Abstract
Current taxonomy indicates a single global species of the Great Barracuda (Sphyraena barracuda) despite differences in color and behavior between Atlantic and Pacific forms. To investigate these differences and qualify the dispersal characteristics of this unique coastal–pelagic teleost (bony fish), we conducted a global phylogeographic survey of 246 specimens from thirteen sampling locations using a 629-base pair fragment of mtDNA cytochrome b. Data indicate high overall gene flow in the Indo-Pacific over large distances (>16,500 km) bridging several biogeographic barriers. The West Atlantic population contains an mtDNA lineage that is divergent from the Indo-Pacific (d = 1.9%), while the East Atlantic (N = 23) has two mutations (d = 0.6%) apart from the Indo-Pacific. While we cannot rule out distinct evolutionary partitions among ocean basins based on behavior, coloration, and near-monophyly between Atlantic and Indo-Pacific subpopulations, more investigation is required before taxonomic status is revised. Overall, the pattern of high global dispersal and connectivity in S. barracuda more closely resembles those reported for large oceanic predators than reef-associated teleosts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Reef-associated fishes predominately disperse during a pelagic larval phase, prior to a sedentary juvenile/adult stage with high-site fidelity to shallow reef habitat (reviewed by Strathmann 1993). The prevalence of a potentially dispersive larval stage in teleost (bony) reef fishes was initially thought to ensure long-distance connectivity (Weersing and Toonen 2009), but phylogeographic surveys sometimes reveal fine-scale barriers to gene flow (e.g., gobies Elactinus spp., damselfish Pomacentrus moluccensis, grouper Plectropomus leopardus, and surgeonfish Zebrasoma flavescens; Taylor and Hellberg 2005; Drew and Barber 2009; Van Herwerden et al. 2009; Eble et al. 2011b). Pelagic (oceanic) species, by comparison, have high vagility as adults (e.g., Carcharadon carcharias, Acanthocybium solandri, and Sphyrna lewini; Pardini et al. 2001; Theisen et al. 2008; Jorgensen et al. 2010; Daly-Engel et al. 2012). These mobile predators exhibit low or no population structure across ocean basins and often have species distributions that bridge substantial biogeographic barriers.
The Great Barracuda, Sphyraena barracuda, is found in all tropical and subtropical waters except the East Pacific (Briggs 1960; Robertson et al. 2004). Despite its geographic ubiquity, notoriety among snorkelers, and commercial importance, relatively little is known about this species’ life history or evolution. The pelagic larval period is unknown, as are most growth parameters and phylogenetic relationships among the 18 members of the genus Sphyraena. Juveniles rely on shallow sea grass and mangrove habitat for shelter during the first year or more of development (Faunce and Serafy 2008). At all life stages, S. barracuda are lie-in-wait predators, feeding primarily on a wide variety of fishes and occasionally on invertebrates, especially cephalopods (de Sylva 1963; Randall 1967). As large-bodied piscivores, adult S. barracuda are highly prone to accumulating ciguatera toxin, though they remain an important human resource in both commercial and artisanal fisheries (Bienfang et al. 2008).
The contemporary range of S. barracuda, which spans approximately 32,000 km and more than 4/5th of the earth’s circumference, indicates that this putative species has bridged nearly every major marine biogeographic barrier in tropical oceans over the course of recent evolutionary history. Aside from continental obstructions, these barriers fall into two categories: distance barriers, like the large span of open ocean that separates Hawaii from the Eastern Tropical Pacific, and temperature barriers, like the cold Benguela Current off southern Africa, which restricts dispersal between the tropical Atlantic and Indian Oceans. These biogeographic barriers have restricted gene flow in the majority of reef fishes (Rocha et al. 2007; Gaither et al. 2010), as well as some of the large predatory species such as tunas, billfishes, and sharks (Graves and McDowell 2003; Duncan et al. 2006; Keeney and Heist 2006).
The near-circumglobal range of S. barracuda is rare for a coastal reef fish (Briggs 1960), though single-species status has not been verified using either morphology or genetic techniques. Color differences between Indo-Pacific (IP) and Caribbean specimens have been noted (J.E.R. pers. obs.), though sample sizes did not allow for thorough morphometric examination. Specifically, two large black spots are often visible on the caudal fin of S. barracuda in the Indo-Pacific, but not on fish in the West Atlantic (Fig. 1). Furthermore, large individuals are often found offshore in the Indo-Pacific, while the same size class tends to remain in shallow waters in the Atlantic (Randall 2007). In terms of dispersal, S. barracuda shows movement patterns characteristic of both coastal reef fishes and pelagic billfishes. Though it is primarily regarded as a reef-associated predator, adult S. barracuda are commonly observed in the open ocean and down to depths of 100 m (de Sylva 1968), and tagged juveniles have been documented moving between Hawaiian Islands (NMFS unpubl. data). Pelagic larval duration is unknown.
To determine the range-wide phylogeography of the Great Barracuda, we collected 246 tissue specimens from thirteen locations throughout the species range (Fig. 2) and resolved 629 base pair of mtDNA cytochrome b. Our goals were twofold: (1) To assess the single-species status of Sphyraena barracuda and the possibility of cryptic evolutionary partitions and (2) to determine whether the phylogeographic architecture of this fish more closely resembles that of sedentary reef fishes or the more mobile pelagic predators.
