Abstract
Type I polyketide synthases (PKSI) are modular multidomain enzymes involved in the biosynthesis of many natural products of industrial interest. PKSI modules are minimally organized in three domains: ketosynthase (KS), acyltransferase (AT), and acyl carrier protein. The KS domain phylogeny of 23 PKSI clusters was determined. The results obtained suggest that many horizontal transfers of PKSI genes have occurred between actinomycetales species. Such gene transfers may explain the homogeneity and the robustness of the actinomycetales group since gene transfers between closely related species could mimic patterns generated by vertical inheritance. We suggest that the linearity and instability of actinomycetales chromosomes associated with their large quantity of genetic mobile elements have favored such horizontal gene transfers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polyketide synthases type I (PKSI) are large enzymes that are responsible for the biosynthesis of polyketides, natural products of pharmaceutical and industrial interest such as erythromycin (antibiotic), rapamycin (immunosuppressant), and soraphen (antifungal). Among the three types of bacterial PKS that have been reported to date (Shen 2003), only type I shows a modular organization, allowing the comparative study of modules within or between enzymes. The PKSI consists of multifunctional enzymes grouped in modules (Fig. 1). A biosynthesis pathway is commonly composed of several clustered PKSI genes. Proteins encoded by these clustered PKSI genes catalyze the linear steps of polyketide biosynthesis. As an example, three PKSI proteins, called eryA, eryB, and eryC, (Fig. 2) required for the biosynthesis of erythromycin. Each PKSI protein loads and condenses two acyl groups to build the erythromycin (Staunton and Weissman 2001). The three required PKSI proteins are called “erythromycin synthase.” In this study, the term “X synthase” refers to any of the PKSI proteins involved in the synthesis of polyketide X.
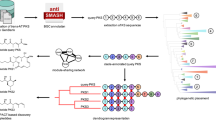
General representation of the PKSI organization. The PKSI are usually composed of a loading module followed by n extension modules building the growing acyl chain. Each substrate can be reduced by an optional reductive loop from one to three steps. A last domain thioesterase, is often present and catalyzes the release of the final product. KS, ketosynthase; AT, acyltransferase; ACP, acyl carrier protein; DH, dehydratase; KR, ketoreductase; ER, enoylreductase and TE, thioesterase.
Compilation of information concerning the published PKSI sequences used in this study. The PKSI protein name and their respective number of modules are indicated. As an example, the amphotericin synthase encompasses 6 PKSI proteins and 19 modules. Specific features in the Remarks column are detailed in the text.
Two types of module exist inside a PKSI protein: the first module is called the loading module, while the following ones are called extender modules (Fig. 1). Each extender module receives the incoming acyl chain from the previous module and catalyzes the condensation of the acyl groups to the growing acyl chain. Then the module transfers the acyl chain to the next module for an additional elongation step (Watanabe et al. 2003). A minimal extender module consists of three functional domains: a ketosynthase domain (KS), an acyltransferase domain (AT), and an acyl carrier protein (ACP). The KS domain catalyzes the decarboxylative condensation of an acyl group onto the growing chain. The AT domain selects and activates the acyl group. The ACP domain loads the growing chain (Staunton and Weissman 2001). The presence of additional domains with enzymatic activities of ketoreductase (KR), dehydratase (DH), and enoylreductase (ER) can modify the growing acyl chain (Hopwood 1997). Usually, the last domain of PKSI is a thioesterase, which catalyzes the release and the cyclization of the completed acyl chain (Fig. 1).
Several synthases combine both PKSI and nonribosomal peptide synthases (NRPS) modules. The NRPS are also modular multienzymes, which build peptides with amino acids as the PKSI build polyketides with acyl groups (Du et al. 2001).
Hopwood (1997) proposed that the PKSI diversity has evolved by gene duplication followed by sequence divergence. In addition to this mechanism, horizontal gene transfer (HGT) has been invoked as a major force in bacterial evolution (Gogarten et al. 2002; Lopez 2003). Even if HGT must be invoked with precaution (Kurland et al. 2003), the specific genetic context of secondary metabolism in actinomycetales supports the view of many HGTs within this order. The complete genome sequence of Streptomyces coelicolor reveals 23 gene clusters devoted to secondary metabolism and representing a significant fraction (4.5%) of the genome (Challis and Hopwood 2003). The extensive genetic diversity of biosynthetic clusters among actinomycetales (especially that of genus Streptomyces) exceeds a theoretical phenotypical potential and, therefore, supports HGT (Weber et al. 2003). The aim of this paper is to highlight the PKSI gene’s evolutionary history within bacteria using phylogenetic reconstructions from well-characterized PKSI clusters.
Materials and Methods
Published Sequences
Polyketide synthases were selected when the produced polyketides were experimentally characterized. Thus, 23 biosynthetic pathways encompassing 203 protein sequences of the KS domain were computed (Fig. 2). The KS domain acts as a generic condensing function, whereas the AT domain is substrate-specific. The KS domain was retained for phylogenetic analyses. The AT and ACP domains were analyzed in the particular case of the close relationship among amphotericin, nystatin, and pimaricin synthases. The ACP domain holds the growing chain whatever its length. This domain would have been useful since, like the KS domain, its function is generic. However, the ACP domain does not conceal enough informative sites due to its short length. Thus, the phylogeny of ACP domains was determined only for the closest polyketide synthases.
