Abstract
Tom40 proteins represent an essential class of molecules which facilitate translocation of unfolded proteins from the cytosol into the mitochondrial intermembrane space. They are part of a high-molecular mass complex that forms the protein-conducting channel in outer mitochondrial membranes. This study concerns the recombinant expression, purification and folding of amino-terminally truncated variants of the two human Tom40 isoforms for structural biology experiments. Both CD and FTIR secondary structure analysis revealed a dominant beta-sheet structure and a short alpha-helical part for both proteins together with a high thermal stability. Two secondary structure elements can be denatured independently. Reconstitution of the recombinant protein into planar lipid bilayers demonstrated ion channel activity similar to Tom40 purified from Neurospora crassa mitochondrial membranes, but conductivity fingerprints differ from the structurally closely related VDAC proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The first step in mitochondrial protein import is mediated by a multisubunit protein-conducting channel (TOM complex) located in the outer membrane, which comprises up to seven different subunits (Chacinska et al. 2009; Endo and Yamano 2009; Endo et al. 2011; Mokranjac and Neupert 2009; Nussberger and Neupert 2002; Prokisch et al. 2002). Various receptor proteins selectively recognize different substrates. Although some subunits differ or are absent among species (Macasev et al. 2000, 2004; Perry et al. 2006; Poynor et al. 2008), all TOM complexes comprise a 40-kDa protein, termed Tom40, as a major component. Based on single-channel measurements (Ahting et al. 2001; Hill et al. 1998; Poynor et al. 2008; Romero-Ruiz et al. 2010) of isolated Tom40, an ion- and peptide-conducting pore in lipid membranes was demonstrated. It may thus also function as the actual protein-conducting channel that facilitates the transfer of virtually all mitochondrial preproteins synthesized in the cytosol. To date, most Tom40 homologs involved in this translocation process have been described for mitochondria of Saccharomyces cerevisiae, Neurospora crassa, Arabidopsis thaliana, Homo sapiens and Rattus norvegicus (Kinoshita et al. 2007; Schwartz and Matouschek 1999; Suzuki et al. 2000; Werhahn et al. 2001, 2003; Perry et al. 2006).
Electron microscopy, electrophysiology and biochemical studies measuring the effect of rigid gold labels introduced into precursor proteins during import into mitochondria of S. cerevisiae indicated Tom40 pore diameters of ~20 Å (Hill et al. 1998; Kinoshita et al. 2007; Schwartz and Matouschek 1999; Suzuki et al. 2000; Werhahn et al. 2001, 2003; Ahting et al. 2001). This diameter is sufficient to accommodate unfolded protein chains or partially folded domains. Detailed structural and functional studies addressing the interaction of preproteins with purified mammalian Tom40 protein, however, have been hampered by the considerable complexity to isolate the protein from higher eukaryotic organisms.
In this report we explored the possibility of expressing human Tom40 isoforms in Escherichia coli and purifying these proteins from inclusion bodies under denaturing conditions. We show that purified, amino-terminally truncated variants of human Tom40A and Tom40B (abbreviated as hTom40AΔN and hTom40BΔN) can functionally be refolded by rapid dilution of denatured protein in different detergent solutions. Secondary structure analyses demonstrated significant β-barrel architecture for both proteins. Reconstitution into planar lipid bilayers and electrophysiology studies showed that both channels are ion-conducting.
Materials and Methods
Cloning
The genes of human Tom40 isoforms A and B were synthesized in truncated form. hTom40A encoding a gene product missing amino acids 1–82 and hTom40B missing amino acids 1–29 were cloned into the NcoI and XhoI cloning sites of expression vector pET24d (Novagen, Darmstadt, Germany) under control of a T7 promotor (Trenyzme Biotechnology, Konstanz, Germany). Both genes encoded proteins with a carboxy-terminal hexahistidinyl tag.
Bacterial Expression and Preparation of Inclusion Bodies
Human Tom40 proteins were expressed as inclusion bodies in E. coli BL21-CodonPlus (DE3) cells (Stratagene, Waldbronn, Germany). Cells were grown in LB medium using shaking flasks at 37°C. Tom40 expression was induced with 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG) at a cell density corresponding to an OD600 of 0.6. After 6 h of growth, cells were harvested by centrifugation. The expression of human Tom40 was confirmed by SDS-PAGE and Western blotting using mammalian Tom40-specific antibodies.
For purification of Tom40 isoforms, cells were washed twice with PBS buffer and resuspended in 10 ml PBS/g cells supplemented with 2 mg/ml DNaseI from bovine pancreas (Sigma-Aldrich, Hamburg, Germany). After incubation on ice for 20 min, cells were lysed in a French pressure cell (AMINCO; American Instrument Exchange, Haverhill, MA). Inclusion bodies were separated from cell debris by centrifugation at 20,000×g for 20 min at 4°C. The inclusion body pellet was washed twice with buffer containing 50 mM Tris and 100 mM NaCl, pH 8. Inclusion bodies were subsequently solubilized in 6 M guanidine hydrochloride, 20 mM Tris, 150 mM NaCl, and 1 mM β-mercaptoethanol (pH 8), using a glass–glass homogenizer. To remove unsolubilized material the homogenate was centrifuged at 30,000×g for 30 min at 4°C, and supernatants were recovered and stored at 4°C for further use.
