Abstract
Colorectal cancer (CRC) is one of the most common causes of cancer-related death worldwide. Emerging evidence has shown that lipid metabolism plays important roles in the occurrence and progression of CRC. The identification of potential biomarkers for CRC progression is critical for precise diagnosis and treatment. Therefore, the aim of this study is to explore the potential lipid markers in relation to CRC progression. The plasma of patients with stage I/II CRC (n = 20) and stage III/IV CRC (n = 20) was collected. Lipidomic screening was performed by ultrahigh-performance liquid chromatography–mass spectrometry. After multivariate data analysis, including orthogonal partial least squares discriminant analysis, determination of the fold change, and the Mann–Whitney U test, eight lipid species with altered levels with p < 0.05 and fold change greater than 2 were selected as potential lipid biomarkers. Compared with patients with early-stage CRC, patients with advanced-stage CRC showed significantly higher levels of cholesteryl ester (20:4) and some triglycerides with a saturated fatty acid chain and a lower level of fatty acid ester of hydroxy fatty acid 27:1 (9:0-18:1) in plasma. Furthermore, the receiver operating characteristic including these potential lipid biomarkers yielded a sensitivity of 85% and specificity of 80% for separation of early-stage CRC patients from advanced-stage CRC patients. In all, this is the first report showing that the levels of triglycerides, the major contents of lipid droplets, increase in plasma of advanced-stage CRC patients compared with early-stage CRC patients. These data indicate that lipid droplets may be target organelles for the study of CRC progression and treatment.

Graphical abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The global incidence of colorectal cancer (CRC) ranks third, but mortality ranks second, according to the global cancer statistics for 2018 [1]. Because of economic development, dietary pattern changes, and lifestyle changes, Chinese incidence rates of CRC, especially for men, morbidity (annual average change of 1.6 percentage points ), and mortality (annual average change of 1.3 percentage points) have increased in the last decade [1, 2]. It takes 10–15 years for a small polyp to grow into CRC. For stage I/II CRC patients, it is necessary to remove it directly. For stage III/IV CRC patients, the treatment procedures are complex, and neoadjuvant chemotherapy is probably necessary. Besides, although some patients are in the early stage of CRC, the metastasis-related symptoms are not obvious. Currently, CRC diagnosis and monitoring are based mainly on colonoscopy, an immunochemical-based fecal occult blood test, a carcinoembryonic antigen test, and other techniques. However, these methods are expensive, lack sensitivity and specificity, or are invasive [3, 4]. Misdiagnosis will lead to undertreatment or overtreatment of CRC patients, thereby resulting in increased mortality. The latest data indicate that the survival rate of CRC patients drastically declines with disease progression, from almost 90% for stage I/II CRC to 13% for stage IV CRC [5]. Therefore, a convenient and effective method is needed to improve the classification accuracy of the stages for CRC diagnosis and treatment.
Lipids have several functions, especially as mediators, which can participate in signal transduction pathways, apoptosis, proliferation, and membrane trafficking in cells [6]. Adipocytes are associated with CRC tumor invasion. Co-culture of colon cancer cells with adipocytes led to the release of free fatty acids (FFAs), causing the growth of cancer cells [7]. The products of lipid oxidation stress, including cholesterol and fatty acids, can produce inflammatory factors that will promote CRC [8]. By inhibiting the expression of genes that encode lipogenesis regulatory enzymes, the proliferation, migration, and invasion characteristics of colon cancer will be repressed [9]
Omics-based methods have exhibited great potential for biomarker discovery, especially in cancer metabolism. Lipidomics, as a branch of metabolomics, reflects the profiling of different kinds of lipids and their related networks. Although there have been several lipidomics studies in CRC, almost all of them attempted to diagnose CRC. Lysophosphatidylcholines (LPCs) and phosphatidylcholines (PCs) are the most explored biomarkers related to the occurrence of CRC [10,11,12]. With regard to CRC staging, Li et al [13] applied direct-infusion electrospray ionization (ESI) Fourier transform ion cyclotron resonance mass spectrometry (MS) to analyze the lipid profile of CRC patients and healthy controls. It was found that some fatty acids were downregulated and some phospholipids were upregulated in plasma of stage III/IV CRC patients compared with stage I/II CRC parents. Another study used a gas chromatography–MS-based method to analyze the lipid levels in plasma of healthy individuals, stage I/II CRC patients, and stage III/IV CRC patients [14]. The results indicated that C6–C12 fatty acids could be deemed as biomarkers to distinguish early-stage CRC patients and advanced-stage CRC patients. Since researchers use one or several lipid standards with similar structure to quantify the corresponding class of lipids, the approach of combining high-performance liquid chromatography (HPLC) with MS has become the mainstream method in lipid analysis [15, 16].