Phylogenetic Reconstruction and Analysis
Domains of public protein sequences were aligned using DbClustal (Thompson et al. 2000). Then the alignments were manually corrected using SeaView (Galtier et al. 1996). Phylogenetic reconstructions were performed via the distance method using neighbor joining (NJ) with the PAM matrix through Phylo_win (Galtier et al. 1996) and the maximum likelihood method (ML) using the Jones–Taylor–Thornton (JTT; 1992) model of substitution through PhyML (Guindon and Gascuel 2003). All trees were built with 500 bootstrap replicates. For the ML reconstruction, 500 data sets were generated using the SEQBOOT program from the PHYLIP package v3.6 (Felsenstein 1993). A tree was built for each replicate with PhyML, then bootstraps were computed with CONSENSE (Felsenstein 1993). All trees were drawn with NJplot (Perrière and Gouy 1996) and rooted with a fatty acid synthase (mas; Uniprot accession No. Q02251), which is a protein family that has evolved apart from PKSI (Hopwood 1997; Zhu et al 2002).
Relative rate tests were performed with RRTree (Robinson-Rechavi and Huchon 2000). This software compares substitution rates between DNA and protein sequences grouped in phylogenetically defined lineages. The Kishino–Hasegawa–Templeton (KHT) test (Kishino and Hasegawa 1989) was computed using ProML (Felsenstein 1993). It allows testing of phylogenetic hypotheses through likelihood comparison of phylogenetic trees.
Results
Three main arguments are assumed to underlie HGT: (i) exceptions to the clustering of KS domains from actinomycetales PKSI; (ii) the specific history of amphotericin, nystatin, and pimaricin synthases; and (iii) the dual origin of the soraphen synthase.
Exceptions to KS Clustering from One Actinomycetale PKSI
The majority of analyzed KS domains are clustered under a reliable node (Fig. 3; marked with an asterisk; 100/97 as ML/NJ bootstrap values). All these KS domains appear to be encoded by actinomycetales species. The rest of the KS domains are located within a blend group including Nostocales, Chroococcales, Myxococcales, and Pseudomonadales species (Fig. 2). In most cases, the KS domains of a given actinomycetale PKSI were clustered as a monophyletic group. As an example, 12 KS domains from the avermectin synthase (aveA1 to aveA4; Fig. 2) are clustered. Such groupings of KS domains are only supported within the actinomycetales group, as previously reported for a shorter set of 92 KS domains (Lopez 2003). Several exceptions to the rule of KS domain clustering in actinomycetales were observed, however. The KS domains of ascomycin (fkb), rapamycin (rap), and rifamycin (rif) synthases showed an imperfect clustering pattern since the KS domains from fkbA4, rapC4, and rifA2 did not fit these groupings (Figs. 3 and 5A). Moreover, in the case of mycinamicin synthase, four KS domains, mycAI2, mycAIII, mycAIII2, and mycAIV, are clustered apart from the other mycA KS domains (Fig. 3). The four KS gene regions exhibit a low GC content (66%) for a Streptomyces gene (Fig. 2). Two other examples of phylogenetic incongruencies concern pikAI3 (pikromycin) and rifA2 (rifamycin) (Fig. 3), which fall outside their PKSI group.
Phylogenetic analysis of ketosynthase (KS) domains. Reconstruction was computed for 204 protein sequences on 413 sites using both the distance method (NJ;PAM matrix) and the maximum likelihood method (BIONJ; JTT matrix) with 500 bootstrap replicates. Both distance and maximum likelihood (ML) methods provided the same tree topology, suggesting that observed grouping did not result from computational artefacts. Bootstrap values are indicated as ML/NJ. Only bootstrap values higher than 70 are indicated, except when necessary. When KS domains of one polyketide synthase are grouped, they are represented as a triangle labeled with the synthesized polyketide. The module names (e.g., A, B, C, etc.) are explicitly written for the amphotericin, nystatin, or pimaricin synthases. The number of amino acid substitutions is proportional to the length of the scale. An asterisk marks the actinomycetales node. The entire tree is available at our Web site (http://web.libragen.com/Phylogeny).
The Particular Case of Amphotericin, Nystatin, and Pimaricin Synthases
The amphotericin (amp) and nystatin (nys) synthases are very similar (Caffrey et al. 2001). Their modular organizations are identical (Fig. 4) and most of the domains are close pairwise (Fig. 5). In other words, considering the KS, AT, and ACP phylogenies, nysA is strongly clustered with ampA and not with other nys domains (Fig. 5). The additional domains of the reductive loop are mostly identical, even when the domains are nonfunctional (Fig. 4). The second to the sixth modules of nysI PKS were originally described with additional dehydratase domains (DH), which were noted to be nonfunctional due to large deletions (Brautaset et al. 2000). However, these domains were probably erroneously annotated as DH domains instead of AT–KR linkers (Yadav et al. 2003). Thus, the reductive loops (i.e., additional domains DH, KR, and ER; Fig. 1) of amp and nys synthases are identical. The amphotericin and nystatin synthases are mostly symmetrical (Fig. 4; top).
Hypothetical evolution for the amphotericin, nystatin, and pimaricin synthases. The three polyketide synthases are aligned at the top and their putative evolutionary history is proposed below. A rectangle represents a PKSI module with the implied necessary KS, AT, and ACP domains, which are not labeled. In other words, the empty rectangle of pimS0 represents one module containing the KS–AT–ACP domains. The additional reductive domains are indicated inside each rectangle as dehydratase (DH), ketoreductase (KR), or enoylreductase (ER). The inactive domains are labeled within lowercase letters. The gray rectangle highlights the domains that are closely associated with the KS, AT, and ACP phylogenies (Fig. 5). The curved arrows indicate hypothetical duplications based on phylogenies, and the plus signs, the addition of units.
Phylogenetic subtrees for the amphotericin, nystatin, and pimaricin PKSI synthases based on (A) KS domains, (B) AT domains, and (C) ACP domains. The two subtrees from KS and AT phylogenies are extracted from the complete reconstruction described in Materials and Methods. The ACP phylogeny was computed (NJ, PAM) with the 51 domains from the three polyketide synthases on 84 sites. The substrate incorporation for AT domains is indicated as malonyl (M) and methylmalonyl (mM).
