Abstract
Cnidarian–dinoflagellate symbioses are not well understood at the molecular level. Observed specificity between partners during initiation, establishment, and maintenance of the relationship strongly implies a role for chemical signaling. This report presents biochemical and immunocytochemical evidence for potential signaling molecules, as large molecular weight glycoproteins, secreted by Symbiodinium dinoflagellates both in culture and in symbiosis. Polyclonal antibodies directed against recovered exudate from S. microadriaticum, the natural endosymbiont of Cassiopea xamachana, the upside–down jellyfish, were highly specific in recognizing exudates from Symbiodinium species that can successfully induce developmental metamorphosis in the host but did not recognize exudates from Symbiodinium species that do not. Immunoblot analyses showed S. microadriaticum exudate to be protease sensitive. Release of antigenic material by symbiotic S. microadriaticum was demonstrated through light and electron microscopy using immunogold-labeled anti-S. microadriaticum (anti-Sm-XuLg) antibodies as probes. These secreted, symbiont-derived glycoconjugates may be candidates for interspecific molecular signals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In many symbioses, the partnership must be established de novo with each new generation of the host. These intimate relationships are often highly specific, containing predictable partner associations. From the host’s perspective, adaptations to accommodate symbionts may leave them vulnerable to opportunistic infection by parasites and/or pathogens (Sachs and Simms 2006; Sachs and Wilcox 2006; Weis and Allemand 2009). From the symbiont’s perspective, opportunistic invaders may offer increased competition for resources. Therefore, achieving and maintaining a highly specific association may be of critical importance to both partners in the relationship (Foster and Wenseleers 2006). The squid–Vibrio and legume–rhizobia associations have been extensively studied, and the molecular signaling events responsible for the onset of symbiosis have been well described (Nyholm and McFall-Ngai 2004; Cooper 2007). In these associations, recognition between free-living partners requires a multitude of signaling events from both partners before, during, and after contact. A successful partnership only occurs after several checkpoint stages have been completed along the way. Molecular research on cnidarian–dinoflagellate mutualisms is revealing that these associations may also undergo a similar “winnowing” or polyphasic selection process during infection and establishment that leads to formation of a stable symbiotic association (Nyholm and McFall-Ngai 2004; Weis et al. 2008).
The cnidarian–dinoflagellate association is unique and of great interest for two important reasons: (1) it is a eukaryote–eukaryote symbiosis, rather than the more commonly observed eukaryote–prokaryote association (see perspective in Schwarz 2008) and (2) symbiotic cnidarians include scleractinian or calcifying corals that form the trophic and structural foundation of all coral reef ecosystems - critical habitats under severe threat from climate change and other anthropogenic impacts (Stat et al. 2006; Hoegh-Guldberg et al. 2007; Weis 2008). Unfortunately, in part, because corals can be difficult to work with, we are only just beginning to understand the molecular dialogue between cnidarian hosts and their algal symbionts (Weis et al. 2008).
Much of our current understanding of cnidarian symbioses relies on work done on non-calcifying cnidarians, including Aiptasia and Cassiopea, which host microalgae of the dinoflagellate genus Symbiodinium, and the freshwater green hydra-chlorophyte system (reviewed in Rodriguez-Lanetty et al. 2006; Weis et al. 2008). This study describes more foundational work with the C. xamachana–S. microadriaticum system that was completed several years ago but never before published (Markell 1995). It demonstrates that Symbiodinium in culture, and at least S. microadriaticum in hospite, release high molecular weight proteinaceous compounds with glycoconjugate characteristics. As these compounds are presented to potential hosts in the environment and directly to hosts while in symbiosis, they make ideal candidates for symbiont-derived molecular signals. This work also challenges previous assumptions that symbiotic microalgae secrete virtually nothing but glycerol, simple sugars, and some amino acids in hospite (Muscatine 1967; Muscatine and Porter 1977), making a case for additional research on carbon and nitrogen exchange in algal–invertebrate symbioses.