The present study aims to develop a nontargeted ultrahigh-performance liquid chromatography (UHPLC)–MS-based lipidomics approach to explore the significant changed lipid metabolites for differentiating early-stage CRC patients from advanced-stage CRC patients. Orthogonal partial least squares discriminant analysis (OPLS-DA), fold change, and the Mann–Whitney U test were used for data analysis. Ultimately, some potential lipid biomarkers were identified that may provide a new approach for further study of the molecular mechanism of CRC progression.
Materials and methods
Chemicals
HPLC-grade methanol, dichloromethane, and 2-propanol were purchased from Merck (Germany). Ultrapure water was prepared by a Milli-Q system (Millipore, Billerica, MA, USA). The SPLASH LIPIDOMIX MS standard (catalog no. 330707) was purchased from Avanti Polar Lipids (Alabaster, AL, USA). Detailed information is presented in Table S1.
Participants and sample collection
The study was approved by the Ethics Committee of the Institution of Clinical Pharmacology, Central South University (registration no. CTXY-150001-2), and was performed as part of a previous clinical trial, which focused on the circulating tumor DNA of CRC patients and healthy people for disease diagnosis and prognosis. (registration no. ChiCTR-DCD-15006289; http://www.chictr.org.cn/showprojen.aspx?proj=10851). Participants provided written informed consent. Plasma samples were collected from CRC patients at Xiangya Hospital from April 2017 to April 2018. Forty patients were selected according to the following criteria: (1) CRC confirmed by pathological examination; (2) no other diseases, such as cancers, hyperlipidemia, and diabetes; (3) no previous operation, chemotherapy, or radiotherapy; (4) availability of complete clinical records; (5) detailed pathological diagnosis of staging according to the TNM clinical staging system. The plasma was collected before surgery after a 10-h fasting period. Venous blood specimens were collected in tubes that contained the anticoagulant EDTA and were centrifuged at 4000g for 10 min at 4 °C. The supernatant was transferred into an Eppendorf tube and stored at -80 °C until analysis.
Sample preparation
For lipid extraction, a modified Bligh and Dyer method was used [17]. Briefly, 40 μL plasma was mixed with 960 μL water and then placed on ice for 10 min. Then 2 mL CH3OH and 900 μL CH2Cl2 were added. After vortex mixing, 10 μL of internal standard was added. This was followed by vortex mixing and placement on ice for 30 min. To achieve phase separation, 1 mL pure water and 900 μL CH2Cl2 were added. After the tube had been inverted ten times, the mixture was centrifuged at 290g for 10 min. Then 800 μL of the lower layer was transferred into a fresh glass. To complete extraction, 2 mL CH2Cl2 was mixed with the residual from the extraction tube and centrifuged, and 2 mL of the lower layer was added to the first extract. A total of 2.8 mL of lipid extract was evaporated under a stream of nitrogen. Finally, the dried extract was reconstituted with 100 μL CH3OH–CH2Cl2 (1:1 v/v), followed by centrifugation at 17,949g for 5 min at 4 °C. The supernatant was used for injection into the UHPLC–MS system. Furthermore, to ensure good data quality for lipidomics profiling, 10 μL plasma from every sample was mixed and processed as described previously to prepare the quality control (QC) samples. To validate the repeatability of the method and the stability of the UHPLC–MS system, the QC samples were inserted six times before the batch and then inserted after every ten injections.
