Abstract
By cDNA sequencing we have achieved the first, and complete, hemocyanin sequence of a bivalve (Nucula nucleus). This extracellular oxygen-binding protein consists of two immunologically distinguishable isoforms, here termed NnH1 and NnH2. They share a mean sequence identity of 61%, both contain a linear arrangement of eight paralogous, ca.50-kDa functional units (FUs a-h), and in both isoforms the C-terminal FU-h possesses an extension of ca. 100 amino acids. The cDNA of NnH1 comprises 11,090 bp, subdivided into a 5′utr of 75 bp, a 3′utr of 791 bp, and an open reading frame for a signal peptide of 19 amino acids plus a polypeptide of 3389 amino acids (M r = 385 kDa). The cDNA of NnH2 comprises 10,849 bp, subdivided into a 5′utr of 47 bp, a 3′utr of 647 bp, and an open reading frame for a signal peptide of 16 amino acids plus a polypeptide of 3369 amino acids (M r = 387 kDa). In contrast to other molluscan hemocyanins, which are highly glycosylated, the bivalve hemocyanin sequence exhibits only four potential N-glycosylation sites, and within both isoforms a peculiar indel is present, surrounding the highly conserved copper-binding site CuA. Phylogenetic analyses of NnH1 and NnH2, compared to the known hemocyanin sequences of gastropods and cephalopods, reveal a statistically sound closer relationship between gastropod and protobranch hemocyanin than to cephalopod hemocyanin. Assuming a molecular clock, the last common ancestor of protobranch and gastropods lived 494 million ± 50 million years ago, in conformity with fossil records from the late Cambrian.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Molluscan hemocyanins are very large extracellular oxygen transporting proteins with a type 3 copper center and a decameric quaternary structure. This decamer is composed of five dimers, which form a dipentameric hollow ring that in many mollusks dimerizes and in some instances, forms elongated multidecamers (Miller et al. 1990; Markl et al. 1991). The protein monomer consists of a polypeptide with a molecular mass of ∼350–400 kDa, that is folded into seven or eight different globular functional units (FUs) of ca. 50 kDa, termed FU-a to FU-h (from N- to C-terminus). The FUs are connected via short linker regions of 10 to 15 amino acids (Lang and van Holde, 1991; Miller et al. 1998; Lieb et al. 2000). While FUs-a to-f build the wall of the decamer, the internal collar complex is formed by FU-g and FU-h (Meissner et al. 2000). Each FU carries an oxygen-binding site containing two copper ions complexed by six strictly conserved histidine residues. The so-called CuA and CuB sites of each FU together reversibly bind 1 dioxygen molecule, leading to a maximum of up to 8 O2 molecules per subunit, or 80 per decammer. Previous studies have shown the variability of hemocyanin quaternary structure in the different molluscan classes. In gastropods and bivalves didecamers predominate, formed by face-to-face assembly of two decamers (reviewed by van Holde and Miller 1995). In many marine gastropods tube-like multidecamers are also present, with decamers added at one, or both ends of a central didecamer (Herskovits and Hamilton 1991; Markl et al. 1991, 2001). In contrast, the hemocyanins of cephalopods and chitons are exclusively decamers. The subunit of Octopus dofleini hemocyanin (OdH) has a molecular mass of only ∼350 kDa, due to the absence of a FU-h (Miller et al. 1990, 1998); FU-h is also absent at the gene level (Lieb et al. 2001). In OdH, FU-g forms the central collar, which corresponds structurally to the arc of gastropod hemocyanins (Lamy et al. 1993). Another feature of molluscan hemocyanins is the presence of more than one isoform in several organisms. Within the abalone Haliotis tuberculata two immunologically distinguishable isoforms occur, which are named HtHl and HtH2 and possess a sequence identity of ∼66% (Altenhein et al. 2002). Contrary to this, the two isoforms of Octopus dofleini hemocyanin (OdHA and OdHG) share a sequence identity of ∼96% (Miller et al. 1998).
These differences raise questions of their phylogenetic origin and of their structural-functional significance. Therefore, it is necessary to study the hemocyanins from the different molluscan classes at different structural levels. Within the cephalopods, the complete cDNA and genomic sequences of Octopus dofleini and Nautilus pompilius hemocyanin are available, and have been compared to the corresponding data from the gastropods Haliotis tuberculata and Aplysia californica (Miller et al. 1998; Lieb et al. 2000, 2001, 2004; Altenhein et al. 2002; Bergmann et al. 2006). In addition, two partial amino acid sequences of Sepia officinalis hemocyanin (SoH) have been published (Declerq et al. 1990). There is some knowledge of the biochemistry of bivalve hemocyanin, but no sequence data are currently available.
The class Bivalvia consists of five subclasses, namely, Protobranchia, Pteriomorpha, Heterodonta, Paleoheterodonta, and Anomalodesmata. The presence of hemocyanin as respiratory protein has only been shown for the protobranch bivalves (Morse et al. 1986). The other subclasses either have (in rare cases) hemoglobin or lack a respiratory protein (van Holde and Millen 1995).Within the bivalves, hemocyanin seems to be a primordial feature because the protobranchs are said to be the most ancient bivalve branch. In this context we examined whether similarities and differences within the hemocyanin quaternary structure between the bivalves and gastropods are also reflected within the gene and protein sequences.