Materials and methods
Maintenance of algal cultures
Cultures were grown in vitro in ASP-8 under axenic conditions, as described previously (Markell et al. 1992). The algae used in this study were Symbiodinium microadriaticum (Freudenthal) emend, S. kawagutii, and S. pilosum (Trench and Blank 1987), S. burmudense nom. nud., and Symbiodinium sp. (#175 - Trench lab collection; Table 1).
Gel filtration of secreted macromolecules from culture media
Algal exudates were recovered from the culture media as described previously (Markell and Trench 1993). Recovered exudates were fractionated by gel filtration as follows: 10–30 mg dry weight of each sample was reconstituted in 1 mL TE buffer (10 mM Tris–HCl pH 8.0, 1 mM EDTA, 200 mM NaCl) with 10% sucrose and layered onto a 2.5 × 22 cm column of Sephadex G-100. The column was eluted at 10 mL h−1, and 5-mL fractions were collected. The column was calibrated using Mouse IgG 160 kDa, bovine serum albumin (BSA) 67 kDa; sperm whale myoglobin 17.8 kDa, and bacitracin 1.5 kDa. The void volume was determined using blue dextran (MW > 2 × 106). Absorbance (280 nm) of the eluent was continuously monitored using an ISCO UA 5 UV detector. Absorbance of individual fractions was measured using a Hewlett Packard 8452A diode array spectrophotometer. Protein content was estimated using BSA as the standard (Lowry et al. 1951).
HPLC analysis of exudates using DEAE ion exchange
Recovered exudates from each algal culture were analyzed using a Showdex 825 DEAE ion exchange HPLC column eluted at 1 mL min−1 under linear gradient of 0.001–0.25 M NaCl developed over 1 h in 20 mM imidazole buffer (pH 6.5). Absorbance (265 nm) was detected using a Waters Model 481 LC Spectrophotometer. Data analyses were performed using BASELINE 810 software (Millipore), and elution profiles were normalized to the highest peak.
Production and characterization of anti-Sm-XuLg
Polyclonal antibodies were produced in rabbit against the S. microadriaticum excluded fraction (>100 kDa) by Berkeley Antibody Co. Initial injection dose (500 μg antigen in RIBI adjuvant) was on June 15, 1990, and followed by a booster (250 μg) on July 6, 1990. Serum was taken on August 6, 27, and September 18, 1990.
To characterize anti-Sm-XuLg, exudates from algal cultures were separated using 7.5% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) and transferred to nitrocellulose, then air-dried, and fixed in acetic acid:2-propanol:water (10:25:65, v/v/v). Blots used to visualize total protein were stained with colloidal gold (BioRad). Antibody blots were rinsed in double deionized water (DDI), immersed in Tris-buffered saline (TBS; 50 mM Tris–HCl pH 7.5, 200 mM NaCl) for 5 min, and transferred to blocking solution (TBS with 0.1% Tween-20 (TBST), 1% BSA, and 2% powdered non-fat dry milk) for 1 h. Blots were incubated in fresh blocking solution with anti-Sm-XuLg serum (1:250 dilution) or preimmune serum (1:50 dilution) and incubated overnight at 4°C. The following day, blots were washed for 30 min in TBST, incubated 1 h with horseradish peroxidase-conjugated goat anti-rabbit antibodies (BioRad, 1:2500 dilution) in blocking solution, and washed for 2 h in TBST then DDI.
Enzymatic digestion of exudates
Exudates containing 10 μg of protein were incubated in digestion buffer (10 mM Tris pH 7.8, 1 mM CaCl2, 0.5% SDS) in 20 μL final volume for 2 h at 52°C with and without proteinase-K (100 μg mL−1, Tritirachium album, Fisher Scientific; Sambrook et al. 1989). Exudates were also digested for 0, 1, 2, 10 min, 1, and 3 h at 25°C with trypsin (10,200U mg−1, final concentration 10 μg mL−1) or 3 h without trypsin. SDS–PAGE buffer (2×) was added at the end of the time point, and the reaction was stopped by boiling. BSA controls were co-incubated to confirm enzyme function. Samples were separated using SDS–PAGE, blotted, and probed with anti-Sm-XuLg or colloidal gold (as above).
