Abstract
In this work, simple, rapid, and low-cost multiplexed detection of tumor-related micro-RNAs (miRNAs) was achieved based on multi-color fluorescence on a microfluidic droplet chip, which simplified the complexity of light path to a half. A four-T-junction structure was fabricated to form uniform nano-volume droplet arrays with customized contents. Multi-color quantum dots (QDs) used as the fluorescence labels were encapsulated into droplets to develop the multi-path fluorescence detection module. We designed an integrated multiplex fluorescence resonance energy transfer system assisted by multiple QDs (four colors) and one quencher to detect four tumor-related miRNAs (miRNA-20a, miRNA-21, miRNA-155, and miRNA-221). The qualitative analysis of miRNAs was realized by the color identification of QDs, while the quantitative detection of miRNAs was achieved based on the linear relationship between the quenching efficiency of QDs and the concentration of miRNAs. The practicability of the multiplex detection device was further confirmed by detecting four tumor-related miRNAs in real human serum samples. The detection limits of four miRNAs ranged from 35 to 39 pmol/L was achieved without any target amplification. And the linear range was from 0.1 nmol/L to 1 μmol/L using 10 nL detection volume (one droplet) under the detection speed of 320 droplets per minute. The multiple detection system for miRNAs is simple, fast, and low-cost and will be a powerful platform for clinical diagnostic analysis.

Graphical abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MiRNAs are a class of tiny, endogenous, non-coding RNAs (approximately 22 nucleotides) that regulate gene expression by modulating the activity of specific miRNA targets in many cellular, physiologic, and pathologic processes [1], including differentiation, proliferation, apoptosis, metabolism, viral defense, and hematopoiesis [1,2,3,4]. Many diseases are proved to correlate to the dysregulation of miRNAs, such as tumors [5], cancers [6], psychiatric syndromes [7], and cardiovascular diseases [8, 9]. So miRNAs are considered as a promising class of tumor-derived biomarkers. Especially since the circulating miRNAs are found to express stably in the human blood [10], they show great potential to be non-invasive biomarkers for early diagnosis and prognosis assessment of cancer [11, 12]. However, it is insufficient for diagnosing the disease in early stage only by detecting the single kind of tumor-derived miRNAs, because the molecular mechanism of the miRNAs is so complicated that one disease usually involves aberrant expression of a group of miRNAs (generally 2–15) [13,14,15]. Hence, it is very necessary to develop a direct and quantitative analysis of multiple miRNAs for distinguishing cancerous tissues and even specific cancer cell subtypes [16,17,18], rising the accuracy of early-stage evaluation of cancers and contributing to investigate the mechanism of miRNAs at molecular and gene level.
Currently, the typical multiplex detection method is the microarray based on hybridization, but it often shows comparatively low sensitivity [33,34,35]. Taking high sensitive methods (such as qRT-PCR or biochemical sensor systems) for the detection of multiple miRNAs will lead to high consumption of labor, material, and time for the multiple amplification processes and enzymatic/chemical modifications [36, 37]. Hence, it is important to develop a simple, low-cost, and rapid method for multiplex detection of miRNAs.
Droplet microfluidic system usually utilizes two immiscible liquid phases to form dispersed microdroplets, which can be used as nano/pico-liter reactors for low-consumption, rapid, and high-throughput analysis [19, 20]. It has been developed for the sensitive and high-throughput analysis of nucleic acids [21]. Furthermore, the multiplex detection of nucleic acids has been reported for different applications. Kim et al. developed a multiplexed micro-RNA detection method by isolating encoded, functional hydrogel microparticles based on microfluidic droplet system and achieved quantitative detection of three miRNAs [22]. Yuan et al. developed a droplet microfluidic system for multi-target foodborne pathogens detection based on loop-mediated isothermal amplification. However, the complexity and throughput still need to be considered in the future [23].