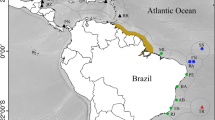
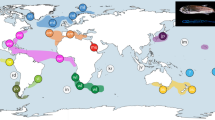
Map showing the global range of Sphyraena barracuda (shaded gray), sampling locations (CM Cameroon, SY Seychelles, MD Maldives, CA Chagos Archipelago, WA West Australia, EA East Australia, MA Mariana Islands, MI Marshall Islands, HI Hawaii, SI Society Islands, BA Bahamas, UV U.S. Virgin Islands, PA Panama), and known marine biogeographic barriers from Rocha et al. 2007 (MAB Mid-Atlantic Barrier, BB Benguela Barrier, OWB Old World Barrier, IPB Indo-Pacific Barrier, EPB East Pacific Barrier, IPA Isthmus of Panama, and AB Amazon Barrier). The question mark refers to the unknown status of barracuda populations in the Tropical East Pacific
Methods
Great barracuda specimens were collected by fishing or purchased from fish markets in thirteen coastal locations in three ocean basins (Table 1; Fig. 2): (1) Pacific (Oahu Hawaii, Guam in the Mariana Islands, Marshall Islands, the Great Barrier Reef in East Australia, and Moorea in the Society Islands); (2) Indian (Seychelles, Maldives, Diego Garcia in the Chagos Archipelago, and Broome Western Australia); and (3) Atlantic (San Blas Panama, Bimini Bahamas, the U.S. Virgin Islands, and Cameroon). Because the actual source of samples collected from fish markets can be uncertain, in all cases where specimens were purchased, special effort was made to ensure that they were locally caught. Fin clips or muscle tissues (approximately 1 cm2) were stored in 20% dimethylsulfoxide (DMSO)-saturated salt buffer (Seutin et al. 1991) or 75% ethanol (EtOH). DNA was extracted from tissue using a salting-out protocol adapted from Sunnucks and Hales (1996). Specimens stored in EtOH were squeezed with a dry paper towel and dried in a speed vacuum for 30 min at 55°C before extraction.
The polymerase chain reaction (PCR) was used to amplify 629 bp of the cytochrome b region of the mitochondrial genome using the primers 01H (5′-GTGACTTGAAAAACCACCGTT-3′) and 02L (5′-AATAGGAAGTATCATTCGGGT-3′; Taberlet et al. 1992; Song et al. 1998). PCRs consisted of 0.1 U Biolase Taq DNA polymerase (Bioline; Randolph, Massachusetts), 1x PCR buffer, 1.0 mM each primer, 200 μm each dNTP, and 2.0 mM MgCl2. PCR amplification on a MyCycler (Bio-Rad; Hercules, California) consisted of an initial denaturation at 95°C for 4 min, followed by 30 cycles of 1 min at 95°C, 30 s at 60°C, and 30 s at 72°C, followed by a final extension at 72°C for 20 min. PCR products were cycle sequenced using Big Dye chemistry on an ABI 3100 automated sequencer (Applied Biosystems, Foster City CA) at the Hawaii Institute of Marine Biology EPSCoR Sequencing Facility, aligned by eye, and edited using Sequencher 4.6 (Gene Codes Corporation, Ann Arbor MI).
Arlequin 3.5 (Excoffier and Lischer 2010) was used to generate nucleotide and haplotype diversities and estimate divergence (ΦST) for the nine sampling locations with N > 5. Patterns of pairwise differentiation were used to group sampling locations, and the magnitude of genetic differentiation between locations was assessed in Arlequin with a hierarchical Analysis of Molecular Variance (AMOVA; 20,000 permutations) using the Tamura-Nei distance method (Tamura and Nei 1993) with 20,000 permutations, which partitions genetic variation within sampling locations, among locations within groups, and among groups.
MrBayes 3.1.2 (Huelsenbeck and Ronquist 2001; Ronquist and Huelsenbeck 2003) was used to construct a Bayesian inference phylogenetic tree for S. barracuda and the nine other Sphyraenid cytochrome b sequences available in GenBank. Analyses of Markov Chain Monte Carlo (MCMC) chains were run for 10,000,000 generations while sampling one tree per 100 generations. Convergence between simultaneous runs was reached when the average standard deviation of split frequencies fell below 0.01 (Ronquist et al. 2005). Overlay plots of the generation versus parameter values for the runs were examined in the program Tracer 1.3 (Rambaut and Drummond 2003) to assess stabilization and mixing of likelihood and parameter values. Following a burn-in phase of 10,000 steps, parameter values were averaged, and posterior clade probabilities were calculated and the likelihood scores for all the topologies averaged. Paup* 4.0b10 (Swofford 2000) was used to construct a neighbor-joining tree with 10,000 bootstrap replicates, which was then visually compared with the Bayesian tree.
Relationships between mitochondrial haplotypes are described with a parsimony network constructed using TCS 1.21 (Fig. 3; Clement et al. 2000). Mismatch analysis was performed in Arlequin to test for demographic expansion with current effective population size (N e) calculated from the equation θ = N e2μt, where θ is estimated from the mean number of segregating sites (θS), μ is the estimated annual fragment mutation rate, and t is the estimated generation time. Population ages in years were estimated from the population age parameter tau (τ), with τ = 2μt, where t is the time since the most recent population bottleneck. We provisionally applied a generation time of 8.8 years for S. barracuda based on age and growth parameters for Sphyraenidae generated by Pauly (1978), which includes S. barracuda. Because divergence estimates for cytochrome b have been obtained for a number of teleost fishes (though not for a Sphyraenid), we employed a range of minimum and maximum mutation rates that encompass those evolutionary rates that have been reported for teleosts: 1% per 106 year within lineages (Bowen et al. 2001; Reece et al. 2010b) to 2.5% per 106 year (Lessios 2008). We tested for population expansion in Arlequin by calculating F S (Fu 1997) and D (Tajima 1989), with population expansion being indicated by significant negative values, and by comparing observed and expected mismatch distributions given a model of recent, sudden population expansion (Rogers and Harpending 1992) in Arlequin with 10,000 replicates.
