After she ate the entire cake, she grew so large that her head struck against the ceiling in the hall.
Alice’s Adventures in Wonderland, Lewis Carroll
Abstract
Good genes, good food, good friends. That is what parents hope will sustain and nurture the harmonious growth of their children. The impact of the genetic background and nutrition on postnatal growth has been in the spot light for long, but the good friends have come to the scene only recently. Among the good friends perhaps the most crucial ones are those that we are carrying within ourselves. They comprise the trillions of microbes that collectively constitute each individual’s intestinal microbiota. Indeed, recent epidemiological and field studies in humans, supported by extensive experimental data on animal models, demonstrate a clear role of the intestinal microbiota on their host’s juvenile growth, especially under suboptimal nutrient conditions. Genuinely integrative approaches applicable to invertebrate and vertebrate systems combine tools from genetics, developmental biology, microbiology, nutrition, and physiology to reveal how gut microbiota affects growth both positively and negatively, in healthy and pathological conditions. It appears that certain natural or engineered gut microbiota communities can positively impact insulin/IGF-1 and steroid hormone signaling, thus contributing to the host juvenile development and maturation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The developmental program of each organism is largely genetically encoded [1]. Still, gene–environment interactions can shape the spectrum of phenotypes, including growth, as we can observe for example by comparing genetically identical siblings raised under different conditions [2]. Growth is defined as an increase in size [1]. Here, it is necessary to mention that growth may have different connotations in adults and in juvenile individuals. In adults, growth is usually an increase in weight (ponderal growth). On the other hand, juvenile growth is defined as increase both in size (longitudinal growth) and weight. Juvenile growth is accompanied or followed by maturation, giving rise to a fertile adult, fit to produce the next generation. Nutrition certainly plays a central role in development, with under- and malnutrition clearly affecting metabolism, as well as the rate of development, the timing of sexual maturation, and the final size reached at adulthood. Nutrient availability impacts cell size and cell division rate, as well as hormone production [3, 4]. Multicellular animals have evolved a specialized organ system for food digestion and absorption of nutrients, the digestive system. In vertebrates, it comprises the gastrointestinal tube and the annexed organs, like the pancreas and liver. Although located within the host, the lumen of the digestive tube is topologically equivalent and directly connected with the outside environment and, in many of its parts, this tube is teeming with life. It is the home of microorganisms (primarily bacteria, but also fungi, viruses, and protists) that collectively comprise an animal’s intestinal microbiota. This microbiota is normally acquired soon after hatching or birth [5]. Generally, intestinal microbes are non-pathogenic to the host, as long as they are contained within their appropriate intestinal compartment. It is now widely recognized that these microbes play a fundamental role in the physiology of the host, with whom they establish complex commensal and mutualistic symbiotic relationships.
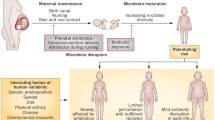
Nutrition exerts a pivotal influence on host physiology. Besides the direct influence on host metabolism, nutrition has also an impact on the resident microbial community in the intestine [6]. And vice versa microbiota is an inseparable part of the host nutritional environment; besides taking its share from the available nutrition pool members of the intestinal microbial community can expand the host’s metabolic potential or, in some cases, they can even serve as nutrition for the host. In line with this notion, several groups have recently reported that besides nutrition, intestinal bacteria are key players influencing host juvenile growth [7,8,9,10,11,12]. Integrative approaches will therefore benefit from seeing the eukaryotic individuals as holobionts [13], taking the intestinal microbiota and its genetic and metabolic potential as an intrinsic factor determining the dynamics of host juvenile growth and maturation (Fig. 1).
Gut Microbiota and Animal Physiology
Animal–microbiota interactions have far reaching effects on the host’s physiology, pathology, ecology, and evolution [14, 15]. In the last two decades, pioneer investigations by the groups of Jeffrey Gordon, Martin Blaser, and Rob Knight on the human microbiota and of Margaret McFall-Ngai on the squid and its symbiotic bioluminescent bacteria, along with work from several laboratories in multiple systems, have expanded our appreciation of the extent to which several developmental and life traits of multicellular organisms are influenced by the microorganisms they are associated with. A comprehensive review of microbiota influence on animal physiology is well beyond the scope of this article, the interested reader is referred to [14, 16, 17]. It is sufficient to remind here a couple of the discoveries that made it to newspaper headlines, such as the demonstrated impact of antibiotic treatment on newborn metabolism and health downstream of microbiota alterations [18]; the link between gut microbiota and autism [19]; the study of skin microbiota said to have caused the ban of nineteen soaps in the USA [20]; or, for a more exotic touch, the dependence of light organ development and camouflage capacity of the squid on the colonization by the luminous bacteria Vibrio fischeri [21,22,23].
Particular attention has been paid to the microbiota associated with the gut and how host physiology can be deranged by modifications of this intestinal microbiota [22, 23]. In most animals, the microorganismal community populating the intestinal tract is by far the largest microbiota, both in quantity and in diversity of species. Next-generation sequencing technology and powerful new bioinformatics tools have allowed the description of intestinal microbiota communities from many species, with high accuracy and independently of the ability to culture the microbes in the laboratory. In parallel, epidemiology has started to look at human microbiota variations in healthy individuals and patients affected by a variety of conditions and to draw suggestive correlations [24, 25]. Manipulations of the human gut microbiota, as those obtained by fecal transplantations or following antibiotic treatments, provide functional data on potential cause–consequence relationships between changes in microbiota and alterations in host physiology [25]. Crucially, work on both vertebrate and invertebrate model organisms has given us rigorous experimental evidence of the causative role of alterations in the microbial communities of the digestive tract on derangements in host’s functions, metabolism, and immunity among others [26, 27]. Furthermore, the use of model organisms has granted the first glimpses on the molecular and cellular details of the dialogue between microbes and hosts in the intestine, the mechanisms that underlie local (intestinal) and systemic host responses and the role played by the host in relation to the ability of the microbe to occupy and expand in certain niches. It has become evident that only multidisciplinary approaches will be able to answer the many questions pertaining the truly integrative physiology of host–microbiota interactions [28]. Similarly, integrating knowledge and advances from diverse model systems will allow major breakthroughs in the field.
Microbiota and the Experimental Use of Axenic and Gnotobiotic Animals
In 1885, Louis Pasteur conceived the idea of rearing animals devoid of microbes on pure sterile diet from birth. In his opinion, the mutualistic relationship and dependence of the host on the bacteria was so profound, that life without microbes should become impossible [29]. It took 10 years before the first animal, a guinea pig, was raised germ free by G. Nuttal and H. Thierfelder [30]. Although it was kept sterile only for a short period of time (8 days), they proved that life without microbes is indeed possible. This pioneering experiment paved the way to germ-free rearing of other animal species, such as flies, chickens, or rodents [31, 32]. With technological advances, the isolators for rearing germ-free animals were improved and the first long-term mouse colonies continuously bred under the germ-free condition were established after the Second World War [33, 34].
The key experimental advantage of germ-free systems is obvious—the possibility to recolonize the intestines of these animals with single microorganismal species or defined mixed populations, thus creating the so-called gnotobiotic animals: mono-associated, oligo-associated, or conventionalized, depending on the complexity of the microorganismal consortia used [35,36,37,38]. One can then ask what the contribution of the microbiota is to a given phenotypic trait of interest in the host and can test whether the trait is transferable by the microbiota. Importantly, the gnotobiotic system allows looking at the microbiota as a whole community or at its different components individually (Fig. 2).
Eukaryotic host can be germ free, gnotobiotic, or conventional. Germ-free animals harbor no detectable living microorganisms and can be seen as an extreme condition of gnotobiotic animal, in which all colonizing microorganisms are known. Conventional animal is colonized by undefined consortium of microorganisms. The host organism and the colonizing microbiota can be in a state of eubiosis, where the host profits from the presence of microorganisms and the juvenile growth velocity is increased. On the other hand, in state of dysbiosis the microbiota drains on the host resources which results in growth depression
Many insects host obligate endosymbionts that are essential for their survival in nature and whose elimination is also lethal for the animal partner in most laboratory conditions [39]. On the other hand, several insects and other invertebrates, including the genetic model organisms Drosophila melanogaster (the fruitfly) and the worm Caenorhabditis elegans, can survive and reproduce in germ-free conditions under a range of environmental settings [40, 41]. The ease to generate, maintain, and manipulate axenic invertebrate model organisms, their short life cycle and powerful genetic amenability, together with the relatively low complexity of their natural microbiota, has made the mechanistic study of invertebrate host–microbiota interactions appealing and tractable, in fruitflies in particular. The economical, medical, and ecological interest in additional invertebrate species, such as the honey bee, malaria mosquito vectors, and aquatic crustaceans and molluscs, has further sustained recent efforts to understand the interesting biology of host–microbe interactions in such systems [42, 43].
Despite the undisputable advantages of invertebrate models, the simplicity of the microbiota, difference in physiological processes, and the lack of adaptive immune system is also a limiting factor for testing hypothesis relevant to humans, where the microbiota comprise both aerobes and anaerobes. In this regard, vertebrate models such as zebrafish (Danio rerio), mouse (Mus musculus), or pig (Sus scrofa) are more relevant. With recent advances in gnotobiology, they can be rendered germ free with each model organism having its specific advantages concerning the generation time, microbiota community structure and scalability, and price of gnotobiotic breeding [28]. Particularly, the mouse is the mammalian model of choice not only in classical biomedical research but also in gnotobiology. Although the fish and the pig can be raised germ free, mouse is currently the most widespread mammalian model organism that has been bred in the isolators for successive generations on the large scale [15]. Moreover, the existence of mouse isogenic lines and the powerful genetic tools enabling the generation of transgenic and knockout mice [44] combined with the long-term gnotobiotic breeding enables to study all aspects of the mammalian development and physiology within the context of host–microbiota interactions.
In the first section of the review, we will summarize advances in invertebrate model systems that asked how growth, and therefore rate of size increase and timing of metamorphosis and sexual maturation is affected by the gut microbiota. We will then move on to the vertebrate models and discuss the lessons learned about the role of the intestinal microbiota in the context of the postnatal growth under germ-free and gnotobiotic conditions. We will see how, by simplifying and stripping down the systems to their essential elements, while preserving biological relevance, these studies could identify specific signaling pathways, metabolites, and genes involved in host–microbiota cross-talk and host’s response, with special attention for the effects on juvenile growth.
Who Does What and How: Understanding the Link Between Gut Microbiota and Growth in Invertebrate Model Systems
Gut Microbiota and Growth in Drosophila melanogaster
About 40 years after the observation that sterilized, germ-free Drosophila melanogaster embryos can grow to adulthood in axenic conditions [40], a few laboratories started taking advantage of the novel sequencing technologies to cast a fresh look at host–bacteria interactions in the fruitfly. Renewed interest in the influence of the intestinal microbiota on human health, the success of the ground-breaking work in Drosophila immunity, and host–microbe interactions and the growing interest of fly geneticists for physiology, metabolism, and translational research, all combined for a rapidly expanding and productive research community studying host–microbiota interactions.
