microbiol Ecol 42 441–449 Occurrence Handle10.1016/S0168-6496(02)00381-1 Occurrence Handle1:CAS:528:DC%2BD38XoslGiur0%3D
R Grosskopf PH Janssen W Liesack (1998) ArticleTitleDiversity and structure of the methanogenic community in anoxic rice paddy soil microcosms as examined by cultivation and direct 16S rRNA gene sequence retrieval. Appl Environ Microbiol 64 960–969 Occurrence Handle1:CAS:528:DyaK1cXhs1art74%3D Occurrence Handle9501436
R Grosskopf S Stubner W Liesack (1998) ArticleTitleNovel euryarchaeotal lineages detected on rice roots and in the anoxic bulk soil of flooded rice microcosms. Appl Environ Microbiol 64 4983–4989 Occurrence Handle1:CAS:528:DyaK1cXnvF2htL8%3D Occurrence Handle9835592
BA Hales C Edwards DA Ritchie G Hall RW Pickup JR Saunders (1996) ArticleTitleIsolation and identification of methanogen-specific DNA from blanket bog feat by PCR amplification and sequence analysis. Appl Environ Microbiol 62 668–675 Occurrence Handle1:CAS:528:DyaK28XovVCqtw%3D%3D Occurrence Handle8593069
MSM Jetten AJM Stams AJB Zehnder (1992) ArticleTitleMethanogenesis from acetate—a comparison of the acetate metabolism in Methanothrix soehngenii and Methanosarcina spp. FEMS Microbiol Rev 88 181–197 Occurrence Handle1:CAS:528:DyaK38Xlt1Ort74%3D
M Krüger G Eller R Conrad P Frenzel (2002) ArticleTitleSeasonal variation in pathways of CH4 production and in CH4 oxidation in rice fields determined by stable carbon isotopes and specific inhibitors. Global Change Biol 8 265–280 Occurrence Handle10.1046/j.1365-2486.2002.00476.x
M Krüger P Frenzel R Conrad (2001) ArticleTitleMicrobial processes influencing methane emission from rice fields. Global Change Biol 7 49–63 Occurrence Handle10.1046/j.1365-2486.2001.00395.x
S Lehmann-Richter R Grosskopf W Liesack P Frenzel R Conrad (1999) ArticleTitleMethanogenic archaea and CO2-dependent methanogenesis on washed rice roots. Environ Microbiol 1 159–166 Occurrence Handle10.1046/j.1462-2920.1999.00019.x Occurrence Handle1:CAS:528:DyaK1MXktFykuro%3D Occurrence Handle11207731
J Lelieveld PJ Crutzen FJ Dentener (1998) ArticleTitleChanging concentrations, lifetime and climate forcing of atmospheric methane. Tellus 50B 128–150 Occurrence Handle1:CAS:528:DyaK1cXivFertbk%3D
D Lloyd et al. (1998) ArticleTitleMicro-ecology of peat—minimally invasive analysis using confocal laser scanning microscopy, membrane inlet mass spectrometry, and PCR amplification of methanogen-specific gene sequences. FEMS Microbiol Ecol 25 179–188 Occurrence Handle10.1016/S0168-6496(97)00094-9 Occurrence Handle1:CAS:528:DyaK1cXht1WgsbY%3D
T Lueders KJ Chin R Conrad M Friedrich (2001) ArticleTitleMolecular analyses of methyl-coenzyme M reductase alpha-subunit (mcrA) genes in rice field soil and enrichment cultures reveal the methanogenic phenotype of a novel archaeal lineage. Environ Microbiol 3 194–204 Occurrence Handle10.1046/j.1462-2920.2001.00179.x Occurrence Handle11321536
T Lueders M Friedrich (2000) ArticleTitleArchaeal population dynamics during sequential reduction processes in rice field soil. Appl Environ Microbiol 66 2732–2742 Occurrence Handle10877762
T Lueders MW Friedrich (2002) ArticleTitleEffects of amendment with ferrihydrite and gypsum on the structure and activity of methanogenic populations in rice field soil. Appl Environ Microbiol 68 2484–2494 Occurrence Handle10.1128/AEM.68.5.2484-2494.2002 Occurrence Handle1:CAS:528:DC%2BD38XjsFGqtbg%3D Occurrence Handle11976125
T Lueders MW Friedrich (2003) ArticleTitleEvaluation of PCR amplification bias by terminal restriction fragment length polymorphism analysis of small-subunit rRNA and mcrA genes by using defined template mixtures of methanogenic oure cultures and soil DNA extracts. Appl Environ Microbiol 69 320–326 Occurrence Handle10.1128/AEM.69.1.320-326.2003 Occurrence Handle1:CAS:528:DC%2BD3sXkvValsw%3D%3D Occurrence Handle12514011