Results
The TCS parsimony network (Fig. 3) shows 31 haplotypes defined by 35 variable sites, the most common of which (ha01) encompasses 110 individuals representing every sampling site within the IP and nine of the ten fish from the Bahamas that carried IP haplotypes. The tenth specimen carried another IP haplotype (ha25), and together these comprised approximately 1/5th of all the fish collected from the Bahamas. Haplotype diversity was greatest in East Australia in the IP, while nucleotide diversity was highest in the Bahamas (Table 1).
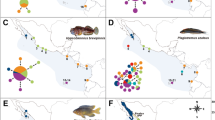
Haplotype network for Sphyraena barracuda constructed by statistical parsimony in TCS 1.21 (Clement et al. 2000) and arranged by biogeographic region. Area of circles is proportional to the abundance of the respective haplotype: the smallest circles represent one individual, while the largest represents 110 individuals. The remaining circles are scaled accordingly. Small black dots indicate missing haplotypes and shades indicate haplotype location (see key). Haplotype numbers as in Fig. 4
The neighbor-joining tree constructed in Paup* (Swofford 2000) and the Bayesian tree from Mr Bayes returned similar topologies; we therefore present only the Bayesian tree in Fig. 4. Both indicate two monophyletic lineages, though Bayesian posterior probabilities are somewhat weak (0.52 and 0.51, respectively): one exclusively composed of fishes from the West Atlantic, and a second containing all Indo-Pacific (IP) fishes, and as an unsupported internal node, all fishes from the East Atlantic (which comprised a single haplotype with N = 23). The IP lineage was separated from the West Atlantic clade by 1.1–1.9% sequence divergence and from the East Atlantic node by 0.6% (uncorrected distances generated in Paup*; Jukes-Cantor and Kimura 2-parameter returned similar results). Since ten individuals from the Caribbean contained an IP haplotype, collecting records and specimens came under additional scrutiny, and DNA from these specimens was re-isolated and re-sequenced, with an identical result. It should be noted that the specimens from Bimini, Bahamas were locally field-caught, so this discrepancy cannot be explained by collection from fish markets.
a Molecular phylogeny of cytochrome b sequences for all Sphyraenid species for which sequences were available, constructed using Bayesian inference in MrBayes. Harmonic mean likelihood = −5,457.09. Numbers before slash denote bootstrap values from 10,000 replicates derived from a neighbor-joining tree with similar topography; numbers after slash denote Pp values. Numbers in parentheses indicate sample size for each haplotype. A# refers to GenBank Accession number. b Detail of molecular phylogeny containing only sequences from Sphyaena barracuda (GenBank accession #JQ088688–JQ088717). Bootstrap and Pp values <50/0.50 are not shown
Pairwise ΦST (Table 2) for the nine sampling locations with N > 5 showed significant structure between the IP and either coast of the Atlantic Ocean. The East Atlantic (Cameroon) was significantly differentiated from every other site, both in the IP and the West Atlantic (ΦST = 0.705–1.000, P < 0.001). The three sampling locations in the West Atlantic were all significantly differentiated from every site in the IP (ΦST = 0.498–0.968, P < 0.001). Within the IP, little pairwise structure was found between locations with the exception of the eastern and western ends of the range. The Seychelles showed low but significant divergences from three other IP locations (West Australia, Mariana Islands, and Hawaii; (ΦST = 0.046–0.234), and Hawaii showed stronger significant divergences from the same locations (Seychelles, West Australia, and Mariana Islands; (ΦST = 0.093–0.207). None of the other IP sites showed significant differentiation. Direct sequencing showed two fixed mutational differences separating the East Atlantic from all other sampling sites, while five fixed mutations differentiate the West Atlantic haplotypes from both the IP and East Atlantic.
Analysis of Molecular Variance (AMOVA) was performed both within and between the four major groupings revealed by pairwise ΦST: (1) Hawaii; (2) the rest of the IP (Mariana Islands, Society Islands, Marshall Islands, East Australia, West Australia, Maldives, Chagos Archipelago, and the Seychelles); (3) West Atlantic (Bahamas, Panama, and the U.S.Virgin Islands); and (4) East Atlantic (Cameroon). Initial AMOVA groupings that separated out the Seychelles from the rest of the IP were found to be non-significant. Overall, structure across the global range of S. barracuda was statistically significant (global ΦST = 0.709, P < 0.001). The nine IP sampling locations showed low but significant structure when grouped together (within group ΦSC = 0.080, P = 0.011), though indications of structure disappeared when Hawaii was removed (within group ΦSC = 0.039, P = 0.165). AMOVA and pairwise tests for differentiation both reveal S. barracuda to be well connected across the IP, the only exceptions being Hawaii in the Central Pacific and (marginally) the Seychelles in the West Indian Ocean (Table 2). There was no evidence of differentiation among the three West Atlantic/Caribbean sampling locations, though together these were found to be significantly different from the IP (W. Atlantic vs. IP, between group ΦST = 0.793, P < 0.001;) and East Atlantic (between group ΦST = 0.759, P < 0.001). Similarly, the East Atlantic was differentiated from both the West Atlantic and IP (ΦST = 0.800, P < 0.001). Overall, AMOVA indicated that 74.6% of the variance was partitioned among groups (ΦCT = 0.746, P > 0.001) and 25.4% within groups (ΦSC = 0.254, P > 0.001).