Purification and Refolding of Tom40 Isoforms
Human Tom40A and B were isolated from inclusion bodies under denaturing conditions in 6 M guanidine hydrochloride, 20 mM Tris (pH 8), 150 mM NaCl, 1 mM β-mercaptoethanol using an Äkta Basic system (GE Healthcare, Uppsala, Sweden) and 1 or 20 ml Ni-Sepharose HiTrap columns (GE Healthcare). Then, the purified proteins were diluted to ~5 mg/ml and kept at 4°C.
For refolding, human Tom40A and B proteins (~5 mg/ml) in 6 M guanidine hydrochloride were diluted dropwise 10- to 20-fold in different refolding buffer solutions at 4°C. After 1, 24 and 170 h, the efficiency of protein folding was assessed by evaluation of the amount of precipitated protein after centrifugation at 15,000×g for 10 min at 4°C and determination of the protein concentration in the supernatant by UV absorption at a wavelength of 280 nm. The buffer substances used were 50 mM citric acid (pH 5), 50 mM sodium-phosphate buffer (pH 6), 50 mM Tris buffer (pH 7) and 50 mM glycine-sodium-hydroxide (pH 10) and contained 1 mM β-mercaptoethanol. The detergents were either n-dodecyl-β-d-maltoside (DDM; Glycon, Luckenwalde, Germany), lauryl-dimethylamine-oxide (LDAO, Sigma-Aldrich), n-octyl-polyoxyethylene (oPOE; Bachem, Weil am Rhein, Germany), Brij35 (Sigma-Aldrich), 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonate (CHAPS; Merck, Darmstadt, Germany) and n-octyl-β-d-glucopyranoside (β-OG, Glycon) at concentrations corresponding to ~5× CMC, respectively.
Optimal refolding of human Tom40A and B was achieved by a 10-fold dilution of 5 mg/ml protein in 6 M guanidine hydrochloride into 20 mM Tris (pH 8), 150 mM NaCl, 1 mM β-mercaptoethanol and 1% (w/v) LDAO at 4°C. For further use the LDAO concentration was reduced to 0.5% by dropwise addition of the same buffer without detergent. Final samples were centrifuged at 100,000×g, and supernatants were subjected to Ni-NTA chromatography to concentrate the refolded protein. Residual amounts of contaminants were removed by gel filtration using a Superose 12 column (GE Healthcare).
All fractions containing Tom40 were merged and concentrated using spin columns with a 30-kDa cut-off (Vivaspin, GE Healthcare) up to 10 mg/ml. The concentrations of hTom40A and hTom40B were determined by UV absorbance spectroscopy using extinction coefficients ε W,280nm ≈ 5,600 M−1 cm−1 for tryptophan and ε Y,280nm ≈ 1,200 M−1 cm−1 for tyrosine (Pace et al. 1995). All samples were filtered (Rotilabo® filter, pore size 0.22 μm; Carl Roth, Karlsruhe, Germany) and centrifuged at 16,000×g for 5 min to remove possible protein aggregates prior to circular dichroism (CD) and Fourier transform infrared spectroscopy (FTIR) measurements.
Far UV-CD Spectroscopy
CD spectroscopic measurements were performed using a J-815 spectrometer (Jasco, Tokyo, Japan) in quartz cuvettes of 0.1 cm path length. Spectra at 20°C were recorded from 185 to 250 nm with a resolution of 1.0 nm and an acquisition time of 20 nm/min. To avoid high-voltage output from the photomultiplier at high temperature measurements, CD spectra were recorded from 195 to 250 nm. Final CD spectra were obtained by averaging five consecutive scans and corrected for background by subtraction of spectra of protein-free samples recorded under the same conditions. Mean residue ellipticity (Θ) was calculated based on the molar protein concentration and the number of amino residues of the Tom40 isoforms. The protein concentration used for CD spectroscopy was adjusted to 0.2–0.5 mg/ml.
The secondary structure content was estimated using the CDpro package, according to Sreerama and Woody (2000, 2003, 2004), namely, CDSSTR, CONTIN/LL and SELCON 3. Melting curves were recorded at constant wavelength at 216 nm for hTom40AΔ1–82 and BΔ1–29 from 20 to 98°C by applying a temperature ramp of 1°C/min. The percentage of unfolded protein content, f U(T), was calculated according to
where ΘU(T) = aT + b and Θ N (T) = cT + d represent the pre- and posttransition baselines, respectively. For evaluating the protein melting temperature, T m, the resulting data f U(T) were fitted by the Boltzmann equation, f U(T) = (f 0 − f max)/(1 + exp (T − T m)/T s) + f max, where f 0, f max and T s are the minimum and maximum percentages of unfolded protein content and the temperature range over which the transition occurs, respectively (Pace et al. 1998; Fersht 1999).
FTIR Spectroscopy
Samples for FTIR with a protein concentration of ~5 mg/ml were dialyzed against 20 mM Tris (pH 8), 150 mM NaCl, 1 mM β-mercaptoethanol and 0.5% LDAO at 4°C for 48 h. FTIR spectra were recorded using a Confocheck TENSOR 27 FTIR spectrometer (Bruker Optics, Ettlingen, Germany) equipped with a linear, photovoltaic, mercury–cadmium–telluride detector (Bruker Optics). All spectra were recorded between 4,000 and 800 cm−1 for 25 s with a wave number resolution of 4 cm−1. To avoid temperature-induced variations of the water signal, the measurement cell was kept at a constant temperature of 20°C. For each spectrum, 32 interferograms were collected and averaged. The aperture setting was 6 mm, and the scanner velocity was at 10 kHz. All procedures were carried out to optimize the quality of the spectrum in the amide I region, between 1,600 and 1,700 cm−1. Secondary structure predictions were calculated from baseline corrected spectra with the multivariant pattern recognition method (Fabian and Schultz 2000) of the OPUS Quant 2 software supplied by Bruker Optics. Spectral data were factorized and compared to reference data from a library of more than 40 proteins with known structure (Bruker Optics, source: Protein Data Bank, http://www.pdb.org).
