Abstract
Serpentine soils are characterized by high levels of heavy metals (Ni, Co, Cr), and low levels of important plant nutrients (P, Ca, N). Because of these inhospitable edaphic conditions, serpentine soils are typically home to a very specialized flora including endemic species as the nickel hyperaccumulator Alyssum bertolonii. Although much is known about the serpentine flora, few researches have investigated the bacterial communities of serpentine areas. In the present study bacterial communities were sampled at various distances from A. bertolonii roots in three different serpentine areas and their genetic diversity was assessed by terminal restriction fragment length polymorphism (T-RFLP) analysis. The obtained results indicated the occurrence of a high genetic diversity and heterogeneity of the bacterial communities present in the different serpentine areas. Moreover, TRFs (terminal restriction fragments) common to all the investigated A. bertolonii rhizosphere samples were found. A new cloning strategy was applied to 27 TRFs that were sequenced and taxonomically interpreted as mainly belonging to Gram-positive and α-Proteobacteria representatives. In particular, cloned TRFs which discriminated between rhizosphere and soil samples were mainly interpreted as belonging to Proteobacteria representatives.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Serpentine (ultramafic) outcrops are distributed all over the world and, for their natural geological origin, are characterized by high levels of cobalt, chromium, and especially nickel [5]. The vegetation adapted to survive in these soils [22] can include the so-called nickel-hyperaccumulating plants [3] that concentrate metal in stems and leaves to levels higher than the substrate concentration and far in excess from any physiological requirement (more than 1000 μg g−1 shoot dry matter). Alyssum bertolonii Desv. (Brassicaceae) is a nickel hyperaccumulator, endemic to the serpentine outcrops of Central Italy [51], belonging to a genus which recently has stirred new and increasing attention due to its practical application for phytoremediation [8, 41].
The study of metal-hyperaccumulating plant effects on soil microorganisms is an important topic. Microorganisms can have a great impact on the performances of revegetation of polluted soils [37]. Studies on serpentine soils may provide new insight into bacterial diversity under unfavorable conditions, new isolates, and probably new genetic information on heavy-metal resistance, which could be exploited.
Bacteria present in serpentine soils and their interaction with hyperaccumulating plants have focused the attention of several investigators in past years [9, 15, 16, 26, 31, 36, 39, 44, 52, 53]. These authors found that serpentine bacterial communities tolerated spiking of metals, such as nickel, more than those from unpolluted soils and that the presence of hyperaccumulating plants as Sebertia acuminata, Thlaspi caerulescens, and Alyssum bertolonii led to an increase in metal-resistant bacteria proportion in the soil samples collected near the plants. Moreover, metal-resistant bacteria present in the plant rhizosphere may play an important role in regulating the availability of metal for the plant [46, 53].
Nickel-resistant rhizosphere bacteria have recently been shown to increase nickel uptake into the shoots of the nickel hyperaccumulator Alyssum murale [1].
DNA-based techniques have become a powerful tool for studying diversity and composition of soil bacterial communities in cultivation-independent ways [49]. One of the most important methods for soil bacteria surveys is the analysis of a 16SrDNA clone library [14, 17]. However, because of the complexity of soil communities and the effort required for this type of analysis, 16SrDNA clone libraries have been restricted to the analysis of a single or a few samples in an environment.
Terminal Restriction Fragment Length Polymorphism (T-RFLP) is a recently introduced PCR-based tool for studying the genetic diversity of bacterial communities [20, 25, 28]. T-RFLP analysis is based on the detection of a single restriction fragment in each sequence amplified directly from the environmental sample of DNA and is capable of surveying dominant members comprising at least 1% of the total community [10]. In recent years T-RFLP has been widely used for the analysis of bacterial communities in different conditions (for examples, see [11, 13, 27, 33, 34, 35, 40]) and to assess spatial heterogeneity of bacterial communities in soils [23], sediments, and water environments [4, 7, 21, 43].
The aim of this work was to investigate the genetic diversity of bacterial communities present in serpentine soils and in the rhizosphere of the nickel-hyperaccumulating plant A. bertolonii by using T-RFLP analysis.