Among the five PKSI of the pimaricin synthase, only the KS domains from pimS2 and pimS3 appear clustered with the amp/nys KS domains. All the KS domains from ampI, nysI, pimS2 and KS domains from ampJ2, nysJ2, pimS3 are clustered as trios (Fig. 5A). These modules are closely associated (Fig. 4) within the KS, AT, and ACP domain phylogenies with high bootstrap values (Fig. 5).
The Soraphen Synthase is Separated Between Two Bacterial Orders
Three KS domains from the soraphen (sor) synthase of a myxococcale (Sorangium cellulosum) were clustered in the actinomycetales group, whereas the four remaining domains were strongly clustered in the blend group (Fig. 3).
KS with Unusual Functions
The active site of KS domains is usually a cysteine. This amino acid can be replaced by a glutamine or a serine, leading to the KSQ or KSS domain, respectively. The KSQ domains lack condensation activity and were observed grouped apart from their PKSI (Fig. 3). The KSs domains also lack condensation activity and their decarboxylase activity is low (Caffrey et al. 2001). They were observed inside their polyketide synthase of origin (Fig. 5A).
The KS associated with 2 AT merged in one loading module have to condense two substrates instead of one (Ligon et al. 2002) and were grouped with KSQ. Lastly, the KS domains involved in hybrid NRPS/PKSI structures condense an acyl group onto an amino acid chain and were all clustered, whatever their origin (Fig. 3).
Discussion
According to the hypothesis of PKSI gene evolution, modular organization and chemical diversity of polyketides produced by PKSI should have arisen from duplication of domains (Hopwood 1997; Zhu et al. 2002). Thus, more sequence similarity is expected between the domains of a given PKSI than those of different ones (Gokhale and Khosla 2000) if the time elapsed between duplication and speciation is not too long. In this last case, the pattern observed would certainly be two closely associated clusters such as the amphotericin and nystatin synthases. Indeed, many PKSIs computed in this study support this expectation (Figs. 3 and 5). This indicates that PKSIs in actinomycetales are more homogeneous than in other orders. This assumption was confirmed by comparison of the two means of pairwise differences between the actinomycetale group (n = 12,403, mean = 0.4788) and the blend group (n = 1081, mean = 0.7979), which were significantly different by Mann–Whitney test (p < 2.2 × 10−16). This homogeneity may result from a high frequency of HGT within the actinomycetales group.
Several possible examples of HGT events were observed, especially between actinomycetales species. HGT would ensure genetic coherence and slow diversification between related bacteria (Torsvik et al. 2002). In the case of closely related species, frequent HGT events might also create similarities as vertical inheritance (Gogarten et al. 2002). Hence, HGT is a likely explanation for the homogeneity of the actinomycetales group.
Phylogenetic Incrongruencies Reveal HGT Events
Evidence for the transfer of single domains was observed as for the KS domains fkbB4 and rapC4, which are separated from their PKSI of origin (Fig. 3). Rapamycin and ascomycin synthases are PKSIs produced by two strains of Streptomyces hygroscopicus and involved in the synthesis of two closely related immunosuppressive macrolides (Wu et al. 2000). Moreover, since fkbA4 and rapC4 are very similar, the rapamycin and ascomycin synthases could have acquired this subunit by a recent HGT (Hopwood 1997).
It was suggested that the domains from the mycinamicin synthase, whose genes display a low GC content, resulted from a HGT (Anzai et al. 2003). Since atypical sequences from HGT tend to homogenize with the receptor genetic features (Gogarten et al. 2002; Lawrence and Roth 1996), this transfer must be recent. Indeed, the GC content (66%) is not actually optimized for an actinomycetale like Micromonospora griseorubida (72%). Nevertheless, this GC deviation was found in the third position of codons (Anzai et al. 2003). Since the phylogenetic reconstructions were based on protein sequences, most of the deviation was blurred due to the genetic code redundancy. Therefore, the taxonomic order of the donor organism cannot be determined.
The KS domains of Streptomyces species were already described as a monophyletic group (Moffitt and Neilan 2003). However, the strongly supported node for the actinomycetales group also contains the KS domains from a myxococcale, Sorangium cellulosum (soraphen producer) (Fig. 3). The monophyletic phylogenetic hypothesis, where all soraphen KS domains would be clustered outside the actinomycetales node (100/97), was tested with the KHT test and rejected (ln L/SD = 6.29 > 1.96), Schupp et al. (1995) suggested that a genetic exchange occurred between actinomycetales and S. cellulosum, based on their GC content and codon usage. Such a clustering pattern may support the hypothesis that half of the soraphen cluster was acquired by HGT from an actinomycetales species.
HGT Versus the Alternative Evolution Hypothesis
The main alternative hypothesis to explain the homogeneity of actinomycetales could rely on a recent origin of PKSI for the actinomycetales, despite the fact that they exhibit great polyketide diversity. Such a hypothesis is ruled out by the analysis of the blend group. The PKSIs appear to be mixed between different bacterial phyla (Fig. 3). The common bacterial ancestor therefore probably carried a large diversity of PKSIs which predated the origin of actinomycetales. Explaining the relative homogeneity within actinomycetales by common ancestry would imply that the initial diversity was lost, then re-created through de novo duplications and specialization — little parsimonious hypothesis. Alternatively, we propose that numerous HGTs between actinomycetales may have secondarily homogenized PKSI genes in this phylum. Eventually, this pattern may be due to a higher evolution rate of the PKSI genes from nonactinomycetales species. However, no significant difference was observed between the relative evolution rates of actinomycetales and nonactinomycetales, thus excluding this hypothesis (p = 0.673).
Another hypothesis would rely on a phylogenetic reconstruction artifact: long branch attraction. First, we computed the reconstruction by two different methods—distance and maximum likelihood—and the same topology was obtained. Second, we computed a new distance-based reconstruction without the longest branches of the hybrid group, the KS from loading modules, and the closest KS from the root: the three KSs from the soraphen synthase. The tree topology remains unchanged and the actinomycetales group is always supported by a bootstrap value of 99.