Materials and Methods
Animals
Nucula nucleus was collected in part by Frank Zal (Roscoff, France) and ordered in part by the Service Mer et Observation (Roscoff, France).
Preparation of RNA and RT-PCR
RNA was extracted from whole animals using the GTC method (Chomczynski and Sacchi 1987; Sambrook et al. 2001). Reverse transcription and subsequent PCR were performed with Expand Reverse Transcriptase and Expand DNA polymerase, respectively (Roche Diagnostics, Mannheim, Germany) following the manufacturer’s instructions. The missing 5′ and 3′ ends were obtained using the 5′ and 3′ Race Kit (Invitrogen, Karlsruhe, Germany), respectively.
Purification and Sequencing of PCR Fragments
PCR samples were analyzed in standard agarose electrophoresis gels in 1 × TBE buffer (89 mM Tris/chloride, 89 mM sodium borate, 2 mM sodium EDTA, pH 8.0) and purified using the gel extraction kit from Qiagen (Hilden, Germany). Prior to sequencing, PCR fragments were either cloned into TOPO pCR 2.1 and pCR-XL-TOPO (Invitrogen) or sequenced (Genterprise, Mainz, Germany) and analysed directly.
Sequence Data Analyses and Phylogenetic Studies
Data obtained from the automated sequencer were computer-analyzed using CHROMAS (http://www.trishul.sci.gu.edu.au/∼conochromas.html), TRANSLATE (http://www.expasy.ch), ALIGN (http://www.expasy.ch), and BLAST Japan (http://www.blast.genome.ad.jp). SIGNALP VI. 1 was used for prediction of the signal peptide (http://www.expasy.ch). Multiple sequence alignments and identity matrices were calculated by CLUSTALX 1.83 (Thompson et al. 1997) after manual optimization with the aid of GENEDOC 2.6 (Nicholas and Nicholas 1997). Phylogenetic analyses were performed with the PHYLIP package (Felsenstein 2001) and MRBAYES 2.01, assuming different matrices (Huelsenbeck and Ronquist 2001). For Baysian analyses, Metropolis-coupled Markov chain Monte Carlo sampling was performed with four chains that were run for 150,000 generations. Prior probabilities for all trees were equal, starting trees were randomly assigned, and tree sampling was done every 10 generations. Posterior probabilities were estimated on 5000 trees (burn-in = 10,000). For molecular clock calculation, the software Protdist, using the JTT matrix of the PHYLIP package, was used.
Preparation of Hemocyanin from the Hemolymph
Hemolymph was taken from about 50 living Nucula nucleus, after putting them on ice for narcosis. The shell was removed manually, the tissue was pierced by the needle of a syringe, and the hemolymph was taken off with a pipette. To remove cellular contaminants, the collected hemolymph was centrifugated for 5 min at 2000g. The high molecular mass hemocyanin in this low-g supernatant was sedimented by ultracentrifugation at 132,000g for 4 h. The blue hemocyanin pellet was suspended in stabilizing buffer (0.05 M Tris, 5 mM CaCl2, 5 mM MgCl2, 0.15 M NaCl, 1 mM PhCH2SO2F, pH 7.4) overnight and stored at 4°C. From 50 individuals, we obtained 250 μl hemolymph to 2mg hemocyanin.
Gel Filtratiom Chromatography
To purify the hemocyanin from Nucula nucleus the samples were applied to an Econo chromatographic column (120 cm height and 1.5 cm in diameter, with a volume of 230 ml) loaded with BioGel A 15m (BioRad, Munich, Germany). After elution with stabilizing buffer the fractions were tested using SDS-PAGE and the hemocyanin-containing fractions were pooled.
SDS-PAGE and Crossed Immunoelectrophoresis
SDS-PAGE was performed according to the procedure of Laemmli (1970). Crossed and crossed-line immunoelectrophoreses were carried out following to the methods described by Weeke (1973) and Kroll (1973).
Amino Acid Analysis
Proteolytic fragments were separated by SDS-PAGE (Laemmli 1970) and subsequently electro transferred to ProBlot membranes (Applied Biosystems, Weiterstadt, Germany) in a vertical blotting chamber. The transferred bands were detected by Ponceau S staining. Polypeptides of interest were cut out and sequenced by a commercial amino acid sequencing service (Dr. Hans Heid, DKFZ, Heidelberg, Germany).
Electron Microscopy and Protein Structure Modeling
Negative staining was performed by W. Gebauer using the single-droplet procedure (Harris and Horne 1991). Carbon support films were initially glow-discharged for 20 s to render them hydrophilic and adsorptive for the protein. Hemocyanin samples (∼0.1 mg/ml) were adsorbed to the carbon film, then washed twice with droplets of water, and, finally, negatively stained with 2% (w/v) uranyl acetate. After drying, the grids were studied in a Zeiss EM 900 transmission electron microscope at 80 kV.
Results
Electron Microscopy, Biochemical Analysis and Immunoelectrophoresis
Nucula nucleus hemocyanin (NnH) was purified from cell-free hemolymph by ultracentrifugation followed by gel filtration chromatography. Electron microscopy of negatively stained native hemocyanin molecules revealed top views and side views, mainly of didecamers but with a small number of tridecamers (Fig. 1).