Materials and Methods
Sampling Procedure and Soil Paramters
Soil samples (at ~10 cm of depth) of ~200 g were taken during winter 2001/2002 with a clean steel spatula sterilized with ethanol. Samples were shaken and kept in sterile plastic boxes at 4°C for a few hours before DNA extraction. Samples were collected from three serpentine outcrops, located in Tuscany (Italy): Galceti, Impruneta, and Pieve Santo Stefano. The three localities had different plant coverage of the serpentine outcrop; in particular, the Galceti outcrop was a rocky hill covered by few pines (Pinus maritima), whereas the Impruneta outcrop was covered by a wood of cypress (Cupressus sempervirens), and Pieve Santo Stefano outcrop was in a cliff covered by a few grasses. Soil chemical characteristics were determined on soil samples taken at 10 cm from the stem of Alyssum bertolonii following the methods described by Sparks [47]. For the metal analysis, samples were dried at 80°C for 1 day, weighed, and digested by wet ashing with HNO3:HClO4 (5:2 v/v). The soil metal extractable fraction was determined following the method of Escarré et al. [12]. Metal concentration in the samples was determined by atomic absorption spectroscopy (PE Biosystems 370, PerkinElmer, Norwalk, CT, USA). All the measurements were performed in triplicate. In each location three soil samples were taken at various distances from the hyperaccumulating plant Alyssum bertolonii Desv. Soil samples were labeled as A, B, and C portions to indicate absence of the plant (soil free from plants in an area of 50 cm of diameter), presence of the plant at a distance of 10 cm, and plant shoot at 5 cm, respectively. No plant species other than A. bertolonii were present in an area of 1 m of diameter from the sampling site. For the collection of the rhizosphere soil particles, the main root of the plant with a diameter of ~0.2 cm was cut from 5 cm to 10 cm under the collar in sterile conditions and the larger soil particles were shaken away. These rhizosphere samples were labeled as D portion.
DNA Extraction and PCR Conditions
DNA was extracted with the Fast Prep DNA Kit for Soil (BIO101, Qbiogene, Carlsbad, CA, USA) following manufacturer’s instructions. For the rhizosphere samples, the roots were mixed in a sterile tube with 40 mL of 10 mM MgSO4 and shaken for 2 h on a slowly rotating plate; then soil particles were collected by centrifugation and used for DNA extraction. Extractions were performed in duplicate.
16S rDNA was amplified in a 50-μL volume with 20 ng of template DNA and 2 U of Taq DNA polymerase (Dynazyme II, Finnzyme, Espoo, Finland) using 27f primer labeled with TET (4,7,2’,7’-tetrachloro-6-carboxyfluorescein) and FAM (6-carboxyfluorescein) and 1495r primer [24], as described in Mengoni et al. [31]. The use of two fluorescent dyes allowed two different restriction digestions to run together in the same capillary, reducing cost and time of the analysis. PCR reactions were repeated three times on each DNA sample (technical replicate).
16S rDNA T-RFLP
The amplified products were purified with a Qiaquick PCR purification kit (Qiagen Inc. Chatsworth, CA, USA), and 600 ng of amplified 16SrDNA was digested with 20 U of MspI, HhaI, RsaI, or HinfI (New England Biolabls, Beverly, MA, USA) for 3 h at 37°C. A 200-ng aliquot of the digested products was resolved by capillary electrophoresis on an ABI310 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA) using TAMRA 500 (Applied Biosystems) as size standard for GenScan (Applied Biosystems) analysis. Fragment sizes from 35 to 500 bp were considered for profile analysis.
Analysis of T-RFLP Profiles
By comparison of T-RFLP profiles from the duplicate DNA samples and the three technical replicates a derivative profile was created following the same criteria used by Dunbar et al. [10]. Only fragments with fluorescence intensity ≥50 arbitrary units of fluorescence were considered, and the total DNA quantity represented by each profile was checked by summing all peak intensities. Alignment of the profiles was performed directly on the output table of the software GenScan and ±0.5 bp was allowed to discriminate peaks of consecutive sizes. Derivative T-RFLP profiles of the different enzymes were then combined together and a binary vector, in which presence or absence of peaks were scored as strings of ones or zeros, was prepared. The vectors of binary profiles of each soil portion and location were then compared to compute the community similarity values using the Dice index as implemented in the software NTSYSpc 2.0 [38], The matrix of Dice similarity values was then used for construction of a UPGMA dendrogram and for Principal Component Analysis using the modules present in NTSYSpc 2.0 [38].
Cloning and Sequencing of TRFs
TRFs of interest were cloned by means of adapter as described in Mengoni et al. [32]. The sequences of double strand adapter and selection primer for the HhaI restriction site were: 5′-GAGCATCTGACGCATGGTTAA-3′, 5′-CCATGCGTCAGATGCTCCG-3′, selection primer: 5′-CCATGCGTCAGATGCTCCGC-3′; and for the HinfI restriction site, 5′-ANTCTCGTAGACTGCGTACC-3′, 5′-GGGGGGTACGCAGTCTACGAG-3′, selection primer: 5′-GGGGGGTACGCAGTCTACGAG-3′ Clones were sequenced from M13 forward primer external to the insertion site of cloning plasmid [32] using the DYEnamic ET Terminator Cycle Sequencing Kit (Amersham Biosciences Europe GmbH, Freiburg, Germany), and sequences were run on an ABI310 genetic analyzer (Applied Biosystems). Sequences were compared with those present in the GenBank database by using the BLASTn tool [2] to obtain similarity matches.