One can argue that transferred genes are stable only under positive selection. Thus, alien genes must be functional and provide a novel adaptive advantage; otherwise they are lost by random mutation (Kurland et al. 2003). Secondary metabolites like polyketides respond to that constraint: they are synthetized under selective pressures and do not modify primary metabolism. The functionality of PKSI in a heterologous environment was demonstrated with the production of a functional megalomicin PKSI in Streptomyces lividans and Saccharopolyspora erythraea, despite the fact that this PKSI gene cluster was originally described in Micromonospora megalomicea (Volchegursky et al. 2000; Shah et al. 2000).
Putative Evolutionary History of Amphotericin,Nystatin, and Pimaricin Synthases
The three KS domains from ampA, nysA, and pimS0 are KSS. However, even if pimS0 and nysA share a KSs, their similarities do not seem sufficient to speculate a common origin (Brautaset et al. 2000). This active site mutation could have been acquired twice independently by convergent evolution.
The close pairwise-associated domains (Fig. 4; gray rectangle) observed with the three phylogenies (Fig. 5) suggest a common origin for these three synthases. The amphotericin and nystatin synthases are highly similar, implying a recent common origin. The pimS2 PKS appears to have been integrated though a HGT from an amp or a nys gene. The pimS2 gene shows a peculiar codon usage and third-codon position GC content compared to other pimaricin synthase genes (Aparicio et al. 2000). This region, which includes seven modules, represents a large HGT event of 34 kb.
Hybrid NRPS/PKSI and Unusual KS Function
The KS domains positioned after a NRPS module possess an unusual function that mediates the condensation of an acyl extender unit onto the amino acid chain from the previous NRPS module. This functional constraint might explain why each KS domain preceded by a NRPS module clustered within the hybrid group and not within its own PKSI.
It has been suggested that epoB was acquired by HGT, based on several rigorous criteria (Lopez 2003). One of them is the phylogenetic incongruence, where epoB is excluded from the epothilone PKS but is clustered within the hybrid NRPS/PKS group. Lopez suggested that this hybrid cluster does not result from a common ancestor but from a convergent evolution, since several hybrids, such as KS domains from the rapamycin and ascomycin synthases, do not cluster in this group. However, these two cases revealed by Lopez (2003) are not exceptions since the hybrid group concerns only the KS preceded by a NRPS module and not positioned after PKS domains as in the rapamycin or ascomycin synthase. Nevertheless, the GC deviation, the presence of flanking transposon sequences, and the great genetic distance consistently support the hypothesis that HGT is responsible for the occurrence of epoB (Lopez 2003). Moreover, the hypothesis of convergent evolution is likely to be true, since the KS domains from hybrid systems are clustered due to a functional constraint. The function of KSS domains seems to be very low processive decarboxylation activity and remains to be explained. These domains do not seem to have evolved under strong functional pressure, unlike the other unusual KS domains.
Polyketide Synthase Genes May Evolve with Mobile Elements
Since large HGTs of PKSI clustered genes have occurred between actinomycetales and not within the other bacterial orders, a biological mechanism must exist, specific to actinomycetales. We suggest that the linearity and the instability of actinomycetales chromosomes could be implicated in this mechanism.
The chromosome of Streptomyces species is linear and its extremities are genetically unstable since large deletions and circularization occur at significant frequencies (up to 1%) (Chen et al. 2002; Weaver et al. 2004). Many of the biosynthetic pathways of Streptomyces coelicolor and Streptomyces avermitilis were found on chromosome arms, whereas the essential genes were located in the core chromosome (Weber et al. 2003; Challis and Hopwood 2003). Linear plasmids can acquire chromosomal end genes (Wiener et al. 1998; Weaver et al. 2004) and reciprocally (Bentley et al. 2004; Weaver et al. 2004). Therefore, these gene clusters might be transferred through extrachromosomal elements within the genus Streptomyces and other species with a high GC content, such as actinomycetales.
The antibiotic gene clusters and their associated resistance genes are likely to be cotransferred, such as in the case of the streptomycin synthase (Egan et al. 2001; Challis and Hopwood 2003) and the tylosin gene cluster (Stonesifer et al. 1986). The streptomycin and its resistance gene were demonstrated to be transferred at a high frequency between Streptomyces species (Wiener et al. 1998; Egan et al. 2001). Entire biosynthetic pathways may have evolved through HGT in a similar way as did pathogenicity islands (Thompson et al. 2002).
The presence of transposon-like sequences near biosynthetic gene clusters may support genetic exchanges in the cell. Direct repeated sequences are present at both extremities of the PKSII mithramycin gene cluster (Lombó et al. 1999) and suggest a transfer through the integration of extrachromosomal DNA. Then a gene cluster similar to the mithramycin cluster was found on the linear plasmid pSLA2-L (Mochizuki et al. 2003). This hypothesis of transfer between genetic structures was also confirmed for the methylenomycin gene cluster, which can be located on a circular plasmid, on a linear plasmid, and integrated into the S. coelicolor chromosome (Wiener et al. 1998; Yamasaki et al. 2003). The gene transfer of the methylenomycin gene cluster from the circular to the linear plasmid was confirmed (Yamasaki et al. 2003). Concerning PKSI genes, the tylosin gene cluster was demonstrated to be highly transferred through conjugation between Streptomyces fradie strains (Stonesifer et al. 1986). Under selective pressures, some S. fradie strains seem to have acquired the tylosin gene cluster from other strains. A Mycobacterium ulcerans PKSI gene cluster, which likely encodes a mycolactone PKSI, is located between two repeated insertion sequences (Jenkin et al. 2003; Stinear et al. 2004). Moreover, subtractive hybridization experiments performed between Mycobacterium marinum and M. ulcerans demonstrated that the most important DNA acquisition by M. ulcerans was this PKSI cluster and the two insertion sequences (Jenkin et al. 2003). This gene cluster is located on a giant plasmid, suggesting evolution through a recent horizontal acquisition followed by duplication (Stinear et al. 2004). Such an association of insertion sequences with PKSI genes is a strong argument for HGT (Lopez 2003). This first insight of PKSI associated with insertion sequences highlights the possible genetic exchanges of these biosynthetic clusters, adding further credibility to the hypothesis of frequent HGT in actinomycetales PKSI.