Transmission electron microscopy of Nucula nucleus hemocyanin. Negatively stained NnH didecamers attached to the carbon support film are shown as side views (rectangular) and as top views (circles). A few tridecamers (black arrow) are present, where a further single decamer is attached to a didecamer.
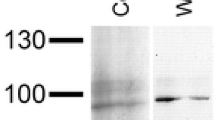
By SDS-PAGE, the NnH polypeptide was shown to possess a single band, migrating at a similar rate to the gastropod hemocyanin subunit RtHl from Rapana thomasiana, which is known to contain eight FUs (Fig. 2) (for references, see Söhngen et al. 1997; Gebauer et al. 1999). From such experiments, a molecular mass of ∼400 kDa was calculated for the NnH polypeptide.
Molecular mass determination of the NnH subunit by SDS-PAGE; 7.5 % polyacrylamide gel was used. As standards, we used commercial molecular mass markers (M: myosin, β-galactosidase, phosphorylase A, bovine serum albumin), and for comparison, the molluscan hemocyanin subunits from Rapana thomasiana, known to contain eight FUs (RtH1, 400 kDa), plus two submit fragments known to contain six and two FUs, respectively (RtH2.1, 300 kDa; RtH2.2, 100 kDa). Note that NnH migrates predominantly as a single band of ∼400 kDa, confirmed by calibration plot of molecular mass versus migration distance (not shown); the small fraction of faster-moving material might result from partial proteolytic cleavage, as often observed in molluscan hemocyanin preparations. For references, see Gebauer et al. (1999), Söhngen et al. (1997), and Idakieva et al. (1993).
Crossed immunoelectrophoresis, using polyclonal rabbit antibodies developed against purified, pH 9.6, dissociated NnH, yielded two precipitation peaks, indicating the presence of two immunologically distinguishable polypeptides; they have been termed NnHl and NnH2 (Fig. 3).
Immunoelectrophoresis (IE) of the subunits of the two NnH isoforms. In the first dimension, the anode was on the left. In the second dimension, the anode was at the bottom. A mixture of different rabbit antibodies against molluscan hemocyanin was applied. Crossed IE of the dissociated 400-kDa subunits, obtained by overnight dialysis of native NnH against alkaline buffer (pH 9.6); the two immuno-precipitates suggest heterogeneity at the subunit level, indicating the presence of the two isoforms, NnHl and NnH2.
cDNA Sequencing
Total RNA was isolated from whole Nucula nucleus tissue, and cDNA encoding the two complete NnH polypeptides was obtained by RT-PCR. First, degenerated primers derived from various known molluscan hemocyanin sequences were used, which were coding for the conserved copper-binding sites. The cDNA fragments so obtained were translated into amino acid sequences and were analyzed by multiple sequence alignments. After the sequences had been assigned to their respective FUs, NnH-specific primers were designed to close the gaps between the fragments. The terminal sequences were completed by 5′ and 3′ RACE. In order to prevent to mix the two sequences, overlaps of 200–300 bp between the various fragments were required.
The complete cDNA sequence of NnHl comprises 11,090 bp (EMBL/GenBank database accession number AJ786639). The first potential start codon is at position 76, yielding a 5′utr (untranslated region) of 75 bp. The first stop codon (TAA) is at position 10,297, followed by a 3′utr of 791 bp. Close to the end of the latter is a polyadenylation signal (AATAAA), and 103 bp further downstream a poly(A) tail is present.
The complete cDNA sequence of NnH2 comprises 10,849 bp (EMBL/GenBank database; accession number AJ786640). The first potential start codon is at position 48 yielding a 5′utr (untranslated region) of 47 bp. The first stop codon (TGA) is at position 10,200, followed by a 3′utr of 647 bp. Close to the end of the latter, a typical polyadenylation signal (AATAAA) is present and 39 bp further downstream a poly(A) tail.
The deduced primary structure of both hemocyanin polypeptides starts with a signal peptide, which is typical for extracellular proteins, including the molluscan hemocyanins (Sminia 1977; Ruth et al. 1988; Albrecht et al. 2001; Lieb et al. 2001). Using the SIGNALP V1.1 software, the lengths of the signal peptides was found to account for 19 amino acids in NnHl and 16 amino acids in NnH2, respectively. The signal peptide is followed by eight paralogous FUs, termed NnH-a to NnH-h, each containin 399–420 amino acids. NnHl-h and NnH2-h show a C-terminal extension of ∼100 amino acids, which has also been found in FU-h from gastropod hemocyanins (Lieb et al. 2004). Within both isoforms three potential N-glycosylation sites have been found, which are located within the functional units NnHl-a, NnH2-a, and NnH2-d (Fig. 4).
Primary structure of the eight FUs of NnHl and NnH2, as predicted from their cDNA sequence. A multiple sequence alignment, in comparison to Haliotis tuberculata hemocyanin isoform 1, is shown (HtHl; Lieb et al. 2000). A consensus numbering system was applied. The position of secondary structure features, as in OdH-g (Cuff et al. 1998), is indicated by blocks. Potential N-glycosylation sites (3 in NnHl/2, 13 in HtHl) are highlighted in dark grey for NnHl/2 and in light grey for HtHl. Putative disulfide and thioether bridges are marked. The alignment was performed with ClustalX and was edited by GeneDoc.