Results
Metal Concentration and Soil Characteristics
The three serpentine locations showed high concentrations of heavy metals as Ni, Co and Cr and a higher level of Mg in relation to Ca (Table 1). The soils had very similar and slightly basic pH values. The CEC values were similar, too, whereas the organic matter concentration varied from 1.66% (Pieve Santo Stefano) to 8.26% (Galceti).
Ribotype Richness of the Soil Samples
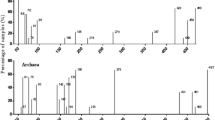
The ribotype richness of serpentine soil samples was estimated determining the number of terminal restriction fragments (TRFs). The four restriction endonucleases applied to 16S rDNAs amplicons yielded a total of 138 different peaks or TRFs. MspI produced the highest number of peaks (48), while HhaI gave the lowest (27); the other two enzymes, RsaI and HinfI, yielded 35 and 28 peaks, respectively. Figure 1 shows the overall trends of the total number of TRFs for the three localities. Impruneta and Galceti showed slightly more peaks in soil portions B, C and D than in A.
Total number of TRFs derived from amplified 16S rDNA genes within the soil portions in the different localities. Plotted values represent the sums of the number of TRFs from the restriction digestions with four endonucleases (MspI, RsaI, HhaI, Hnf I) obtained in soil portions A, B, C, and D for the localities of Impruneta (I), Galceti (G), and Pieve Santo Stefano (P).
Similarities of T-RFLP Community Profiles
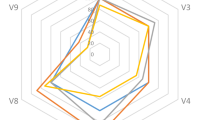
The similarity of the 12 samples based on UPGMA analysis of T-RFLP profiles is shown in Fig. 2. Samples from the same locality did not form homogeneous clusters. Comparison of the samples, with respect to the soil portion they were derived from, revealed that soil portions A and B of the different localities were the most variable (average similarity: 0.71 and 0.74, respectively). Soil portions C were more similar (average similarity: 0.82) and rhizosphere samples (portions D) were the most similar (average similarity: 0.91), forming a homogeneous cluster. Principal component analysis (Fig. 3) illustrated, on the first two components (explaining the 42.6% of total variance), a similar pattern with the samples from the same locality being interspersed, while the same soil portions (especially D and C) formed homogeneous groups. In particular the first component mainly separated the free soil samples (A) from samples nearer to the plant (B, C, and D). The second component separated the rhizosphere samples (D) from B and C samples.
Taxonomic Interpretation of TRFs and Dominant Eubacterial Groups
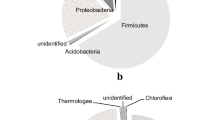
Among the 138 TRFs obtained, 27 were selected on the basis of their distribution. In particular, eight TRFs present in all samples, seven TRFs exclusive of rhizosphere samples, and 12 distributed in more than one soil portion were chosen for the analysis. These TRFs had a fluorescence intensity >200 fluorescence units in our experimental conditions. Nineteen TRFs derived from MspI digestion, six from HhaI digestion, and two from HinfI digestion were sequenced using a previously described procedure [29], and the sequences compared to sequences present in GenBank to obtain a taxonomic interpretation of T-RFLP profiles. Sequences are presented in the Appendix. The results of the analysis are shown in Table 2. Most of the sequenced TRFs showed similarities with Gram-positive bacteria (10) and α-Proteobacteria (6). The distribution of the sequenced TRFs indicated that most of the TRFs common to all portions could be interpreted as belonging to non-Proteobacteria representatives, while most of TRFs exclusive of all portions D (rhizosphere) were interpreted as belonging to Proteobacteria representatives, mainly from the Alpha subdivision.
Discussion
The studies on bacterial communities present in heavy-metal-contaminated ecosystems have mainly focused on the effect of anthropogenic metal pollution in shaping the genetic structure of the community [42]. In recent years the attention has been focused on metal-resistant bacterial communities present in the serpentine soil naturally rich in heavy metal and on bacteria associated with the roots of serpentine-endemic metal hyperaccumulating plants [9, 15, 26, 31, 44, 52, 53]. In this work, by using the T-RFLP technique, we provided a first description of the total bacterial communities isolated from serpentine soils along a distance gradient from the nickel-hyperaccumulating plant Alyssum bertolonii. In particular, the same rationale and the same soil samples which we previously used for the analysis of nickel-resistant bacteria [31] were investigated to possibly correlate the results obtained from the previous analysis with those from the total community profiles.