Conclusion
Several observations based on KS domain phylogeny suggest that frequent HGTs of PKSIs genes have occurred between actinomycetales species. The homogeneity and the robustness of the actinomycetales group may result in part from these genetic transfers. The linearity and instability of actinomycetales chromosome and the many possibilities for natural conjugation and for the transfer of genetic elements could explain the high frequency of HGT. The suggestion is advanced here that their homogeneity may rely on the genetic specificity of actinomycetales.
The Selfish Operon Theory (Lawrence and Roth 1996) proposes that genes evolved clustered in operon through HGT to maintain their frequencies in the gene pool. In this theory, a selective advantage for the host is not required since genes from operons are clustered for themselves to avoid genetic drift. This model is well adapted to the PKSI gene clusters (Lopez 2003). HGT maintains the capability for numerous genetic responses to environmental stress in bacterial populations without strong environmental pressures. Therefore, numerous genetic exchanges would enable rapid adaptation if the environmental pressures appear again.
References
Y Anzai N Saito M Tanaka K Kinoshita Y Koyama K Kato (2003) ArticleTitleOrganization of the biosynthetic gene cluster for the polyketide macrolide mycinamicin in Micromonospora griseorubida FEMS Microbiol Lett 218 135–341 Occurrence Handle10.1016/S0378-1097(02)01123-0 Occurrence Handle1:CAS:528:DC%2BD3sXhtVGjs7Y%3D Occurrence Handle12583909
JF Aparicio R Fouces MV Mendes N Olivera JF Martin (2000) ArticleTitleA complex multienzyme system encoded by five polyketide synthase genes is involved in the biosynthesis of the 26-membered polyene macrolide pimaricin in Streptomyces natalensis Chem Biol 7 895–905 Occurrence Handle1:CAS:528:DC%2BD3cXosVWitLo%3D Occurrence Handle11094342
SD Bentley S Brown LD Murphy DE Harris MA Quail J Parkhill BG Barrell JR McCormick RI Santamaria R Losick M Yamasaki H Kinashi CW Chen G Chandra D Jakimowicz HM Kieser T Kieser KF Chater (2004) ArticleTitleSCP1, a 356 023 bp linear plasmid adapted to the ecology and developmental biology of its host, Streptomyces coelicolor A3(2) Mol Microbiol 51 1615–1628 Occurrence Handle10.1111/j.1365-2958.2003.03949.x Occurrence Handle1:CAS:528:DC%2BD2cXivFCrs74%3D Occurrence Handle15009889
T Brautaset ON Sekurova H Sletta TE Ellingsen AR Strom S Valla SB Zotchev (2000) ArticleTitleBiosynthesis of the polyene antifungal antibiotic nystatin in Streptomyces noursei ATCC 11455: analysis of the gene cluster and deduction of the biosynthetic pathway Chem Biol 7 395–403 Occurrence Handle1:CAS:528:DC%2BD3cXktl2nsLk%3D Occurrence Handle10873841
P Caffrey S Lynch E Flood S Finnan M Oliynyk (2001) ArticleTitleAmphotericin biosynthesis in Streptomyces nodosus Deductions from analysis of polyketide synthase and late genes. Chem Biol 8 713–723 Occurrence Handle1:CAS:528:DC%2BD3MXltVGju7s%3D
GL Challis DA Hopwood (2003) ArticleTitleSynergy and contingency as driving forces for the evolution of multiple secondary metabolite production by Streptomyces species Proc Natl Acad Sci USA 100 14555–14561 Occurrence Handle10.1073/pnas.1934677100 Occurrence Handle1:CAS:528:DC%2BD3sXpsFaku78%3D Occurrence Handle12970466
CW Chen CH Huang HH Lee HH Tsai R Kirby (2002) ArticleTitleOnce the circle has been broken: dynamics and evolution of Streptomyces chromosomes Trends Genet 18 522–529
J Cortes SF Haydock GA Roberts DJ Bevitt PF Leadlay (1990) ArticleTitleAn unusually large multifunctional polypeptide in the erythromycin-producing polyketide synthase of Saccharopolyspora erythraea Nature 348 176–178
E Cundliffe N Bate A Butler S Fish A Gandecha L Merson- Davies (2001) ArticleTitleThe tylosin-biosynthetic genes of Streptomyces fradiae Antonie Van Leeuwenhoek 79 229–234 Occurrence Handle10.1023/A:1012065300116 Occurrence Handle1:CAS:528:DC%2BD3MXpt12lsrw%3D Occurrence Handle11816964
L Du C Sanchez B Shen (2001) ArticleTitleHybrid peptide-polyketide natural products: biosynthesis and prospects toward engineering novel molecules Metab Eng 3 78–95 Occurrence Handle10.1006/mben.2000.0171 Occurrence Handle1:CAS:528:DC%2BD3MXnvFOnsg%3D%3D Occurrence Handle11162234
S Egan P Wiener D Kallifidas EM Wellington (2001) ArticleTitlePhylogeny of Streptomyces species and evidence for horizontal transfer of entire and partial antibiotic gene clusters Antonie Van Leeuwenhoek 79 127–133 Occurrence Handle10.1023/A:1010296220929 Occurrence Handle1:CAS:528:DC%2BD3MXmslWkt7w%3D Occurrence Handle11519998
Felsenstein J (1993) PHYLIP (Phylogeny Inference Package) version 3.5c. Distributed by the author. Department of Genetics, University of Washington, Seattle