To start, the species and strains of bacteria associated with the Drosophila body, and with the gut in particular, were identified in samples collected from laboratories or in the wild. The main findings are (i) an overall small number of different bacterial species isolated from fly guts, ranging in the order of tens (therefore much less than the hundreds of different species isolated in a given human gut microbiota); (ii) the exact gut microbiota makeup varies according to the life stage at which specimens were gathered and primarily to the food substrate on which flies lived, pointing at a close link between gut microbiota and diet; (iii) a handful of recurrent and dominating species can be named including species from the Acetobacteraceae and Lactobacillaceae families, such as Acetobacter pomorum, Acetobacter tropicalis, Lactobacillus brevis, and Lactobacillus plantarum [45,46,47]. Similar studies have followed to describe Drosophila-associated yeast/fungi and viral communities [48, 49]. For the sake of this review, we focus on gut-associated bacteria, whose roles in host growth have been more extensively studied. For an informative yet concise overview of the parallels between Drosophila and mammalian intestinal cell biology and physiology, relevant to host–microbiota interactions, we refer the reader to a recent review from the Watnick’s laboratory [50].
The next step has been the identification of life traits demonstrably affected by the microbiota (for a recent overview, see for example [51]). A link with growth emerged: a clear difference becomes apparent when embryos and larvae are left to grow on suboptimal food substrates, namely protein-poor food. Here, conventional larvae grow more slowly than on protein-rich food because of a decrease in TOR signaling and the consequent disarray in the insulin and steroid hormonal signaling ([52]; reviewed by [53]). Germ-free larvae are much more delayed: at any given time, their size is smaller and they start metamorphosis later, though eventually they give rise to adults indistinguishable from their conventional siblings. The developmental delay of germ-free larvae is apparent only on suboptimal nutritional conditions; lack of microbiota has a small impact on growth on rich food [54]. Re-association of germ-free embryos with gut-derived bacteria restores development to the control level [10, 12]. The delayed growth and maturation of axenic larvae on poor food, incidentally, appear not to have a fitness cost for the enclosing adults, at least under the many conditions tested in Tefit and Leulier [55]. A crucial finding in those studies [10, 12] was the identification of single bacterial species from the microbiota pool that can recapitulate the beneficial effects of the whole gut microbiota when mono-associated with axenic embryos. On diets containing free sugars, it is A. pomorum (one of the widespread fruitfly gut bacteria in the wild), while L. plantarum, another common Drosophila commensal which is also found in the human intestine, fits the bill when no simple sugars are present in the food. Simplified one host-one bacterium systems were thus established [10, 12]. Moreover, not only was it possible to identify species that perform better than others in recapitulating the growth advantage imparted by the whole bacterial microbiota pool, but it became apparent that even different strains of the same bacterial species can vary greatly in their growth-promoting efficacy.
In these simplified systems, both bacterial and fly genetics can be applied to unravel the mechanisms underlying the observed microbiota-dependent growth acceleration. Combined with microbiological and metabolic investigations, gene gain- and loss-of-function analyses of genetically engineered microbes and animals have now identified key features of interactions allowing beneficial effects on host’s growth. The findings are transposable to, and in some cases already validated in, vertebrate systems.
There are several lessons learned from the gnotobiotic Drosophila-bacteria cross-talk. First, the intestinal microbiota impacts developmental rate by modulating the production and/or release of insulin/insulin-like growth factors from the insulin-producing cells of the host. This occurs downstream of a relay signal from the fat body (an organ serving the combined function of the liver and adipose tissues in flies), which is itself dependent on cell-autonomous TOR signaling and the ability of the fat body to sense the availability of branched amino-acids via the membrane amino-acid transporter encoded by the slimfast gene. Modulation of systemic insulin signaling influences the mobilization and utilization of sugar and fat storage to be made available for anabolic purposes [10, 12]. Maturation (for flies, the transition into the pupal stage and the eclosion of sexually mature adults) is also delayed in germ-free larvae. Microbiota re-association rescues the delay by stimulating an earlier production of the steroid hormone ecdysone from endocrine glands, downstream of cell-autonomous TOR signaling in the endocrine glands themselves [12]. As discussed in the following section on vertebrates, the growth hormone/IGF-1 axis has been also implicated in microbiota-dependent growth of malnourished mice [9]. Moreover, the same L. plantarum strains impart similar degrees of growth advantages in flies and mice [9], making a strong case for the relevance of the results from the Drosophila system to mammals.
Second, increased amino-acid uptake lies at the root of the higher and earlier hormonal production observed in larvae grown in the presence of microbiota. Transcriptomics and subsequent genetic functional analysis have identified various intestinal proteases whose genes are upregulated by microbiota association and which are important for the faster growth of L. plantarum mono-associated larvae. Increased proteolytic activity is detected in mono-associated larval guts and metabolic analysis shows an increased level of single and di-amino-acids in mono-associated larvae compared to germ-free ones [56]. Further, this transcriptional upregulation of several protease genes triggered by L. plantarum association depends on the host’s immune system: it relies in part on bacterial peptidoglycan recognition and signaling cascade of the Imd/Relish pathway [56].
Third, identified bacterial products required to stimulate growth in the host under nutrient deprivation include a short chain fatty acid (acetate) and bacterial cell walls bearing d-alanine-modified teichoic acids [10, 57]. The two discoveries were accomplished in two distinct mono-association experimental setups, with two different bacteria. Conceptually very similar, the setups differ for the food substrate where larvae grow and the bacterial species that can recapitulate the beneficial growth effect of the whole gut microbiota community. As mentioned above, both diets are protein poor, but diverge in the carbohydrate composition and the identity of the bacterial species capable of accelerating growth on their own. Acetate must be produced by bacteria or be added to the medium for A. pomorum to promote growth in the presence of simple sugars [10]. It might also have a more direct action: it could boost insulin signaling by repressing transcription of IMPL2, a negative regulator of insulin-like factors, as suggested by studies of fly infection with an intestinal pathogen [58]. Cell wall-bearing D-alanine-modified teichoic acids from L. plantarum impact protease upregulation and larval growth on poor medium where starch is essentially the only carbohydrate source [57]. In the larva, D-alanylated teichoic acids are likely sensed by an alternative, yet unidentified microorganism detection system that is independent of the Imd/Relish pathway [57]. How have these bacterial molecules been identified? Thanks to unbiased screens performed with libraries of thousands of mutagenised bacteria: each library clone was cultured and mono-associated with axenic larvae, while larval growth was used as a phenotypic read-out. Such a strategy is permitted by the short life cycle of the fly and the ease to handle germ-free and mono-associated animals. Together with comparative genomic analysis of more versus less beneficial strains/bacteria and the gene-trait matching interrogations of the bacteria–host symbiotic phenotypes [59], bacterial genetics holds the key to unveil what makes a microbe a desirable commensal. We can expect that these approaches will uncover additional bacterial products that can induce beneficial microbiota-mediated effects on their own, with potential applications beyond basic research.
Fourth, upon bacterial association, the physiology of the intestine changes, including proteolytic activity, pH, and electric properties of the gut epithelium. The mere over-expression of the proteases, whose transcription is boosted by the gut microbiota, can accelerate the growth of germ-free larvae on protein-poor food in the absence of bacteria [56]. This argues that, with the correct cues, axenic larvae can deploy an appropriate digestive machinery that can extract enough energy to sustain such a growth rate. The microorganisms used for mono-association experiments do not serve as simple food supplements [60]; rather, they modulate intestinal physiology. In line with these observations, it has been described that the luminal pH profile of larval intestines is dependent on microbiota presence, possibly via changes in the expression or activity of ion channels and pumps [61]. Moreover, Shanbhag, Tripathi, and colleagues recently showed by microperfusion-associated electrophysiology that the intestinal epithelia of germ-free larvae lose the asymmetrical membrane conductance between apical and basal membranes typical of epithelia of conventional larvae: again, the trafficking or regulation of ion channels and pumps must be deranged in germ-free larvae [62]. While overall apico-basal cell membrane asymmetry of enterocytes is not altered in germ-free larvae [10, 60] and transcellular permeability is not detectably altered [62], it nonetheless appears that enterocytes might be modified in some aspects of their cell biology and digestive properties. Whether changes in pH and in the electric properties of the gut epithelium impact growth has yet to be determined.
Fifth, DNA replication and proliferation of intestinal enterocytes and stem cells is increased by microbiota, both in larvae and adults. This was observed in L. plantarum mono-associated flies as well as L. rhamnosus mono-associated mice [63]. Further, the same group showed that an additional response to Lactobacilli mono-association, again observed both in flies and mice, is the activation of the transcription factor Nrf2-dependent cyto-protective pathway, relying on glutathione S-transferases and Cytochrome P450 family genes [64]. It seems that microbial contact-induced epithelial ROS generation is a conserved and universal phenomenon by which bacteria can modulate a variety of signaling and homeostatic processes in the host [65]. It will be interesting to see whether these bacteria-induced changes, occurring downstream of the NADPH oxidase (NOX)-dependent stress-response pathway, are also involved in host juvenile growth promotion.
Sixth, feeding and nutrient uptake might be affected by the gut microbiota. Yamada, Deshpande, and colleagues in the Ja’s laboratory looked at how flies obtain nutrients from a given food source. Microorganisms modify the food substrate, including the pH, which is typically lower in their presence; acidic food positively impacts palatability and thus feeding, hence the overall energy and nutrients made available, at least in adults [66]. Furthermore, some of the yeasts often associated with fruitflies in the wild can favor amino-acid harvesting from a poor food substrate, which they share with their host: while growing on the substrate they might transform the food proteins into more easily digestible and absorbable forms, as shown in elegant nutrient radio-labeling experiments [67]. Although not performed on growing larvae, their findings are very interesting. Shin et al. [10] and Storelli et al. [12] reported no detectable impact of microbiota on overall larval feeding with the assays they employed. Radio-labeled nutrient tracing has not been reported for larvae and their mono-associated bacteria yet, a path that is worth pursuing. Wong et al. [68] recently reported that microbiota composition might influence olfactory-guided microbial preferences and substrate foraging: the choice of ingested food must integrate and balance the needs for an optimized source of necessary nutrients and of beneficial microbes.
Finally, host genetic makeup is bound to be important in determining whether, and to what extent, association with a given bacterial species will accelerate growth. Although no extensive published data on the topic are available, researchers are addressing these and similar questions in the context of growth and other microbiota-dependent phenotypes. On the one hand, host response to microbiota removal and manipulation are overall robust and independent from the genetic background of the host: results are largely reproducible when different “wild type” and transgenic strains of D. melanogaster are used in published work. On the other hand, host gene polymorphic differences associated with differential host response could be appreciated in at least one Genome-Wise Association Study (GWAS). This study employed a panel of more than a hundred isogenic D. melanogaster lines (i.e., highly inbred lines, each composed of essentially genetically identical individuals) [69]. Scored parameters consisted of a handful of nutritional indices (such as weight and lipid content) and they were assessed for each isogenic line before and after elimination of the gut microbiota. Indeed, a variation in the magnitude and sign (increase/decrease) of microbiota-dependent effects could be detected [69]. In principle, loci and Single-Nucleotide Polymorphisms (SNPs) associated with larval response to growth-promoting bacteria could be identified with a similar approach. Validation of candidate genes and the exact nature of the changes in gene activity responsible for variation will have to follow.