Chemical characteristics of the sampled soils were similar to those of other serpentine soils [5].
A high genetic diversity was detected, especially in the samplings at 10 cm (B) and 5 cm (C) distant from the plant. Actually, average numbers of TRFs for the four different enzymes were consistent, or slightly less, with numbers reported for forest soil [10] and marine sediments [4, 50], but higher than numbers reported for copper-contaminated lake sediments [21]. The number of TRFs in Galceti and Impruneta samples showed a slight increase from A to D, particularly from A to B, suggesting an effect of the plant in increasing the microbial diversity. A plant-driven effect on bacterial community was also suggested by the comparison of T-RFLP profiles. Both UPGMA and principal component analyses indicated plant, more than locality, as the main factor in shaping the community profile. In particular, both first and second principal components divided the overall variance mainly in relation with the proximity to the plant. Furthermore, it was possible to recognize a gradient of similarities among samples belonging to different localities, from the most different soil portions A, to the more similar B and C portions, to the rhizosphere in which differences between samples of the three localities appeared very low. The effect of plant roots on bacterial communities is a well-established topic [19]. Plant rhizosphere has been recognized by several authors [18, 30, 45] as harboring a complex and differentiated bacterial community. Recently the rhizosphere bacteria of the Zn hyperaccumulator Thlaspi caerulescens have stirred much attention about their possible role in increasing metal accumulation by the plant. In particular, two papers [9, 26] reported that T. caerulescens rhizosphere was rich in metal-resistant bacteria, also exhibiting multiple heavy-metal resistances. Recently a cultivation-independent approach was used for the analysis of T. caerulescens rhizosphere (gremion) showing a high proportion of Actinobacteria in the metabolically active bacterial community. Finally, Whiting et al. [53] demonstrated that the addition of rhizosphere bacteria to axenic plant cultures increased plant Zn uptake. For the nickel hyperaccumulator A. bertolonii an effect of the rhizosphere was previously shown for the culturable fraction of microbial community [31], showing that as well as for T. caerulescens, a high number of metal-resistant isolates harbored multiple metal resistances (Ni, Co, Cr). Recently, bacterial strains isolated from the rhizosphere of the congeneric nickel hyperaccumulator Alyssum murale have been shown to increase plant nickel uptake up to 32% [1]. These strains seemed to facilitate the release of nickel from the nonlabile phase in the soil (by organic acids or siderophore production and phosphate solubilization), thus enhancing the availability of nickel to A. murale.
T-RFLP analysis has mainly been applied as a tool for the comparison of microbial communities [20, 25, 28], limiting the taxonomic interpretation of the profile either to clone libraries screening (for a review see [20]) or to in silico prediction [29]. In this work we applied a recently developed technique [32] which provides a partial taxonomic interpretation of TRFs by their sequencing. Taxonomic interpretation of some of the common and exclusive TRFs by this technique showed an uneven distribution of bacterial groups in the samples. Rhizosphere-exclusive TRFs were mainly interpreted as due to Proteobacteria (mainly α-Proteobacteria), whereas TRFs present in all samples were mainly due to non-Proteobacteria representatives. In a previous analysis on nickel-resistant bacteria present in the soil and in the rhizosphere of A. bertolonii [31], we found that several Pseudomonas strains were present in the plant rhizosphere, whereas on free soil several exclusive Streptomyces strains were found. It could be possible that most rhizosphere-specific α-Proteobacteria representatives found in T-RFLP profiles did not harbor Ni-resistant determinants or were not culturable under the aerobic and heterotrophic conditions applied. A similar result was found for the rhizosphere of other members of the family Brassicaceae (Brassica napus and Thlaspi caerulescens) in which rhizosphere representatives from α-Proteobacteria and Cytophaga–Flavobacterium–Bacteroides subdivisions were found in 16SrDNA clone libraries, while representatives from β- and γ-Proteobacteria subdivisions were found among cultured isolates [15, 18]. Several studies have provided evidence that heavy-metal-resistant Proteobacteria may protect plants or other bacteria from the toxic effects of heavy metals or even enhance metal uptake by hyperaccumulator plants [6, 48, 53]. Recently, an α-Proteobacteria representative (Sphingomonas macrogoltabidus) was isolated from the rhizosphere of the nickel hyperaccumulator Alyssum murale and has been shown to increase plant nickel uptake [1].
Summarizing, our results indicated that (i) the bacterial community present in serpentine soils is highly differentiated with members from Gram-positive species, Proteobacteria, Acidobacteria, Verrucomicrobia, and Chlorobi groups; (ii) the presence of the Ni-hyperaccumulating plant A. bertolonii seems to shape the community composition along a distance gradient at least at a 5-cm distance; and (iii) most of the genetic diversity distinguishing plant roots from free soil appears to be related to Proteobacteria strains.