A radial phylogenetic tree was constructed using a multiple sequence alignment of the eight FUs of both N. nucleus hemocyanin isoforms, for comparison with the completely sequenced FUs of the gastropods H. tuberculata and A. californica and the cephalopoda N. pompilius, O. dofleini, and S. officinalis (Fig. 5). To construct this tree, the Baysian inference method was applied. A relative rate test according to Tajima (1993) revealed that the two NnH subunits possess the relatively low evolutionary rate of ∼10−9 mutations per site per year and follow an approximately linear evolution. This allowed calculation of a rather robust molecular clock based on divergence rates, as presented in Table 1. The time scale thus obtained suggests that the Protobranchia Gastropoda split occurred 494 million ± 50 million years ago and that the gene duplication yielding NnHl and NnH2 happened 396 million ± 55 million years ago (Fig. 6).
Phylogenetic analysis of molluscan hemocyanin functional units. A radial phylogenetic tree with hemocyanin sequences from Nucula nucleus (two isoforms: NnHl, NnH2), Nautilus pompilius (NpH), Octopus dofleini (OdH), Aplysia californica (AcH), and Haliotis tuberculata (two isoforms: HtHl, HtH2), constructed assuming the Dayhoff substitution matrix. The values at the branches represent Bayesian posterior probabilities. Note that eight major branches are obtained, corresponding to the eight different FUs.
Molecular clock calculated from the distance values of various molluscan hemocyanin sequences (bars represent standard deviation of these mean values). On the basis of fossil records (Benton 1993), the splitting point of gastropods and cephalopods was used for calibration (520 million years ago). Note that the Protobranchia-Gastropoda split occurred about 20 million years after the calibration point and was followed by the duplication of NnH subunit after a further 100 million years (396 ± 55 MYA). Distance values are given in Table 1.
Discussion
Quaternary Structure and Biochemical Studies
Native Nucula nucleus hemocyanin is present in the hemolymph mainly as didecamers that resemble the quaternary structure of the hemocyanin from the closely related species Nucula hanleyi (Mangum et al. 1987; Lambert et al. 1995) Additionally, a few tridecamers are shown in the electron micrographs of NnH, resulting from the association of a further decamer with one didecamer. Thus, the quaternary structure of Nucula nucleus hemocyanin corresponds to gastropod hemocyanin in general, rather than to cephalopod hemocyanin, in that the latter occurs only as decamers. These similar properties of the quaternary structure fits the present result that the bivalve hemocyanin FUs group phylogenetically along with the gastropod hemocyanin FUs rather than the cephalopod FUs, which would have been expected from the current hypothesis of the phylogenetic realtionship of these taxa.
Sequence Analysis
The most conserved features within each NnH FU are the six histidines, forming the copper-binding sites CuA and CuB, and two potential disulfide bridges between the positions Cys66-Cys77 and Cys199-Cys266. As, the oxygen-binding capacity of hemocyanins is completely lost after disulfide bond reduction (Topham et al. 1999), these two disulfide bridges might stabilize the core domain of each FU. A third potential disulfide bridge lies between Cys358 and Cys370 at the surface of the β-sandwich domain (Fig. 4). A potential disulfide bridge exists within the extension of FU-h in both isoforms of Nucula nucleus hemocyanin. These two conserved cysteine residues are also found in FU-h from Haliotis tuberculata, Megathura crenulata, and Aplysia californica, which supports this hypothesis, but for verification, the high-resolution X-ray structure of a FU-h is required. Another structural feature of NnH shared with other molluscan hemocyanins is a potential thioether bridge between Cys78 and His80. However, both isoforms of NnH also show specific properties. Between the β-sheets β2 and β3, four of the eight FUs show an indel, comprising two amino acids for NnH-c and six amino acids for NnH-d, NnH-f, and NnH-g (Figs. 4 and 7A). It is conspicuous that a glycine is positioned in start of the indels. The small size of this amino acid possibly allows a narrow coil between both β-sheets. Within other sequence areas further indels are found which, however, are mainly located within connecting loops of α-helices or β-strands. In NnH-h, the sequence between strands β8 and β9 seems to be significantly different compared to the FU-h of Haliotis tuberculata. For both isoforms NnH-h lacks about seven amino acids, which might be expected to protrude into the surrounding hydration shell (Figs. 4 and 7; arrowhead), thus possibly providing a slightly more condensed quarternary structure.
A A three-dimensional model of Odh-g. Potential carbohydrate side chains of NnH1/2-a and NnH1-e are present between β11 and β12 (star). Indels observed within FU-c, -d, -f, and -g cause a lack of β2/β3, on which a prominent sugar side chain is present in FU OdH-g (arrow, bracket). FU-h-specific indels between β8 and β9 are also marked (arrowhead; see text for detailed discussion). B, C A usually conserved tyrosine residue (Tyr217) between α8 and α9 protrudes into a pouch build by β4 and β5. Note that within NnH2-c this tyrosine might be substituted by a “newly” inserted tyrosine positioned within β4 (not shown).