The same haplotype was observed in all 23 fish collected from the East Atlantic, precluding tests for demographic expansion and indicating a likely genetic bottleneck. Tests for demographic expansion were run on the remaining three populations delineated in the AMOVA (Table 3) with no significant deviation between simulated and observed mismatch distributions, suggesting recent population expansion at all three locations. F S and D were, however, significantly negative in the IP only. Mismatch analyses indicate the age of the most recent population expansion to be on the order of 87,000–219,000 years in the IP and roughly half that in Hawaii (32,000–130,000 years) and the W. Atlantic (55,000–137,000 years).
Discussion
Phylogeographic analyses across the nearly circumglobal range of Sphyraena barracuda revealed four distinct populations that align with major marine biogeographic provinces (Briggs 1974b): (1) the Hawaiian Islands; (2) Indo-Pacific (IP, Society Islands to the Seychelles); (3) the West Atlantic/Caribbean (Bahamas, Panama, and the U.S. Virgin Islands); and (4) the East Atlantic (Cameroon). Parsimony and mismatch analyses indicate that the IP hosts the oldest population, invoking an IP origin for contemporary S. barracuda populations with subsequent dispersal eastward into the Central Pacific and westward into the Atlantic.
The longitudinal distribution of S. barracuda traverses more than 4/5th of the Earth’s surface and includes such diverse habitats as estuaries, archipelagos, continental shelves, and pelagic blue-water environments. Within this range, there are several natural biogeographic barriers that can inhibit connectivity (Fig. 2). Over evolutionary time, these barriers have yielded a complex phylogeographic landscape (Rocha et al. 2007). Concordant measures of genetic distance in S. barracuda show both high connectivity across thousands of kilometers as well as strong population breaks between some biographic provinces. Phylogeographic patterns indicate that, while well-defined biogeographic barriers clearly present an obstacle to dispersal in S. barracuda, some barriers are more permeable than others.
The most prominent obstacles to gene flow in S. barracuda appear to be the East Pacific Barrier and the Mid-Atlantic Barrier, both large stretches of open water that obstruct gene flow in many marine species (Briggs 1974b). The East Pacific Barrier (EPB), a 4000 km stretch of open ocean separating the islands of the central Pacific (including Hawaii and French Polynesia) from the Tropical East Pacific, represents a substantial barrier to many reef fishes, including S. barracuda (Briggs 1974a; Briggs 1974b). Conversely, the Mid-Atlantic Barrier (MAB), which entails the 3,000 km stretch of open ocean separating East and West Atlantic (Briggs 1974b; Rocha et al. 2007), is apparently more of a filter than an absolute barrier to S. barracuda dispersal, because individuals carrying IP haplotypes clearly had to cross the Atlantic recently to account for the haplotype distribution in the Caribbean. However, given the paucity of S. barracuda on the Mid-Atlantic Ridge Islands (Edwards 1980) and the reciprocal monophyly of East and West Atlantic populations (ΦST = 0.759, P < 0.001), this barrier is clearly substantial. Bowen et al. (2006a) note that 8 of 15 trans-Atlantic comparisons of reef fishes reveal cryptic evolutionary (and possibly taxonomic) separations. The influence of distance on gene flow in S. barracuda can also be seen in comparisons between Hawaii and other locations within the IP, with both pairwise comparisons and AMOVA clearly revealing isolation of Hawaiian S. barracuda (between group ΦST = 0.115, P = 0.011). Though some haplotypes are shared across the IP (including Hawaii), the TCS network and Bayesian tree show two types (ha07 and ha14) that are private to Hawaii, further indicating limited gene flow between these regions.
With the exception of Hawaii, IP sampling locations are well connected across the 16,500 km of islands and open ocean from the Society Islands in French Polynesia to the Seychelles in the Indian Ocean, a pattern matched by few other reef fishes (Horne et al. 2008; Reece et al. 2010a; Eble et al. 2011a). In particular, this region encompasses the Indo-Polynesian Province, a biogeographic region that spans half the planet, from French Polynesia to the central Indian Ocean (Briggs 1974b; Briggs and Bowen 2012). As noted by Schultz et al. (2008), most of this vast province can be traversed without crossing more than 800 km of open water. Hence, the open ocean distance barriers that are so effective elsewhere in the range of S. barracuda are absent here. The isolation of Hawaii is well established in both biogeography and phylogeography (Randall 1998; Randall 2007; DiBattista et al. 2011).
The Indo-Pacific Barrier (IPB, Fig. 2), the shallow continental shelf between the Pacific and Indian Oceans, is a substantial obstacle to gene flow in both tropical invertebrates and fishes (Barber et al. 2006; Gaither et al. 2010, 2011). At glacial maxima, the IPB is exposed and forms a nearly impenetrable land bridge between Pacific and Indian Oceans (Briggs 1974b). Gaither et al. (2010) noted that repeated cycles of shoaling and submersion of the IPB has resulted in strong population-level or evolutionary separations in the majority of reef fishes that straddle this region. Though regarded as an obligate reef species, S. barracuda shows no genetic structure across the IPB, a trait it shares with numerous pelagic teleosts and elasmobranchs (Chow and Ushiama 1995; Reeb et al. 2000; Keeney and Heist 2006; Castro et al. 2007; Theisen et al. 2008).