References
RA Abou-Shanab JS Angle TA Delorme RL Chancy P van Berkum H Moawad K Ghanem HA Ghozlan (2003) ArticleTitleRhizobacterial effects on nickel extraction from soil and uptake by Alysum murale. New Phytol 158 219–224 Occurrence Handle1:CAS:528:DC%2BD3sXjt1Wltrg%3D
SF Altschul TL Madden AA Schäffer J Zhang Z Zhang W Miller DJ Lipman (1997) ArticleTitleGapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25 3389–3402 Occurrence Handle1:CAS:528:DyaK2sXlvFyhu7w%3D Occurrence Handle9254694
AMJ Baker (1981) ArticleTitleAccumulators and excluders: strategies in the response of plants to heavy-metals. J Plant Nutr 3 643–654 Occurrence Handle1:CAS:528:DyaL3MXhtlemsb8%3D
G Braker HL Ayala-del-Rio AH Devol A Fesefeldt JM Tiedje (2001) ArticleTitleCommunity structure of denitrifiers, Bacteria, and Archaea along redox gradients in Pacific Northwest marine sediments by Terminal Restriction Fragment Length Polymorphism analysis of amplified nitrite reductase (nirS) and 16S rRNA genes. Appl Environ Microbiol 67 1893–1901 Occurrence Handle10.1128/AEM.67.4.1893-1901.2001 Occurrence Handle1:CAS:528:DC%2BD3MXis1egtbc%3D Occurrence Handle11282647
RR Brooks (1987) Serpentine and Its Vegetation. A Multidisciplinary Approach Croom Helm London
GI Burd DG Dixon BR Click (1998) ArticleTitleA plant growth-promoting bacterium that decreases nickel toxicity in seedlings. Appl Environ Microbiol 64 3663–3668 Occurrence Handle1:CAS:528:DyaK1cXms1entr4%3D Occurrence Handle9758782
EO Casamayor R Massana S Benlloch L Øvreås B Díez VJ Goddard JM Gasol I Joint F Rodríguez-Valera C Pedrós-Alió (2002) ArticleTitleChanges in archaeal, bacterial and eukaryal assemblages along a salinity gradient by comparison of genetic fingerprinting methods in a multipond solar saltern. Environ Microbiol 4 338–348 Occurrence Handle10.1046/j.1462-2920.2002.00297.x Occurrence Handle12071979
Chaney, RL, Angle, JS, Baker, AJM, Li, Y-M (1998) Method for phytomining of nickel, cobalt and other metals from soils. US Patent No. 5,711,784
TA Delorme JV Gagliardi JS Angle RL Chaney (2001) ArticleTitleInfluence of the zinc hyperaccumulator Thlaspi caerulescens J. & C. Presl. and the nonmetal accumulator Trifolium pratense L. on soil microbial population. Can J Microbiol 47 773–776 Occurrence Handle10.1139/cjm-47-8-773 Occurrence Handle1:CAS:528:DC%2BD3MXnt12gu70%3D Occurrence Handle11575505
J Dunbar LO Ticknor CR Kuske (2000) ArticleTitleAssessment of microbial diversity in four southwestern United States soils by 16S rRNA gene terminal restriction fragment analysis. Appl Environ Microbiol 66 2943–2950
J Dunbar LO Ticknor CR Kuske (2001) ArticleTitlePhylogenetic specificity and reproducibility and new method for analysis of terminal restriction fragment profiles of 16S rRNA genes from bacterial communities. Appl Environ Microbiol 67 190–197
J Escarré C Lefebvre W Gruber M Leblanc J Lepart Y Riviere B Delay (2000) ArticleTitleZinc and cadmium hyperaccumulation by Thlaspi caerulescens from metalliferous and nonmetalliferous sites in the Mediterranean area: implications for phytoremediation. New Phytologist 145 429–437 Occurrence Handle10.1046/j.1469-8137.2000.00599.x
N Fierer JP Schimel PA Holden (2003) ArticleTitleInfluence of drying–rewetting frequency on soil bacterial community structure. Microb Ecol 45 63–71 Occurrence Handle10.1007/s00248-002-1007-2 Occurrence Handle1:STN:280:DC%2BD3s%2FmsVWksg%3D%3D Occurrence Handle12469245
SJ Giovannoni TB Britschgi CL Moyer KG Field (1990) ArticleTitleGenetic diversity in Sargasso Sea bacterioplankton. Nature 345 60–63 Occurrence Handle10.1038/345060a0 Occurrence Handle1:CAS:528:DyaK3cXktFymu7s%3D Occurrence Handle2330053