N Gaitatzis B Silakowski B Kunze G Nordsiek H Blocker G Hofle R Muller (2002) ArticleTitleThe biosynthesis of the aromatic myxobacterial electron transport inhibitor stigmatellin is directed by a novel type of modular polyketide synthase J Biol Chem 277 13082–13090 Occurrence Handle10.1074/jbc.M111738200 Occurrence Handle1:CAS:528:DC%2BD38XivFSjtrw%3D Occurrence Handle11809757
N Galtier M Gouy C Gautier (1996) ArticleTitleSEAVIEW and PHYLO_ WIN: two graphic tools for sequence alignment and molecular phylogeny Comput Appl Biosci 12 543–548 Occurrence Handle1:CAS:528:DyaK2sXhtlWktLw%3D Occurrence Handle9021275
JP Gogarten WF Doolittle JG Lawrence (2002) ArticleTitleProkaryotic evolution in light of gene transfer Mol Biol Evol 19 2226–2238 Occurrence Handle1:CAS:528:DC%2BD38Xps12hsL4%3D Occurrence Handle12446813
RS Gokhale C Khosla (2000) ArticleTitleRole of linkers in communication between protein modules Curr Opin Chem Biol 4 22–27 Occurrence Handle10.1016/S1367-5931(99)00046-0 Occurrence Handle1:CAS:528:DC%2BD3cXhtlygtrw%3D Occurrence Handle10679375
S Guindon O Gascuel (2003) ArticleTitleA simple, fast, and accurate algorithm to estimate large phytogenies by maximum likelihood Syst Biol 52 696–704 Occurrence Handle10.1080/10635150390235520 Occurrence Handle14530136
D Hoffmann JM Hevel RE Moore BS Moore (2003) ArticleTitleSequence analysis and biochemical characterization of the nostopeptolide A biosynthetic gene cluster from Nostac sp. GSV224 Gene 311 171–180 Occurrence Handle10.1016/S0378-1119(03)00587-0 Occurrence Handle1:CAS:528:DC%2BD3sXltFGjsbc%3D Occurrence Handle12853152
DA Hopwood (1997) ArticleTitleGenetic contributions to understanding polyketide synthases Chem Rev 97 2465–2497 Occurrence Handle10.1021/cr960034i Occurrence Handle1:CAS:528:DyaK2sXntFWks7o%3D Occurrence Handle11851466
H Ikeda T Nonomiya M Usami T Ohta S Omura (1999) ArticleTitleOrganization of the biosynthetic gene cluster for the polyketide anthelmintic macrolide avermectin in Streptomyces avermitilis Proc Natl Acad Sci USA 96 9509–9514 Occurrence Handle1:CAS:528:DyaK1MXlsVyms74%3D Occurrence Handle10449723
GA Jenkin TP Stinear PDR Johnson JK Davies (2003) ArticleTitleSubtractive hybridization reveals a type I polyketide synthase locus specific to Mycobacterium ulcerans J Bacteriol 185 6870–6882 Occurrence Handle10.1128/JB.185.23.6870-6882.2003 Occurrence Handle1:CAS:528:DC%2BD3sXpt1ynu7Y%3D Occurrence Handle14617651
DT Jones WR Taylor JM Thornton (1992) ArticleTitleThe rapid generation of mutation data matrices from protein sequences CABIOS 8 275–282 Occurrence Handle1633570
SJ Kakavas L Katz D Stassi (1997) ArticleTitleIdentification and characterization of the niddamycin polyketide synthase genes from Streptomyces caelestis J Bacteriol 179 7515–7522 Occurrence Handle1:CAS:528:DyaK2sXnvFeksbw%3D Occurrence Handle9393718
H Kishino M Hasegawa (1989) ArticleTitleEvaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in hominoidea J Mol Evol 29 170–179 Occurrence Handle1:CAS:528:DyaL1MXkvFCnsbc%3D Occurrence Handle2509717
CG Kurland B Canback OG Berg (2003) ArticleTitleHorizontal gene transfer: a critical view Proc Natl Acad Sci USA 100 9658–9662 Occurrence Handle10.1073/pnas.1632870100 Occurrence Handle1:CAS:528:DC%2BD3sXmvVejtrw%3D Occurrence Handle12902542
JG Lawrence JR Roth (1996) ArticleTitleSelfish operon theory: Horizontal gene transfer may drive the evolution of gene clusters Genetics 143 1843–1860 Occurrence Handle1:CAS:528:DyaK28XlvVSqt78%3D Occurrence Handle8844169
J Ligon S Hill J Beck R Zirkle I Molnar J Zawodny S Money T Schupp (2002) ArticleTitleCharacterization of the biosynthetic gene cluster for the antifungal polyketide soraphen A from Sorangium cellulosum So ce26 Gene 285 257–267 Occurrence Handle10.1016/S0378-1119(02)00396-7 Occurrence Handle1:CAS:528:DC%2BD38XitlWmu7g%3D Occurrence Handle12039053
F Lombó AF Braña C Méndez JA Salas (1999) ArticleTitleThe mithramycin gene cluster of Streptomyces argillaceus contains a positive regulatory gene and two repeated DNA sequences that are located at both ends of the cluster J Bacteriol 181 642–647 Occurrence Handle9882681
JV Lopez (2003) ArticleTitleNaturally mosaic operons for secondary metabolite biosynthesis; variability and putative horizontal transfer of discrete catalytic domains of the epothilone polyketide synthase locus Mol Gen Genomics 270 420–431 Occurrence Handle10.1007/s00438-003-0937-9 Occurrence Handle1:CAS:528:DC%2BD3sXps1Kjs7k%3D
S Mochizuki K Hiratsu M Suwa T Ishii F Sugino K Yamada H Kinashi (2003) ArticleTitleThe large linear plasmid pSLA2-L of Streptomyces rochei has an unusually condensed gene organization for secondary metabolism Mol Microbiol 48 1501–1510