Electrophysiology
Human Tom40AΔ1–82 and BΔ1–29 in LDAO or o-POE were reconstituted into black lipid membranes, and channel currents were recorded according to standard protocols (Arnold et al. 2007; Poynor et al. 2008; Romero-Ruiz et al. 2010). A 0.5% solution of diphytanoyl phosphatidylcholine (Avanti Polar Lipids, Alabaster, AL) in n-decane/butanol (9:1) was spread over a circular hole of 250 μm diameter in the partitioning wall of a cylindrical Delrin cup of a bilayer chamber BCH-13A (Warner Instruments, Hamden, CT). The chamber was filled symmetrically with 1 M KCl and 20 mM Hepes (pH 7.2). Protein solution enriched with cholesterol was added to the cis side of the membrane to a final concentration of 10–50 μg/ml. Current fluctuations through single and multiple channels were recorded in voltage-clamp mode using an EPC-8 patch-clamp amplifier (HEKA Electronik, Lambrecht, Germany) connected to the bilayer chambers by a pair of Ag/AgCl pellet electrodes (WPI, Sarasota, FL). Current signals were low pass–filtered at 3 kHz using the built-in Bessel-filter of the amplifier and monitored for channel insertion using an analog oscilloscope. Current and voltage signals from the amplifier were digitized at a sampling rate of 10 kHz per channel using a NI-USB-6251 interface (National Instruments, Austin, TX) controlled by a program of the Strathclyde electrophysiology suite (WinEDR 2.8 or WinWCP 3.6, J. Dempster).
Results
Expression and Purification
Tom40 isoforms exhibit sequence identities between mammalian species of >91% for Tom40A and 96% for Tom40B. To obtain large amounts of human Tom40 for biochemical and structural studies, human Tom40A and B were cloned as N-terminally truncated proteins (hTom40AΔ1–82 and hTom40BΔ1–29, termed hTom40AΔN and hTom40BΔN; Fig. 1a). Since the N termini of both proteins are predicted to be disordered, we rationalized that they are not essential for formation of the mitochondrial protein translocation pore (see also supplementary Fig. S1 for the secondary structure prediction of hTom40AΔN). Our construction of hTom40AΔ1-82 and hTom40BΔ1-29 was in line with studies on truncated rat Tom40AΔΝ (Suzuki et al. 2004) and the published mammalian voltage-dependent anion channel 1 (VDAC1) structures (Bayrhuber et al. 2008; Hiller et al. 2008; Ujwal et al. 2008; Zeth 2010). Hexahistidine-tagged hTom40AΔN and hTom40BΔN were expressed in E. coli as inclusion bodies. Bacterial cells grew to a mass of 5 g/l of culture. The wet yield of inclusion bodies ranged between 2 and 3 g/l of culture. Furthermore, testing the functional refolding of recombinant full-length bovine Tom40A (supplementary Fig. S2) revealed a yield of ~5 mg protein/l of E. coli culture, which is insufficient for functional and structural studies.
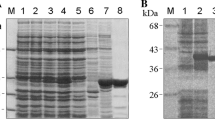
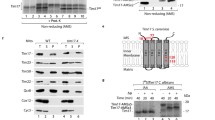
Secondary structure prediction and purification of hTom40A and B. a PSIPRED-predicted secondary structure of full-length hTom40A and hTom40B. hTom40A and B were truncated by 82 and 29 amino acids, respectively. b, c Purification of human Tom40A and B. Inclusion bodies isolated from E. coli containing recombinant hTom40AΔ1–82 (hTom40AΔN) and hTom40BΔ1–29 (hTom40BΔN) with C-terminal hexahistidinyl tags were solubilized by 6 M guanidine hydrochloride and loaded onto an Ni-NTA affinity column. hTom40A and B were eluted under denaturing conditions with 300 mM imidazole (Ni-NTA I) and refolded by rapid dilution into 0.5% LDAO. Refolded proteins were subjected to Ni-NTA chromatography (Ni-NTA II) and eventually passed over a Superose12 size-exclusion column (SEC). Aliquots of the resulting column fractions were analyzed by SDS-polyacrylamide gel electrophoresis and Coomassie blue staining. The Ni-NTA II eluate containing hTom40B shows an additional strong band (*), which has been characterized by mass spectrometry as a degradation product. Lane 1, E. coli cells expressing hTom40A or B; lane 2, purified inclusion bodies containing hTom40A or B; lanes 2–5, Ni-NTA column fractions and imidazole eluates; lane 6, peak fraction of size exclusion column
Human Tom40A (hTom40AΔN) was already highly enriched in inclusion bodies (Fig. 1b). The first purification of unfolded protein in chaotropic buffer revealed a rather pure protein. The yield starting from 1 l of culture was ~200 mg of protein. Optimized refolding was achieved using LDAO detergent containing buffer at pH 8 (see Table 1). No or little refolded protein was observed at only slightly decreased pH below 7 or in the presence of detergents such as DDM, oPOE or n-octyl-β-d-glucoside (Tables 1, 2). To concentrate refolded human Tom40, the samples were again applied to Ni-NTA affinity and size exclusion chromatography (Fig. 1b, c) for further purification was performed, if needed. When using affinity chromatography before and after refolding, unspecifically bound proteins have been efficiently removed with low imidazole concentrations. After size exclusion chromatography the protein was virtually pure (see Fig. 1b, c).