In this work, we aim at setting up a simple, rapid, and low-cost multiplexed detection method of miRNAs based on a four-T-junction droplet chip with integrated multiplex laser-induced fluorescence detection module. Compared with traditional methods, miRNA detection in droplets can effectively reduce the consumption of samples. At the same time, the detection speed can be greatly improved by the flow detection mode. The most characteristic is that multiplexed detection can be realized based on our droplet detection system. The concentrations of four targets can be measured based on the effect of fluorescence resonance energy transfer (FRET) between four color quantum dots (QDs) and quenchers. The qualitative analysis of four miRNAs was realized by the color identification of QDs, while the quantitative detection of miRNAs was achieved based on the proportional relationship between quenching efficiency and miRNAs concentration. Four miRNAs, miRNA-21, miR-221, miR-155, and miR-20a, were chosen as model targets. MiRNA 21 is a broad-spectrum tumor marker, and its over expression is found in 80% of tumor samples [24, 25]. Otherwise, miR-221, miR-155, and miR-20a associate with various cancers, such as breast, liver, lung, and B cell and Hodgkin’s lymphomas [26,27,28,29]. The simultaneous detection of four miRNAs will greatly increase the accuracy of early non-invasive measurement for tumors or cancers.
Experimental
Regents and materials
Carboxyl water-soluble quantum dots (QDs, the maximum fluorescence emission wavelengths of 525 ± 5 nm, 565 ± 5 nm, 605 ± 5 nm, and 650 ± 5 nm, 8 μmol/L) were purchased from Jiayuan Quantum Dots Co., Ltd., (Wuhan, China). All single-stranded oligonucleotides were purchased from Sangon Biotech Co., Ltd. (Shanghai, China). The sequences of primer probes and targets are shown in Table 1. 1-Ethyl-(3-dimethyl amino propyl) carbonyl two imine hydrochloride (EDC·HCl, 250 g, 98%) was purchased from Shanghai Da Rui Fine Chemical Co., Ltd. (Shanghai, China). Before the experiment, all operating platforms need to be cleaned with solid phase RNase-be-gone purchased from Sangon Biotech Co., Ltd. (Shanghai, China). All the water used in the experiment is diethylpyrocarbonate (DEPC) treated water.
Fabrication of microfluidic droplet chip
The microfluidic droplet chip was fabricated by traditional soft lithography technology with poly(dimethylsiloxane) (PDMS, RTV615, Momentive Corporation, USA) as described previously [11, 30, 31]. High precision mask (2400 dpi) was printed to make a SU-82100 photoresist positive mold (Microchem Corporation, USA). After cure of PDMS on the mold at 70 °C for 40 min, the PDMS layer was peeled from the mold. Then, the PDMS layer was punched to form inlet and outlet holes and bonded to a thin PDMS slide (about 150-μm thickness) by oxygen plasma (PDC-32G, Harrick Scientific Corporation, USA). At last, the chip was heated at 120 °C for 2 h. The aqueous branch channels are 150 μm in height and 100 μm in width. The main channel (oil channel) is 150 μm in height and 270 μm in width.
Fabrication of multiplexed detection module
A simple multiplexed fluorescence detection module was fabricated to match up with the microfluidic droplet chip. The detection system is consisted of a laser (MDL-III, 405 nm, 10 mW, Changchun New Industries Optoelectronics Technology Co., Ltd., Changchun, China), four silica fibers (UVHCS, 380-μm inner diameter, Nanjing Chunhui Science and Technology Industrial Co., Ltd., Nanjing, China), and four photomultiplier tubes (H5784, Hamamatsu Photonics Co., Ltd., Japan). An orthogonal optical path mode was adopted. In addition, narrow-band filters (half wave bandwidth 8 nm, peak transmittance 50%, Huibo Optics Technology Co., Ltd., Shenyang, China) were placed between each of the optical fiber outlets and each window of the photomultiplier tubes. Fluorescent signals collected from different photomultiplier tubes were recorded and analyzed by using chromatographic signal acquisition units (CT-22, Shanghai Spectrum Peak Software Co., Ltd., Shanghai, China).