F Gremion A Chatzinotas H Harms (2003) ArticleTitleComparative 16S rDNA and 16S rRNA sequence analysis indicates that Actinobacteria might be a dominant part of the metabolically active bacteria in heavy-metal-contaminated bulk and rhizosphere soil. Env Microbiol 5 896–907 Occurrence Handle10.1046/j.1462-2920.2003.00484.x Occurrence Handle1:CAS:528:DC%2BD3sXos1CltLg%3D
M Héry S Nazaret T Jaffré P Normand E Navarro (2003) ArticleTitleAdaptation to nickel spiking of bacterial communities in neocaledonian soils. Environ Microbiol 5 3–12 Occurrence Handle10.1046/j.1462-2920.2003.00380.x Occurrence Handle12542708
P Hugenholtz BM Goebel NR Pace (1998) ArticleTitleImpact of culture-independent studies on the emerging phylogenetic view of bacterial diversity. J Bacteriol 180 4765–4774 Occurrence Handle1:CAS:528:DyaK1cXmt1Sgu7k%3D Occurrence Handle9733676
O Kaiser A Pulher W Selbitschka (2001) ArticleTitlePhylogenetic analysis of microbial diversity in the rhizoplane of oilseed rape (Brassica napus cv. Westar) employing cultivation-dependent and cultivation-independent approaches. Microb Ecol 42 136–149 Occurrence Handle1:CAS:528:DC%2BD3MXmslOqtLo%3D Occurrence Handle12024277
AD Kent EW Triplett (2002) ArticleTitleMicrobial communities and their interactions in soil and rhizosphere ecosystems. Ann Rev Microbiol 56 211–236 Occurrence Handle10.1146/annurev.micro.56.012302.161120 Occurrence Handle1:CAS:528:DC%2BD38Xos1Gis7s%3D
CL Kitts (2001) ArticleTitleTerminal restriction fragment patterns: a tool for comparing microbial communities an assessing community dynamics. Curr Issues Intest Microbiol 2 17–25 Occurrence Handle1:CAS:528:DC%2BD3MXjvVyiu7o%3D Occurrence Handle11709853
KT Kostantinidis N Isaaca J Fett S Simpson DT Long TL Marsh (2003) ArticleTitleMicrobial diversity and resistance to copper in metal-contaminated lake sediment. Microb Ecol 45 191–202 Occurrence Handle10.1007/s00248-002-1035-y Occurrence Handle12545313
AR Kruckeberg (1954) ArticleTitleThe ecology of serpentine soils. III. Plant species in relation to serpentine soils. Ecology 35 267–274
CR Kuske LO Ticknor ME Miller JM Dunbar JA Davis SM Barns J Belnap (2002) ArticleTitleComparison of soil bacterial communities in rhizospheres of three plant species and the interspaces in an arid grassland. Appl Environ Microbiol 68 1854–63 Occurrence Handle10.1128/AEM.68.4.1854-1863.2002 Occurrence Handle1:CAS:528:DC%2BD38XivFGlur4%3D Occurrence Handle11916705
DJ Lane (1991) 16S/23S rRNA sequencing. E Stackebrandt M Goodfellow (Eds) Nucleic Acid Techniques in Bacterial Systematics John Wiley and Sons New York 115–175
W-T Liu TL Marsh H Cheng LJ Forney (1997) ArticleTitleCharacterization of microbial diversity by determining terminal restriction fragment length polymorphism of genes encoding 16 rRNA. Appl Env Microbiol 63 4516–4522 Occurrence Handle1:CAS:528:DyaK2sXnt12ntbs%3D
C Lodewyckx M Mergeay J Vangronsveld H Clijsters D Van Der Lelie (2002) ArticleTitleIsolation, characterization and identification of bacteria associated with the zinc hyperaccumulator Thlaspi caerulescens subsp. calaminaria. Int J Phytoremediation 4 101–115 Occurrence Handle1:CAS:528:DC%2BD38XlvVKrt7g%3D Occurrence Handle12655804
T Lueders MW Friedrich (2003) ArticleTitleEvaluation of PCR amplification bias by terminal restriction fragment length polymorphism analysis of small-subunit rRNA and mcrA genes by using defined template mixtures of methanogenic pure cultures and soil DNA extracts. Appl Environ Microbiol 69 320–326 Occurrence Handle10.1128/AEM.69.1.320-326.2003 Occurrence Handle1:CAS:528:DC%2BD3sXkvValsw%3D%3D Occurrence Handle12514011
TL Marsh (1999) ArticleTitleTerminal restriction fragment length polymorphism (T-RFLP): an emerging method for characterizing diversity among homologous populations of amplification products. Curr Opin Microbiol 2 323–327 Occurrence Handle10.1016/S1369-5274(99)80056-3 Occurrence Handle1:CAS:528:DyaK1MXkt1Cms70%3D Occurrence Handle10383864