Fractionation of S. microadriaticum cells
Culture medium was centrifuged at 7,500 g. The cell pellet was washed in TE buffer, spun at 15,000 g, and supernatant was discarded. Cells were resuspended in 200 μL 1% SDS, pelleted at 15,000 g, and the supernatant was saved. The pellet was resuspended in 10 mL TE and passed three times through a French pressure cell. The slurry was centrifuged; the supernatant was serial precipitated with ammonium sulfate (25, 50, and 100% saturation at 4°C), and the pellet containing cell wall and other insoluble cellular components was dialyzed against DDI and dried under partial vacuum in a SpeedVac. The pellet was reconstituted in 1 mL TE, mixed with 9 mL 80% sucrose, and spun at 160,000 g for 2 h. The top and middle third of the sucrose fractions were dialyzed and dried as above. The bottom third (cell wall fraction) was washed 10× with DDI, then chloroform and methanol to remove residual membrane. Purified cell walls were boiled in 1% SDS for 30 min to extract surface components (Markell and Trench 1993). Fractions were separated using SDS–PAGE, blotted, and probed with anti-Sm-XuLg.
Fixation, embedding, probing
Cassiopea xamachana scyphistomae (oral disk diameter 0.5–1.0 mm) were infected with S. microadriaticum and fed sparingly for 30–90 days. Infected and un-infected controls were fixed for 2 h in high osmolality cacodylate buffered formaldehyde-glutaraldehyde solution (Karnovsky 1965) at 23°C, transferred to 0.1 M cacodylate buffer (pH 7.5) for 1 h, and rinsed in DDI for 1 h. Samples were dehydrated at 4°C in an ethanol series with 2% (w/v) uranyl acetate (15, 30, 50, 70, 85, 95, and 100% v/v ethanol, 20 min per step) and washed twice in 100% ethanol. Dehydrated samples were transferred to 50:50 LR-White and ethanol, capped, and rotated overnight at 4°C. The following day, samples were uncapped and rotated at 23°C for 4 h, then transferred to LR-White (100%) and cured in bullet molds at 56°C for 2 days. Trimmed blocks were sectioned on a LKB Ultratome using a diamond knife: thick sections (ca. 10 μm) for light microscope (LM) immunohistochemistry on ProbeOn Plus slides (Fisher Scientific) with plastic wells for buffer solutions and thin sections (ca. 10 nm) on nickel grids for electron microscopy (EM).
Light microscope (LM) sections were prewetted with blocking solution for 5 min at 37°C, then incubated for 30 min in blocking solution with normal goat serum (1:20 dilution). Samples were incubated for 2 h with anti-Sm-XuLg primary antibodies (1:1,000 dilution) or preimmune serum (1:100 dilution, control for non-specific binding), rinsed 3× in TBST with 10 min soaks in between. Samples were incubated for 1 h with 10 nm gold-conjugated goat anti-rabbit antibodies (1:100 dilution) and washed as before with addition of DDI. Gold labeling was enhanced using a silver enhancing kit (Ted Pella, Inc.) Sections were stained with basic fuchsin and imaged with an Olympus Vanox microscope using Kodak Ektachrome 160T color slide film.
Electron microscopy (EM) sections were probed using the same methods, with omission of silver enhancement. Labeled sections were stained by floating on aqueous 1% uranyl acetate for 10 min, rinsed with DDI, floated on Reynolds lead citrate for 10 min, rinsed with DDI, and dried before placing in a parafilm lined dish for osmication with a 2-h exposure to vapors from a drop of 4% OsO4. Sections were photographed using a Seimens Elmisop I.
Results
Gel filtration and analysis of secreted macromolecules
In order to determine the size distribution of secreted macromolecules, algal culture media was analyzed by gel filtration on Sephadex G-100 under non-denaturing conditions. An excluded (void) peak of >100 kDa was consistently observed (Supplemental Fig. 1a, representative elution profile of Symbiodinium microadriaticum with calibration curve). A second peak containing small molecules (<1.5 kDa) or molecules that interacted strongly with the column was also observed. This peak likely contained photosynthetically derived molecules, such as glucose and glycerol, as well as amino acids like alanine, which are commonly secreted from symbiotic algae (Trench 1971; Muscatine and Porter 1977). Therefore, the high molecular weight fraction was selected for further study. HPLC analysis of DEAE ion exchange elution profiles from the five Symbiodinium cultures revealed a net negative charge was common in all exudates, but the charge profiles varied between exudates (Supplemental Fig. 1b-g).