Preparation of QDs-DNA probes
Carboxyl-modified QDs were attached with amino-modified capture DNAs based on condensation reaction between carboxyl and amino group. Briefly, borate buffer solution (20 mmol/L, pH 7.4) was used to dilute the quantum dot solution, and DEPC water was used to dilute the capture DNAs. Four quantum dots (525 nm, 565 nm, 605 nm, and 650 nm) were respectively modified with capture DNA-21, capture DNA-20a, capture DNA-155, and capture DNA-221 by adding moderate EDC. The mixture was firstly incubated for 2 h at room temperature in the thermostatic oscillator, centrifuged for 5 min with 6000 rpm in the ultrafiltration tube (the interception molecular weight of 50 KD), and washed three times using the PBS buffer. At last, the four QD-DNA conjugates were dissolved in PBS buffer and kept in 4 °C dark storage.
In order to determinate the connection of QDs and DNA, the absorption spectra of QDs and QDs-DNA probes were measured firstly (see Electronic Supplementary Material (ESM) Fig. S1). And gel electrophoresis was carried out for investigating the optimal molar ratio of QD to DNA [32]. In brief, the QD solution (10 μL) and the QDs-DNA conjugate solutions (10 μL for each) with different molar ratios of QD to DNA were simultaneously tested by agarose gel electrophoresis. The agarose gel was dissolved in a dipotassium hydrogen phosphate solution (10 mmol/L) with the final concentration of 1.5%. The electric field intensity was set up at 50 V. A UV lamp was used to illuminate the gel and the gel images were taken for analysis.
MiRNA hybridization
The QD-DNA conjugates (10 μmol/L, 10 μL) were mixed with BHQ-DNA (10 μmol/L, 10 μL) and target miRNA standard solutions (10 μL, a set of gradient concentrations from 0.1 to 1 μmol/L), and the mixture was stirred at 25 °C for 1.5 h in 30 μL PBS (pH 7.4, containing 20 mmol/L NaCl). In order to improve the hybridization efficiency, the experimental conditions, such as the molar ratio of reactants, hybridization time, and hybridization temperature, were investigated. The hybridization process for four target miRNAs were tried to use uniform conditions for simplicity of operation.
Process of multiple detection of target miRNAs
The four dispersed phase solutions (hybridization mixtures) and continuous phase liquid (mineral oil containing 4% Span 80) were injected into the chip by micro-injection pumps (PHD2000, Harvard, USA) with the flow of 0.5 μL/min and 4 μL/min. The dispersed phase solutions flowed along four branch channels and met mineral oil solution at the T-junctions. Then, the four dispersed phase solutions were separately sheared into droplets. Finally, the droplets were pushed into the downstream of the main channel and flowed through detection zone in sequence. The multi-color QDs encapsulated in different droplets were excited by the laser and emitted different fluorescent signals. The fluorescent signals were respectively collected by four photomultiplier tubes and recorded by the chromatographic signal acquisition unit. Fluorescence quenching efficiency is used to indicate the degree of fluorescence attenuation. The expression for the fluorescence quenching efficiency is
where F is the fluorescence intensity with target miRNA and F0 is the fluorescence intensity without target miRNA.
Results and discussion
Design of microfluidic droplet-based multiplex detection system
We aim to develop a simple, rapid, and low-consumption multiplex miRNAs detection system. The microfluidic multi-color droplet chip integrated with a multiplexed fluorescence detection module was designed (Fig. 1). Firstly, a simple droplet chip with one oil channel and four aqueous branch channels were designed (Fig. 1a). Four T-junctions were fabricated on the upstream of the oil channel one after another to achieve simultaneous formation of four kinds of aqueous droplets. Secondly, the multiplexed fluorescence detection module was designed to monitor the fluorescence signals of droplets in real time. To assemble an effective and simple multiplex fluorescence detection module, orthogonal optical path mode and finite optical elements were chosen in this work. As the volume of droplets is nano-liter scale, the light spot of the laser can cover the zone containing more than four droplets in line (Fig. 1b). So the laser can excite multiple droplets at the same time in this design, avoiding the use of multiple excitation light sources in conventional fluorescence detection systems, which simplified the complexity of light path to a half. Thirdly, silica fibers were chosen to collect the fluorescence signals of droplets based on the matching between their cross section and the droplet size. All designs mentioned above simplified the multiplex fluorescence detection module to a great degree (Fig. 1c). For realizing the multiplexed fluorescence detection based on the microfluidic multi-color droplet chip integrated with a multiplexed fluorescence detection module, four QDs in four colors were chosen in this work. The aqueous solutions of the four QDs were introduced into four branch channels, separately sheared into droplets in the four T-junctions, excited and detected by the multiplex fluorescence detection module (Fig. 1d and e). The average volume of the droplets was 10 nL, and the frequency of droplet generation was 320 droplets per minute.