By comparing both NnH isoforms, some significant differences can be observed. NnHl contains 20 additional amino acids in the coding region and the indels are scattered throughout the eight FUs. Another feature is the lack of the third disulfide bridge in NnH2-c, whereas in NnHl-c it is present. Probably, this missing structural element is compensated for by the deletion of six amino acids in this region, which facilitates the loop between strands β8 and β9. In both NnH isoforms together a total of only 4 potential N-glycosylation sites is found, 1 for NnHl and 3 for NnH2, whereas there are, for example, 13 found in HtHl. Indeed, by measuring the glycosylation level in lectin-binding assays, we found a two times lower sugar content in NnH compared to Nautilus pompilius hemocyanin (NpH) (data not shown). Also, this is in good agreement with the observation that NpH possess 12 potential N-glycosylation sites (Bergmann et al. 2006). Furthermore, one potential N-glycosylation site (in NnH2-e) has the sequence NPS and therefore might be blocked for sugar addition due to the central proline (Bause 1983). This site is located between the helices αl1 and α12 within NnH2-e (Fig. 4). The remaining three motifs are placed between the sheets β11 and β12 in NnHl-a, NnH2-a, and NnH2-d (Figs. 4 and 7A; star). This is the most frequent potential N-glycosylation site in molluscan hemocyanin FUs (Lieb et al. 2000) and might stabilize their tertiary structure. No potential N-glycosylation site is present in NnHl-d in this position. The unequal distribution of the potential N-glycosylation sites in both isoforms may have an influence on their biological functions or may act as a signal to distinguish between the two isoforms during regulation processes. The question of glycosylation as a differentiating factor was already raised in studies on the hemocyanin isoforms KLH1 and KLH2 of Megathura crenulata (Markl et al. 1991; Gebauer et al. 1994; Harris and Markl 2000).
Another interisoform- and also interspecies-specific difference which might shed further light on essential amino acids required for correctly folding the FUs can be found within the regions of helices α8 and α9 and the strands β4 and β5 of NpH2-c (Figs. 4 and 7B and C). Only NnH2-c possesses a four-amino acid indel and, thus, lacks a conserved tyrosine residue which usually, when seen within the X-ray structure context (Figs. 7B and C), comes into close contact with the β4/β5-strands, forming a platform that probably stabilizes the tertiary structure of the FUs. However, this tyrosine might be functionally substituted by another tyrosine, which is “newly” inserted by a second indel located 63 amino acids toward the N-terminus, directly positioned between β4 and β5. Additionally, and due to the fact that all FUs had evolved before the complete hemocyanin gene duplicated, we can assume an evolutionary scenario, at least for this region, where (i) two amino acids were inserted into β4, and (ii) thereafter four amino acids were lost from a location behind α8 only in NnH2-c, but (iii) this was compensated by a mutation further upstream which replaced an unknown amino acid by a tyrosine. Although all these suggestions are highly speculative and need to be proven by further functional studies, our cDNA sequence data provide some insight as to how the folding of the tertiary structure of the molluscan hemocyanin FUs is maintained.
Phylogeny
To study the molecular evolution of the molluscan hemocyanins seven complete sequences were used, which stem from the vetigastropod Haliotis tuberculata (HtHl and HtH2), the opisthobranch gastropod Aplysia californica (AcH), the dibranchiate cephalopod Octopus dofleini (OdH), the tetrabranchiate cephalopod Nautilus pompilius (NpH), and the two isoforms of Nucula nucleus hemocyanin (NnHl and NnH2) presented here. Table 1 shows that both isoforms of NnH have a sequence identity of ∼62%, which is slightly lower compared to the two isoforms of HtH (∼66%). NnHl exhibits higher identity values to all gastropod hemocyanins than does NnH2, which might be due to the numerous indels, especially in NnH2-c. However, in the phylogenetic tree NnHl and NnH2 clearly group together. Compared to the cephalopod hemocyanins, the gastropod and the bivalve hemocyanin sequences show a greater sequence identity. This observation becomes more clearly apparent by analyzing the phylogenetic tree, which was constructed using the Baysian inference method and included the completely sequenced FUs of all the aforementioned molluscan hemocyanins (Fig. 5). We choose an unrooted representation of the phylogenetic tree because no suitable outgroup is available. Despite the phylogenetical relationship among tyrosinase, catechol oxidase, and hemocyanin (e.g., Klabunde et al. 1998; van Gelder et al. 1997), which is also indicated by the very similar active site in these proteins, and the weak phenol oxidase activity of molluscan hemocyanin (Salvato et al. 1998), the relationship of tyrosinases and catechol oxidases to the primary structure of the molluscan hemocyanins is too remote. Hence the tree shows eight stable branches; no statistically sound support could be achieved resolving the phylogentic relationship of the single functional units. However, this tree clearly shows that the complete hemocyanin subunit evolved before all molluscan classes split and radiated (cf. Lieb and Markl 2004). This strongly supports the well-accepted monophyly of the phylum Mollusca and suggests that the last common ancestor of all living mollusks already possessed the multisubunit hemocyanin. In most bivalves, however, this feature has been lost.