The Benguela Barrier (BB), the cold-water upwelling of the Benguela Current in southern Africa, appears to be intermittently permeable to S. barracuda dispersal. This barrier closed off a warm water connection between the Indian and Atlantic Oceans about 2.5 million years ago (Shannon 1985). However, the warm water corridor resumes at the end of each glacial epoch, allowing tropical faunal colonization from the Indian to Atlantic Ocean (Peeters et al. 2004; Rocha et al. 2005; Bowen et al. 2006b), and rarely in the reverse direction (Bourjea et al. 2007). The fact that East and West Atlantic populations originated independently from the IP, with close connections to specifically Indian Ocean haplotypes and no intervening gene flow across the Atlantic, supports the Agulhas current via the Benguela corridor as the most likely mechanism for dispersal of Great Barracuda into the Atlantic Ocean. Although colonization between the East Pacific and Atlantic prior to shoaling of the Isthmus of Panama has been demonstrated in some reef species (Bowen et al. 2001; Muss et al. 2001; Lessios 2008; Schultz et al. 2008), it is an unlikely explanation for S. barracuda given the shallow IP-Atlantic divergences (<1 MY) and lack of an East Pacific population.
Though precise colonization time cannot be pinpointed, rough estimates based on percent sequence divergence and coalescence analysis between Atlantic and IP clades are contemporaneous with the period of Pleistocene cessations (2,588,000–11,700 years before present) of the cold-water Benguela upwelling. We hypothesize that an early Benguelan cessation allowed warm water from the Indian Ocean to carry S. barracuda adults, juveniles, or larvae first to the West Atlantic coast of South America via the Southern Equatorial Current, with subsequent northward dispersal to the Caribbean (though we were unable to acquire tissue samples from Brazil, a genetic survey of Brazilian S. barracuda would be informative on this point). A subsequent cessation event allowed dispersal along the West African coast to the Gulf of Guinea, though elapsed time has not been enough for the magnitude of molecular evolutionary separation observed in the West Atlantic. The contemporary presence of IP haplotypes in the Caribbean is likely a remnant of the last Pleistocene glaciation.
Given the responses of S. barracuda to these biogeographic barriers, does the genetic architecture of this species resemble that of reef fishes or pelagic fishes? While the genetic cohesion across the Indo-Polynesian Province is a common pattern in reef fishes, most aspects of barracuda phylogeography more closely resemble that of the tunas, billfishes, and pelagic sharks. Particularly telling is the relationship between Atlantic and IP barracuda populations. When reef fishes colonize the Atlantic, the observed pattern indicates a single colonization event from the Indian Ocean to the West Atlantic, followed by range expansion and diversification (Rocha et al. 2005; Bowen et al. 2006a). In contrast, the pelagic wanderers often show evidence of multiple independent colonizations from Indian to Atlantic Ocean, as indicated by two or more divergent mtDNA lineages, with one confined to the Atlantic and others widespread in the IP but also detected in the Atlantic. This is usually interpreted as evidence for long-term isolation of tropical Atlantic and IP faunas, followed by an influx of IP individuals into Atlantic populations, a conclusion that applies to many highly vagile marine taxa such as swordfish (Xiphias gladius; Alvarado Bremer et al. 1996), blue marlin (Makaira spp.; Graves and McDowell 1995), sailfish (Istiophorus spp.; Graves and McDowell 1995; Graves and McDowell 2003), bluefin tuna (Thunnus thynnus; Alvarado Bremer et al. 2005), bigeye tuna (Thunnus obesus; Durand et al. 2005), the white/striped marlin complex (Tetrapturus spp.; Graves and McDowell 2003), several sea turtles (Bowen and Karl 2007), scalloped hammerhead sharks (Sphyrna lewini; Daly-Engel et al. 2012), and the short-finned pilot whale (Globicephala macrorhynchus; Oremus et al. 2009).
How does intermittent dispersal between the Indian and Atlantic oceans influence decisions about evolutionary partitions and the taxonomy of S. barracuda? Most of the West Atlantic/Caribbean population is differentiated from the rest of S. barracuda by cytochrome b sequence divergences of d = 1.1–1.9%, well within the range of sister species in other fish genera (Johns and Avise 1998) but far below the mtDNA differentiation that separates S. barracuda from available congeners (Fig. 4). Based on morphology, no definitive case exists for taxonomic distinction, but de Sylva (1963, p. 28) wrote “Although measurements for large fish are too scarce to draw any critical conclusions, the preanal and snout lengths seem to present real differences which are far removed from the regression slope.” We therefore recognize the possibility that IP and West Atlantic populations may merit taxonomic revision, perhaps at the subspecific level. However, the Caribbean individuals with IP haplotypes invoke the possibility that interbreeding may dissolve any taxonomic/evolutionary differences that now exist. Further investigations with nuclear loci and morphology are necessary to determine whether or not West Atlantic and IP barracuda are interbreeding and whether these two populations are distinct evolutionary entities. Testing with nuclear markers would also be beneficial for illuminating any potential sex bias in the dispersal of adult S. barracuda and to help clarify contemporary dispersal events.
In the interim, the pelagic fishes as well as the sea turtles, shark, and marine mammal cited above are instructive. In three cases, blue marlin (Makaira spp.), sailfish (Istiophorus spp.), and white/striped marlin (Tetrapturus spp.), the Atlantic and IP populations were described as separate species (Graves and McDowell 1995; Graves and McDowell 2003), but they share identical or closely related mtDNA haplotypes (but see Nakamura 1985; Collette et al. 2006). Graves and McDowell (1995) conclude, on the basis of these genetic results and broad overlap in morphological and meristic characters, that the species-level distinctions between ocean basins should be eliminated. Likewise, the loggerhead sea turtle (Caretta caretta) previously was assigned to IP and Atlantic species or subspecies, but these designations were discarded after morphological and mtDNA comparisons (Hughes 1974; Bowen et al. 1994). We conclude that despite deep evolutionary breaks, the Great Barracuda has shown sufficient dispersal ability to maintain genetic and evolutionary cohesiveness across a global range. Like the pelagic billfish and tunas and unlike reef fishes, this species has demonstrated a dispersal potential that may inhibit or retard regional isolation over evolutionary timescales.