Given that in nature fruitflies, like most animals, harbor a more complex microbiota, simplified mono-association systems must be complemented by those employing more complex communities. As the number of bacterial species included increases, the questions and methodological approaches shift. To start, bacterial population dynamics might differ; effects on host might change between mono- and bi-, oligo-, or multi-associations, and matrix-like combinatorial association schemes might be used to investigate the simpler cases [70]. As an additional example, the team of Nichole Broderick recently showed that emergent microbiome metabolites—produced by the cooperation of the enzymatic activities of two distinct microorganisms, but not by each of them separately—can affect behavior: oviposition site and food choice. Indeed, Drosophila larvae and adults seem capable to select a multispecies, interactive microbiota in a way capable of increasing their fitness [71]. Hence, inter-species competition and interactions determine microorganismal physiology, metabolite production, and niche occupation. All this is under the influence of environmental parameters, including nutrient source and host, and it can impact host growth, metabolism, and behavior.
We have summarized here (and in Fig. 3) the main outcomes of studies on growth and host–gut microbiota interactions in Drosophila. What else have other invertebrate systems taught us?
Gut Microbiota and Growth in Other Invertebrates
Caenorhabditis elegans, a free-living roundworm, is an extremely powerful genetic model to study development, metabolism, and aging, but it has been until recently at the fringe of gut microbiota research. In the wild, this type of nematodes dwells in the soil, where they eat bacteria. For biomedical research, the bacterivorous C. elegans is traditionally grown on agar plates in “sterility,” with pure cultures of auxotrophic Escherichia coli strains making a bacterial lawn on the agar surface as the only food source for the worms (reviewed in [72]). Perhaps inspired by studies in Drosophila, researchers have now taken advantage of the de facto mono-association of C. elegans with bacterial cultures and screened mutagenized bacteria to study host–microbiota interactions. They have shown that certain E. coli strains and E. coli mutants can influence worm life history traits, including metabolism, growth, and development. They have identified bacterial and worm gene functions (like the gene cya, interfering with the production of cyclic AMP in the bacteria and interfering with the worm TGF-beta pathway controlling developmental transitions). They revealed the importance of gene–diet interactions ([73]; reviewed in [72, 74]). Mono-association with another bacterial species led to the identification of a pathway that promotes growth and molting in the worm and is activated by bacteria. Unexpectedly, this is not the insulin pathway, but one downstream of the novel hormone receptor NHR-23, and independent of both TOR and insulin [75]. NHR23 regulates cyclic genes during molting. Interestingly, NHR-23 is homologous to the fly gene DHR3 (itself target of the molting steroid hormone ecdysone) and to vertebrate ROR-a (a circadian oscillating gene). Thus, a mechanism linking nutrition, intestinal microbiota, and oscillatory gene expression to pace developmental transitions might exist and be evolutionarily conserved. Another exciting paper recently showed that, in the worm, microbiota-produced vitamin B2 critically affects food uptake and feeding behavior. It does so by controlling specific protease gene expression and protease activity in the intestine, downstream of FAD-dependent ATP sensing, channeled via signaling of the TOR Complex 1 [76]. Would this explain why B-vitamins supplementation compensates microbiota elimination in Drosophila larvae [77]? The fast pace at which research in C. elegans can proceed promises many more exciting discoveries in the near future.
Results of research on the microbiota of crustaceans do not deviate from these main lines. The water flea Daphnia magna, an environmental indicator of water quality and an important model system for research in ecology and evolution, also grows smaller when germ free [78]. The difference between symbiotic and germ-free animals is more pronounced under intermediate and high food levels than under low food levels, contrary to what was seen in Drosophila [79]. In alternative aquatic arthropod-microbiota systems using related sets of crustaceans and microbes, gut bacteria appeared to be parasitic under a high-quality diet and mutualistic under a poor diet, while in other cases, bacteria were parasitic under a poor diet. As Callens and coworkers comment [79], it is not easy to predict the strength and direction in which host–microbiota interactions change under different dietary conditions. This echoes the variation in host response to gut microbiota alterations observed among different isogenic fruitfly lines (see point ten above) and warrants caution when claiming the universal applicability of results from specific experimental systems. In addition, given that food quality and quantity is highly variable in natural settings, the ecological relevance of this is clear.
Finally, a lot could be learnt on the impact of microbiota on development from the now classic squid-Vibrio fischeri system, though here the effects are seen on the morphogenesis and differentiation of a specific light organ, rather than growth in size of the animal (reviewed in [80]). For example, Aschtgen et al. [81, 82] showed that outer membrane vesicles, shed from the bioluminescent symbiotic bacteria by blebbing or when rotating their flagella, can signal to host tissue and contribute to organ development. They carry peptidoglycans that trigger haemocyte infiltration in the tissue, one of the host responses contributing to early organ morphogenesis and differentiation. They also contain lipopolysaccharides (LPS), an instructive signal for later differentiation events. Will outer membrane vesicles also be the messengers sent by growth inducing microbiota to enterocytes and other target intestinal and immune cells in flies, mice, and humans? Will they be the carriers of membrane and cell wall macromolecules, like modified teichoic acids? Or would their content (soluble proteins, micro-RNAs, metabolites) be meaningful to bacteria–host communication?
Conversely, host-derived molecules can be important for the maintenance of beneficial microbiota. First, V. fischeri join the developing eye by chemotaxis, following a gradient of chitin molecules (an N-Acetyl-glucosamine polymer found in fungi, insects, and arthropods and some other marine animals) released from the squid light organ epithelium. Second, they can survive in the optic crypt thanks to the circadian release of chitin that they can digest [83]. This is reminiscent of Akkermansia muciniphila feeding on mucus glycoproteins and fucose in human and murine intestines (reviewed in [84]).
We repeatedly underlined that the nutritional and chemico-physical properties of the food substrate influence how and which bacterial species are beneficial for larval growth. In the Drosophila-microbiota symbiosis paradigm for example, both host and bacteria share the same food source and both can modify its properties: chemically by digestion and secretions, or physically by burrowing through it (larvae) or creating biofilm-lined microenvironments (microbes). Both bacteria and host act on the food and their actions are not devoid of consequences for the other symbiotic partner, thus contributing to the mutualistic character of their relationship [60, 78]. Indeed, it is the triad food–host–microbiota that must be considered to fully appreciate the contribution of intestinal microorganisms to animal growth.
A first general conclusion from the studies carried out in invertebrates and summarized above is that the gut microbiota can influence the gut physiology and systemic hormonal systems of its host during growth, in a way the host can take better advantage of the available food for metabolic and anabolic purposes. Regulation of protease expression and modulation of insulin-like and growth hormone pathways appear to be common themes. Having constantly evolved in the presence of both stably and transiently (but recurrently) associated microorganisms, multicellular organisms may have learnt to use their tiny partners not only as sources of macro- and micro-nutrients, or as providers of the most varied enzymatic digestive functions that they lack, but also as environmental indicators. Microbiota, with their capacity to grow more or less well on a given substrate, might be used by animals as quality indicators of the food substrate, to choose between kicking-off or reining-in dispendious digestive and metabolic responses—responses implying massive production of intestinal enzymes and mucus, visceral muscle peristaltic activity, and inter-organ communication with energy-storing organs. The presence of products and metabolites derived from (growing) live bacteria must mean that an adequate food substrate is locally available. This in turn signifies that it is energetically sensible for the animal to stop searching for food: it can start to feed and activate dispendious digestion programs. Otherwise these processes should be shut-off to allow for exploration and search for better nutritional horizons farther away. The ability to detect bacteria for the purpose of immunological defense might share some molecular features with the ability to digest bacteria as food [85]. One can speculate that organisms also evolved a gene regulatory branch, downstream of the recognition of certain microbes, to control energy metabolism and the digestion of foods where those microbes can grow.
The leads offered by the above-mentioned studies in Drosophila and other systems can be extended to murine and clinical work. We next dissect the mechanisms underlying the effects of the gut microbiota on growth in vertebrates.
Gut Microbiota and Growth in Vertebrates
In vertebrates, the hormonal regulation of juvenile growth is governed by the activity of the somatotropic axis [86], consisting genuinely of growth hormone (GH), insulin-like growth factors (IGF-I and -II), and their associated carrier proteins and receptors. GH is released from the anterior lobe of pituitary gland in a pulsatile pattern and acts by binding to its receptor in the membrane of the target cells in the liver or peripheral organs [87]. The binding leads to the initiation of a signaling cascade, leading to phosphorylation of STATs proteins and induction of transcription of GH-regulated genes, such as insulin-like growth factor-1 (IGF-1) [88]. IGF-1 is the main, though not exclusive, mediator of GH actions and it also inhibits GH release by classical negative feedback loop.
Microbiota and Growth in Fish
In 1942, when platyfish (Xiphophorus maculatus) was made and kept germ free for several weeks, fish entered the world of gnotobiology [89]. Soon after other species followed, confirming that fish can survive under germ-free condition for a certain period of time and also highlighting the greatest challenge yet to be conquered in fish gnotobiotic research—the formulation of nutritionally adequate diets to support optimal growth and survival of different germ-free larval stages and germ-free fish in general [90].
The zebrafish (Danio rerio) later established itself as a valuable model in developmental genetics and toxicology [91] and recently it has emerged as a model in physiology [92]. Zebrafish gut bacterial communities have similar phylogenetic diversity compared to those of humans or other mammals, but with little similarity on deeper taxonomic levels [93]. Similarly to what has been observed in other gnotobiotic models, gut bacteria impact fish physiology, namely by increasing intestinal epithelial renewal [94], optimizing the uptake of dietary nutrients [95] and stimulating the innate immune system [96]. No dedicated studies comparing the long-term growth of germ-free and conventional zebrafish have been conducted so far. To our knowledge, only Melancon et al. mentioned that after successfully raising germ-free zebrafish for up to 1 month, the animals were significantly smaller compared to their counterparts raised on conventional diets [90]. But whether this is a direct effect of the lack of bacteria or inadequate food formulation remains to be determined.
More research aims to improve the growth performance in industrial aquacultures via the manipulation of fish-associated microbial communities [97]. Aquaculture represents one of the most important branches of the food supply sector in the world. Similar to Drosophila larvae or other invertebrate water model systems, in the fish, bacteria can reach high densities not only in the host intestine but also in the surrounding environment, where high bacteria burden impacts negatively on host growth. The management of proper water quality poses a challenge to rearing the fish at high densities and at large scale. Fish, and fish larvae in particular, are prone to bacterial infections. As a solution, prophylactic antibiotics have been used to control the bacteria and enhance the overall fish yield and growth performance [98]. However, the unrestricted use of antibiotics has grave negative effects on the environment, animal, and human health and increases the spreading of antibiotic-resistant bacteria [99]. As an alternative, supplementation of fish food with lactic acid bacteria (LAB) is emerging as an alternative concept with promising results [97]. Supplementing the fish feed with LAB can be efficient not only for the management of pathogenic bacteria, but also to enhance the ponderal growth of conventionally reared fish [98, 100, 101]. These findings were further confirmed using the zebrafish model. Avella et al. [102] showed that supplementation with Lactobacillus rhamnosus strain increased both body length and weight of conventionally raised zebrafish at day 20 post egg fertilization. On the molecular level, they showed that administration of L. rhamnosus stimulated the IGF system, which correlated with faster backbone calcification. L. rhamnosus also increased the expression of gonadotropin-releasing hormone, which led to faster gonadal development and sex differentiation. Whether the underlying molecular mechanisms of LAB-mediated growth promotion in fish include improved absorption of nutrients and/or vitamins or the direct impact of LAB on host somatotropic axis requires further experiments.