Within each branch of this tree shown in Fig. 5, a strict grouping into the different molluscan classes is found. The two isoforms of Nucula nucleus hemocyanin are arranged constantly and well supported in a common twig. From this it follows that the duplication of the isoform occurred after the separation of the class Bivalvia. The location of NnH within the phylogenetic tree indicates a closer relationship between gastropod and bivalve hemocyanin, which is well supported by the posterior probability values. In contrast, the cephalopod hemocyanin FUs branch off in a basal position. This observation does not fit the morphology-based concepts suggesting that the cephalopods and gastropods form a sister group (Stasek 1972).
Within the major branches of the phylogenetic tree, we found a very stable separation into gastropod and cephalopod hemocyanin, which has previously been used to calibrate a molecular clock based on the fossil records of the gastropod-cephalopod split, in the late Cambrian (Benton 1993; Lieb et al. 2000; Altenhein et al. 2002). Accordingly, the Protobranchia-Gastropoda split occurred 494 million ± 50 million years ago (Fig. 6). This is in good agreement with fossil data of the first Protobranchia appearing in the late Cambrian and the Nuculidae were assigned to the Ordovician, ∼485 million years ago (Keen 1969; Cope 2000). The duplication of the subunit of N. nucleus hemocyanin is then estimated as occurring 396 million ± 55 million years ago (Fig. 6).
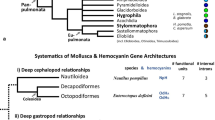
Previous phylogenetic relationship investigations on bivalves based on molecular data show many discrepancies, both to other molluscan classes and within the bivalve subclasses. Analysis of the 18S rRNA sequences revealed gastropods and bivalves to be monophyletic, forming a sister group (Winnepenninckx et al. 1994). But a similar analysis including more bivalve taxa suggests that the bivalves are polyphyletic (Kenchington et al. 1994). A current arrangement of the Protobranchia within the outgroups contributed to the poorly supported monophyly of the class Bivalvia in molecular studies (Adamkewicz et al. 1997; Campell et al. 1998). A similar result was shown in an extensive study by Giribet and Wheeler (2002) that was based on both molecular and morphological data. Here the monophyly of the Protobranchia is questionable and the results tend to indicate paraphyly. The Protobranchia are morphologically classified in the two orders Solemyoida and Nuculoida, but the monophyletic origin of the Nuculoida (with the superfamilies Nuculoidea and Nuculanoidea) is uncertain (Morton 1996; Waller 1998). Members of the Nuculoidea are said to be the most primordial bivalves within the Protobranchia, and ancestral to the Lamellibranchiata. It is supposed that the Protobranchia are a sister group to the remaining bivalves. Waller (1998) divided the Bivalvia into two subclasses, namely, Protobranchia and Autobranchia (Autolamellibranchiata). It was assumed that both these taxa have a different origin and therefore the bivalves might be diphyletic. Characteristic features of the Protobranchia and Lamellibranchiata support this assumption: In contrast to the lamellibranchiate veliger larvae, the Protobranchia possess the pericalymma larvae (Gustafson and Reid 1986, 1988). Furthermore, the stomach structure of the Protobranchia is similar to that of the gastropods (Purchon 1956). The fact that hemocyanin has only been found in protobranchiate bivalves does not permit the determination of a deeper and more accurate phylogenetic resolution of the Bivalvia based on this protein, but the assumption that the Bivalvia are biphyletic might explain the unexpected close relationship between gastropod and protobranch hemocyanins.
References
Adamkevicz SL, Harasewych MG, Blake J, Saudek D, Bult CJ (1997) A molecular phylogeny of the Bivalve mollusks. Mol Biol Evol 14:619–629
Albrecht U, Keller H, Gebauer W, Markl J (2001) Rhogocytes (pore cells) as the site of hemocyanin biosynthesis in the marine gastropod Haliotis tuberculata. Cell Tissue Res 304:455–462
Altenhein B, Markl J, Lieb B (2002) Gene structure and hemocyanin isoform HtH2 from the mollusc Haliotis tuberculata indicate early and late intron hot spots. Gene 301:53–60
Bause E (1983) Structural requirements of N-glycosylation of proteins. Studies with proline peptides as conformational probes. Biochem J 209:331–336
Benton MJ (1993) The fossil record 2. Chapman & Hall, London, pp 125–270
Bergmann S, Lieb B, Ruth P, Markl J (2006). The hemocyanin from a living fossil, the cephalopod Nautilus pompilius: protein structure, gene organization, and evolution. J Mol Evol 62:362–374