UHPLC–MS analysis
UHPLC–MS was performed with an ExionLC AD instrument (AB SCIEX, USA) coupled to a time-of-flight (TOF) mass spectrometer (Triple TOF 5600, AB SCIEX, USA). A Kinetex C18 column (2.6 μm, 2.1-mm inner diameter, 100 mm-length; Phenomenex, USA) was used for lipid separation. Mobile phases A and B were water–methanol–acetonitrile (1:1:1 v/v/v) and 2-propanol, respectively. Both mobile phases contained 5 mM ammonium acetate. The gradient conditions were as follows: 0.5–1.5 min, linear gradient from 20% mobile phase B to 40% mobile phase B; 1.5–3.0 min, 40% mobile phase B to 60% mobile phase B; 3.0–13.0 min, 60% mobile phase B to 98% mobile phase B; 13.0–13.1 min, 98% mobile phase B to 20% mobile phase B and held for 3.9 min at 20% mobile phase B. The flow rate was 0.3 mL/min. The temperatures of the column and the autosampler were set at 45 and 4 °C, respectively. The tandem MS (MS/MS) and high-resolution MS data for injection of 1 μL of sample for UHPLC–MS analysis were acquired by time-of-flight MS full scan and information-dependent acquisition ranging from 200 to 1300 m/z. Each sample was injected in positive ESI and negative ESI mode. The mass spectrometer was operated with the ion source gas 1 and gas 2 of both 60 psi. The current gas was set at 35 psi, the temperature was set at 600 °C. IonSpray voltage floating was set at 5500 V in positive mode and -4500 V in negative mode. Declustering potential was set at 50 V. Collision energy was set at 45 V.
Data processing and statistical analyses
For data acquisition and processing, Analyst TF (version 1.7.1, AB SCIEX, USA) was used. Raw data files were exported to MS-DIAL (version 2.9.4) for lipid extraction and identification [18]. Then MultiQuant (version 3.0, AB SCIEX, USA) was used for further data processing. Some parameters set in MultiQuant were as follows: Gaussian smoothing width, 1.0 points; RT (retention time) half windows, 10.0 s; minimum peak width, 5 points; minimum peak height, 500; noise percentage, 90%; baseline substraction, window, 0.1 min; peak splitting, 2 points. Finally, the peak area of each metabolite was normalized by the internal standard of the corresponding category. After removal of peaks with a coefficient of variation greater than 30% among QC samples and missing value (the peak intensity was zero ) greater than 20% among all samples, the remaining peaks were included for further analysis. However, there is no FFA species in the mixed lipid standard, and thus lysophosphatidylethanolamine (LPE), which has a similar retention time, was used as a substitute standard for FFAs.
The statistical results were input into SIMCA-P version 14.1 for OPLS-DA with mean centering and unit square root of variance scaling (pareto scaled). The quality of the OPLS-DA model was evaluated by the parameters R2 (explained parameter) and Q2 (predictive parameter). Additionally, the model was further validated by a 200 random permutation test. The Mann–Whitney U test (SPSS Statistics version 17.0, SPSS, Chicago, IL, USA) was performed for stage I/II CRC patients and stage III/IV CRC patients. The trend (fold change) of upregulation or downregulation showed the variation of the differential metabolites between patients with early-stage CRC and patients with advanced-stage CRC. Only the metabolites with p < 0.05 and fold change of the peak area greater than 2 were defined as potential biomarkers. Finally, the receiver operating characteristic was used to predicate the accuracy of the potential biomarkers.
Results
Study participant characteristics
Forty participants were enrolled, including 20 stage I/II CRC patients and 20 stage III/IV CRC patients. CRC stage was diagnosed and graded by pathological examination and TNM level. Detailed clinical data are provided in Table 1. Age and triglyceride (TG) plasma level were matched by the Mann Whitney U test. Sex, degrees of differentiation, and tumor location were matched by the chi-square test (SPSS Statistics version 17.0, SPSS, Chicago, IL, USA) with p > 0.05.
Analysis and identification of lipids
The original files were imported into MS-DIAL to identify the lipids by our referring to the accurate m/z, retention time, and branched-chain fragments of metabolites. Then the peak area of each metabolite was obtained by our importing the file into MultiQuant with the retention time and m/z limited. After normalization by the internal standard and removal of the abnormal peaks, 250 lipids were identified in positive mode and 162 were identified in negative mode. The detected lipid species included PC, phosphatidic acid, sphingomyelin (SM), LPC, phosphatidylethanolamine (PE), LPE, FFA, fatty acid ester of hydroxy fatty acid (FAHFA), phosphatidylinositol (PI), oxidized PI, oxidized PC, cholesteryl ester (CE), and oxidized PE (Table S2). The total ion chromatograms of plasma lipid extracts are presented in Fig. S1.