By contrast, hTom40BΔN was expressed at a lower level (Fig. 1c), and its yield ranged between 20 and 30 mg/l of culture. Nevertheless, the amount of protein obtained was sufficient for electrophysiology, secondary structure determination experiments like CD and FTIR spectroscopy and first crystallization trials. The protocol for pure and refolded protein essentially followed the protocol applied for the Tom40A version (see Fig. 1b).
Secondary Structure Determination by CD Spectroscopy
The secondary structure content and thermal stability of hTom40AΔN and hTom40BΔN were analyzed in LDAO by CD spectroscopy and compared to spectra of known β-barrel proteins. The spectra showed a clear dominance of beta-sheet content (Fig. 2; Table 3). The minimum of the spectra consistently clustered around 216 nm, with a crossover of the baseline at around 203 nm. At wavelengths >245 nm, the CD spectrum approached ellipticity values close to zero, indicating that the hTom40ΔN preparations were virtually free of any higher-order aggregates which would cause light-scattering effects and interfere with the interpretation of the data. Deconvolution of the data points via CDpro revealed a predominance of β-sheet secondary structure (>30%) for human Tom40s with low α-helical content (<24%) (Table 3). Hence, efficient refolding of truncated hTom40s can be assumed under the conditions chosen. This is further supported by the high thermal stability of hTom40AΔN and BΔN (Tm of 73.1 and 74.1°C; Fig. 2c, d). Thus, the truncated amino acids of the Tom40 isoforms might not be involved in the overall barrel structure. In addition, hTom40AΔN and BΔN exhibit similar CD spectral characteristics with slightly left-shifted spectra for hTom40 isoform B.
Far UV CD spectra and thermal stability of hTom40AΔN and hTom40BΔN. a, b hTom40AΔN (~0.2 mg/ml) and hTom40BΔN (~0.2 mg/ml) were solubilized in 0.5% LDAO and 20 mM Tris (pH 8.0). For each CD spectrum, five scans were accumulated at 20°C and background-corrected. Mean residue ellipticity was calculated based on the molar protein concentration and the number of amino acid residues of the according protein. The spectra of hTom40A and B indicate a high ratio of β-sheet with a minimum at 216 nm and a crossover of the baseline at 203 nm. c, d Melting curves of hTom40A and hTom40B determined from corrected CD signals measured at a wavelength of 216 nm. The melting temperatures of hTom40AΔN and hTom40BΔN were at 73 and 74°C, respectively. Insets represent spectra recorded at 20°C (solid line), 50°C (dashed line) and 70°C (dashed line)
An interesting observation was obtained during the analysis of the CD spectra of hTom40s when recorded just below the melting point of the protein at 70°C. We found that both CD spectra of hTom40 isoforms underwent a remarkable right-shift in combination with decreased ellipticity values in comparison to CD spectra recorded at 20°C (Fig. 2a, b) and 50°C (Fig. 2c, d). Determination of secondary structure content displayed a substantial loss in the α-helical content with an increase in β-sheet portion (see supplementary Table 1). For hTom40AΔN and BΔN the α-helical content decreased from 20 to 5% and from 24 to 12%, respectively. The corresponding relative β-sheet content increased from 36 to 43% and from 32 to 37%. The relative amount of random coil and turn structure did not change significantly with temperature. Secondary structure development of hTom40AΔΝ (Fig. 2c) and BΔN (Fig. 2d) during melting showed that, first, the α-helical portion is reduced during temperature increase, which indicates that the N-terminal helix denatures before the barrel portion follows. Again, hTom40BΔN showed small differences, whereas the overall transition was similar to hTom40AΔN (Fig. 2a, b). This might be due to a higher melting point of hTom40BΔN compared to hTom40AΔN. It is noteworthy that similar observations were gained for Tom40 from Aspergillus fumigatus in Brij35 and LDAO (data not shown).
Secondary Structure Determination by FTIR Spectroscopy
To determine a complementary number for the secondary structure and to support the CD spectroscopic data, hTom40AΔN and BΔN were analyzed by FTIR. The amide I band (1,700–1,600 cm−1) of the IR spectra of both isoforms is shown in Fig. 3 as it represents the region in which α-helical, β-sheets and random coil structures are activated. The maximum was at 1,632 cm−1, which is indicative of β-sheet content. The spectra show a clear dominance of β-sheet contribution, which is comparable to measurements of other β-barrel proteins like human VDAC1 (Engelhardt et al. 2007). As calculated from reference data of more than 40 proteins with known 3D structure, the amount of β-sheet was calculated to be >55% (Table 3). At wave numbers of 1,650 cm−1 the spectra showed no significant local maximum. Comparisons with reference spectra indicated an α-helical content of <10%. The additional maximum at 1,656 cm−1 indicates antiparallel β-sheet and hints at the organization of the β-sheets in the protein. These measurements support the model of a β-barrel protein with an α-helical elongation at the N terminus. In agreement with CD spectroscopic data, the content of β-sheet is dominant in the structure, while the α-helical amount is relatively low.