MC Moffitt BA Neilan (2003) ArticleTitleEvolutionary affiliations within the superfamily of ketosynthases reflect complex pathway associations J Mol Evol 56 446–457 Occurrence Handle10.1007/s00239-002-2415-0 Occurrence Handle12664164
I Molnar T Schupp M Ono R Zirkle M Milnamow B Nowak-Thompson N Engel C Toupet A Stratmann DD Cyr J Gorlach JM Mayo A Hu S Goff J Schmid JM Ligon (2000) ArticleTitleThe biosynthetic gene cluster for the microtubule-stabilizing agents epothilones A and B from Sorangium cellulosum So ce90 Chem Biol 7 97–109 Occurrence Handle10.1016/S1074-5521(00)00075-2 Occurrence Handle10662695
B Nowak-Thompson SJ Gould JE Loper (1997) ArticleTitleIdentification and sequence analysis of the genes encoding a polyketide synthase required for pyoluteorin biosynthesis in Pseudomonas fluorescens Pf-5 Gene 204 17–24 Occurrence Handle10.1016/S0378-1119(97)00501-5 Occurrence Handle9434161
G Perrière M Gouy (1996) ArticleTitleWWW-Query: an on-line retrieval system for biological sequence banks Biochimie 78 364–369 Occurrence Handle10.1016/0300-9084(96)84768-7 Occurrence Handle1:CAS:528:DyaK28XmvVCmuro%3D Occurrence Handle8905155
M Robinson-Rechavi D Huchon (2000) ArticleTitleRRTree: Relative-rate tests between groups of sequences on a phylogenetic tree Bioinformatics 16 296–297 Occurrence Handle10.1093/bioinformatics/16.3.296 Occurrence Handle10869026
T Schupp C Toupet B Cluzel S Neff S Hill JJ Beck JM Ligon (1995) ArticleTitleA Sorangium cellulosum (myxobacterium) gene cluster for the biosynthesis of the macrolide antibiotic soraphen A: cloning, characterization, and homology to polyketide synthase genes from actinomycetes J Bacteriol 177 3673–3679 Occurrence Handle7601830
T Schwecke JF Aparicio I Molnar A Konig LE Khaw SF Haydock M Oliynyk P Caffrey J Cortes JB Lester GA Bohm J Stauton PF Leadlay (1995) ArticleTitleThe biosynthetic gene cluster for the polyketide immunosuppressant rapamycin Proc Natl Acad Sci USA 92 7839–7843 Occurrence Handle7644502
S Shah Q Xue L Tang JR Carney M Betlach R McDaniel (2000) ArticleTitleCloning, characterization and heterologous expression of a polyketide synthase and P-450 oxidase involved in the biosynthesis of the antibiotic oleandomycin J Antibiot 53 502–508 Occurrence Handle1:CAS:528:DC%2BD3cXjslejurg%3D Occurrence Handle10908114
B Shen (2003) ArticleTitlePolyketide biosynthesis beyond the type I, II and III polyketide synthase paradigms Curr Opin Chem Biol 7 285–295 Occurrence Handle10.1016/S1367-5931(03)00020-6 Occurrence Handle12714063
B Silakowski HU Schairer H Ehret B Kunze S Weinig G Nordsiek P Brandt H Blocker G Hofle S Beyer R Muller (1999) ArticleTitleNew lessons for combinatorial biosynthesis from myxobacteria The myxothiazol biosynthetic gene cluster of Stigmatella aurantiaca DW4/3-1 J Biol Chem 274 37391–37399 Occurrence Handle10.1074/jbc.274.52.37391 Occurrence Handle10601310
B Silakowski G Nordsiek B Kunze H Blocker R Muller (2001) ArticleTitleNovel features in a combined polyketide synthase/non-ribosomal peptide synthetase; the myxalamid biosynthetic gene cluster of the myxobacterium Stigmatella aurantiaca Sga15 Chem Biol 8 59–69 Occurrence Handle10.1016/S1074-5521(00)00056-9 Occurrence Handle11182319
J Staunton KJ Weissman (2001) ArticleTitlePolyketide biosynthesis: a millennium review Nat Prod Rep 18 380–416 Occurrence Handle10.1039/a909079g Occurrence Handle1:CAS:528:DC%2BD3MXltlyqsbo%3D Occurrence Handle11548049
TP Stinear A Mve-Obiang PLC Small W Frigui MJ Pryor R Brosch GA Jenkin PDR Johnson JK Davies RE Lee S Adusumilli T Gamier SF Haydock PF Leadlay ST Cole (2004) ArticleTitleGiant plasmid-encoded polyketide synthases produce the macrolide toxin of Mycobacterium ulcerans Proc Natl Acad Sci USA 101 1345–1349 Occurrence Handle10.1073/pnas.0305877101 Occurrence Handle1:CAS:528:DC%2BD2cXhtlWjtbs%3D Occurrence Handle14736915
J Stonesifer P Matsushima RH Baltz (1986) ArticleTitleHigh frequency conjugal transfer of tylosin genes and amplifiable DNA in Streptomyces fradiae Mol Gen Genet 202 348–355 Occurrence Handle10.1007/BF00333261 Occurrence Handle1:CAS:528:DyaL28XktVWnurs%3D Occurrence Handle3458997
L Tang YJ Yoon CY Choi CR Hutchinson (1998) ArticleTitleCharacterization of the enzymatic domains in the modular polyketide synthase involved in rifamycin B biosynthesis by Amycolatopsis mediterranei Gene 216 255–265 Occurrence Handle10.1016/S0378-1119(98)00338-2 Occurrence Handle1:CAS:528:DyaK1cXlvFOjsbw%3D Occurrence Handle9729415
CJ Thompson D Fink LD Nguyen (2002) ArticleTitlePrinciples of microbial alchemy: insights from the Streptomyces coelicolor genome sequence Genome Biol 3 1–4 Occurrence Handle10.1186/gb-2002-3-7-reviews1020