Schematic illustration of microfluidic multi-color droplet chip integrated with a multiplexed fluorescence detection module. a Overall schematic illustration of the device. b Schematic illustration of detection zone. c Diagram of signals collected from the four detection paths. d Image of four T-junctions on the chip. e Image of droplets generated in the main channel. Scale bar, 500 μm
Mechanism of multiplex miRNAs detection
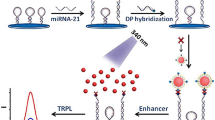
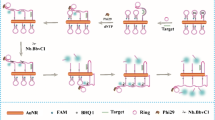
The QDs-DNA conjugate has the excellent optical properties of QDs and the good recognition properties of DNA, and the fluorescence intensity of QDs can be quenched by organic quencher Black Hole Quencher (BHQ) via fluorescence resonance energy transfer (FRET) [32]. We tried to extend the single FRET mode into a multiplex FRET mode, as shown in Fig. 2. To avoid the interference among the four detection paths, the multi-color QDs without obvious spectral overlap was chosen as the fluorescence labels in the detection system. Briefly, carboxyl-modified QDs were respectively attached with four amino-modified DNAs to form QDs-DNA conjugates. The four capture DNAs respectively were partially complementary strands of four target miRNAs strands. Meantime, four BHQ-modified DNAs were complementary strands of the other parts of the four target miRNAs strands. When the target miRNA is absent, the capture DNA strand cannot hybridize with BHQ-DNA strand. It means that the quenching agent BHQ is far away from QDs, and FRET cannot occur. Once the target miRNA is present in the test environment, the capture DNA strand, BHQ-DNA strand, and target miRNA will form a sandwich structure based on the hybridization reaction. The structure leads to the appropriate distance between BHQ and QDs, which triggers the fluorescence quenching of QDs by BHQ. As the absorption spectra of BHQs are completely covered by the emission spectra of the four capture QDs, obvious spectral overlap ensures sensitivity of the four quenching systems to their target miRNAs. As a consequence, four miRNAs can be quantitatively determined at the same time without interference.
Schematic diagram of the designed multiplex miRNAs detection system based on the multiplex FRET mode. a Quenching systems for four target miRNAs. b The absorption of BHQ-DNA (green, orange, and blue lines), the fluorescence emission spectra of QDs-DNA probes (red line), and QDs-BHQ-DNA complex (black line)
The experiment results also demonstrate the feasibility of multiplex miRNAs detection based on the multiplex FRET mode. According to the absorption and emission spectra of BHQ-DNA and QDs-DNA probes in Fig. 2b, the emission spectra of the four quantum dots show a maximum wavelength at 525 nm, 565 nm, 605 nm, and 650 nm. Furthermore, four quantum dots have broad absorption spectra and long fluorescence life, which provides great convenience to simplify the multiple excitation sources to a single laser. The absorption spectra of BHQ-DNA strands (modified with BHQ1, BHQ2, and BHQ3) were also tested and showed good spectral overlap with the emission spectra of four capture QDs-DNA conjugates, which greatly supports the occurrence of the multiplex FRET. In our experiments, it was observed (Fig. 2b) that the fluorescence intensity was significantly reduced after the addition of the target miRNA (curve named as QDs-BHQ-DNA). It meant that QDs-DNA (the capture DNA strand), BHQ-DNA strand, and target miRNA formed a sandwich structure (QDs-BHQ-DNA complex) based on the hybridization reaction and led to the appropriate distance between BHQ and QDs, which triggers the fluorescence quenching of QDs by BHQ. Therefore, the feasibility of the experiment was verified.