References
Alvarado Bremer JR, Mejuto J, Greig TW, Ely B (1996) Global population structure of the swordfish (Xiphias gladius L.) as revealed by analysis of the mitochondrial DNA control region. J Exp Mar Biol Ecol 197:295–310
Alvarado Bremer JR, Vinas J, Mejuto J, Ely B, Pla C (2005) Comparative phylogeography of Atlantic bluefin tuna and swordfish: the combined effects of vicariance, secondary contact, introgression, and population expansion on the regional phylogenies of two highly migratory pelagic fishes. Mol Phylogenet Evol 36:169–187
Barber PH, Erdmann MV, Palumbi SR (2006) Comparative phylogeography of three codistributed stomatopods: origins and timing of regional diversification in the coral triangle. Evolution 60:1825–1839
Bienfang P, Oben B, DeFelice P, Moeller K, Huncik P, Oben R, Toonen R, Daly-Engel T, Bowen B (2008) Ciguatera: the detection of neurotoxins in carnivorous reef fish from the coast of Cameroon, West Africa. Afr J Mar Sci 30:533–540
Bourjea J, Lapegue S, Gagnevin L, Broderick D, Mortimer JA, Ciccione S, Roos D, Taquet C, Grizel H (2007) Phylogeography of the green turtle, Chelonia mydas, in the Southwest Indian Ocean. Mol Ecol 16:175–186
Bowen BW, Karl SA (2007) Population genetics and phylogeography of sea turtles. Mol Ecol 16:4886–4907
Bowen BW, Kamezaki N, Limpus CJ, Hughes GR, Meylan AB, Avise JC (1994) Global phylogeography of the loggerhead turtle (Caretta caretta) as indicated by mitochondrial DNA haplotypes. Evolution 48:1820–1828
Bowen BW, Bass AL, Rocha LA, Grant WS, Robertson DR (2001) Phylogeography of the trumpetfishes (Aulostomus): ring species complex on a global scale. Evolution 55:1029–1039
Bowen BW, Bass AL, Muss AJ, Carlin J, Robertson DR (2006a) Phylogeography of two Atlantic squirrelfishes (family Holocentridae): exploring pelagic larval duration and population connectivity. Mar Biol 149:899–913
Bowen BW, Muss A, Rocha LA, Grant WS (2006b) Shallow mtDNA coalescence in Atlantic pygmy angelfishes (Genus Centropyge) indicates a recent invasion from the Indian Ocean. J Hered 97:1–12
Briggs JC (1960) Fishes of worldwide (Circumtropical) distribution. Copeia 3:171–180
Briggs JC (1974a) Marine zoogeography. McGraw-Hill, New York
Briggs JC (1974b) Operation of zoogeographic barriers. Syst Zool 23:248–256
Briggs JC, Bowen BW (2012) A realignment of marine biogeographic provinces with particular reference to fish distributions. J Biogeogr 39:12–30
Castro ALF, Stewart BS, Wilson SG, Hueter RE, Meekan MG, Motta PJ, Bowen BW, Karl SA (2007) Population genetic structure of Earth’s largest fish, the whale shark (Rhincodon typus). Mol Ecol 16:5183–5192
Chow S, Ushiama H (1995) Global population structure of albacore (Thunnus alalunga) inferred by RFLP analysis of the mitochondrial ATPase gene. Mar Biol 123:39–45
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1660
Collette BB, McDowell JR, Graves JE (2006) Phylogeny of recent billfishes (Xiphioidei). Bull Mar Sci 79:455–468
Daly-Engel TS, Duncan KM, Holland KN, Coffey JP, Nance HA, Toonen RJ, Bowen BW (2012) Global molecular phylogeography reveals male-mediated dispersal in the endangered scalloped hammerhead shark (Sphyrna lewini). PLoS ONE 7(1):e29986
de Sylva DP (1963) Systematics and life history of the great barracuda Sphyraena barracuda Walbaum. University of Miami Press, Coral Gables
de Sylva DP (1968) Systematics and life history of the great barracuda Sphyraena barracuda (Walbaum). Toxicon 5:227
DiBattista JD, Wilcox C, Craig MT, Rocha CR, Bowen BW (2011) Phylogeography of the Pacific blueline surgeonfish Acanthurus nigroris reveals a cryptic species in the Hawaiian archipelago. J Mar Biol, Article ID 839134
Drew JA, Barber PH (2009) Sequential cladogenesis of Pomacentrus moluccensis (Bleeker, 1853) supports the peripheral origin of marine biodiversity in the Indo-Australian archipelago. Mol Phylogenet Evol 53:355–359
Duncan KM, Martin AP, Bowen BW, De Couet HG (2006) Global phylogeography of the scalloped hammerhead (Sphyrna lewini). Mol Ecol 15:2239–2251
Durand JD, Collet A, Chow S, Guinand B, Borsa P (2005) Nuclear and mitochondrial DNA markers indicate unidirectional gene flow of Indo-Pacific to Atlantic bigeye tuna (Thunnus obesus) populations, and their admixture off southern Africa. Mar Biol 147:313–322
Eble JA, Rocha LA, Craig MT, Bowen BW (2011a) Not all larvae stay close to home: insights into marine population connectivity with a focus on the brown surgeonfish (Acanthurus nigrofuscus). J Mar Biol, Article ID 518516
Eble JA, Sorenson LS, Papastamatiou YP, Basch L, Toonen RJ, Bowen BW (2011b) Escaping paradise: larval export from Hawaii in an Indo-Pacific reef fish, the yellow tang. Mar Ecol Prog Ser 426:245–258
Edwards A (1980) Fish and Fisheries of Saint Helena Island. Center for Tropical Coastal Management Studies, University of Newcastle upon Tyne, U.K.