FTIR spectra of hTom40AΔN and hTom40BΔN. The IR spectra of hTom40AΔN (~7 mg/ml) and hTom40BΔN (~5 mg/ml) represent an average of three independent spectra with 32 consecutive scans each. Spectral bands assigned to the α-helical structure are centered at a wave number of about 1,650 cm−1, random components at 1,645–1,640 cm−1 and β-sheet at 1,630–1,625 cm−1. The shoulder at 1,695 cm−1 indicates an antiparallel β-sheet with particularly short turns. For all spectra, the baseline was subtracted. The main peak at 1,625 cm−1 for hTom40AΔN and hTom40BΔN, respectively, indicates high β-sheet content
Pore-Forming Activity of hTom40AΔN and hTom40BΔN
To test whether purified hTom40ΔN is active and capable of conducting ion flux, the isolated and refolded hTom40AΔN was inserted in an artificial planar lipid bilayer. The channel-forming activity of the protein was tested upon application of different voltages, and the subsequent current traces were recorded. The traces were measured with single channels as well as multiple channels inserted in the artificial lipid layer. For single-channel characterization a planar membrane was reconstituted and the zero current was measured as baseline. Protein was added to the cis side of the chamber, while a voltage of 30 mV was applied and current traces were recorded. Single-channel insertions have been observed (Fig. 4a). For human Tom40AΔN the channel showed a conductivity of ~1 nS at 1 M KCl (Fig. 4b). For human Tom40BΔN the single-channel conductivity could not be determined yet.
Channel conductance of human Tom40AΔN and BΔN. a Purified hTom40AΔN was added to both sides of a planar lipid membrane formed by DiphPC/n-decane/butanol, and single-channel conductance was measured in 1 M KCl and 50 mM HEPES (pH 7.2) at a membrane potential of 30 mV. b Histogram of channel conductance measured at voltages between −50 and +50 mV. A total of 82 single-channel insertions were analyzed. Most channel insertions showed a conductance of 1.2 nS. c, d Voltage dependence of hTom40A and hTom40B channel conductance: hTom40AΔN and hTom40BΔN were reconstituted into black lipid membranes, and current traces were recorded at constant voltage between −160 and +160 mV at 10-mV increments. The membrane conductance, Gnorm, was calculated according to Ohm’s law and normalized to the mean conductance at +10 mV
To test whether the inserted channels show voltage-dependent gating, several Tom40AΔN and Tom40BΔN channels were allowed to reconstitute into planar lipid membranes and steps of different voltages, starting from ±10 mV and ending at ±160 mV, were applied. Baseline currents at 0 mV were always measured in between. For human Tom40AΔN and BΔN the recorded currents showed no voltage-dependent gating at voltages below 120 mV. The conductivity of multiple channels, which was measured at different voltages, remained the same. For hTom40AΔN the measurements showed a slight decrease of conductivity at voltages higher than 120 mV (Fig. 4c). For hTom40BΔN no voltage-dependent gating was observed even at voltages above 120 mV (Fig. 4d). Predominantly, a voltage-independent behavior was observed for both isoforms.
Discussion
The TOM complex of the outer mitochondrial membrane is responsible for the translocation of proteins from the cytosol to the mitochondrial inner membrane space. From electron microscopic analysis it is well established that TOM complexes can show up to three pores (Künkele et al. 1998; Model et al. 2002, 2008), which are based on the presence of single Tom40 molecules. We suppose this holo TOM complex to incorporate three Tom40 monomers to form a trimer. The core complex may incorporate two Tom40 monomers to form a dimer (Mager et al. 2010). How these monomers assemble with the other components of the holo complex; the receptors Tom70, Tom20 and Tom22; as well as the small proteins Tom5, Tom6 and Tom7 to the high-molecular mass complex of 400–500 kDa (Künkele et al. 1998; Model et al. 2002, 2008) is not yet clear.
The presence of several Tom40 isoforms has been shown for other mammalian organisms like Rattus norvegicus (Kinoshita et al. 2007) and Bos taurus (unpublished data). The C-terminal part of the proteins, which forms the barrel walling, shows an identity of >60% for both human isoforms. Both isoforms can occur in the same tissue in different amounts and form part of the TOM complex (Kinoshita et al. 2007). Whether both isoforms can be present in the same complex is not yet determined.
The N-terminus of Tom40 isoforms differs significantly among several species. In Arabidopsis thaliana, the N-terminus is extended to 50 amino acids and in human Tom40A a proline-rich region presents a rather unfolded N-terminal extension. Due to the specific sequence contribution, this extension may contribute to actin binding (Quinlan and Kerkhoff 2008). However, it was reported not to be responsible for targeting and subsequent assembly with preexisting Tom subunits (Humphries et al. 2005). Generally, polyproline-rich sequences result in unstructured parts of proteins and are known to form the basis, e.g., domains known to polymerize actin (FH1, WH2 domains) (Quinlan and Kerkhoff 2008). Nevertheless, the function of the N-terminal helix may take part in the SAM-mediated assembly of the monomer in the complex (Humphries et al. 2005) or may play a role in the interaction with other components of the TOM complex, like the Tom22 receptor (Taylor et al. 2003).