Characterization of anti-Sm-XuLg
Polyclonal antibodies against the >100-kDa secreted fraction from S. microadriaticum were characterized by immunoblotting exudates from five Symbiodinium cultures (Fig. 1). Anti-Sm-XuLg antibodies had high affinity to S. microadriaticum exudates for compounds >30 kDa (denatured; Fig. 1, lane C’) and some labeling of exudates from Symbiodinium spp. (#175), which is also able to infect Cassiopea xamachana (Fig. 1, lane E’). The antibodies did not cross-react with exudates from any other Symbiodinium culture.
Characterization of polyclonal antibodies directed against S. microadriaticum exudate fraction (>100 kDa) and hybridized to SDS–PAGE of exudate from six algal cultures. Total protein visualized with colloidal gold (lanes A–F, ~10 μg protein per lane as determined by (Lowry et al. 1951)) or probed with anti-Sm-XuLg antibodies (lanes A’–F’, ~30 μg protein per lane). (A, A’) Homogenized aposymbiotic C. xamachana; (B, B’) S. burmudense; (C, C’) S. microadriaticum; (D, D’) S. kawagutii; (E, E’) Symbiodinium spp. #175 [from T. maxima]; (F, F’) S. pilosum
To determine the chemical nature of the antigen(s) recognized by anti-Sm-XuLg, exudate from S. microadriaticum was digested with proteolytic enzymes. Undigested exudate, analyzed under denaturing conditions, contained several components recognized by the antibodies (Fig. 2a, no trypsin). Immediately upon addition of trypsin, bands ranging from ~40 to 100 kDa (Fig. 2a No Trypsin bracket) began to diminish in staining intensity or disappeared altogether, and a band at ~30 kDa began to appear (Fig. 2a arrow). No other bands increased in intensity as a result of digestion, and bands >110 kDa did not change in staining intensity. S. microadriaticum exudate digested with and without proteinase-K demonstrated that anti-Sm-XuLg antibody–antigen binding required an intact polypeptide, suggesting that the epitope is either proteinaceous in nature or may contain post-translational modifications that require a partially intact protein sequence (Fig. 2b).
Digestion and immunoblot analysis of S. microadriaticum exudates. a Trypsin digestion was monitored over 180 min and b proteinase-K was incubated for complete digest. The 40–100 kDa range is bracketed, and an arrow highlights the 30-kDa band. BSA was digested to confirm enzyme activity. The scale is appropriate for trypsin digest data only
Algal fractionation and immunochemical localization of Sm-XuLg antigens
In order to conclude that antigenic products in the exudates were truly algal derived, cultured S. microadriaticum were washed and fractionated to release internal antigens. Ammonium sulfate precipitation separated intracellular proteins by ionic charge, and the sucrose gradient separated the cell wall components by density; fractions were screened with anti-Sm-XuLg. Immunoblot analysis revealed high molecular weight antigens in the water-soluble, intracellular fraction precipitated by 25% ammonium sulfate (Fig. 3, lane D bracket), indicating that the molecules had a relatively low ionic charge. Antigens were also present in the top the layers of the sucrose gradient suggesting a low buoyant density relative to cell wall material (Fig. 3, lane G bracket). No Sm-XuLg antigens were recovered in the SDS-wash of whole cells or the SDS-soluble fraction of the cell walls.