Characterization of the formation of QDs-DNA probes
In order to demonstrate the formation of QDs-DNA probes, the absorption spectra of four QDs solutions and QDs-DNA conjugates (ESM Fig. S1) were measured. Compared with the absorption spectra of QDs solutions, obvious absorption peaks near the wavelength of 260 nm (characteristic peak of DNA) occur in the absorption spectra of QDs-DNA conjugates solutions, which demonstrate the formation of QDs-DNA probes. When the electrophoresis movement speed of QDs-DNA conjugates is placid, the DNA loaded on the quantum dot surface tends to be saturated [32]. According to the results of gel electrophoresis in our work, QDs-DNA conjugates with the molar ratio of 1:30 were chosen as the optimal ratio of QDs to DNA. Consequently, the QDs-DNA conjugates with ratio of 1:30 were used in subsequent experiments. In addition, the fluorescence spectra of QDs modified with capture DNAs (QDs-DNA probes) were compared with the mixture of QDs-DNA probes and DNA-BHQs probes. Figure 3 shows that the fluorescence intensity of four QDs were reduced by 2.4%, 3.7%, 3.5%, and 2.1%, respectively. It meant that the quencher BHQ could hardly quench the fluorescence of QDs when QDS-DNA is mixed with BHQ-DNA without target miRNA. This phenomenon indicates that fluorescence resonance energy transfer can only occur in the presence of target miRNA.
Multiplex detection of four target miRNAs
During the process of detection, the hybridization mixtures of four target miRNAs were introduced into the four branch channels and sheared into droplets by the oil phase solution at the four T-junctions. When the droplets passed the detection zone in order, the fluorescence signals of the four QDs-DNA conjugates from different droplets were nearly simultaneously collected by different PMTs with the help of fibers and filters. And the fluorescence signals of each QDs-DNA conjugate were recorded respectively. As the fluorescence of QDs-DNA conjugates in droplets was quenched by BHQs, the fluorescence signal values of QDs-DNA conjugates were reduced with the increase of the concentrations of target miRNAs in droplets (Fig. 4). The signal detected by the device is stable, and the RSD value of fluorescence signal intensities is 1.41% for the continuous measurement of 65 droplets in 50s for the same target (see ESM Fig. S2).
In the concentration range (0.1 nmol/L to 1 μmol/L), the fluorescence quenching efficiency of QDs shows the linear relationship with the concentration of target miRNAs, as shown in Fig. 5, and their correlation coefficients all reach 0.99. It indicates that the multiplex detection system is effective for simultaneous detection of four miRNAs in the concentration range from 0.1 nmol/L to 1 μmol/L with low detection limits (36.0 pmol/L for miRNA-21, 35.1 pmol/L for miRNA-20a, 36.5 pmol/L for miRNA-155, and 38.3 pmol/L for miRNA-221). The detection speed is equal to the generation frequency of droplets, which is about 320 droplets per minute.
