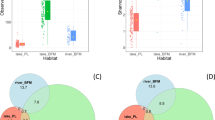
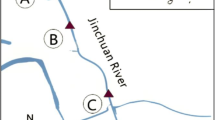
We compared the species composition in phytobenthic communities at different sampling sites in a small French river presenting polluted and unpolluted areas. For each sampling point, the total DNA was extracted and used to construct an 18S rRNA gene clone library after PCR amplification of a ca 400 bp fragment. Phytobenthic community composition was estimated by random sequencing of several clones per library. Most of the sequences corresponded to the Bacillariophyceae and Chlorophyceae groups. By combining phylogenetic and correspondence analyses, we showed that our molecular approach is able to estimate and compare the species composition at different sampling sites in order to assess the environmental impact of xenobiotics on phytobenthic communities. Changes in species composition of these communities were found, but no evident decrease in the diversity. We discuss the significance of these changes with regard to the existing level of pollution and their impact on the functionality of the ecosystem. Our findings suggest that it is now possible to use faster molecular methods (DGGE, ARISA...) to test large numbers of samples in the context of ecotoxicological studies, and thus to assess the impact of pollution in an aquatic ecosystem.
Article PDF
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Dorigo, U., Bérard, A. & Humbert, J. Comparison of Eukaryotic Phytobenthic Community Composition in a Polluted River by Partial 18S rRNA Gene Cloning and Sequencing . Microb Ecol 44, 372–380 (2002). https://doi.org/10.1007/s00248-002-2024-x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00248-002-2024-x