The lipid identification was based on the accurate m/z, retention time, and characteristic fragment ions through use of MS-DIAL and the LIPID MAPS database (http://www.lipidmaps.org/). The mass accuracy was well below 5 ppm for all included lipid species. For almost all lipid classes identified, the positive and negative MS/MS spectra provided specific fragments. At a retention time of 5.273 min, the LIPID MAPS search results suggest m/z 740.5202 likely corresponded to [M + H]+ of PE (16:0-20:4) in positive mode (Fig. 1a), with an acceptable error of -3.11 ppm, and m/z 599.5014 likely corresponded to the fragment ion of neutral loss of ethanolamine phosphate. In negative mode, the precursor ion with m/z 738.5111 at a retention time of 5.28 min with 4.29-ppm error was likely [M − H]- of PE (16:0-20:4) (Fig. 1b). In addition, it can be inferred that m/z 303.2331 and m/z 255.2333 corresponded to the fragment ions of 20:4-carboxylate anion and 16:0-carboxylate anion, respectively. Structural information on this lipid, including the number of carbon atoms and double bonds per fatty acyl chain, suggested it was PE (16:0-20:4). However, the positions of the double bonds marked in the dashed boxes in Fig. 1a, b were still uncertain. Figure 1c shows the MS/MS spectrum of TAG (16:0-18:0-16:0). The precursor ion with m/z 852.8017 at a retention time of 10.763 min with 0.23-ppm error was probably the quasimolecular ion [M + NH4]+ according to LIPID MAPS. In addition, it can be inferred that the fragment ions with m/z 579.5340 and m/z 551.5030 corresponded to the loss of palmitic acid (256 Da) and stearic acid (284Da), respectively. It was deduced that the peak at m/z 852.8017 at a retention time of 10.763 min corresponded to TAG (16:0-18:0-16:0).
Evaluation of the method
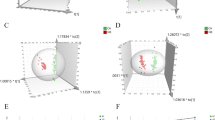
As Fig. 2 shows, the principal component analysis (PCA) model and the control chart of 40 samples, including 11 QC, samples were built. The 11 QC samples were tightly clustered and generally separated from other samples in the score plot (Fig. 2b). The QC samples were almost stable throughout the analysis in the control chart (Fig. 2c), which indicated that the differences in the test samples were due to biological variation [19, 20]. In addition, the relative standard deviations for all metabolites in the QC samples were mostly within 0.15 (Fig. 2d). The data processing results indicated that the method of lipid profiling was stable and reproducible.
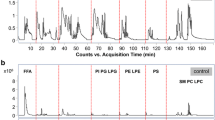
a The main species distribution of lipid species in plasma by nontargeted lipidomics. b PCA score plot of the distribution of quality control (QC) and other samples. c Control chart for the distribution of QC and other samples. The red circles are QC samples and the blue circles are the other samples. The x-axis is the sample run order and the y-axis is the first principal component t[1] with 2σ and 3σ limits. d The coefficient of variation (CV) distribution of metabolites around QC samples. CE cholesteryl ester, DAG diacylglycerol, LPC lysophosphatidylcholine, LPE lysophosphatidylethanolamine, PA phosphatidic acid, PC phosphatidylcholine, PE phosphatidylethanolamine, PA phosphatidic acid, PI phosphatidylinositol, SM sphingomyelin, TAG triacylglycerol.
Discovery of potential lipid biomarkers
Lipid species of PC, LPC, PE, LPE, phosphatidic acid, sphingomyelin, FFA, PI, FAHFA, oxidized PE, oxidized PC, and oxidized PI in negative mode and TAG, diacylglycerol, and CE in positive mode were found (Table S2, Fig. 2a). After normalization with internal standards, each relative peak area of residual metabolites was imported into SIMCA-P version 14.1 to perform the OPLS-DA model with unit square root of variance scaling (par) transformed. The score plot of the OPLS-DA model displayed a distinct separation trend between stage I/II CRC patients and stage III/IV CRC patients (Fig. 3a). As expected, the stage I/II CRC group and the stage III/IV CRC group were completely separated in the direction of the [t1] component with the explained parameters R2X = 0.326 and R2Y = 0.693 and the predictive parameter Q2 = 0.452, respectively. For avoiding overfitting of the OPLS-DA model, 200 permutations tests were provided with the Q2-intercept of −0.417 (Fig. 3b). It indicated that the model was not overfitting the data. In addition, the Mann–Whitney U test was performed to validate the significance of the lipids by the fold change. Metabolites with fold change greater than 2 and p<0.05 were selected as potential biomarkers (Fig. 3c). Finally, eight lipid species, including CE (20:4), FAHFA 27:1 (9:0-18:1), and some TAGs with a saturated fatty acid chain, were included (Table 2). To optimize the diagnosis model, the receiver operating characteristic curve of these significant metabolites was obtained. The area under the curve of 89.8%, with 85% sensitivity and 80% specificity, demonstrated the reliability of the model (Fig. 3d).