Specificity of multiplex detection
The specificity of multiplex detection of target miRNAs was also investigated in this work. In the miRNA-21 test, the QD525-DNA-21 conjugate (1 μmol/L) was mixed with the BHQ-DNA-21 probe (1 μmol/L), and then respectively incubated with the non-cognate RNA (500 nmol/L) and three other target miRNAs (500 nmol/L) at 25 °C for 1.5 h in PBS at pH 7.4 (containing 20 mmol/L NaCl). The interference effect was investigated based on the comparison of their quenching signals. The quenching efficiency of miRNA-21 was obviously higher than those of miRNA-20a, miRNA-155, miRNA-221, and non-cognate RNA. In theory, the target miRNA-21 can specially hybridize with the QD-525-DNA-21 probes and BHQ-DNA-21 probes based on complementary pairing principle, but the other three of the four target miRNAs and the non-complementary single strand miRNA (non-cognate RNA) cannot hybridize with the probes for miRNA-21, which is consistent with the experiment result in our work. So the specificity test demonstrates that the interference of other miRNAs to miRNA-21 can be ignored in this work. In addition, similar phenomena were observed in the specificity tests of the other three target miRNAs. The data (Fig. 6) shows that there is no obvious cross interference in our multiple miRNA detection. It is demonstrated that the multiplex detection system can efficiently differentiate target miRNAs from other miRNAs benefiting from the multiplex FRET mode based on microfluidic multi-color droplets.
Detection of four target miRNAs in human serum
To test the feasibility of the practical application of the droplet-based multiplex detection system, we carried out the multiplex detection of four miRNAs mentioned above in healthy human serum (obtained from Northeastern University Hospital, China). Four target miRNAs (miRNA-21, miRNA-20a, miRNA-155, and miRNA-221) were not detected in 20-fold dilution of human serum samples. The standard solutions of four target miRNAs (miRNA-21, miRNA-20a, miRNA-155, and miRNA-221) with different concentrations were spiked into the diluted (20-fold) human serum samples to measure the recoveries. With the addition of target miRNAs, the fluorescent intensities of droplets from a set of serum samples spiked with different concentration targets (concentrations of 1.00 nmol/L and 100.0 nmol/L) were reduced correspondingly. As shown in Table 2, the recoveries for the spiked miRNA-21, miRNA-20a, miRNA-155, and miRNA-221 are respectively 101% and 102%, 88.1% and 102%, 94.1% and 98.6%, and 97.0% and 95.8% (n = 3). The relative standard deviations (RSDs) are from 2.1 to 12.2% for the four target miRNAs. These results indicate that the multiplex miRNA detection can be realized by using the microfluidic multi-color droplet chip integrated with a multiplexed fluorescence detection module, and the simple device has great potential for rapid and low-consumption detection of multiple miRNAs in real samples without nucleic acid amplification.
Conclusions
We developed a microfluidic multi-color droplet-based fluorescence detection system for the multiplex detection of miRNAs based on the effect of FRET between multi-color QDs and BHQ quenchers. The system focuses on the development of a small multi-color droplet generation chip integrated with a simple multi-path fluorescence detection module for the application of multiplex detection of miRNAs related with tumor or cancer. The device can be used for simple, rapid, and sensitive multiplex detection of four miRNAs in human serum. In addition, the consumption was reduced greatly owing to the use of the nano-liter droplets (average volume 10 nL). Meanwhile, four droplets were simultaneously excited in detection zone by one single laser, which makes the multi-path fluorescence detection module simplified greatly. The detection speed was highly enhanced by the flow measurement mode based on the microfluidic droplets and integrated fluorescence detection module (320 droplets per min). The hybridization reaction between the target miRNAs and their probes ensures the specificity of multiplex detection. And the quenching system of QDs and BHQ quenchers ensures the high sensitivity of the method. Therefore, a simple and low-consumption multiplex detection method of miRNAs in human serum was developed successfully. Moreover, the microfluidic measurement system exhibits excellent sensitivity and stability of multiplex miRNAs detection, which demonstrates potential application prospect in early-stage diagnosis and prognosis of cancers.
References
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 116(2):281–97. https://doi.org/10.1016/S0092-8674(04)00045-5.
Bartel DP, Chen CZ. Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat Rev Genet. 5(5):396–400.
Chan JA, Krichevsky AM, Kosik KS. MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res. 65(14):6029–33.
Tian T, Xiao H, Zhang Z, Long Y, Peng S, Wang S, et al. Sensitive and convenient detection of microRNAs based on cascade amplification by catalytic DNAzymes. Chemistry. 19(1):92–5. https://doi.org/10.1002/chem.201203344.