PE Luton JM Wayne RJ Sharp PW Riley (2002) ArticleTitleThe mcrA gene as an alternative to 16S rRNA in the phylogenetic analysis of methanogen populations in landfill. Microbiology 148 3521–3530 Occurrence Handle1:CAS:528:DC%2BD38Xpt1ekt7o%3D Occurrence Handle12427943
D Nercessian M Upton D Lloyd C Edwards (1999) ArticleTitlePhylogenetic analysis of peat bog methanogen populations. FEMS Microbiol Lett 173 425–429 Occurrence Handle10.1016/S0378-1097(99)00073-7 Occurrence Handle1:CAS:528:DyaK1MXitFWrtbs%3D
B Nüsslein KJ Chin W Eckert R Conrad (2001) ArticleTitleEvidence for anaerobic syntrophic acetate oxidation during methane production in the profundal sediment of subtropical Lake Kinneret (Israel). Environ Microbiol 3 460–470 Occurrence Handle10.1046/j.1462-2920.2001.00215.x Occurrence Handle11553236
M Ohkuma S Noda K Horikoshi T Kudo (1995) ArticleTitlePhylogeny of symbiotic methanogens in the gut of the termite Reticulitermes speratus. FEMS Microbiol Lett 134 45–50 Occurrence Handle10.1016/0378-1097(95)00379-J Occurrence Handle1:CAS:528:DyaK2MXps1WqsLw%3D Occurrence Handle8593954
B Ramakrishnan T Lueders PF Dunfield R Conrad MW Friedrich (2001) ArticleTitleArchaeal community structures in rice soils from different geographical regions before and after initiation of methane production. FEMS Microbiol Ecol 37 175–186 Occurrence Handle10.1016/S0168-6496(01)00158-1 Occurrence Handle1:CAS:528:DC%2BD3MXnslCisbg%3D
D Scheid S Stubner R Conrad (2003) ArticleTitleEffects of nitrate- and sulfate-amendment on the methanogenic populations in rice roots incubations. FEMS Microbiol Ecol 43 309–315 Occurrence Handle10.1016/S0168-6496(02)00432-4 Occurrence Handle1:CAS:528:DC%2BD3sXhvFeqtr0%3D
RD Shannon JR White (1996) ArticleTitleThe effects of spatial and temporal variations in acetate and sulfate on methane cycling in two Michigan peatlands. Limnol Oceanogr 41 435–443 Occurrence Handle1:CAS:528:DyaK28XkvFKht7w%3D
LK Sigren ST Lewis FM Fisher RL Sass (1997) ArticleTitleEffects of field drainage on soil parameters related to methane production and emission from rice paddies. Global Biogeochem Cycles 11 151–162 Occurrence Handle1:CAS:528:DyaK2sXjslSmsr8%3D
E Springer MS Sachs CR Woese DR Boone (1995) ArticleTitlePartial gene sequences for the A subunit of methyl-coenzyme M reductase (mcrI) as a phylogenetic tool for the family Methanosarcinaceae. Int J Syst Bacteriol 45 554–559
A Watanabe T Takeda M Kimura (1999) ArticleTitleEvaluation of origins of CH4 carbon emitted from rice paddies. J Geophys Res 104 23623–23629 Occurrence Handle10.1029/1999JD900467 Occurrence Handle1:CAS:528:DyaK1MXntleht78%3D
S Weber T Lueders MW Friedrich R Conrad (2001) ArticleTitleMethanogenic populations involved in the degradation of rice straw in anoxic paddy soil. FEMS Microbiol Ecol 38 11–20 Occurrence Handle10.1016/S0168-6496(01)00168-4 Occurrence Handle1:CAS:528:DC%2BD38XitVyi
XL Wu KJ Chin R Conrad (2002) ArticleTitleEffect of temperature stress on structure and function of the methanogenic archaeal community in a rice field soil. FEMS Microbiol Ecol 39 211–218 Occurrence Handle10.1016/S0168-6496(01)00216-1 Occurrence Handle1:CAS:528:DC%2BD38XjsFOjurw%3D
Acknowledgements
We thank Peter Claus for analysis of 13C-isotopic data and the Fonds der Chemischen Industrie, Germany, for financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chin, KJ., Lueders, T., Friedrich, M. et al. Archaeal Community Structure and Pathway of Methane Formation on Rice Roots . Microb Ecol 47, 59–67 (2004). https://doi.org/10.1007/s00248-003-2014-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-003-2014-7