Fractionation of cultured S. microadriaticum by ammonium sulfate precipitation or sucrose gradient. Fractions were separated by SDS–PAGE gels and probed with anti-Sm-XuLg antibodies. A Total exudate; B extracellular SDS-wash; C total soluble cell extract; D–F precipitated by ammonium sulfate (D = 25%, E = 50%, F = 100%); (G–I) recovered sucrose gradient (G = top 3rd, H = middle 3rd, I = bottom 3rd); J SDS-extract of purified cell walls. No size markers are shown. Brackets highlight high-molecular weight antigens
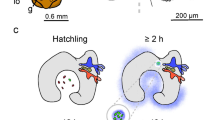
Production of Sm-XuLg antigens in hospite was demonstrated by immunohistochemical localization of Sm-XuLg in symbiotic C. xamachana. Under LM, silver enhanced-gold spheres were observed in association with algal cells and also dispersed nearby in the mesoglea and ectoderm (Fig. 4a, b). Non-specific labeling with pre-immune serum was not observed in symbiotic C. xamachana (Fig. 4c), and no labeling was seen in non-infected scyphistomae incubated with anti-Sm-XuLg (Fig. 4d). Similar results were seen using EM; Sm-XuLg antigens were present inside an algal cell (Fig. 4f), immediately surrounding algal cells and dispersed in the mesoglea (Fig. 4e), with no labeling observed in controls (Fig. 4g–h).
Immunohistochemical localization of anti-Sm-XuLg in hospite. Light micrographs (a–d) and electron micrographs (e–h) of anti-Sm-XuLg labeling. C. xamachana infected with S. microadriaticum (a, b, e, and f) showed strong labeling of anti-Sm-XuLg (visible as gold spheres, silver enhanced for LM, highlighted by arrows) near the host–algal interface and in the surrounding mesoglea (a, b, e) as well as within the algal cell (f). Infected C. xamachana labeled with pre-immune serum (c, g) and uninfected hosts (d, h) did not label indicating no cross-reactivity to Sm-XuLg and no antigens present, respectively. (Ch, chloroplast; CW, cell wall; Ec, ectoderm; En, endoderm; Mg, mesoglea; PM, plasma membrane; Sm, S. microadriaticum)
Discussion
This and prior studies (Markell et al. 1992; Markell and Trench 1993) demonstrate that there are significant biochemical and biophysical differences between exudates of various Symbiodinium species in culture. These exudates are hypothesized to function as species-specific signaling molecules during recognition and uptake by a potential host, and, when secreted in hospite, may continue to function in communication between symbiotic partners.
Preliminary characterization of antigens from S. microadriaticum (large exudate fraction, Sm-XuLg) through proteolytic digestion revealed they either have very few cleavage sites for trypsin or the cleavage sites were post-translationally modified (i.e., glycosylation), blocking trypsin activity. Intermediate buoyant density of these compounds, relative to proteins and carbohydrates, also suggests they were glycoconjugates (Beeley 1985). Cellular fractionation confirmed the antigens were of algal origin and not an artifact of the culture medium as the antibodies identified high molecular weight compounds present in the intracellular water-soluble fraction of S. microadriaticum cells. The lack of detectable antigens in the SDS-wash of whole cells or purified cell wall fractions suggests the algae actively secreted the antigens into the culture medium and were not released as a product of cell death (Markell and Trench 1993). It remains unknown whether the antigen pool in culture precisely reflects that in symbiosis, or if the pool changes during the algal cell cycle, which has been seen for dinoflagellate cell surface antigens (Aguilera and Gonzalez-Gil 2001). However, at least some, if not all of the components in the >100 kDa-fraction were released while in symbiosis as anti-Sm-XuLg labeling was detected in symbiotic C. xamachana hosts immediately surrounding the symbionts and diffused into the mesoglea. It is unclear whether antigens released into the cnidarian host remain unchanged, especially since the molecules retained antigenicity even after experimental digestion with trypsin.