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Faunce CH, Serafy JE (2008) Selective use of mangrove shorelines by snappers, grunts, and great barracuda. Mar Ecol Prog Ser 356:153–162
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Gaither MR, Toonen RJ, Robertson DR, Planes S, Bowen BW (2010) Genetic evaluation of marine biogeographical barriers: perspectives from two widespread Indo-Pacific snappers (Lutjanus kasmira and Lutjanus fulvus). J Biogeogr 37:133–147
Gaither MR, Bowen BW, Bordenave T-R, Rocha CR, Newman SJ, Gomez JA, Van Herwerden L, Craig MT (2011) Phylogeography of the reef fish Cephalopholus argus (Epinephelidae) indicates Pleistocene isolation across the Indo-Pacific barrier with contemporary overlap in the Coral Triangle. BMC Evol Biol 11:89
Graves JE, McDowell JR (1995) Inter-ocean genetic divergence of istiophorid billfishes. Mar Biol 122:193–203
Graves JE, McDowell JR (2003) Stock structure of the world’s istiophorid billfishes: a genetic perspective. Mar Freshw Res 54:287–298
Horne JB, Van Herwerden L, Choat JH, Robertson DR (2008) High population connectivity across the Indo-Pacific: congruent lack of phylogeographic structure in three reef fish congeners. Mol Phylogenet Evol 49:629–638
Huelsenbeck JP, Ronquist F (2001) MrBayes: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Hughes GR (1974) The sea turtles of southeast Africa. I. Status, morphology, and distributions. Oceanographic Research Institute Investigational Report 35:1–144
Johns GC, Avise JC (1998) A comparative summary of genetic distances in the vertebrates from the mitochondrial cytochrome b gene. Mol Biol Evol 15:1481–1490
Jorgensen SJ, Reeb CA, Chapple TK, Anderson S, Perle C, Van Sommeran SR, Fritz-Cope C, Brown AC, Klimley AP, Block BA (2010) Philopatry and migration of Pacific white sharks. Proc R Soc B 277:679–688
Keeney DB, Heist EJ (2006) Worldwide phylogeography of the blacktip shark (Carcharhinus limbatus) inferred from mitochondrial DNA reveals isolation of western Atlantic populations coupled with recent Pacific dispersal. Mol Ecol 15:3669–3679
Lessios HA (2008) The great American schism: divergence of marine organisms after the rise of the Central American isthmus. Annu Rev Ecol Evol Syst 39:63–91
Muss A, Robertson DR, Stepien CA, Wirtz P, Bowen BW (2001) Phylogeography of the genus Ophioblennius: the role of ocean currents and geography in reef fish evolution. Evolution 55:561–572
Nakamura I (1985) FAO species catalogue, vol 5. Billfishes of the world: an annotated and illustrated catalogue of marlins, sailfishes, spearfishes and swordfishes known to date FAO Fisheries Synopsis, Rome
Oremus M, Gales R, Dalebout ML, Funahashi N, Endo T, Kage T, Steel D, Baker SC (2009) Worldwide mitochondrial DNA diversity and phylogeography of pilot whales (Globicephala spp.). Biol J Linn Soc 98:729–744
Pardini AT, Jones CS, Noble LR, Kreiser B, Malcolm H, Bruce BD, Stevens JD, Cliff G, Scholl MC, Francis M, Duffy CAJ, Martin AP (2001) Sex-biased dispersal of great white sharks. Nature 412:139–140
Pauly D (1978) A preliminary compilation of fish length growth parameters. Ber Inst Meereskd Christian-Albrechts-Univ Kiel 55:1–200
Peeters FJC, Acheson R, Brummer GJA, de Ruijter WPM, Schneider RR, Ganssen GM, Ufkes E, Kroon D (2004) Vigorous exchange between the Indian and Atlantic Oceans at the end of the past five glacial periods. Nature 430:661–665
Rambaut A, Drummond AJ (2003) Tracer version 1.3. http://evolve.zoo.ox.ac.uk/
Randall JE (1967) Food habits of reef fishes of the West Indies. Stud Trop Oceanogr 5:665–847
Randall JE (1998) Zoogeography of shore fishes of the Indo-Pacific region. Zool Stud 37:227–268
Randall JE (2007) Reef and shore fishes of the Hawaiian Islands. University of Hawaii Press, Honolulu
Reeb CA, Arcangeli L, Block BA (2000) Structure and migration corridors in Pacific populations of the Swordfish Xiphius gladius, as inferred through analyses of mitochondrial DNA. Mar Biol 136:1123–1131
Reece JS, Bowen BW, Joshi K, Goz V, Larson AF (2010a) Phylogeography of two moray eels indicates high dispersal throughout the Indo-Pacific. J Hered 101:391–402
Reece JS, Bowen BW, Smith DG, Larson AF (2010b) Molecular phylogenetics of moray eels (Murenidae) demonstrates multiple origins of a shell-crushing jaw (Gymnomuraena, Echidna) and multiple colonizations of the Atlantic Ocean. Mol Phylogenet Evol 57:829–835
Robertson DR, Grove JS, Jack S, McCosker JE (2004) Tropical transpacific shore fishes. Pac Sci 58:507–565
Rocha LA, Robertson DR, Rocha CR, Van Tassel JL, Craig MT, Bowen BW (2005) Recent invasion of the tropical Atlantic by an Indo-Pacific coral reef fish. Mol Ecol 14:3921–3928