JD Thompson F Plewniak J-C Thierry O Poch (2000) ArticleTitleDbClustal: rapid and reliable global multiple alignments of protein sequences detected by database searches Nucleic Acids Res 28 2919–2926 Occurrence Handle10.1093/nar/28.15.2919 Occurrence Handle10908355
D Tillett E Dittmann M Erhard H Dohren Particlevon T Borner BA Neilan (2000) ArticleTitleStructural organization of microcystin biosynthesis in Microcystis aeruginosa PCC7806: an integrated peptide-polyketide synthetase system Chem Biol 7 753–764 Occurrence Handle10.1016/S1074-5521(00)00021-1 Occurrence Handle1:CAS:528:DC%2BD3cXotVWitrw%3D Occurrence Handle11033079
V Torsvik L Ovreas TF Thingstad (2002) ArticleTitleProkaiyotic Diversity–Magnitude, dynamics, and controlling factors Science 296 1064–1066 Occurrence Handle10.1126/science.1071698 Occurrence Handle12004116
Y Volchegursky Z Hu L Katz R McDaniel (2000) ArticleTitleBiosynthesis of the anti-parasitic agent megalomicin: transformation of erythromycin to megalomicin in Saccharopolyspora erythraea Mol Microbiol 37 752–762
C Waldron P Matsushima PR Rosteck SuffixJr MC Broughton J Turner K Madduri KP Crawford DJ Merlo RH Baltz (2001) ArticleTitleCloning and analysis of the spinosad biosynthetic gene cluster of Saccharopolyspora spinosa Chem Biol 8 487–499 Occurrence Handle10.1016/S1074-5521(01)00029-1 Occurrence Handle11358695
K Watanabe CCC Wang CN Boddy DE Cane C Khosla (2003) ArticleTitleUnderstanding substrate specificity of polyketide synthase modules by generating hybrid multimodular synthases J Biol Chem 278 42020–42026 Occurrence Handle10.1074/jbc.M305339200 Occurrence Handle12923197
D Weaver N Karoonuthaisiri HH Tsai CH Huang ML Ho S Gai KG Patel J Huang SN Cohen DA Hopwood CW Chen CM Kao (2004) ArticleTitleGenome plasticity in Streptomyces: identification of 1 Mb TIRs in the S. coelicolor A3(2) chromosome Mol Microbiol 51 1535–1550 Occurrence Handle10.1111/j.1365-2958.2003.03920.x Occurrence Handle15009883
T Weber K Welzel S Pelzer A Vente W Wohlleben (2003) ArticleTitleExploiting the genetic potential of polyketide producing Streptomyces J Biotechnol 106 221–223 Occurrence Handle10.1016/j.jbiotec.2003.08.004 Occurrence Handle1:CAS:528:DC%2BD3sXpsVSrtLo%3D Occurrence Handle14651864
P Wiener S Egan MH Wellington (1998) ArticleTitleEvidence for transfer of antibiotic-resistance genes in soil populations of Streptomycetes Mol Ecol 7 1205–1216 Occurrence Handle10.1046/j.1365-294x.1998.00450.x Occurrence Handle1:CAS:528:DyaK1cXmtFOisb0%3D Occurrence Handle9734076
K Wu L Chung P Revill L Katz CD Reeves (2000) ArticleTitleThe FK520 gene cluster of Streptomyces hygroscopicus var. ascomyceticus (ATCC 14891) contains genes for biosynthetis of unusual polyketide extender units Gene 251 81–90 Occurrence Handle1:CAS:528:DC%2BD3cXktV2itLk%3D Occurrence Handle10863099
Y Xue D Wilson DH Sherman (2000) ArticleTitleGenetic architecture of the polyketide synthases for methymycin and pikromycin series macrolides Gene 245 203–211 Occurrence Handle1:CAS:528:DC%2BD3cXhsFanurs%3D Occurrence Handle10713461
G Yadav RS Gokhale D Mohanty (2003) ArticleTitleComputational approach for prediction of domain organization and substrate specificity of modular polyketide synthases J Mol Biol 328 335–363 Occurrence Handle10.1016/S0022-2836(03)00232-8 Occurrence Handle1:CAS:528:DC%2BD3sXislehtr0%3D Occurrence Handle12691745
M Yamasaki Y Ikuto A Ohira K Chater H Kinashi (2003) ArticleTitleLimited regions of homology between linear and circular plasmids encoding methylenomycin biosynthesis in two independently isolated streptomycetes Microbiology 149 1351–1356
G Zhu MJ LaGier F Stejskal JJ Millership X Cai JS Keithly (2002) ArticleTitleCryptosporidium parvum: the first protist known to encode a putative polyketide synthase Gene 298 79–89 Occurrence Handle1:CAS:528:DC%2BD38XotFOmur8%3D Occurrence Handle12406578
Acknowledgment
We are indebted to the Pôle Bioinformatique Lyonnais for the kind access to their databanks and informatic resources. We gratefully acknowledge Jean-Luc Pernodet and Ludovic Orlando for the fruitful discussions regarding the manuscript. This contribution was supported by European project METACONTROL QLK3-CT-2002-02068.
Author information
Authors and Affiliations
Corresponding author
Additional information
Reviewing Editor : Dr. Nicolas Galtier
Rights and permissions
About this article
Cite this article
Ginolhac, A., Jarrin, C., Robe, P. et al. Type I Polyketide Synthases May Have Evolved Through Horizontal Gene Transfer. J Mol Evol 60, 716–725 (2005). https://doi.org/10.1007/s00239-004-0161-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-004-0161-1