a Orthogonal partial least square discriminant analysis (OPLS-DA) model for discrimination between stage I/II colorectal cancer patients and stage III/IV colorectal cancer patients [R2X (cum) = 0.326, R2Y (cum) = 0.693, Q2 (cum) = 0.452]. b Permutation test of the OPLS-DA model (n = 200; Q2 intercept -0.417). c Volcano plot of lipid metabolites in plasma by nontargeted lipidomics. d Prediction through the combination of eight lipid biomarkers, including cholesteryl ester (CE) (20:4), fatty acid ester of hydroxy fatty acid (FAHFA) 27:1 (9:0-18:1), triglyceride (TAG) 40:0 (12:0-12:0-16:0), TAG 42:0 (12:0-14:0-16:0), TAG 44:0 (14:0-14:0-16:0), TAG 46:0 (14:0-16:0-16:0), TAG 48:0 (16:0-16:0-16:0), and TAG 54:0 (16:0-18:0-20:0) (area under the curve 89.8%, sensitivity 85%, specificity 80%). FC (fold chang of stage III/IV to stage I/II)
Discussion
This study analyzed the plasma lipid profile of CRC patients with early and advanced stage CRC by UHPLC–TOF MS in positive ESI mode and negative ESI mode. After an efficient and robust analysis, eight lipid species, including CE (20:4), FAHFA 27:1 (9:0-18:1), and some TAGs with a saturated fatty acid chain, were identified for differentiation of disease states.
A number of lipidomics-based studies have provided evidence of lipid metabolites in plasma and tissue associated with the presence of CRC. However, few studies have focused on the progression of CRC. LPCs are the most explored biomarkers for the occurrence of CRC [21, 22]. Changes in Phospholipid levels in tissue was related to CRC progression [23]. This is the first time that TAGs have been shown to be the major altered lipids in the different states of CRC. A high level of TAGs in human plasma has been reported to be associated with cardiovascular disease. Changes in TAG levels was also found to be associated with some cancers, such as non-small-cell lung cancer [24] and hepatocellular carcinoma [25, 26]. It is noteworthy that the TAGs with saturated fatty acids showed significant differences between early-stage CRC patients and advanced-stage CRC patients. However, our clinical data showed that the total TAG content in the plasma of patients did not differ between the two groups.
TAGs exist mainly in lipid droplets, which are fat-storage organelles regulating the production, trafficking, cycling, and signaling of lipids (Fig. 4). Recent studies found that the alteration of lipid droplets was related to the life activity of cancer cells [27, 28]. Some genes that play a significant role in the production of lipid droplets were demonstrated to be related to cancer cell presence and cancer progression. As shown in the pathway diagram in Fig. 4, the diacylglycerol acyltransferase (DGAT) gene family is the main producer of TAG. A study demonstrated that positive regulation of DGAT1 can promote prostate cancer cell growth [29]. In addition, adipose triglyceride lipase (ATGL), the rate-limiting enzyme of TAG hydrolysis, was studied with regard to tumor-suppressor function in hepatocellular carcinoma [26]. When ATGL was inhibited, the fatty acids in lipid droplets were sequestered from mitochondrial pathways for reactive oxygen species generation, establishing an anaerobic environment for cancer cell survival [30]. However, in a study of breast cancer, the overexpression of ATGL was positively correlated with cancer cell migration and invasion [31]. Additionally, in CRC cells, knockdown of ATGL might impede the proliferation and invasion of cells [32]. These studies suggested that there are probably diverse mechanisms of lipid metabolism among different cancers. However, there is no doubt that the alteration of lipid droplets, especially TAGs, plays a vital role in cancer presence and progression.