TL Marsh P Saxman J Cole J Tiedje (2000) ArticleTitleTerminal restriction fragment length polymorphism analysis program, a Web-based research tool for microbial community analysis. Appl Environ Microbiol 66 3616–3620
P Marschner C-H Yang R Lieberei DE Crowley (2001) ArticleTitleSoil and plant specific effects on bacterial community composition in the rhizosphere. Soil Biol Biochem 33 1437–1445 Occurrence Handle10.1016/S0038-0717(01)00052-9 Occurrence Handle1:CAS:528:DC%2BD3MXmtFSlsrk%3D
A Mengoni R Barzanti C Gonnelli R Gabbrielli M Bazzicalupo (2001) ArticleTitleCharacterization of nickel-resistant bacteria isolated from serpentine soil. Environ Microbiol 3 691–698 Occurrence Handle10.1046/j.1462-2920.2001.00243.x Occurrence Handle1:CAS:528:DC%2BD38XlvVKmsw%3D%3D Occurrence Handle11846759
A Mengoni E Grassi M Bazzicalupo (2002) ArticleTitleA cloning method for the taxonomic interpretation of T-RFLP patterns. Biotechniques 33 990–992
MM Moeseneder JM Arrieta G Muyzer C Winter GJ Herndl (1999) ArticleTitleOptimization of terminal-restriction fragment length polymorphism analysis for complex marine bacterioplankton communities and comparison with denaturing gradient gel electrophoresis. Appl Environ Microbiol 65 3518–25 Occurrence Handle10427043
K Nagashima T Hisada M Sato J Mochizuki (2003) ArticleTitleApplication of new primer–enzyme combinations to terminal restriction fragment length polymorphism profiling of bacterial populations in human feces. Appl Environ Microbiol 69 1251–62 Occurrence Handle10.1128/AEM.69.2.1251-1262.2003 Occurrence Handle1:CAS:528:DC%2BD3sXhtF2itLY%3D Occurrence Handle12571054
AM Osborn ER Moore KN Timmis (2000) ArticleTitleAn evaluation of terminal-restriction fragment length polymorphism (T-RFLP) analysis for the study of microbial community structure and dynamics. Environ Microbiol 2 39–50 Occurrence Handle10.1046/j.1462-2920.2000.00081.x Occurrence Handle1:CAS:528:DC%2BD3cXhs12ht7k%3D Occurrence Handle11243261
JE Park KE Young HG Schlegel HG Rhie HS Lee (2003) ArticleTitleConjugative plasmid mediated inducible nickel resistance in Hafnia alvei 5-5. Int Microbiol 6 57–64 Occurrence Handle1:CAS:528:DC%2BD3sXjtlWruro%3D Occurrence Handle12730713
TE Pawloska RL Chaney M Chin I Charvat (2000) ArticleTitleEffects of metal phytoextraction practices on the indigenous community of Arbuscular mycorrhizal fungi at a metal-contaminated landfill. Appl Environ Microbiol 66 2526–2530 Occurrence Handle10.1128/AEM.66.6.2526-2530.2000 Occurrence Handle10831433
FJ Rohlf (1990) NTSYS-pc. Numerical Taxonomy and Multivariate Analysis System. Version 2.02 Exeter Software New York
D Saintpierre H Amir R Pineau L Sembiring M Goodfellow (2003) ArticleTitleStreptomyces yatensis sp. nov., a novel bioactive streptomycete isolated from a New-Caledonian ultramafic soil. Antonie van Leeuwenhoek 83 21–26 Occurrence Handle10.1023/A:1022906325397 Occurrence Handle1:CAS:528:DC%2BD3sXisV2hsr8%3D Occurrence Handle12755476
Y Sakano KD Picketing PF Strom LJ Kerkhof (2002) ArticleTitleSpatial distribution of total, ammonia-oxidizing, and denitrifying bacteria in biological wastewater treatment reactors for bioregenerative life support. Appl Environ Microbiol 68 2285–93 Occurrence Handle10.1128/AEM.68.5.2285-2293.2002 Occurrence Handle1:CAS:528:DC%2BD38XjsFGqt74%3D Occurrence Handle11976099
DE Salt RD Smith I Raskin (1998) ArticleTitlePhytoremediation. Ann Rev Plant Physiol Plant Mol Biol 49 643–668 Occurrence Handle10.1146/annurev.arplant.49.1.643 Occurrence Handle1:CAS:528:DyaK1cXjvVSgs7g%3D