Ahir BK, Ozer H, Engelhard HH, Lakka SS. MicroRNAs in glioblastoma pathogenesis and therapy: a comprehensive review. Crit Rev Oncol Hematol. 120:22–33. https://doi.org/10.1016/j.critrevonc.2017.10.003.
Qiu X, Hildebrandt N. Rapid and multiplexed microRNA diagnostic assay using quantum dot-based forster resonance energy transfer. ACS Nano. 9(8):8449–57.
Geaghan M, Cairns MJ. MicroRNA and posttranscriptional dysregulation in psychiatry. Biol Psychiatry. 78(4):231–9. https://doi.org/10.1016/j.biopsych.2014.12.009.
Small EM, Olson EN. Pervasive roles of microRNAs in cardiovascular biology. Nature. 469(7330):336–42. https://doi.org/10.1038/nature09783.
Thum T, Mayr M. Review focus on the role of microRNA in cardiovascular biology and disease. Cardiovasc Res. 93(4):543–4. https://doi.org/10.1093/cvr/cvs085.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 105(30):10513–8. https://doi.org/10.1073/pnas.0804549105.
Kroh EM, Parkin RK, Mitchell PS, Tewari M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods. 50(4):298–301. https://doi.org/10.1016/j.ymeth.2010.01.032.
Li LM, Hu ZB, Zhou ZX, Chen X, Liu FY, Zhang JF, et al. Serum microRNA profiles serve as novel biomarkers for HBV infection and diagnosis of HBV-positive hepatocarcinoma. Cancer Res. 70(23):9798–807. https://doi.org/10.1158/0008-5472.CAN-10-1001.
Mandiannasser M, Karami Z. An innovative paradigm of methods in microRNAs detection: highlighting DNAzymes, the illuminators. Biosens Bioelectron. 107:123–44. https://doi.org/10.1016/j.bios.2018.02.020.
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 65(16):7065–70. https://doi.org/10.1158/0008-5472.CAN-05-1783.
Wegman DW, Cherney LT, Yousef GM, Krylov SN. Universal drag tag for direct quantitative analysis of multiple microRNAs. Anal Chem. 85(13):6518–23. https://doi.org/10.1021/ac401185g.
Michael MZ, O'Connor SM, Pellekaan NGV, Young GP, James RJ. Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res. 1(12):882–91.
Wegman DW, Krylov SN. Direct quantitative analysis of multiple miRNAs (DQAMmiR). Angew Chem Int Ed. 50(44):10335–9. https://doi.org/10.1002/anie.201104693.
Wegman DW, Krylov SN. Direct miRNA-hybridization assays and their potential in diagnostics. Trac-Trend Anal Chem. 44:121–30. https://doi.org/10.1016/j.trac.2012.10.014.
Choi JW, Min KM, Hengoju S, Kim GJ, Chang SI, deMello AJ, et al. A droplet-based microfluidic immunosensor for high efficiency melamine analysis. Biosens Bioelectron. 80:182–6. https://doi.org/10.1016/j.bios.2015.12.023.
Duan K, Ghosh G, Lo JF. Optimizing multiplexed detections of diabetes antibodies via quantitative microfluidic droplet array. Small 13 (46). doi:https://doi.org/10.1002/smll.201702323
Ding Y, Choo J, deMello AJ. From single-molecule detection to next-generation sequencing: microfluidic droplets for high-throughput nucleic acid analysis. Microfluid Nanofluid. 21(3). https://doi.org/10.1007/s10404-017-1889-4.
Kim JJ, Chen L, Doyle PS. Microparticle parking and isolation for highly sensitive microRNA detection. Lab Chip. 17(18):3120–8. https://doi.org/10.1039/c7lc00653e.
Yuan H, Chao Y, Li S, Tang MYH, Huang Y, Che Y, et al. Picoinjection-enabled multitarget loop-mediated isothermal amplification for detection of foodborne pathogens. Anal Chem. 90(22):13173–7. https://doi.org/10.1021/acs.analchem.8b03673.