HPLC elution profiles of recovered exudates from the five Symbiodinium were unique, but all showed a net negative charge as determined by DEAE ion exchange. A previous study by Markell and Trench (1993) revealed significant differences among exudates with respect to amino acid and sugar composition. They also found a correlation between high levels of uronic acid in exudates from cultured algae and the capacity to infect C. xamachana scyphistomae. However, graphical normalization of HPLC elution profiles in this study showed no apparent correlation between either elution profile patterns or net quantity of negative charge (as determined by elution time) and the ability of algae to infect C. xamachana (S. microadriaticum and Symbiodinium #175, data not shown). Symbiodinium species used in this study likely produced exudates containing isoforms of one or more components that may have a role in host–symbiont specificity, but those potential differences were not detected by HPLC. However, antibodies to Sm-XuLg were demonstrated to be specific to recovered exudates from S. microadriaticum and Symbiodinium sp. #175 (>30 kDa denatured). Anti-Sm-XuLg did not show binding to control, uninfected C. xamachana, to exudates from S. bermudense, which infects at a much slower rate than S. microadriaticum or Symbiodinium spp. #175 (DAM personal observation), or cultured algae that do not infect C. xamachana (S. kawagutii and S. pilosum; Fig. 1). These observations support the hypothesis that secreted macromolecules may also play a potential role in specificity following initial infection and/or could be involved in triggering developmental metamorphosis in the host. The release of unique chemical isoforms by the symbiont, both for recognition by a potential host and induction of morphological changes in the host, is a well-studied strategy used in establishing mutualisms, such as legume–rhizobia associations (Oldroyd et al. 2005; Fauvart and Michiels 2008).
Studies examining the molecular dialogue of cnidarian–dinoflagellate partnerships are beginning to implicate mechanisms of specificity similar to those observed in other, more well-studied mutualistic associations. In addition to this study, there is evidence that cell surface lectin/glycan interactions play a role during the onset of infection (Reisser et al. 1982; Lin et al. 2000; Wood-Charlson et al. 2006; Kvennefors et al. 2008), and previous studies by Markell et al. (1992) and Markell and Trench (1993) showed that Symbiodinium exudates contained high molecular weight compounds that differ between Symbiodinium species. The research documented in this manuscript provides new evidence for soluble, species-specific glycoconjugates secreted by Symbiodinium in culture and, more importantly, also secreted into tissues of the host in an intact association that theoretically could be used for communicating with their cnidarian host.
References
Aguilera A, Gonzalez-Gil S (2001) Lectin analysis of surface saccharides during the cell cycle in four dinoflagellate species. J Exp Mar Biol Ecol 256:149–166
Beeley JG (1985) Glycoprotein and proteoglycan techniques. Elsevier, Amsterdam
Cooper JE (2007) Early interactions between legumes and rhizobia: disclosing complexity in a molecular dialogue. J Appl Microbiol 103:1355–1365
Fauvart M, Michiels J (2008) Rhizobial secreted proteins as determinants of host specificity in the rhizobium-legume symbiosis. FEMS Microbiol Lett 285:1–9
Foster KR, Wenseleers T (2006) A general model for the evolution of mutualisms. J Evol Biol 19:1283–1293
Hoegh-Guldberg O, Mumby PJ, Hooten AJ, Steneck RS, Greenfield P, Gomez E, Harvell CD, Sale PF, Edwards AJ, Caldeira K, Knowlton N, Eakin CM, Iglesias-Prieto R, Muthiga N, Bradbury RH, Dubi A, Hatziolos ME (2007) Coral reefs under rapid climate change and ocean acidification. Science 318:1737–1742
Karnovsky MJ (1965) A formaldehyde-glutaraldehyde fixative of high osmolarity for use in electron microscopy. J Cell Biol 27:137–138A
Kvennefors EC, Leggat W, Hoegh-Guldberg O, Degnan BM, Barnes AC (2008) An ancient and variable mannose-binding lectin from the coral Acropora millepora binds both pathogens and symbionts. Dev Comp Immunol 32:1582–1592