Campbell DC, Hoekstra KJ, Carter JG (1998) 18S ribosomal DNA and evolutionary relationships within the Bivalvia. In: Johnston PA, Haggart JW (eds) Bivalves: an eon of evolution — Palaeobiological studies honouring Norman D. Newell. University of Calgary Press, Calgary, pp 1–45
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162:156–159
Cope JCW (2000) A new look at early bivalve phylogeny. In: Harper EM, Taylor JD, Crame JA (eds) Evolutionary Biology of the Bivalvia Vol 177. Special Publication of the Geological Society, London, pp 81–95
Cuff ME, Miller KI, van Holde KE, Hendrickson WA (1998) Crystal structure of a functional unit from Octopus hemocyanin. J Mol Biol 278:855–870
Declerq L, Witters R, Preaux G (1990) Partial sequence determination of Sepia officinalis hemocyanin via cDNA. In: Preaux G, Lontie R (eds) Invertebrate dioxygen carriers. Leuven University Press, euven, Belgium, pp 131–134
Felsenstein J (2001) PHYLIP (Phylogeny Inference Package), version 3.6alpha2. Distributed by the author, Department of Genetics, University of Washington, Seattle
Gebauer W, Harris JR, Heid H, Stiling M, Hillenbrand R, Sohngen S, Wegener-Strake A, Markl J (1994) Quaternary structure, subunits and domain patterns of two dicrete forms of keyhole limpet hemocyanin: KLH1 and KLH2. Zoology 98:51–68
Gebauer W, Stoeva S, Voelter W, Danese E, Salvato B, Beltramini M, Markl J (1999) Hemocyanin subunit organization of the gastropod Rapana thomasiana. Arch Biochem Biophys 372:128–134
Giribet G, Wheeler W (2002) On bivalve phylogeny: a high-level analysis of the Bivalvia (Mollusca) based on combined morphology and DNA sequence data. Invert Biol 121:271–324
Gustafsen RG, Reid RGB (1986) Development of the pericalymma larva of Solemya reidi Bernard 1980. (Bivalvia, Cryptodonta, Solemyidae) as revealed by light and electron microscopy. Mar Biol 93:411–427
Gustafsen RG, Reid RGB (1988) Larval and post-larval morphogenesis in the gutless protobranch bivalve Solemya reidi (Cryptodonta, Solemyidae). Mar Biol 97:373–387
Harris JR, Home RW (1991) Negative staining. In: Harris JR (ed) Electron microscopy in biology. IRL Press, Oxford, pp 203–228
Harris JR, Markl J (2000) Keyhole limpet hemocyanin: molecular structure of a potent marine immunoactivator. A review. Eur Urol 37:24–33
Herskovits TT, Hamilton MG (1991) Higher order assemblies of molluscan hemocyanins. Comp Biochem Physiol B 99:19–34
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. ioinformatics 17:754–755
Idakieva K, Severov S, Svendsen I, Genov N, Stoeva S, Beltramini M, Tognon G, Di Muro P, Salvato B (1993) Structural properties of Rapana thomasiana grosse hemocyanin: isolation, characterization and N-terminal amino acid sequence of two different dissociation products. Comp Biochem Physiol B 106:53–59
Keen AM (1969) Family Nuculidae Gray 1824. In: Moore RC, Teichert C (eds) Treatise on invertebrate paleontology. Part N. Mollusca 6. Vol 1. Bivalvia. Geological Society of America, Boulder, CO/University of Kansas Press, Lawrence pp N230–N231
Kenchington EL, Roddick DL, Singh RK, Bird CJ (1994) Analysis of the small-subunit rRNA gene sequences from six families of molluscs. J Mar Biotechnol 1:215–217
Klabunde T, Eicken C, Sacchettini JC, Krebs B (1998) Crystal structure of a plant catechol oxidase containing a discopper center. Nat Struct Biol 5:1084–1090
Kroll J (1973) Crossed-line immunoelectrophoresis. Scand J Immunol 2:79–81
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lambert O, Taveau JC, Boisset N, Lamy JN (1995) Three-dimensional reconstruction of the hemocyanin of the protobranch bivalve mollusc Nucula hanleyi from frozen-hydrated specimens. Arch Biochem Biophys 319:231–243
Lamy J, Gielens C, Lambert O, Taveau JC, Motta G, Loncke P, De Geest N, Preaux G, Lamy J (1993) Further approaches to the quaternary structure of Octopus hemocyanin: a model based on immunoelectron microscopy and image processing. Arch Biochem Biophys 305:17–29
Lang WH, van Holde KE (1991) Cloning and sequencing of Octopus dofleini hemocyanin cDNA: derived sequences of functional units Ode and Odf. Proc Natl Acad Sci USA 88:244–248
Lieb B, Markl J (2004) Evolution of molluscan hemocyanins as deduced from DNA sequencing. Micron 35:117–119
Lieb B, Altenhein B, Markl J (2000) The sequence of a gastropod hemocyanin (HtHl from Haliotis tuberculatd). J Biol Chem 275:5675–5681
Lieb B, Altenhein B, Markl J, Vincent A, van Olden E, van Holde KE, Miller KI (2001) Structures of two molluscan hemocyanin genes: significance for gene evolution. Proc Natl Acad Sci USA 98:4546–4551
Lieb B, Boisguerin V, Gebauer W, Markl J (2004) cDNA sequence, protein structure, and evolution of the single hemocyanin from Aplysia californica, an ophistobranch gastropod. J Mol Evol 59:1–10
Mangum CP, Miller KI, Scott JL, van Holde KE, Morse MP (1987). Bivalve hemocyanin structural, functional, and phylogenetic relationships. J Biol Bull 173:205–221