Krichevsky AM, Gabriely G. miR-21: a small multi-faceted RNA. J Cell Mol Med. 13(1):39–53. https://doi.org/10.1111/j.1582-4934.2008.00556.x.
Si ML, Zhu S, Wu H, Lu Z, Wu F, Mo YY. miR-21-mediated tumor growth. Oncogene. 26(19):2799–803. https://doi.org/10.1038/sj.onc.1210083.
Zaleska K, Przybyla A, Kulcenty K, Wichtowski M, Mackiewicz A, Suchorska W, et al. Wound fluids affect miR-21, miR-155 and miR-221 expression in breast cancer cell lines, and this effect is partially abrogated by intraoperative radiation therapy treatment. Oncol Lett. 14(4):4029–36. https://doi.org/10.3892/ol.2017.6718.
Esquela-Kerscher A, Slack FJ. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer. 6(4):259–69. https://doi.org/10.1038/nrc1840.
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 6(11):857–66. https://doi.org/10.1038/nrc1997.
Pencheva N, Tavazoie SF. Control of metastatic progression by microRNA regulatory networks. Nat Cell Biol. 15(6):546–54. https://doi.org/10.1038/ncb2769.
Beitzinger M, Meister G. Preview. MicroRNAs: from decay to decoy. Cell. 140(5):612–4. https://doi.org/10.1016/j.cell.2010.02.020.
Fu Z, Qian F, Yang X, Jiang H, Chen Y, Liu S. Circulating miR-222 in plasma and its potential diagnostic and prognostic value in gastric cancer. Med Oncol. 31(9):164. https://doi.org/10.1007/s12032-014-0164-8.
Su S, Fan J, Xue B, Yuwen L, Liu X, Pan D, et al. DNA-conjugated quantum dot nanoprobe for high-sensitivity fluorescent detection of DNA and micro-RNA. ACS Appl Mater Interfaces. 6(2):1152–7. https://doi.org/10.1021/am404811j.
Weishaupt SU, Rupp S, Lemuth K. Simultaneous detection of different microRNA types using the ZIP-code array system. J Nucleic Acids. 2013;2013:496425. https://doi.org/10.1155/2013/496425.
Li D, Wang Y, Lau C, Lu J. xMAP array microspheres based stem-loop structured probes as conformational switches for multiplexing detection of miRNAs. Anal Chem. 2014;86(20):10148–56. https://doi.org/10.1021/ac501989b.
Jiang L, Shen Y, Zheng K, Li J. Rapid and multiplex microRNA detection on graphically encoded silica suspension array. Biosens Bioelectron. 2014;61:222–6. https://doi.org/10.1016/j.bios.2014.05.020.
Liu Y, Zhang J, Tian J, Fan X, Geng H, Cheng Y. Multiplex detection of microRNAs by combining molecular beacon probes with T7 exonuclease-assisted cyclic amplification reaction. Anal Bioanal Chem. 2017;409(1):107–14. https://doi.org/10.1007/s00216-016-0027-6.
Azzouzi S, Fredj Z, Turner APF, Ali MB, Mak WC. Generic neutravidin biosensor for simultaneous multiplex detection of microRNAs via electrochemically encoded responsive nanolabels. ACS Sensors. 2019;4(2):326–34. https://doi.org/10.1021/acssensors.8b00942.
Funding
This study received financial support from the National Natural Science Foundation of China (21874015 and 21675020).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest, and this manuscript is approved by all authors for publication.
Ethical approval
This study was approved by the Ethics Review Committee of Northeastern University Hospital (Shenyang, China). Informed consent of all volunteers involved in serum experiments.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 459 kb)
Rights and permissions
About this article
Cite this article
Ye, WQ., Wei, YX., Zhang, YZ. et al. Multiplexed detection of micro-RNAs based on microfluidic multi-color fluorescence droplets. Anal Bioanal Chem 412, 647–655 (2020). https://doi.org/10.1007/s00216-019-02266-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-02266-3