Gut Microbiota and Growth in Chickens
Although not widely used as a laboratory model animal, historically, chicken has played an important role for the field of gnotobiology. Around the year 1910, Cohendy managed to derive germ-free chickens that thrived well, thus definitely sealing the notion that life without microbes is possible [103]. Cohendy was also the first to realize that germ-free animals may become a powerful experimental tool and can be used beyond the original motive for determining whether life without microorganisms would be possible [104]. Besides rats, mice, and guinea pigs, chicken is the only vertebrate that has successfully completed its full life cycle under germ-free conditions [105]. Unlike other experimental models, the germ-free chicks grow consistently better than their microbiota colonized counterparts, especially on nutritionally marginally adequate diets or on diets with lower protein content [106,107,108,109]. For example, on a low-protein diet, the grow-rate of germ-free chicks can be increased by an astonishing 80% compared to that of the conventionally reared chicks [110]. However, if the general caloric value of the diet is increased, the growth rate of the conventional chicks becomes equal to that of the germ-free controls, even if the dietary protein levels are moderately decreased [111]. Why would bacteria cause growth depression in chickens? So far, several possible explanations have been proposed. The gut microbiota could act as a drain on the utilizable energy or nutrients in the diet, leaving fewer nutrients for the growing chicken host. Alternatively, the CV chicks may suffer from toxic products such as ammonia and endotoxin generated by intestinal microbial activities. Finally, conventional chicks might generally need to invest more energy into the maintenance of intestinal bacterial burden within the gastrointestinal tract: this ‘chronic microbial stress’ would consume energy that could be utilized for enhanced growth and development in germ-free chicks [111].
Supporting the notion that intestinal microbiota represents a burden that demands allocation of protein and energy from the growing host is the observation that, when fed antibiotics, chickens often show improved growth performance and better feed efficiency (conversion of food calories into body mass) [112]. The improved growth performance after subtherapeutic antibiotic treatment (STAT) is especially true for the chicks raised in unsanitary environments and little to no growth stimulation is reported in the clean environments [107, 113]. Similarly, antibiotics have been reported to improve growth of gnotobiotic chickens monocolonized with pathogenic bacteria, such as Enterococcus faecium [114]. These results indeed suggest that the antibiotic modified microbiota poses reduced antigenic and/or toxic challenges to the growing organism and, possibly, that the food nutrients could be protected from the bacterial destruction resulting in their better absorption by the host [115]. However, the growth-promoting actions of antibiotics go probably beyond the intestinal microbiota alterations as seen from experiments in germ-free chicks in 1960s. Pioneering experiments had shown that feeding antibiotics at doses of 50 mg/kg of diet to germ-free chicks resulted in no increase in growth compared to the no antibiotic controls [106, 107]. However, subsequent work with lower levels of antibiotics (8–25 mg/kg of diet) led to entirely different results with significantly increased growth of germ-free chicks fed the supplemented diets [116]. These surprising results imply that, at least in chicken, part of the antibiotic growth-promoting effects are mediated directly through acting on the host tissues.
Extensive use of antibiotics as growth promoters poses health and environmental problems as mentioned previously for the industrial aquacultures. And again, as a promising alternative, lactic acid bacteria are tested as a way to manipulate the intestinal microbiota and to increase animal growth performance. Along these lines, administration of probiotics to chicken has been shown to impact the intestinal microbiota through competitive exclusion and antimicrobial activity, to improve digestion through increased digestive enzymes activity and to modulate the activity of the immune system [117]. A recent study has shown increased growth and food efficiency after Lactobacillus plantarum and inulin administration, which was correlated with increased expression of IGF-1 and GHR in the liver, suggesting the possible impact of Lactobacillus on the somatotropic axis activity [118].
Gut Microbiota and Growth in Mammalian Models and Humans
For practical reasons, guinea pigs were the first mammals to be reared germ free [30]. Subsequently, they were used by most investigators, because as stated by Nuttal and Thierfelder, ‘Of all the mammals concerned, it alone was usable’, referring to the nidifugous status of newborn guinea pigs without a prolonged suckling period [119]. However, it took almost half a century before the problem with appropriate diet was solved and germ-free guinea pigs could be maintained for the required period of time and even brought to reproduction [120]. The growth of germ-free guinea pigs has been consistently reported 20–25% lower compared to the growth rates of conventionally reared animals fed the same diet [119]. Whether this means that the microbiota compensates for a yet unidentified factor in the diet formulation or that it must directly interact with the host to improve growth is still unsolved.
Germ-free rats and mice can be obtained in large quantities from a dam, prior to parturition, by Cesarean operation. Unfortunately, these animals are quite helpless and vulnerable at birth. Therefore, the establishment of long-term breeding colonies (but even rearing till weaning) was impossible before the proper techniques of hand-feeding and appropriate diet were developed [119]. The first long-term colony of germ-free rats was established in 1950 and mice followed soon after [119, 121]. In the initial reports the growth rates of germ-free mice and rats were reported lower (around 80%) compared to those of the conventional animals, with more pronounced differences seen in males [109, 122]. On the contrary, subsequent work showed that germ-free rats can achieve the same growth rates as conventional controls [119]. There are many drawbacks that hinder us from properly assessing the results of these early works. First, the growth of the reference CV animals may differ from one laboratory to another due to the unreported sanitary status. This may result in the same situation as observed in chickens in unsanitary environment, where bacterial burden causes growth depression [123]. Second, as suggested by Gordon, the growth depression observed in germ-free animals may be caused by the crowding effect [119], thus bringing forth the variable of not standardized germ-free animal housing. Third, authors concentrated only on the ponderal growth, the length gain, and the somatotropic axis activity have not been reported.
Intrigued by these discrepancies, we recently assessed the impact of microbiota on the postnatal growth kinetics in specific-pathogen free (SPF) and germ-free mice [9]. We observed that germ-free male BALB/c mice weaned on nutritionally adequate diet showed inhibited ponderal and longitudinal growth, which was not apparent before weaning. The absence of microbiota resulted in the depressed levels of circulating IGF-1, which we showed to be the main mediator of post-weaning growth dynamics. The post-weaning growth differences were even more stunning on a suboptimal isocaloric diet, low in proteins, and fats. On this protein-poor diet, the growth of germ-free mice was completely arrested, while SPF mice continued to grow, albeit at much lower pace compared to control animals on nutritionally adequate diet. These experiments gave the unequivocal evidence that intestinal microbiota is necessary for proper postnatal growth of mammals and, on the molecular level, for maximizing the activity of the somatotropic axis both under adequate and suboptimal nutritional conditions.
As mentioned in the invertebrate section, certain Drosophila-associated Acetobacter and Lactobacillus bacterial strains can promote host larval growth upon nutrient scarcity [10, 12]. Given that lactobacilli can be found in different habitats including the intestinal tracts of vertebrates [124], we tested the possibility that they can improve the growth dynamics also in mice. Indeed, mice monocolonized with the growth-promoting Lactobacillus plantarum WJL strain weaned on the suboptimal diet showed growth rates comparable to the SPF mice weaned on the same diet. Also on the molecular level, L. plantarum WJL improved the activity of the somatotropic axis and the circulating levels of IGF-1 to the extent observed in SPF animals. Similarly to the fly results, the growth-promoting effects were strictly strain specific as other L. plantarum isolates were shown to be less efficient ([9] and Schwarzer et al., unpublished data). Although we still do not know the exact mechanism underlying the observed physical and functional alterations imposed by the bacteria on their host, the fact that the same bacterial strain was able to promote growth in invertebrate and mammalian hosts suggests an evolutionary conserved cross-talk mechanism. Further, our recent data suggest that L. plantarum WJL retains the ability to promote the host growth after supplementation of the conventionally reared mice, making it a strong case for using selected and tested probiotic strains as a complementary strategy to buffer the adverse effects of undernutrition on growth [Schwarzer et al., unpublished data].
In the last decade, the revolution in the appreciation of the fundamental role of intestinal microbiome regarding nearly all aspects of individual’s biology has sparked new interest in germ-free mice breeding [15]. Germ-free mice have been established as an ultimate tool to prove the role of the microbiota in the etiology of the human diseases such as obesity or depression [125, 126]. Particularly, the group of J. I. Gordon has used the gnotobiotic mouse model to establish the link between microbiota alterations and childhood undernutrition [127]. In humans, the consequences of undernutrition during childhood are both short and long term, including stunted growth, cognitive development deficits, underweight, and wasting. Childhood undernutrition is a complex syndrome and current therapies including nutritional interventions and therapeutic food have only limited efficacy [127]. In tackling the question of the role of microbiota in this condition, Smith et al. transplanted into germ-free mice the fecal microbiota of monozygotic Malawian twins, who had become discordant for kwashiorkor (severe acute form of undernutrition). After feeding these mice with the suboptimal diet they observed more severe weight loss in mice colonized with Kwashiorkor-associated microbiota. This microbiota also showed different metabolic profile and more labile and short-lived nature of the responses to the re-nutrition regime compared to the microbiota from healthy twin donor [11]. In the subsequent work, Blanton et al. identified several bacterial taxa in the fecal microbiota of healthy Malawian infants that in the recipient young mice fed the suboptimal diet correlated with lean body mass gain, improved metabolism, and improved bone morphology. Interestingly, when mice receiving microbiota from healthy or severely stunted infants were co-housed together, the growth discriminatory taxa from the microbiota of the former were able to invade that of the latter and prevent the growth impairments in the recipient animals. They identified strains of two bacterial species, Ruminococcus gnavus and Clostridium symbiosum, that were able to stably invade the undernourished donor’s microbiota and ameliorate the impaired growth phenotype [127].
Charboneau et al. used a slightly different approach. They concentrated on sialylated human milk oligosaccharides as a possible mean to modulate the infant’s intestinal microbiota. They colonized 5-week-old germ-free mice with a consortium of bacterial strains cultured from the fecal microbiota of a 6-month-old stunted Malawian infant. Based on the finding that certain milk oligosaccharides were more abundant in the breast milk of Malawian mothers with healthy infants, they fed the colonized mice with a suboptimal diet with or without sialyated milk oligosaccharides. The supplementation led to a microbiota-dependent amelioration of lean body mass gain, improved bone morphology, and a shift in metabolism, suggesting the host’s greater ability to utilize nutrients for anabolism [8]. This suggests that prebiotics like the tested milk oligosaccharides can shape the composition and metabolic output of intestinal microbiota, leading to the amelioration of the adverse effects of malnutrition. Interestingly, mice colonized with the isolated consortium of bacterial strains lost weight in the first week after colonization and showed no lean mass growth over four weeks of the experiment. On the other hand, germ-free mice gained nearly 15% of lean mass during the same time period. This would suggest that also gnotobiotic dysbiotic microbiota, especially under poor nutritional condition, is responsible for growth depression, a phenomenon which we discussed earlier in the part related to growth and intestinal microbiota in the chicken.
A major contributor to childhood malnutrition is environmental enteropathy, a poorly understood inflammatory disorder of the small intestine. Its endemicity in the regions with low sanitation suggests that, besides poor nutrition, microbial exposure or the intestinal microbiota play a role in its development [128]. This was recently confirmed by Brown et al., who induced stunting in SPF mice by feeding a diet low in proteins and fat. Malnourished mice showed profound changes in small intestinal microbiota with bacterial overgrowth in duodenum and jejunum. Repetitive oral exposure of the malnourished mice to commensal Bacteroidales species and Escherichia coli, mimicking the situation of oro-fecal exposure, resulted in intestinal inflammation, villous blunting, and increased intestinal permeability, resembling the features of environmental enteropathy observed in humans [7]. Interestingly, the dysbiotic duodenal microbiota in malnourished mice showed a striking decrease in abundance of the Lactobacillaceae family. This finding, together with our data about the ability of selected lactobacilli strains to support host growth, further advocates for the use of probiotic supplementation together with the re-nutritional therapeutic strategies for treatment of malnutrition.