Refolding of the protein was induced by methods which were already successfully applied to, e.g., hVDAC isoforms 1 and 2 (Engelhardt et al. 2007). Several detergents induced refolding but at differing amounts. As for the VDAC proteins, LDAO turned out to be the most successful detergent in terms of protein remaining in solution. Another parameter which was clearly important is the pH of the refolding solution. As for VDAC and, e.g., the BamA receptor protein from E. coli, the basic pH increased the refolding excess dramatically, while pH values lower than 7 did in part not reveal any refolded portion of the protein (data on BamA are not published). This behavior may be explained by the close to neutral isoelectric point (pI) of the protein, which is ~6.8 for the truncated version of hTom40A and ~6.4 for the truncated hTom40B protein.
Structural analysis of Tom40 in comparison to VDAC1 with several secondary structure predicting programs revealed a similar pattern of β-sheets. A comparison of the experimentally determined secondary structure of human Tom40 and human VDAC by CD spectroscopy, FTIR and prediction by Psipred (Jones 1999) is given in Table 3 and shows the similarity between the three proteins. This leads to the hypothesis of a similar structure and therefore a similar evolutionary origin, as recently suggested (Bayrhuber et al. 2008; Pusnik et al. 2009; Zeth 2010). According to this topological similarity, Tom40 is predicted to form a β-barrel with 19 strands and an α-helical part at the N-terminus of the protein facing the cytosol (see Fig. S1). Previous studies on Tom40 from fungi, plants and mammals did not address the role of the putative N-terminal α-helical domain of the protein specifically.
This study underlines the predicted β-sheet structure by CD spectroscopy and FTIR and demonstrates the formation of intact channels for both human Tom40 isoforms, which are capable of conducting an ion flux through a lipid membrane. Values for secondary structure are supported by the crystal structure of human VDAC1 and with the secondary structure prediction of other Tom40 proteins. These results are in perfect agreement with former studies on fungal and mammalian Tom40 proteins, suggesting a β-barrel structure for Tom40 in all species (Becker et al. 2005; Hill et al. 1998; Kinoshita et al. 2007; Ahting et al. 2001; Suzuki et al. 2000) and an N-terminal α-helical segment adjacent to the β-barrel (Suzuki et al. 2004). Further, previous studies on mammalian VDAC1 determined similar CD spectrum characteristics: a minimum at ~216 nm, a zero-crossing wavelength at 206 nm and 39.7% β-sheet, together with 17.5% α-helix, 13.3% turn and 28.4% random coil (Malia and Wagner 2007).
Together, these findings suggest the existence of an N-terminal α-helical domain in the 3D structure of human Tom40s that is not necessary for the stabilization of the overall Tom40 β-barrel structure. This fits perfectly with the observed decrease in ellipticity values at increased temperatures as α-helical structures result in much stronger signals than β-sheet structures. Taking into account secondary structure predictions of Tom40 from various organisms, an N-terminal α-helix is predicted in all species in addition to a β-sheet-rich core. In addition, Suzuki et al. (2004) have revealed that the recombinant membrane-embedded C-terminal half of rat Tom40 (Tom40∆1–165) constitutes the preprotein recognition domain with an enriched β-sheet structure. However, this part of the protein clone consisted of only 13 β-strands and not 19, as for the constructs used in this work. Thus, the location of the α-helix at the N-terminus of the Tom40 protein can be suggested. This is in good agreement with the crystal/NMR structure of mammalian VDAC1, featuring a flexible N-terminal α-helix located inside the VDAC channel (Abu-Hamad et al. 2009; Bayrhuber et al. 2008; Hiller et al. 2008; Ujwal et al. 2008).
Assessment of the secondary structure of Tom40 from N. crassa and S. cerevisiae suggested about 16 β-strands for the formation of a functional pore (Becker et al. 2005). Studies addressing the location of functional domains in rat Tom40A indicated that the pore is restricted to the C-terminal segment of the protein, which lacks the first three putative β-strands (Suzuki et al. 2004). Our results predict that Tom40 belongs to the “mitochondrial porin” superfamily with 19 β-strands (Pusnik et al. 2009) that incorporates both the VDACs and Tom40 proteins from mitochondrial outer membranes.
The channel conductance of hTom40AΔN was in the same range as the recombinant yeast Tom40 measured at 250 mM KCl, with states at ~210, ~260, ~440 and ~430 pS (Harsman et al. 2010) as well as native Tom40 from N. crassa investigated in 1 M KCl with +0.4, ~0.8, ~1.4, ~2.0 and ~2.9 nS (Ahting et al. 2001; Poynor et al. 2008; Romero-Ruiz et al. 2010). The channel properties of hTom40AΔN could not yet be compared with those of rat Tom40 (Suzuki et al. 2004) or any other Tom40 protein as they remain to be described.
The electrophysiologic data of both hTom40 isoforms showed no significant voltage dependence. For hTom40AΔN, channel closure has been observed in some experiments and only at voltages above 100 mV. This stands in contrast to electrophysiologic data of hVDAC1, where movement of the N-terminal helix inside the channel has been predicted to result in reduced conductivity of >30% at voltages above 50 mV (Engelhardt et al. 2007; Ujwal et al. 2008). Since our measurements for this publication were performed with protein from different purifications and therefore several refolding experiments, it is possible that the orientation of the flexible N-terminal helix of hTom40 is not definite. If the N-terminal helix is facing into the barrel, it may affect the ion flux and show flickering currents similar to hVDAC1. However, when the N-terminal helix rises out of the barrel, it has no or limited effect on conductance. This would lead to a different behavior of the channel conductance regarding voltage dependence. The channel measurements of isoform B show only little conducting difference. But overall, isoform BΔN shows similar conducting behavior as isoform A and underlines the efficient refolding strategy.