Lin KL, Wang JT, Fang LS (2000) Participation of glycoproteins on zooxanthellal cell walls in the establishment of a symbiotic relationship with the sea anemone, Aiptasia pulchella. Zool Stud 39:172–178
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275
Markell DA (1995) Macromolecules exuded by dinoflagellates in symbiosis: a biochemical and cellular analysis of specificity. Dissertation, University of California, Santa Barbara
Markell DA, Trench RK (1993) Macromolecules exuded by symbiotic dinoflagellates in culture: amino acid and sugar composition. J Phycol 29:64–68
Markell DA, Trench RK, Iglesias-Prieto R (1992) Macromolecules associated with the cell-walls of symbiotic dinoflagellates. Symbiosis 12:19–31
Muscatine L (1967) Glycerol excretion by symbiotic algae from corals and Tridacna and its control by the host. Science 156:516–519
Muscatine L, Porter JW (1977) Reef corals: mutualistic symbioses adapted to nutrient-poor environments. Bioscience 27:454–460
Nyholm SV, McFall-Ngai M (2004) The winnowing: establishing the squid-Vibrio symbiosis. Nature Rev Microbiol 2:632–642
Oldroyd GE, Harrison MJ, Udvardi M (2005) Peace talks and trade deals. Keys to long-term harmony in legume-microbe symbioses. Plant Physiol 137:1205–1210
Reisser W, Radunz A, Wiessner W (1982) Participation of algal surface structures in the cell recognition process during infection of aposymbiotic Paramecium bursaria with symbiotic chlorellae. Cytobios 33:39–50
Rodriguez-Lanetty M, Wood-Charlson EM, Hollingsworth L, Krupp D, Weis V (2006) Temporal and spatial infection dynamics indicate recognition events in the early hours of a dinoflagellate/coral symbiosis. Mar Biol 149:713–719
Sachs JL, Simms EL (2006) Pathways to mutualism breakdown. Trends Ecol Evol 21:585–592
Sachs JL, Wilcox TP (2006) A shift to parasitism in the jellyfish symbiont Symbiodinium microadriaticum. Proc Biol Sci 273:425–429
Sambrook J, Fritsche EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schwarz JA (2008) Understanding the intracellular niche in cnidarian-Symbiodinium symbioses: parasites lead the way. Life and Environment 58:141–151
Stat M, Carter D, Hoegh-Guldberg O (2006) The evolutionary history of Symbiodinium and scleractinian hosts—Symbiosis, diversity, and the effect of climate change. Perspect Plant Ecol Evol Syst 8:23–43
Trench RK (1971) The physiology and biochemistry of zooxanthellae symbiotic with marine coelenterates. II. Liberation of fixed 14C by zooxanthellae in vitro. Proc R Soc London B 177:237–250
Trench RK, Blank RJ (1987) Symbiodinium microadriaticum Freudenthal, S. goreauii sp. nov., S. kawagutii sp. nov., and S. pilosum sp. nov.: Gymnodinioid dinoflagellate symbionts of marine invertebrates. J Phycol 23:469–481
Weis VM (2008) Cellular mechanisms of Cnidarian bleaching: stress causes the collapse of symbiosis. J Exp Biol 211:3059–3066
Weis VM, Allemand D (2009) What determines coral health? Science 324:1153–1155
Weis VM, Davy SK, Hoegh-Guldberg O, Rodriguez-Lanetty M, Pringle JR (2008) Cell biology in model systems as the key to understanding corals. Trends Ecol Evol 23:369–376
Wood-Charlson EM, Hollingsworth LL, Krupp DA, Weis VM (2006) Lectin/glycan interactions play a role in recognition in a coral/dinoflagellate symbiosis. Cell Microbiol 8:1985–1993
Acknowledgments
Douglas A. Markell would like to thank Robert K. Trench, Roberto Iglesias-Prieto, Jim Cooper and Dennis Clegg, and extend his thanks to Patricia E. Thomé for maintenance of the cultures of symbiotic dinoflagellates. EWC would like to thank Todd LaJeunesse for clarification on the status of the algal cultures used in this study, and Robert K. Trench, William K. Fitt, and an anonymous reviewer for their insights and extensive improvements on this manuscript. Support for this study was provided by the Office of Naval Research (ONR NOOO14-92-J-I LSI) to Robert K. Trench. Contribution No. 206 of the Center for Microbial Oceanography: Research and Education (C-MORE) at the University of Hawai’i at Manoa.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. L. Goulet.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Markell, D.A., Wood-Charlson, E.M. Immunocytochemical evidence that symbiotic algae secrete potential recognition signal molecules in hospite . Mar Biol 157, 1105–1111 (2010). https://doi.org/10.1007/s00227-010-1392-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00227-010-1392-x