RA Sandaa V Torsvik O Enger FL Daae T Castberg D Hahn (1999) ArticleTitleAnalysis of bacterial communities in heavy metal-contaminated soils at different levels of resolution. FEMS Microb Ecol 30 237–251 Occurrence Handle10.1016/S0168-6496(99)00062-8 Occurrence Handle1:CAS:528:DyaK1MXmsFahs78%3D
DJ Scala LJ Kerkoff (2000) ArticleTitleHorizontal heterogeneity of denitrifying bacterial communities in marine sediments by terminal restriction fragment length polymorphism analysis. Appl Environ Microbiol 66 1980–1986 Occurrence Handle10.1128/AEM.66.5.1980-1986.2000 Occurrence Handle1:CAS:528:DC%2BD3cXjtV2ltb0%3D Occurrence Handle10788370
HG Schlegel JP Cosson AJM Baker (1991) ArticleTitleNickel-hyperaccumulating plants provide a niche for nickel-resistant bacteria. Bot Acta 194 18–25
K Smalla G Wieland A Buchner A Zock J Parzy S Kaiser N Roskot H Heuer G Berg (2001) ArticleTitleBulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: plant-dependent enrichment and seasonal shifts revealed. Appl Environ Microbiol 67 4742–4751
MP de Souza CPA Huang N Chee N Terry (1999) ArticleTitleRhizosphere bacteria enhance that accumulation of selenium and mercury in wetland plants. Planta 209 259–263 Occurrence Handle10.1007/s004250050630 Occurrence Handle1:CAS:528:DyaK1MXlt1Gjtrc%3D Occurrence Handle10436229
DL Sparks (1996) Methods of Soil Analysis. Part 3. Chemical Methods Soils Science Society of America Madison, WI
JR Stephen YJ Chang SJ Macnaughton GA Kowalchuk KT Leung CA Flemming DC White (1999) ArticleTitleEffect of toxic metals on indigenous soil β-subgroup proteobacterium ammonia oxidizer community structure and protection against toxicity by inoculated metal-resistant bacteria. Appl Environ Microbiol 65 95–101 Occurrence Handle1:CAS:528:DyaK1MXjvVyiuw%3D%3D Occurrence Handle9872765
V Torsvik L Øvreås (2002) ArticleTitleMicrobial diversity and function in soil: from genes to ecosystems. Curr Opin Microbiol 5 240–245 Occurrence Handle10.1016/S1369-5274(02)00324-7 Occurrence Handle1:CAS:528:DC%2BD38XktFynsL4%3D Occurrence Handle12057676
H Urakawa T Yoshida M Nishimura K Ohwada (2000) ArticleTitleCharacterization of depth-related population variation in microbial communities of a coastal marine sediment using 16S rDNA-based approaches and quinone profiling. Environ Microbiol 2 542–554 Occurrence Handle10.1046/j.1462-2920.2000.00137.x Occurrence Handle1:CAS:528:DC%2BD3cXoslGmur4%3D Occurrence Handle11233162
O Vergnano Gambi (1992) The distribution and ecology of the vegetation of ultramafic soils in Italy. BA Roberts J Proctor (Eds) The Ecology of Areas with Serpentinized Rocks—A World View Kluwer Academic Dordrechts, The Netherlands 217–241
JP White (2001) ArticleTitlePhytoremediation assisted by microorganisms. Trends Plant Sci 6 502 Occurrence Handle10.1016/S1360-1385(01)02093-3 Occurrence Handle1:CAS:528:DC%2BD3MXovFGks7k%3D Occurrence Handle11701356
SN Whiting MP de Souza N Terry (2001) ArticleTitleRhizosphere bacteria mobilize Zn for hyperaccumulation by Thlaspi caerulescens. Environ Sci Technol 35 3144–3150 Occurrence Handle10.1021/es001938v Occurrence Handle1:CAS:528:DC%2BD3MXks1eltLY%3D Occurrence Handle11505990
Acknowledgments
We are grateful to Prof. R. Gabbrielli for helpful discussions and suggestions on the physiology and ecology of hyperaccumulating plants, and to Dr. M. Barbafieri and Dr. C. Mastretta for the analyses of soil pH, CEC, and organic matter.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mengoni, A., Grassi, E., Barzanti, R. et al. Genetic Diversity of Bacterial Communities of Serpentine Soil and of Rhizosphere of the Nickel-Hyperaccumulator Plant Alyssum bertolonii. Microb Ecol 48, 209–217 (2004). https://doi.org/10.1007/s00248-003-0149-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-003-0149-1