More efforts are necessary to characterize the channel properties of both human Tom40 constructs in more detail. Nevertheless, our method for the expression, purification and refolding of mammalian Tom40 isoforms thus provides the basis for further structural and physiological investigations.
References
Abu-Hamad S, Arbel N, Calo D, Arzoine L, Israelson A, Keinan N, Ben-Romano R, Friedman O, Shoshan-Barmatz V (2009) The VDAC1 N-terminus is essential both for apoptosis and the protective effect of anti-apoptotic proteins. J Cell Sci 122:1906–1916
Ahting U, Thieffry M, Engelhardt H, Hegerl R, Neupert W, Nussberger S (2001) Tom40, the pore-forming component of the protein-conducting TOM channel in the outer membrane of mitochondria. J Cell Biol 153:1151–1160
Arnold T, Poynor M, Nussberger S, Lupas AN, Linke D (2007) Gene duplication of the eight-stranded β-barrel protein OmpX produces a functional pore: a scenario for the evolution of transmembrane β-barrels. J Mol Biol 336:1174–1184
Bayrhuber M, Meins T, Habeck M, Becker S, Giller K, Villinger S, Vonrhein C, Griesinger C, Zweckstetter M, Zeth K (2008) Structure of the human voltage-dependent anion channel. Proc Natl Acad Sci USA 105:15370–15375
Becker L, Bannwarth M, Meisinger C, Hill K, Model K, Krimmer T, Casadio R, Truscott KN, Schulz GE, Pfanner N, Wagner R (2005) Preprotein translocase of the outer mitochondrial membrane: reconstituted Tom40 forms a characteristic TOM pore. J Mol Biol 353:1011–1020
Chacinska A, Koehler CM, Milenkovic D, Lithgow T, Pfanner N (2009) Importing mitochondrial proteins: machineries and mechanisms. Cell 138:628–644
Endo T, Yamano K (2009) Multiple pathways for mitochondrial protein traffic. Biol Chem 390:723–730
Endo T, Yamano K, Kawano S (2011) Structural insight into the mitochondrial protein import system. Biochim Biophys Acta 1808:955–970
Engelhardt H, Meins T, Poynor M, Adams V, Nussberger S, Welte W, Zeth K (2007) High level expression, refolding and probing the natural fold of the human voltage-dependent anion channel isoforms I and II. J Membr Biol 216:93–105
Fabian H, Schultz CP (2000) Fourier transform infrared spectroscopy in peptide and protein analysis. In: Meyers RA (ed) Encyclopedia of analytical chemistry. Wiley, Chichester, pp 5779–5803
Fersht A (1999) Structure and mechanism in protein science: a guide to enzyme catalysis and protein folding. W.H. Freeman and Company, pp 1–631
Harsman A, Krüger V, Bartsch P, Honigmann A, Schmidt O, Rao S, Meisinger C, Wagner R (2010) Protein conducting nanopores. J Phys Condens Matter 22:454102
Hill K, Model K, Ryan MT, Dietmeier K, Martin F, Wagner R, Pfanner N (1998) Tom40 forms the hydrophilic channel of the mitochondrial import pore for preproteins. Nature 395:516–521
Hiller S, Garces RG, Malia TJ, Orekhov VY, Colombini M, Wagner G (2008) Solution structure of the integral human membrane protein VDAC-1 in detergent micelles. Science 321:1206–1210
Humphries AD, Streimann IC, Stojanovski D, Johnston AJ, Yano M, Hoogenraad NJ, Ryan MT (2005) Dissection of the mitochondrial import and assembly pathway for human Tom40. J Biol Chem 280:11535–11543
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202
Kinoshita J, Mihara K, Oka T (2007) Identification and characterization of a new Tom40 isoform, a central component of mitochondrial outer membrane translocase. J Biochem 141:897–906
Künkele KP, Heins S, Dembowski M, Nargang FE, Benz R, Thieffry M, Walz J, Lill R, Nussberger S, Neupert W (1998) The preprotein translocation channel of the outer membrane of mitochondria. Cell 93:1009–1019
Macasev D, Newbigin E, Whelan J, Lithgow T (2000) How do plant mitochondria avoid importing chloroplast proteins? Components of the import apparatus Tom20 and Tom22 from Arabidopsis differ from their fungal counterparts. Plant Physiol 123:811–816
Macasev D, Whelan J, Newbigin E, Silva-Filho MC, Mulhern TD, Lithgow T (2004) Tom22′, an 8-kDa trans-site receptor in plants and protozoans, is a conserved feature of the TOM complex that appeared early in the evolution of eukaryotes. Mol Biol Evol 21:1557–1564
Mager F, Sokolova L, Lintzel J, Brutschy B, Nussberger S (2010) LILBID-mass spectrometry of the mitochondrial preprotein translocase TOM. J Phys Condens Matter 22:454132
Malia TJ, Wagner G (2007) NMR structural investigation of the mitochondrial outer membrane protein VDAC and its interaction with antiapoptotic Bcl-xL. Biochemistry 46:514–525
Model K, Prinz T, Ruiz T, Radermacher M, Krimmer T, Kühlbrandt W, Pfanner N, Meisinger C (2002) Protein translocase of the outer mitochondrial membrane: role of import receptors in the structural organization of the TOM complex. J Mol Biol 316:657–666