Rocha LA, Craig MT, Bowen BW (2007) Phylogeography and the conservation genetics of coral reef fishes. Coral Reefs 26:501–512
Rogers AR, Harpending H (1992) Population-growth makes waves in the distribution of pairwise genetic-differences. Mol Biol Evol 9:552–569
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Ronquist F, Huelsenbeck JP, van der Mark P (2005) MrBayes 3.1 manual. pp 1–69
Schultz JK, Feldheim KA, Gruber SH, Ashley MV, McGovern TM, Bowen BW (2008) Global phylogeography and seascape genetics of the lemon sharks (genus Negaprion). Mol Ecol 17:5336–5348
Seutin G, White BN, Boag PT (1991) Preservation of avian blood and tissue samples for DNA analyses. Can J Zool 69:82–90
Shannon LV (1985) The Benguela ecosystem. I. Evolution of the Benguela physical features and processes. Oceanogr Mar Biol Annu Rev 23:105–182
Song CB, Near TJ, Page JM (1998) Phylogenetic relations among Percid fishes as inferred from mitochondrial cytochrome b DNA sequence data. Mol Phylogenet Evol 10:343–353
Strathmann RR (1993) Hypotheses on the origins of marine larvae. Ann Rev Ecol Syst 24:89–117
Sunnucks P, Hales DF (1996) Numerous transposed sequences of mitochondrial cytochrome oxidase I–II in aphids of the genus Sitobion (Hemiptera: Aphididae). Mol Biol Evol 13:510–524
Swofford DL (2000) PAUP*. Phylogeny analysis using parsimony (*and Other Methods), Version 4. Sinauer Associates, Sunderland
Taberlet P, Meyer A, Bouvet J (1992) Unusually large mitochondrial variation in populations of the blue tit, Parus caeruleus. Mol Ecol 1:27–36
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tamura K, Nei M (1993) Estimation of the number of nucelotide substitutions in the control region of mitiochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Taylor MS, Hellberg ME (2005) Marine radiations at small geographic scales: speciation in neotropical reef gobies (Elacatinus). Evolution 59:374–385
Theisen TC, Bowen BW, Lanier W, Baldwin JD (2008) High connectivity on a global scale in the pelagic wahoo, Acanthocybium solandri (tuna family Scombridae). Mol Ecol 17:4233–4247
Van Herwerden L, Choat JH, Newman SJ, Leray M, Hillersoy G (2009) Complex patterns of population structure and recruitment of Plectropomus leopardus (Pisces: Epinephelidae) in the Indo-West Pacific: implications for fisheries management. Mar Biol 156:1595–1607
Weersing KA, Toonen RJ (2009) Population genetics, larval dispersal, and demographic connectivity in marine systems. Mar Ecol Prog Ser 393:1–12
Acknowledgments
The authors gratefully acknowledge Jeff Eble, Greg Concepcion, Marc Crepeau, Paul Marek, and Rob Toonen for laboratory support and help with genetic analyses. For help with sample collection, we thank Kim Holland, Dean Grubbs, Robert Kikuta, Paul Bienfang, Benedicta Oben, David Itano, Michelle Gaither, Mindy Mizobe, Lori O’Hara, Zoltan Szabo, Luiz Rocha, Matt Craig, Ross Newton and Tracey Rushford of Reel Teaser Charters in Western Australia, Anne Gasc of EnCoRe inc, David Rowat and the Marine Conservation Society of the Seychelles, Mike Trevor, Lenore Litherland, Monica Mwale, Ross Robertson, Ben Victor, Ross Langston, Barry Alty, Junior Sasis, Samuel Gruber, Charles Sheppard, Bill Signs, Craig Henn, Jimmy Bijoux, Tom Vely, Jon Puritz, Tristan Guttridge, Johann Mourier, John Coffey, Carl Meyer, Yannis Papastamatiou, Nick Whitney, R. Newton, Jeff Johnson, R. Charles Anderson, J. Beets, Hannelore Waska, Emily Marcus, Sean Williams, Bill Signs, the Guam Fisherman’s Cooperative Association, British Indian Ocean Territory Administration, and the Toonen, Bowen, and Holland Labs at the Hawaii Institute of Marine Biology. Funding came from the Graduate Student Organization at the University of Hawaii, National Science Foundation (OCE-0453167, OCE-0623678, and OCE-0929031 to B.W.B. and R.J. Toonen, and EPSCoR EPS-0554657 to University of Hawaii). We thank Sarah Daley, Rajesh Shrestha, and Mindy Mizobe and the HIMB EPSCoR core facility for assistance with DNA sequencing. The lead author is supported by grant No. 2 K12 GM000708 to the PERT Program at the University of Arizona from the National Institute of General Medical Sciences division of NIH. This is contribution #1454 from the Hawaii Institute of Marine Biology and contribution #8211 from the School of Ocean and Earth Science and Technology at the University of Hawaii.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. I. Taylor.
Rights and permissions
About this article
Cite this article
Daly-Engel, T.S., Randall, J.E. & Bowen, B.W. Is the Great Barracuda (Sphyraena barracuda) a reef fish or a pelagic fish? The phylogeographic perspective. Mar Biol 159, 975–985 (2012). https://doi.org/10.1007/s00227-012-1878-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00227-012-1878-9