Markl J, Savel-Niemann A, Wegener-Strake A, Silling M, Schneider A, Gebauer W, Harris R (1991) The role of two distinct subunit types in the architecture of keyhole limpet hemocyanin (KLH). Naturwissenschaften 78:512–514
Markl J, Lieb B, Gebauer W, Altenhein B, Meissner U, Harris JR (2001) Marine tumor vaccine carriers: structure of the molluscan hemocyanins KLH and HtH. J Cancer Res Clin Oncol 127:R3–R9
Meissner U, Dube P, Harris JR, Stark H, Markl J (2000) Structure of a molluscan hemocyanin didecamer (HtHl irom Hanotis tuberculata) at 12 Å resolution by cryoelectron microscopy. J Mol Biol 298:2l–34
Miller KI, Schabtach E, van Holde KE (1990) Arrangement of subunits and domains within the Octopus dofleini hemocyanin molecule. Proc Natl Acad Sci USA 87:1496–1500
Miller KI, Cuff ME, Lang WF, Varga-Weisz P, Field KG, van Holde KE (1998) Sequence of the Octopus dofleini hemocyanin subunit: structural and evolutionary implications. J Mol Biol 278:827–842
Morse MP, Meyhofer E, Otto JJ, Kuzirian AM (1986) Hemocyanin respiratory pigment in bivalve mollusks. Science 231:1302–1304
Morton BS (1996) The evolutionary history of the Bivalvia. In: Taylor JD (ed) Origin and evolutionary radiation of the Mollusca. Oxford University Press, Oxford, pp 337–359
Nicholas KB, Nicholas HB Jr (1997) GeneDoc: analysis and visualization of genetic variation. Distributed by the authors
Perbandt M, Guthohrlein EW, Rypniewski W, Idakieva K, Stoeva S, Voelter W, Genov N, Betzel C (2003) The structure of a functional unit from the wall of a gastropod hemocyanin offers a possible mechanism for cooperativity. Biochemistry 42:6341–6346
Purchon RD (1956) The stomach in the Protobranchia and Septibranchia (Lamellibranchia). Proc Zool Soc London 127:511–525
Ruth P, Schipp R, Kllissendorf B (1988) Cytomorphology and copper content of the basal cells in the midgut-gland of Nautilus (Cephalopoda, Tetrabranchiata). A contribution to the localization of hemocyanin synthesis. Zoomorphology 108:1–11
Salvato B, Santamaria M, Beltramini M, Alzuet G, Casella L (1998) The enzymatic properties of Octopus vulgaris hemocyanin: o-diphenol oxidase activity. Biochemistry 37:14065–14077
Sambrook J, Russel DW (2001) Molecular cloning. A laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Sminia T (1977) Haemocyanin-producing cells in gastropod molluscs. In: Bannister JV (ed) Structure ancLfunction of haemocyanin. Berlin, Springer, pp 279–288
Söhngen SM, Stahlmann A, Harris JR, Muller SA, Engel A, Markl J (1997) Mass determination subunit organization and control of oligomerization states of keyhole limpet hemocyanin (KLH). Eur J Biochem 248:602–614
Stasek CR (1972) The molluscan framework. In: Florkin M, Scheer BT (eds) Chemical zoology. Vol 7. Mollusca. Academic Press, New York, pp 1–44
Tajima F (1993) Simple methods for testing molecular clock hypothesis. Genetics 135:599–607
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Topham R, Tesh S, Westcott A, Cole G, Mercatante D, Kaufman G, Bonaventura C (1999) Disulfide bond reduction: A powerful, chemical probe for the study of structure-function relationships in the hemocyanins. Arch Biochem Biophys 369:261–266
van Gelder CWG, Flurkey WH, Wichers HJ (1997) Sequence and structural features of plant and fungal tyrosinases. Phytochemistry 45:1309–1323
van Holde KE, Miller KI (1995) Hemocyanins. Adv Protein Chem 47:1–81
Waller TR (1998) Origin of the molluscan class Bivalvia and a phylogeny of major groups. In: Johnston PA, Haggart JW (eds) Bivalves: an eon of evolution — Palaeobiological studies honouring Norman D. Newell. University of Calgary Press, Calgary, pp 1–45
Weeke B (1973) Crossed immunoelectrophoresis. Scand J Immunol 2:47–56
Winnepenninckx B, Backeljau T, De Wachter R (1994) Small ribosomal subunit RNA and the phylogeny of Mollusca. Nautilus 2:98–110
Acknowledgments
We thank Prof. Dr. J. Robin Harris (Institute of Zoology, University of Mainz) for critically, reading the manuscript and correcting the language and our colleague Dr. Wolfgang Gebauer for providing the electron micrographs. We also thank Dr. Frank Zal (Roscoff, France) for providing animals. This work was financially supported by DFG grants to J.M. (Ma843), the biosyn company (Fellbach, Germany), and the Feldbausch foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Dr. Rüdiger Cerff]
The sequence reported in this paper has been deposited in the EMBL/GenBank database under accession number AJ786639 for NnH1 and AJ786640 for NnH2.
Rights and permissions
About this article
Cite this article
Bergmann, S., Markl, J. & Lieb, B. The First Complete cDNA Sequence of the Hemocyanin from a Bivalve, the Protobranch Nucula nucleus . J Mol Evol 64, 500–510 (2007). https://doi.org/10.1007/s00239-006-0036-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-006-0036-8