The fact that the unsanitary conditions negatively affect juvenile growth stands true not only for humans but also for other mammals. Besides the well-documented administration of low doses of antibiotics as growth promoters (discussed in chicken section), attempts have been made to establish ‘specific pathogen-free’ pig farms where the new pig stocks were derived sterilely by Cesarean techniques. This led to the general improvement in production and eradication of common diseases and parasites. Indeed, SPF piglets were reported to grow faster, but they were more prone to infections (reviewed in [129]). Same results concerning superior growth were obtained for aseptically delivered mice kept in clean environment compared to the conventional animals from which they were derived [123]. It thus seems plausible, that getting rid of pathogens or some members of microbiota normally inhabiting the animal intestine, though they need not be strictly pathogenic, alleviates the microbial stress and results in the improved host juvenile growth in mammals.
To shed light on the still enigmatic mode of action of low doses of antimicrobial agents in growth promotion, Cho et al. submitted mice to subtherapeutic antibiotic treatment (STAT). The mice receiving antibiotics showed increased adiposity and alterations in lipid metabolism homeostasis. The increased growth rate was observed only when the treatment was started at weaning and not at later time points. The STAT treatment did not change the absolute bacterial number, but imposed substantial taxonomic changes with increased short-chain fatty acids production [130]. Further, the STAT treatment starting at birth resulted in increased juvenile weight accrual, with both the fat and the lean mass increased at 4 weeks of age. The fact that the STAT-modified microbiota aggravated diet induced obesity phenotype and, after transfer to the germ-free host, induced higher fat-weight gain suggests as possible mechanism the improved diet energy extraction capacity. It seems plausible that this extra energy is used to boost the growth rate of juvenile host and in adults results in the fat-mass gain [18]. Humans, and especially children, are not exposed to the STAT, but the disruption of the microbiota by the short-term therapeutical doses of antibiotics is very common. By treating young mice with therapeutical doses of antibiotics, Nobel et al. showed accelerated total mass and bone growth linked with long-lasting changes in intestinal microbiome diversity [131]. However, here the causality between observed growth phenotypes and altered bacterial taxa was not established. Thus, the direct effects of antibiotic treatment on the host tissues cannot be ruled out.
Presented data from animal models are intriguing in light of the development of adult human height. There has been general trend in continuous height increase across the globe in the second half of the twentieth century. Certainly, increased standard of living together with the improved food security is a factor responsible for this phenomena. Yet, the onset of increasing adult height overlaps with the beginning of the antibiotic era and important changes in public health, especially with better health care for children [132]. This might lead to the reduction in the overall bacterial diversity and disappearance of ancestral indigenous microorganisms from the human microbiome, as recently suggested by the Blaser and Falkow [133]. Together with the findings that the trend for the final adult height is already in place during the very earliest phase of childhood [134], we can speculate that the trend in the adult height is a result of the alleviation of growth depression imposed by intestinal microbiota. Removing pathogens and modifying the diversity of intestinal microbiota with improved hygiene and widespread antibiotic therapeutic use might lead to the same growth-increase phenomenon as observed for the SPF or antibiotic-treated mice.
Taken together, we have seen that in vertebrates the intestinal microbiota represents key partner in the juvenile growth. With the only exception of chicken, which is consistently reported to grow better under germ-free conditions, growing fish and mammals benefit from the presence of the intestinal bacteria. Yet, the presence or absence and amount of certain bacteria are crucial for the positive or negative impacts on the juvenile growth. By modulating the early microbiota composition and metabolic capacities through administration of antibiotics, probiotics or prebiotics, the growth rate of the juvenile vertebrate individual can be increased.
Conclusions
Right Amount, Right Composition, Right Place
The field of microbiota research might be perceived by some as promising more than it could ever deliver: an explanation for the secrets of inner body workings and a universal cure for everything. Yet, the long-neglected influences of the trillions of body-associated microorganisms on physiology and health are hard to overstate. Understanding their basis will not only reveal missing key pieces in the puzzle of biological phenomena, but it will also suggest new, complementary and alternative ways to treat and prevent certain conditions.
In the context of growth, evidence collected in diverse experimental systems points to a clear influence of microbiota composition on the rate of development and time of sexual maturation, both in positive and negative connotations. A similar message is derived from epidemiological and field trials. From the mechanistic point of view, the possibility to genetically manipulate animals and microbes in model systems, combined with a multi-pronged approach to analyze host and microbe transcription, metabolism, and physiology, has provided decisive insights into the genetic and molecular basis of how the gut microbiota affects host’s growth: microorganisms provide key enzymatic digestive functions and essential micro-nutrients; they contribute to the maturation and homeostasis of the intestinal epithelium and associated cell types, including neuronal and immune cells; furthermore, bacterial metabolites and membrane-derived molecules are sensed by the host’s enterocytes to modulate digestive and, possibly, barrier functions. By acting on ingested food, influencing intestinal physiology locally and impacting nutrient flux, the gut microbiota appears to entertain a privileged dialogue with the host’s endocrine and metabolic networks, notably the insulin/IGF-1 and steroid hormone systems, and so to get in charge, at least partially, of animal development and maturation. On the other hand, there is enough evidence showing that some members of the microbiota normally inhabiting the animal intestine, though not always or strictly pathogenic, can cause a depression in juvenile growth. The microbiota community may exert growth-inhibitory effects (i) via the alteration of nutrient absorption in the developing host, both by utilizing nutrients for their own growth and by decreasing the host’s absorptive capacities; (ii) as a consequence of the higher energy demand for the management of this ‘chronic microbial stress.’ Juvenile hosts might then be unable to reach their full genetic growth potential. This can be reversed by modulating the intestinal microbiota composition with low doses antibiotics, prebiotics, or probiotic administration.
Ultimately, the eucaryotic host can influence the growth of its intestinal microbiota too, by providing nutrients, modifying the flow of nutrients in the intestinal tract, and/or modifying the environment that both host and microorganisms share as food substrate [60]. Therefore, the ecological ramifications and relevance of the complex interactions among hosts, microbes, and food now need to be further explored. Transferred to the debate on human nutrition and its long-term personal and social impact, a recent perspective article has called for the combined efforts of nutrition and microbiota researchers, plant scientists, and food and waste engineers for our society to be able to supply the increasing world population with affordable and sustainable food products that can ensure children’s growth and life-long health [135].
Continuing microbiome sequencing efforts, epidemiology, and prospective clinical and field studies on children and adolescents will produce a wealth of data to be interrogated, to validate bench-generated ideas and their applicability. They will also deliver new hypothesis on how nutrition impacts growth via alterations of the microbiota and how pre- and probiotics can restore growth in malnourished subjects: hypothesis to be further tested in animal models. If this is successful, perhaps a future Alice, by optimally managing trillions of her intestinal microscopic friends, will not strike her head against the ceiling after eating all that cake.
References
Efstratiadis A (1998) Genetics of mouse growth. Int J Dev Biol 42(7):955–976
Wilson RS (1979) Twin growth: initial deficit, recovery, and trends in concordance from birth to nine years. Ann Hum Biol 6(3):205–220
Cooke L, Llewellyn C (2016) Nature and nurture in early feeding behavior. Nestle Nutr Inst Workshop Ser 85:155–165. https://doi.org/10.1159/000439507
Nijhout HF, Riddiford LM, Mirth C, Shingleton AW, Suzuki Y, Callier V (2014) The developmental control of size in insects. Wiley Interdiscip Rev Dev Biol 3(1):113–134. https://doi.org/10.1002/wdev.124
Dominguez-Bello MG, Costello EK, Contreras M, Magris M, Hidalgo G, Fierer N, Knight R (2010) Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci USA 107(26):11971–11975. https://doi.org/10.1073/pnas.1002601107
David LA, Maurice CF, Carmody RN, Gootenberg DB, Button JE, Wolfe BE, Ling AV, Devlin AS, Varma Y, Fischbach MA, Biddinger SB, Dutton RJ, Turnbaugh PJ (2014) Diet rapidly and reproducibly alters the human gut microbiome. Nature 505(7484):559–563. https://doi.org/10.1038/nature12820
Brown EM, Wlodarska M, Willing BP, Vonaesch P, Han J, Reynolds LA, Arrieta MC, Uhrig M, Scholz R, Partida O, Borchers CH, Sansonetti PJ, Finlay BB (2015) Diet and specific microbial exposure trigger features of environmental enteropathy in a novel murine model. Nat Commun 6:7806. https://doi.org/10.1038/ncomms8806
