Abstract.
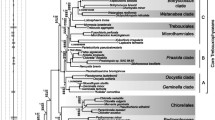
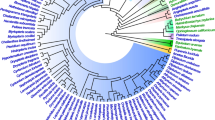
Maximum likelihood (ML) phylogenies based on 9,957 amino acid (AA) sites of 45 proteins encoded in the plastid genomes of Cyanophora, a diatom, a rhodophyte (red algae), a euglenophyte, and five land plants are compared with respect to several properties of the data, including between-site rate variation and aberrant amino acid composition in individual species. Neighbor-joining trees from AA LogDet distances and ML analyses are seen to be congruent when site rate variability was taken into account. Four feasible trees are identified in these analyses, one of which is preferred, and one of which is almost excluded by statistical criteria. A transition probability matrix for the general reversible Markov model of amino acid substitutions is estimated from the data, assuming each of these four trees. In all cases, the tree with diatom and rhodophyte as sister taxa was clearly favored. The new transition matrix based on the best tree, called cpREV, takes into account distinct substitution patterns in plastid-encoded proteins and should be useful in future ML inferences using such data. A second rate matrix, called cpREV*, based on a weighted sum of rate matrices from different trees, is also considered.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received: 3 June 1999 / Accepted: 26 November 1999
Rights and permissions
About this article
Cite this article
Adachi, J., Waddell, P., Martin, W. et al. Plastid Genome Phylogeny and a Model of Amino Acid Substitution for Proteins Encoded by Chloroplast DNA. J Mol Evol 50, 348–358 (2000). https://doi.org/10.1007/s002399910038
Issue Date:
DOI: https://doi.org/10.1007/s002399910038