Model K, Meisinger C, Kuhlbrandt W (2008) Cryo-electron microscopy structure of a yeast mitochondrial preprotein translocase. J Mol Biol 383:1049–1057
Mokranjac D, Neupert W (2009) Thirty years of protein translocation into mitochondria: unexpectedly complex and still puzzling. Biochim Biophys Acta 1793:33–41
Nussberger S, Neupert W (2002) Protein translocation across the outer membrane of mitochondria: structure and function of the TOM complex. In: Kasianowicz JJ, Kellermayer MSZ, Deamer DW (eds) Structure and dynamics of confined polymers. NATO science series. Kluwer Academic, Dordrecht, pp 67–84
Pace CN, Vajdos F, Fee L, Grimsley G, Gray T (1995) How to measure and predict the molar absorption coefficient of a protein. Protein Sci 4:2411–2423
Pace CN, Hebert EJ, Shaw KL, Schell D, Both V, Krajcikova D, Sevcik J, Wilson KS, Dauter Z, Hartley RW, Grimsley GR (1998) Conformational stability and thermodynamics of folding of ribonucleases Sa, Sa2 and Sa3. J Mol Biol 279:271–286
Perry AJ, Hulett JM, Likic VA, Lithgow T, Gooley PR (2006) Convergent evolution of receptors for protein import into mitochondria. Curr Biol 16:221–229
Poynor M, Eckert R, Nussberger S (2008) Dynamics of the preprotein translocation channel of the outer membrane of mitochondria. Biophys J 95:1511–1522
Prokisch H, Nussberger S, Westermann B (2002) Protein import into mitochondria of Neurospora crassa. Fungal Genet Biol 36:85–90
Pusnik M, Charriere F, Maser P, Waller RF, Dagley MJ, Lithgow T, Schneider A (2009) The single mitochondrial porin of Trypanosoma brucei is the main metabolite transporter in the outer mitochondrial membrane. Mol Biol Evol 26:671–680
Quinlan ME, Kerkhoff E (2008) Actin nucleation: bacteria get in-Spired. Nat Cell Biol 10:13–15
Romero-Ruiz M, Mahendran KR, Eckert R, Winterhalter M, Nussberger S (2010) Interactions of mitochondrial presequence peptides with the mitochondrial outer membrane preprotein translocase TOM. Biophys J 99:774–781
Schwartz M, Matouschek A (1999) The dimensions of the protein import channels in the outer and inner mitochondrial membranes. Proc Natl Acad Sci USA 9:13086–13090
Sreerama N, Woody RW (2000) Estimation of protein secondary structure from circular dichroism spectra: comparison of CONTIN, SELCON, and CDSSTR methods with an expanded reference set. Anal Biochem 287:252–260
Sreerama N, Woody RW (2003) Structural composition of betaI- and betaII-proteins. Protein Sci 12:384–388
Sreerama N, Woody RW (2004) Computation and analysis of protein circular dichroism spectra. Methods Enzymol 383:318–351
Suzuki H, Okazawa Y, Komiya T, Saeki K, Mekada E, Kitada S, Ito A, Mihara K (2000) Characterization of Rat TOM40, a central component of the preprotein translocase of the mitochondrial outer membrane. J Biol Chem 275:37930–37936
Suzuki H, Kadowaki T, Maeda M, Sasaki H, Nabekura J, Sakaguchi M, Mihara K (2004) Membrane-embedded C-terminal segment of rat mitochondrial TOM40 constitutes protein-conducting pore with enriched beta-structure. J Biol Chem 279:50619–50629
Taylor RD, McHale BJ, Nargang FE (2003) Characterization of Neurospora crassa Tom40-deficient mutants and effect of specific mutations on Tom40 assembly. J Biol Chem 278:765–775
Ujwal R, Cascio D, Colletier JP, Faham S, Zhang J, Toro L, Ping P, Abramson J (2008) The crystal structure of mouse VDAC1 at 2.3 A resolution reveals mechanistic insights into metabolite gating. Proc Natl Acad Sci USA 105:17742–17747
Werhahn W, Niemeyer A, Jansch L, Kruft VV, Schmitz UK, Braun HP (2001) Purification and characterization of the preprotein translocase of the outer mitochondrial membrane from Arabidopsis. Identification of multiple forms of TOM20. Plant Physiol 125:943–954
Werhahn W, Jänsch L, Braun HP (2003) Identification of novel subunits of the TOM complex of Arabidopsis thaliana. Plant Physiol Biochem 41:407–416
Zeth K (2010) Structure and evolution of mitochondrial outer membrane proteins of beta-barrel topology. Biochim Biophys Acta 1797:1292–1299
Acknowledgments
The authors thank Beate Nitschke for expert technical assistance and Tobias Kulschewski and Dr. Simon Stutz for their contributions at the early stage of this work. The work was supported by a Competence Network on Functional Nanostructures grant of the Baden-Württemberg Foundation and the Human Frontier Science Program to S. N. and K. Z., respectively.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mager, F., Gessmann, D., Nussberger, S. et al. Functional Refolding and Characterization of Two Tom40 Isoforms from Human Mitochondria. J Membrane Biol 242, 11–21 (2011). https://doi.org/10.1007/s00232-011-9372-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-011-9372-8