Charbonneau MR, O’Donnell D, Blanton LV, Totten SM, Davis JC, Barratt MJ, Cheng J, Guruge J, Talcott M, Bain JR, Muehlbauer MJ, Ilkayeva O, Wu C, Struckmeyer T, Barile D, Mangani C, Jorgensen J, Fan YM, Maleta K, Dewey KG, Ashorn P, Newgard CB, Lebrilla C, Mills DA, Gordon JI (2016) Sialylated milk oligosaccharides promote microbiota-dependent growth in models of infant undernutrition. Cell 164(5):859–871. https://doi.org/10.1016/j.cell.2016.01.024
Schwarzer M, Makki K, Storelli G, Machuca-Gayet I, Srutkova D, Hermanova P, Martino ME, Balmand S, Hudcovic T, Heddi A, Rieusset J, Kozakova H, Vidal H, Leulier F (2016) Lactobacillus plantarum strain maintains growth of infant mice during chronic undernutrition. Science 351(6275):854–857
Shin SC, Kim S-H, You H, Kim B, Kim AC, Lee K-A, Yoon J-H, Ryu J-H, Lee W-J (2011) Drosophila microbiome modulates host developmental and metabolic homeostasis via insulin signaling. Science 334(6056):670–674
Smith MI, Yatsunenko T, Manary MJ, Trehan I, Mkakosya R, Cheng J, Kau AL, Rich SS, Concannon P, Mychaleckyj JC, Liu J, Houpt E, Li JV, Holmes E, Nicholson J, Knights D, Ursell LK, Knight R, Gordon JI (2013) Gut microbiomes of Malawian twin pairs discordant for kwashiorkor. Science 339(6119):548–554. https://doi.org/10.1126/science.1229000
Storelli G, Defaye A, Erkosar B, Hols P, Royet J, Leulier F (2011) Lactobacillus plantarum promotes Drosophila systemic growth by modulating hormonal signals through TOR-dependent nutrient sensing. Cell Metab 14(3):403–414
Mindell DP (1992) Phylogenetic consequences of symbioses: eukarya and eubacteria are not monophyletic taxa. Bio Syst 27(1):53–62
McFall-Ngai M, Hadfield MG, Bosch TC, Carey HV, Domazet-Loso T, Douglas AE, Dubilier N, Eberl G, Fukami T, Gilbert SF, Hentschel U, King N, Kjelleberg S, Knoll AH, Kremer N, Mazmanian SK, Metcalf JL, Nealson K, Pierce NE, Rawls JF, Reid A, Ruby EG, Rumpho M, Sanders JG, Tautz D, Wernegreen JJ (2013) Animals in a bacterial world, a new imperative for the life sciences. Proc Natl Acad Sci USA 110(9):3229–3236. https://doi.org/10.1073/pnas.1218525110
Smith K, McCoy KD, Macpherson AJ (2007) Use of axenic animals in studying the adaptation of mammals to their commensal intestinal microbiota. Semin Immunol 19(2):59–69. https://doi.org/10.1016/j.smim.2006.10.002
Blaser M, Bork P, Fraser C, Knight R, Wang J (2013) The microbiome explored: recent insights and future challenges. Nat Rev Microbiol 11(3):213–217. https://doi.org/10.1038/nrmicro2973
Kau AL, Ahern PP, Griffin NW, Goodman AL, Gordon JI (2011) Human nutrition, the gut microbiome and the immune system. Nature 474(7351):327–336. https://doi.org/10.1038/nature10213
Cox LM, Yamanishi S, Sohn J, Alekseyenko AV, Leung JM, Cho I, Kim SG, Li H, Gao Z, Mahana D, Zarate Rodriguez JG, Rogers AB, Robine N, Loke P, Blaser MJ (2014) Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 158(4):705–721. https://doi.org/10.1016/j.cell.2014.05.052
Hsiao EY, McBride SW, Hsien S, Sharon G, Hyde ER, McCue T, Codelli JA, Chow J, Reisman SE, Petrosino JF, Patterson PH, Mazmanian SK (2013) Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155(7):1451–1463. https://doi.org/10.1016/j.cell.2013.11.024
Naik S, Bouladoux N, Wilhelm C, Molloy MJ, Salcedo R, Kastenmuller W, Deming C, Quinones M, Koo L, Conlan S, Spencer S, Hall JA, Dzutsev A, Kong H, Campbell DJ, Trinchieri G, Segre JA, Belkaid Y (2012) Compartmentalized control of skin immunity by resident commensals. Science 337(6098):1115–1119. https://doi.org/10.1126/science.1225152
McFall-Ngai MJ, Ruby EG (1991) Symbiont recognition and subsequent morphogenesis as early events in an animal-bacterial mutualism. Science 254(5037):1491–1494
Backhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI (2005) Host-bacterial mutualism in the human intestine. Science 307(5717):1915–1920. https://doi.org/10.1126/science.1104816
Buchon N, Broderick NA, Lemaitre B (2013) Gut homeostasis in a microbial world: insights from Drosophila melanogaster. Nat Rev Microbiol 11(9):615–626. https://doi.org/10.1038/nrmicro3074
Sekirov I, Russell SL, Antunes LC, Finlay BB (2010) Gut microbiota in health and disease. Physiol Rev 90(3):859–904. https://doi.org/10.1152/physrev.00045.2009
Human Microbiome Project C (2012) Structure, function and diversity of the healthy human microbiome. Nature 486(7402):207–214. https://doi.org/10.1038/nature11234
Ivanov II, Atarashi K, Manel N, Brodie EL, Shima T, Karaoz U, Wei D, Goldfarb KC, Santee CA, Lynch SV, Tanoue T, Imaoka A, Itoh K, Takeda K, Umesaki Y, Honda K, Littman DR (2009) Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139(3):485–498. https://doi.org/10.1016/j.cell.2009.09.033
Vijay-Kumar M, Aitken JD, Carvalho FA, Cullender TC, Mwangi S, Srinivasan S, Sitaraman SV, Knight R, Ley RE, Gewirtz AT (2010) Metabolic syndrome and altered gut microbiota in mice lacking Toll-like receptor 5. Science 328(5975):228–231. https://doi.org/10.1126/science.1179721
Leulier F, MacNeil LT, Lee WJ, Rawls JF, Cani PD, Schwarzer M, Zhao L, Simpson SJ (2017) Integrative physiology: at the crossroads of nutrition, microbiota, animal physiology, and human health. Cell Metab 25(3):522–534. https://doi.org/10.1016/j.cmet.2017.02.001
Pasteur L (1885) Observations related to the previous notes from M. Duclaux. CR Acad Sci 100:68
Nuttal GHF, Thierfelder H (1895) Thierishes Leben ohne Bakterien im Verdauungskanal. Hoppe-Seyler’s Z Physiol Chem 21:109–121
Gordon HA (1960) The germ-free animal. Its use in the study of “physiologic” effects of the normal microbial flora on the animal host. Am J Dig Dis 5:841–867
Wollman E (1911) Sur l’élevage des mouches stériles. Ann Inst Pasteur 25:79–88
Reyniers JA (1957) The production and use of germ-free animals in experimental biology and medicine. Am J Vet Res 18(68):678–687
Trexler PC, Reynolds LI (1957) Flexible film apparatus for the rearing and use of germfree animals. Appl Microbiol 5(6):406–412
Faith JJ, McNulty NP, Rey FE, Gordon JI (2011) Predicting a human gut microbiota’s response to diet in gnotobiotic mice. Science 333(6038):101–104. https://doi.org/10.1126/science.1206025
Falk PG, Hooper LV, Midtvedt T, Gordon JI (1998) Creating and maintaining the gastrointestinal ecosystem: what we know and need to know from gnotobiology. Microbiol Mol Biol Rev 62(4):1157–1170
Ma D, Storelli G, Mitchell M, Leulier F (2015) Studying host-microbiota mutualism in Drosophila: harnessing the power of gnotobiotic flies. Biomed J 38(4):285–293. https://doi.org/10.4103/2319-4170.158620
Tlaskalova-Hogenova H, Stepankova R, Kozakova H, Hudcovic T, Vannucci L, Tuckova L, Rossmann P, Hrncir T, Kverka M, Zakostelska Z, Klimesova K, Pribylova J, Bartova J, Sanchez D, Fundova P, Borovska D, Srutkova D, Zidek Z, Schwarzer M, Drastich P, Funda DP (2011) The role of gut microbiota (commensal bacteria) and the mucosal barrier in the pathogenesis of inflammatory and autoimmune diseases and cancer: contribution of germ-free and gnotobiotic animal models of human diseases. Cell Mol Immunol 8(2):110–120. https://doi.org/10.1038/cmi.2010.67
Douglas AE (2015) Multiorganismal insects: diversity and function of resident microorganisms. Annu Rev Entomol 60:17–34. https://doi.org/10.1146/annurev-ento-010814-020822
Bakula M (1969) The persistence of a microbial flora during postembryogenesis of Drosophila melanogaster. J Invertebr Pathol 14(3):365–374
Houthoofd K, Braeckman BP, Lenaerts I, Brys K, De Vreese A, Van Eygen S, Vanfleteren JR (2002) Axenic growth up-regulates mass-specific metabolic rate, stress resistance, and extends life span in Caenorhabditis elegans. Exp Gerontol 37(12):1371–1378
Kwong WK, Moran NA (2016) Gut microbial communities of social bees. Nat Rev Microbiol 14(6):374–384. https://doi.org/10.1038/nrmicro.2016.43
McFall-Ngai M, Hadfield MG, Bosch TCG, Carey HV, Domazet-Loso T, Douglas AE, Dubilier N, Eberl G, Fukami T, Gilbert SF, Hentschel U, King N, Kjelleberg S, Knoll AH, Kremer N, Mazmanian SK, Metcalf JL, Nealson K, Pierce NE, Rawls JF, Reid A, Ruby EG, Rumpho M, Sanders JG, Tautz D, Wernegreen JJ (2013) Animals in a bacterial world, a new imperative for the life sciences. Proc Natl Acad Sci USA 110(9):3229–3236
Brown SD, Moore MW (2012) The International Mouse Phenotyping Consortium: past and future perspectives on mouse phenotyping. Mamm Genome 23(9–10):632–640. https://doi.org/10.1007/s00335-012-9427-x
Chandler JA, Lang JM, Bhatnagar S, Eisen JA, Kopp A (2011) Bacterial communities of diverse Drosophila species: ecological context of a host-microbe model system. PLoS Genet 7(9):e1002272. https://doi.org/10.1371/journal.pgen.1002272
Ryu JH, Kim SH, Lee HY, Bai JY, Nam YD, Bae JW, Lee DG, Shin SC, Ha EM, Lee WJ (2008) Innate immune homeostasis by the homeobox gene caudal and commensal-gut mutualism in Drosophila. Science 319(5864):777–782. https://doi.org/10.1126/science.1149357
Wong CN, Ng P, Douglas AE (2011) Low-diversity bacterial community in the gut of the fruitfly Drosophila melanogaster. Environ Microbiol 13(7):1889–1900. https://doi.org/10.1111/j.1462-2920.2011.02511.x
Hoang D, Kopp A, Chandler JA (2015) Interactions between Drosophila and its natural yeast symbionts-Is Saccharomyces cerevisiae a good model for studying the fly-yeast relationship? PeerJ 3:e1116. https://doi.org/10.7717/peerj.1116
Webster CL, Waldron FM, Robertson S, Crowson D, Ferrari G, Quintana JF, Brouqui JM, Bayne EH, Longdon B, Buck AH, Lazzaro BP, Akorli J, Haddrill PR, Obbard DJ (2015) The discovery, distribution, and evolution of viruses associated with Drosophila melanogaster. PLoS Biol 13(7):e1002210. https://doi.org/10.1371/journal.pbio.1002210
Wong AC, Vanhove AS, Watnick PI (2016) The interplay between intestinal bacteria and host metabolism in health and disease: lessons from Drosophila melanogaster. Dis Model Mech 9(3):271–281. https://doi.org/10.1242/dmm.023408
Martino ME, Ma D, Leulier F (2017) Microbial influence on Drosophila biology. Curr Opin Microbiol 38:165–170. https://doi.org/10.1016/j.mib.2017.06.004
Layalle S, Arquier N, Leopold P (2008) The TOR pathway couples nutrition and developmental timing in Drosophila. Dev Cell 15(4):568–577. https://doi.org/10.1016/j.devcel.2008.08.003
Boulan L, Milan M, Leopold P (2015) The systemic control of growth. Cold Spring Harbor Perspect Biol. https://doi.org/10.1101/cshperspect.a019117
Ridley EV, Wong AC, Westmiller S, Douglas AE (2012) Impact of the resident microbiota on the nutritional phenotype of Drosophila melanogaster. PloS ONE 7(5):e36765. https://doi.org/10.1371/journal.pone.0036765
Tefit MA, Leulier F (2017) Lactobacillus plantarum favors the early emergence of fit and fertile adult Drosophila upon chronic undernutrition. J Exp Biol 220(Pt 5):900–907. https://doi.org/10.1242/jeb.151522
Erkosar B, Storelli G, Mitchell M, Bozonnet L, Bozonnet N, Leulier F (2015) Pathogen virulence impedes mutualist-mediated enhancement of host juvenile growth via inhibition of protein digestion. Cell Host Microbe 18(4):445–455. https://doi.org/10.1016/j.chom.2015.09.001
Matos R, Schwarzer M, Gervais H, Courtin P, Joncour P, Gillet B, Ma D, Bulteau AL, Martino ME, Hughes S, Chapot-Chartier MP, Leulier F (2017) D-Alanylation of teichoic acids contributes to Lactobacillus plantarum-mediated Drosophila growth during chronic undernutrition. Nature microbiol. https://doi.org/10.1038/s41564-017-0038-x
Hang S, Purdy AE, Robins WP, Wang Z, Mandal M, Chang S, Mekalanos JJ, Watnick PI (2014) The acetate switch of an intestinal pathogen disrupts host insulin signaling and lipid metabolism. Cell Host Microbe 16(5):592–604. https://doi.org/10.1016/j.chom.2014.10.006
Chaston JM, Newell PD, Douglas AE (2014) Metagenome-wide association of microbial determinants of host phenotype in Drosophila melanogaster. mBio 5(5):e01631–e01614. https://doi.org/10.1128/mBio.01631-14
Storelli G, Strigini M, Grenier T, Bozonnet L, Schwarzer M, Daniel C, Matos R, Leulier F (2017) Drosophila perpetuates nutritional mutualism by promoting the fitness of its intestinal symbiont Lactobacillus plantarum. Cell Metab. https://doi.org/10.1016/j.cmet.2017.11.011
Overend G, Luo Y, Henderson L, Douglas AE, Davies SA, Dow JA (2016) Molecular mechanism and functional significance of acid generation in the Drosophila midgut. Sci Rep 6:27242. https://doi.org/10.1038/srep27242
Shanbhag SR, Vazhappilly AT, Sane A, D’Silva NM, Tripathi S (2017) Electrolyte transport pathways induced in the midgut epithelium of Drosophila melanogaster larvae by commensal gut microbiota and pathogens. J Physiol 595(2):523–539. https://doi.org/10.1113/JP272617
Jones RM, Luo L, Ardita CS, Richardson AN, Kwon YM, Mercante JW, Alam A, Gates CL, Wu H, Swanson PA, Lambeth JD, Denning PW, Neish AS (2013) Symbiotic lactobacilli stimulate gut epithelial proliferation via Nox-mediated generation of reactive oxygen species. EMBO J 32(23):3017–3028. https://doi.org/10.1038/emboj.2013.224
Jones RM, Desai C, Darby TM, Luo L, Wolfarth AA, Scharer CD, Ardita CS, Reedy AR, Keebaugh ES, Neish AS (2015) Lactobacilli modulate epithelial cytoprotection through the Nrf2 pathway. Cell Rep 12(8):1217–1225. https://doi.org/10.1016/j.celrep.2015.07.042
Lee WJ (2008) Bacterial-modulated signaling pathways in gut homeostasis. Sci Signal 1(21):pe24. https://doi.org/10.1126/stke.121pe24
Deshpande SA, Yamada R, Mak CM, Hunter B, Soto Obando A, Hoxha S, Ja WW (2015) Acidic food pH increases palatability and consumption and extends drosophila lifespan. J Nutr 145(12):2789–2796. https://doi.org/10.3945/jn.115.222380
Yamada R, Deshpande SA, Bruce KD, Mak EM, Ja WW (2015) Microbes promote amino acid harvest to rescue undernutrition in Drosophila. Cell Rep. https://doi.org/10.1016/j.celrep.2015.01.018
Wong AC, Wang QP, Morimoto J, Senior AM, Lihoreau M, Neely GG, Simpson SJ, Ponton F (2017) Gut microbiota modifies olfactory-guided microbial preferences and foraging decisions in Drosophila. Curr Biol 27(15):2397–2404 e2394. https://doi.org/10.1016/j.cub.2017.07.022
Dobson AJ, Chaston JM, Newell PD, Donahue L, Hermann SL, Sannino DR, Westmiller S, Wong AC, Clark AG, Lazzaro BP, Douglas AE (2015) Host genetic determinants of microbiota-dependent nutrition revealed by genome-wide analysis of Drosophila melanogaster. Nat Commun 6:6312. https://doi.org/10.1038/ncomms7312
Newell PD, Douglas AE (2014) Interspecies interactions determine the impact of the gut microbiota on nutrient allocation in Drosophila melanogaster. Appl Environ Microbiol 80(2):788–796. https://doi.org/10.1128/AEM.02742-13
Fischer CN, Trautman EP, Crawford JM, Stabb EV, Handelsman J, Broderick NA (2017) Metabolite exchange between microbiome members produces compounds that influence Drosophila behavior. https://doi.org/10.7554/eLife.18855
Shapira M (2017) Host-microbiota interactions in Caenorhabditis elegans and their significance. Curr Opin Microbiol 38:142–147. https://doi.org/10.1016/j.mib.2017.05.012
Khanna A, Kumar J, Vargas MA, Barrett L, Katewa S, Li P, McCloskey T, Sharma A, Naude N, Nelson C, Brem R, Killilea DW, Mooney SD, Gill M, Kapahi P (2016) A genome-wide screen of bacterial mutants that enhance dauer formation in C. elegans. Sci Rep 6:38764. https://doi.org/10.1038/srep38764
Yen CA, Curran SP (2016) Gene-diet interactions and aging in C. elegans. Exp Gerontol 86:106–112. https://doi.org/10.1016/j.exger.2016.02.012
MacNeil LT, Watson E, Arda HE, Zhu LJ, Walhout AJ (2013) Diet-induced developmental acceleration independent of TOR and insulin in C. elegans. Cell 153(1):240–252. https://doi.org/10.1016/j.cell.2013.02.049
Qi B, Kniazeva M, Han M (2017) A vitamin-B2-sensing mechanism that regulates gut protease activity to impact animal’s food behavior and growth. https://doi.org/10.7554/eLife.26243
Wong AC, Dobson AJ, Douglas AE (2014) Gut microbiota dictates the metabolic response of Drosophila to diet. J Exp Biol 217(Pt 11):1894–1901. https://doi.org/10.1242/jeb.101725
Sison-Mangus MP, Mushegian AA, Ebert D (2015) Water fleas require microbiota for survival, growth and reproduction. ISME J 9(1):59–67. https://doi.org/10.1038/ismej.2014.116
Callens M, Macke E, Muylaert K, Bossier P, Lievens B, Waud M, Decaestecker E (2016) Food availability affects the strength of mutualistic host-microbiota interactions in Daphnia magna. ISME J 10(4):911–920. https://doi.org/10.1038/ismej.2015.166
McFall-Ngai MJ (2014) The importance of microbes in animal development: lessons from the squid-vibrio symbiosis. Annu Rev Microbiol 68:177–194. https://doi.org/10.1146/annurev-micro-091313-103654
Aschtgen MS, Lynch JB, Koch E, Schwartzman J, McFall-Ngai M, Ruby E (2016) Rotation of Vibrio fischeri flagella produces outer membrane vesicles that induce host development. J Bacteriol 198(16):2156–2165. https://doi.org/10.1128/JB.00101-16
Aschtgen MS, Wetzel K, Goldman W, McFall-Ngai M, Ruby E (2016) Vibrio fischeri-derived outer membrane vesicles trigger host development. Cell Microbiol 18(4):488–499. https://doi.org/10.1111/cmi.12525
Schwartzman JA, Koch E, Heath-Heckman EA, Zhou L, Kremer N, McFall-Ngai MJ, Ruby EG (2015) The chemistry of negotiation: rhythmic, glycan-driven acidification in a symbiotic conversation. Proc Natl Acad Sci USA 112(2):566–571. https://doi.org/10.1073/pnas.1418580112
Pickard JM, Chervonsky AV (2015) Intestinal fucose as a mediator of host-microbe symbiosis. J Immunol 194(12):5588–5593. https://doi.org/10.4049/jimmunol.1500395
Broderick NA (2016) Friend, foe or food? Recognition and the role of antimicrobial peptides in gut immunity and Drosophila-microbe interactions. Philos Trans R Soc London Ser B. https://doi.org/10.1098/rstb.2015.0295
Breier BH (1999) Regulation of protein and energy metabolism by the somatotropic axis. Domest Anim Endocrinol 17(2–3):209–218
Hartman ML, Veldhuis JD, Thorner MO (1993) Normal control of growth hormone secretion. Horm Res 40(1–3):37–47
Bartke A, Sun LY, Longo V (2013) Somatotropic signaling: trade-offs between growth, reproductive development, and longevity. Physiol Rev 93(2):571–598
Baker JA, Ferguson MS, TenBroeck C (1942) Growth of platyfish (Platypoecilus maculatus) free from bacteria and other micro’organisms. Proc Soc Exp Biol Med 51(1):116–119. https://doi.org/10.3181/00379727-51-13854
Melancon E, Gomez De La Torre Canny S, Sichel S, Kelly M, Wiles TJ, Rawls JF, Eisen JS, Guillemin K (2017) Best practices for germ-free derivation and gnotobiotic zebrafish husbandry. Methods Cell Biol 138:61–100. https://doi.org/10.1016/bs.mcb.2016.11.005
Grunwald DJ, Eisen JS (2002) Headwaters of the zebrafish—emergence of a new model vertebrate. Nat Rev Genet 3(9):717–724. https://doi.org/10.1038/nrg892
Briggs JP (2002) The zebrafish: a new model organism for integrative physiology. Am J Physiol Regul Integr Comp Physiol 282(1):R3–R9. https://doi.org/10.1152/ajpregu.00589.2001
Rawls JF, Mahowald MA, Ley RE, Gordon JI (2006) Reciprocal gut microbiota transplants from zebrafish and mice to germ-free recipients reveal host habitat selection. Cell 127(2):423–433. https://doi.org/10.1016/j.cell.2006.08.043
Cheesman SE, Neal JT, Mittge E, Seredick BM, Guillemin K (2011) Epithelial cell proliferation in the developing zebrafish intestine is regulated by the Wnt pathway and microbial signaling via Myd88. Proc Natl Acad Sci USA 108(Suppl 1):4570–4577. https://doi.org/10.1073/pnas.1000072107
Semova I, Carten JD, Stombaugh J, Mackey LC, Knight R, Farber SA, Rawls JF (2012) Microbiota regulate intestinal absorption and metabolism of fatty acids in the zebrafish. Cell Host Microbe 12(3):277–288. https://doi.org/10.1016/j.chom.2012.08.003
Kanther M, Tomkovich S, Xiaolun S, Grosser MR, Koo J, Flynn EJ 3rd, Jobin C, Rawls JF (2014) Commensal microbiota stimulate systemic neutrophil migration through induction of serum amyloid A. Cell Microbiol 16(7):1053–1067. https://doi.org/10.1111/cmi.12257
Bentzon-Tilia M, Sonnenschein EC, Gram L (2016) Monitoring and managing microbes in aquaculture—towards a sustainable industry. Microb Biotechnol 9(5):576–584. https://doi.org/10.1111/1751-7915.12392
Cabello FC (2006) Heavy use of prophylactic antibiotics in aquaculture: a growing problem for human and animal health and for the environment. Environ Microbiol 8(7):1137–1144. https://doi.org/10.1111/j.1462-2920.2006.01054.x
Cabello FC, Godfrey HP, Tomova A, Ivanova L, Dolz H, Millanao A, Buschmann AH (2013) Antimicrobial use in aquaculture re-examined: its relevance to antimicrobial resistance and to animal and human health. Environ Microbiol 15(7):1917–1942. https://doi.org/10.1111/1462-2920.12134
Dhanaraj M, Haniffa MA, Singh SVA, Arockiaraj AJ, Ramakrishanan CM, Seetharaman S, Arthimanju R (2010) Effect of probiotics on growth performance of Koi Carp (Cyprinus carpio). J Appl Aquac 22(3):202–209. https://doi.org/10.1080/10454438.2010.497739
Suzer C, Çoban D, Kamaci HO, Saka Ş, Firat K, Otgucuoğlu Ö, Küçüksari H (2008) Lactobacillus spp. bacteria as probiotics in gilthead sea bream (Sparus aurata, L.) larvae: effects on growth performance and digestive enzyme activities. Aquaculture 280(1):140–145. https://doi.org/10.1016/j.aquaculture.2008.04.020
Avella MA, Place A, Du SJ, Williams E, Silvi S, Zohar Y, Carnevali O (2012) Lactobacillus rhamnosus accelerates zebrafish backbone calcification and gonadal differentiation through effects on the GnRH and IGF systems. PloS ONE 7(9):e45572. https://doi.org/10.1371/journal.pone.0045572
Cohendy M (1912) Experiences sur la vie sans microbes. Ann inst Pasteur 26:106–137
Reyniers JA (1959) The pure-culture concept and gnotobiotics. Ann N Y Acad Sci 78(1):3–16. https://doi.org/10.1111/j.1749-6632.1959.tb53091.x
Reyniers JA, Trexler PC et al (1948) A complete life-cycle in the germ-free bantam chicken. Nature 162(4132):67
Coates ME, Fuller R, Harrison GF, Lev M, Suffolk SF (1963) A comparison of the growth of chicks in the Gustafsson germ-free apparatus and in a conventional environment, with and without dietary supplements of penicillin. Br J Nutr 17:141–150
Forbes M, Park JT (1959) Growth of germ-free and conventional chicks: effect of diet, dietary penicillin and bacterial environment. J Nutr 67(1):69–84
Furuse M, Yokota H (1985) Effect of the gut microflora on chick growth and utilisation of protein and energy at different concentrations of dietary protein. Br Poult Sci 26(1):97–104. https://doi.org/10.1080/00071668508416791
Gordon HA (1959) Morphological and physiological characterization of germfree life*. Ann N Y Acad Sci 78 (1):208–220. https://doi.org/10.1111/j.1749-6632.1959.tb53104.x
Furuse M, Yokota H (1984) Protein and energy utilization in germ-free and conventional chicks given diets containing different levels of dietary protein. Br J Nutr 51(2):255–264
Furuse M, Okumura J (1994) Nutritional and physiological characteristics in germ-free chickens. Compar Biochem Physiol A 109(3):547–556
Lochmiller RL, Deerenberg C (2000) Trade-offs in evolutionary immunology: just what is the cost of immunity? Oikos 88(1):87–98. https://doi.org/10.1034/j.1600-0706.2000.880110.x
Roura E, Homedes J, Klasing KC (1992) Prevention of immunologic stress contributes to the growth-permitting ability of dietary antibiotics in chicks. J Nutr 122(12):2383–2390
Fuller R, Houghton SB, Coates ME (1983) The effect of dietary penicillin on the growth of gnotobiotic chickens monoassociated with streptococcus faecium. Br Poult Sci 24(1):111–114. https://doi.org/10.1080/00071668308416719
Butaye P, Devriese LA, Haesebrouck F (2003) Antimicrobial growth promoters used in animal feed: effects of less well known antibiotics on gram-positive bacteria. Clin Microbiol Rev 16(2):175–188
Luckey TD (1956) Mode of action of antibiotics—evidence from germfree birds. In: Proceedings, First International Conference on the Use of Antibiotics in Agriculture. The National Academies Press, Washington, DC. https://doi.org/10.17226/21265
Lutful Kabir SM (2009) The role of probiotics in the poultry industry. Int J Mol Sci 10(8):3531–3546. https://doi.org/10.3390/ijms10083531
Kareem KY, Loh TC, Foo HL, Akit H, Samsudin AA (2016) Effects of dietary postbiotic and inulin on growth performance, IGF1 and GHR mRNA expression, faecal microbiota and volatile fatty acids in broilers. BMC Vet Res 12(1):163. https://doi.org/10.1186/s12917-016-0790-9
Luckey TD (1963) Germfree life and gnotobiology. Academic Press, Cambridge
Teah BA (1960) Germ-free animal production at Lobund Institute. In: Proceedings of the 2nd symposium on gnotobiotic technology. University of Notre Dame Press, p 25
Pleasants JR (1959) Rearing germfree cesarean-born rats, mice, and rabbits through weaning. Ann N Y Acad Sci 78:116–126
Wostmann BS (1959) Nutrition of the germfree mammal*. Ann N Y Acad Sci 78(1):175–182. https://doi.org/10.1111/j.1749-6632.1959.tb53102.x
Dubos RJ, Schaedler RW (1960) The effect of the intestinal flora on the growth rate of mice, and on their susceptibility to experimental infections. J Exp Med 111:407–417
Martino ME, Bayjanov JR, Caffrey BE, Wels M, Joncour P, Hughes S, Gillet B, Kleerebezem M, van Hijum SA, Leulier F (2016) Nomadic lifestyle of Lactobacillus plantarum revealed by comparative genomics of 54 strains isolated from different habitats. Environ Microbiol 18(12):4974–4989. https://doi.org/10.1111/1462-2920.13455
De Palma G, Lynch MDJ, Lu J, Dang VT, Deng Y, Jury J, Umeh G, Miranda PM, Pigrau Pastor M, Sidani S, Pinto-Sanchez MI, Philip V, McLean PG, Hagelsieb M-G, Surette MG, Bergonzelli GE, Verdu EF, Britz-McKibbin P, Neufeld JD, Collins SM, Bercik P (2017) Transplantation of fecal microbiota from patients with irritable bowel syndrome alters gut function and behavior in recipient mice. Sci Transl Med 9(379):eaaf6397. https://doi.org/10.1126/scitranslmed.aaf6397
Ridaura VK, Faith JJ, Rey FE, Cheng J, Duncan AE, Kau AL, Griffin NW, Lombard V, Henrissat B, Bain JR, Muehlbauer MJ, Ilkayeva O, Semenkovich CF, Funai K, Hayashi DK, Lyle BJ, Martini MC, Ursell LK, Clemente JC, Van Treuren W, Walters WA, Knight R, Newgard CB, Heath AC, Gordon JI (2013) Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 341(6150):1241214. https://doi.org/10.1126/science.1241214
Blanton LV, Charbonneau MR, Salih T, Barratt MJ, Venkatesh S, Ilkaveya O, Subramanian S, Manary MJ, Trehan I, Jorgensen JM, Fan YM, Henrissat B, Leyn SA, Rodionov DA, Osterman AL, Maleta KM, Newgard CB, Ashorn P, Dewey KG, Gordon JI (2016) Gut bacteria that prevent growth impairments transmitted by microbiota from malnourished children. Science. https://doi.org/10.1126/science.aad3311
Ahmed T, Hossain M, Mahfuz M, Choudhury N, Hossain MM, Bhandari N, Lin MM, Joshi PC, Angdembe MR, Wickramasinghe VP, Hossain SMM, Shahjahan M, Irianto SE, Soofi S, Bhutta Z (2014) Severe acute malnutrition in Asia. Food Nutrition Bulletin 35(2_suppl1):S14–S26. https://doi.org/10.1177/15648265140352s103
Kirk RG (2012) “Life in a germ-free world”: isolating life from the laboratory animal to the bubble boy. Bull History Med 86(2):237–275. https://doi.org/10.1353/bhm.2012.0028
Cho I, Yamanishi S, Cox L, Methe BA, Zavadil J, Li K, Gao Z, Mahana D, Raju K, Teitler I, Li H, Alekseyenko AV, Blaser MJ (2012) Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 488(7413):621–626. https://doi.org/10.1038/nature11400
Nobel YR, Cox LM, Kirigin FF, Bokulich NA, Yamanishi S, Teitler I, Chung J, Sohn J, Barber CM, Goldfarb DS, Raju K, Abubucker S, Zhou Y, Ruiz VE, Li H, Mitreva M, Alekseyenko AV, Weinstock GM, Sodergren E, Blaser MJ (2015) Metabolic and metagenomic outcomes from early-life pulsed antibiotic treatment. Nat Commun 6:7486. https://doi.org/10.1038/ncomms8486
Stulp G, Barrett L (2016) Evolutionary perspectives on human height variation. Biol Rev Camb Philos Soc 91(1):206–234. https://doi.org/10.1111/brv.12165
Blaser MJ, Falkow S (2009) What are the consequences of the disappearing human microbiota? Nat Rev Microbiol 7(12):887–894. https://doi.org/10.1038/nrmicro2245
Cole TJ (2000) Secular trends in growth. Proc Nutr Soc 59(2):317–324
Barratt MJ, Lebrilla C, Shapiro HY, Gordon JI (2017) The gut microbiota, food science, and human nutrition: a timely marriage. Cell Host Microbe 22(2):134–141. https://doi.org/10.1016/j.chom.2017.07.006
Acknowledgements
FL lab is supported by an ERC starting grant (FP7/2007-2013-N°309704), FINOVI foundation and the EMBO Young Investigator Program. MSch would like to acknowledge financial support from The Neuron Fund for the Support of Science (Neuron Fund). MSt was supported by Actions d’appui à la Recherche 2017 from UJM Saint-Etienne, University of Lyon.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Martin Schwarzer, Maura Strigini and François Leulier declare that they have no conflict of interest.
Glossary (Based on Encyclopedia Britannica and Wikipedia, with Modifications)
- Dysbiosis
-
Microbial imbalance in the gastrointestinal tract. Change in numbers or proportion of different members of microbiome resulting in the adverse effects on the host
- Environmental enteropathy
-
Chronic disease of small intestine characterized by gut inflammation and barrier disruption, malabsorption, and systemic inflammation in the absence of diarrhea. Endemic in the areas with poor sanitation and high enteropathogen burden
- Gnotobiosis
-
A condition in which all the forms of life associated with an organism can be accounted for. An extreme case is germ-free (axenic) animal which means organism with no associated living microbiota detectable by the up-to-date techniques
- Holobiont
-
The assembly of different species that form an ecological unit. For the purpose of this review, it is used as the eukaryotic host plus all of its symbiotic microbes
- Kwashiorkor
-
Severe form of undernutrition when protein intake is insufficient
- Microbiome
-
The collective genomes of the microorganisms that reside in an environmental niche
- Microbiota
-
An ecological community of commensal, symbiotic, and pathogenic microorganisms found in and on a multicellular organism. It includes bacteria, archaea, protists, fungi, and viruses
- Nidifugous
-
Nidifugous organisms are those that leave the nest shortly after hatching or birth. They are born with open eyes and are capable of independent locomotion
- Prebiotics
-
Non-digestible oligo- and polysaccharide compounds that induce the growth or activity of certain microorganisms
- Probiotics
-
Live microorganisms that, when administered in adequate amounts, confer a health benefit on the host
- Somatotropic axis
-
One of the major hormonal systems regulating postnatal growth in vertebrates. It refers to the hormonal signaling from hypothalamus to anterior pituitary gland, resulting in the release of growth hormone, which in turn stimulates the production of insulin-like growth factor-1 in the liver and peripheral organs
- Subtherapeutical antibiotic treatment
-
Subtherapeutic use of antibiotics in animal feed, as opposed to therapeutic or disease-treating use, enhances efficiency of livestock production by promoting growth. Specifically, through a still unknown mechanism, an animal on subtherapeutic doses of antibiotics will, on a lesser quantity of feed, gain an equal amount of weight as an untreated animal
Rights and permissions
About this article
Cite this article
Schwarzer, M., Strigini, M. & Leulier, F. Gut Microbiota and Host Juvenile Growth. Calcif Tissue Int 102, 387–405 (2018). https://doi.org/10.1007/s00223-017-0368-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00223-017-0368-y