The metabolic pathway of lipid species related to lipid droplets. ACAT acylcoenzyme A:cholesterol acyltransferase, ATGL adipose triglyceride lipase, CE cholesteryl ester, DAG diacylglycerol, DGAT diacylglycerol acyltransferase, FA, fatty acid, FA-COA acetoacetylCoA, HSL hormone-sensitive lipase, LPA lysophosphatidic acid, MAG monoacylglycerol, MGL monoacylglycerol lipase, PA phosphatidic acid, PL phospholipids , TAG triglyceride, TCA tricarboxylic acid
The acylcoenzyme A:cholesterol acyltransferase (ACAT) gene family was associated with CE synthesis in ester droplets, with ACAT1 being found to be upregulated in CRC tissues and cells [33]. Besides, increase of the severity of the disease with higher gene expression in the tissues of patients with prostate cancer has been proved [34].
FAHFA is a lipid species produced through the esterification of long-chain fatty acids with hydroxylated fatty acids. FAHFA is associated with antidiabetic and anti-inflammatory effects in mammals [35]. In the plasma of breast cancer patients, the level of FAHFA was significantly lower than that of normal people [36], which was in accordance with our study in CRC. This likely indicates that the low FAHFA level in plasma of patients with advanced-stage CRC enhanced the proinflammatory effect of the body.
In general, the identified circulating lipid biomarkers can be used to increase sensitivity and convenience in the assessment of the status of patients for precise treatment. Monitoring of the concentration of these biomarkers in plasma would probably be helpful in preventing cancer progression in a specific way in the clinic. In addition, this study proposed that lipid species related to lipid droplets play an important role in the occurrence and progression of CRC. Lipid droplets and the related genes are expected to be targets for the treatment of CRC. Moreover, the cellular mechanisms of lipid droplets in CRC progression need to be studied further.
Conclusion
In this study, we investigated the lipid profile in human plasma of early-stage CRC patients and advanced-stage CRC patients using a UHPLC–TOF MS system. CE (20:4), FAHFA 27:1 (9:0-18:1), and TAGs with a saturated fatty acid chain were significantly altered in advanced-stage CRC patients compared with early-stage CRC patients. This study demonstrated for the first time that TAGs as the main component of lipid droplets can be used to classify CRC stage. It also showed that lipids possess great potential as biomarkers for disease progression. Although the sample size was small, we believe that the approach will be a powerful tool for the discovery of potential lipid biomarkers and classification of cancer stages. Still, further studies, such as studies of the genes that affect the expression of lipid droplets and the lipid profile of tumor tissues or cells, are needed. This study suggested that lipid droplets may be target organelles for exploring the molecular mechanism in connection with CRC progression for precise diagnosis and treatment.
Abbreviations
- ACAT:
-
Acylcoenzyme A:cholesterol acyltransferase
- ATGL:
-
Adipose triglyceride lipase
- CE:
-
Cholesteryl ester
- CRC:
-
Colorectal cancer
- DGAT:
-
Diacylglycerol acyltransferase
- ESI:
-
Electrospray ionization
- FAHFA:
-
Fatty acid ester of hydroxy fatty acid
- FFA:
-
Free fatty acid
- HPLC:
-
High-performance liquid chromatography
- LPC:
-
Lysophosphatidylcholine
- LPE:
-
Lysophosphatidylethanolamine
- MS:
-
Mass spectrometry
- OPLS-DA:
-
Orthogonal partial least squares discriminant analysis
- PC:
-
Phosphatidylcholine
- PE:
-
Phosphatidylethanolamine
- PI:
-
Phosphatidylinositol
- QC:
-
Quality control
- TAG:
-
Triacylglycerol
- TOF:
-
Time of flight
- UHPLC:
-
Ultrahigh-performance liquid chromatography
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
Arnold M, Sierra MS, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global patterns and trends in colorectal cancer incidence and mortality. Gut. 2017;66(4):683–91.
Allison JE. The effect of fecal occult-blood screening on the incidence of colorectal cancer. N Engl J Med. 2001;344(13):1022–3.
Taylor DP, Cannon-Albright LA, Sweeney C, Williams MS, Haug PJ, Mitchell JA, Burt RW. Comparison of compliance for colorectal cancer screening and surveillance by colonoscopy based on risk. Genet Med. 2011;13(8):737−43.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30.
Grunt TW. Interacting cancer machineries: cell signaling, lipid metabolism, and epigenetics. Trends Endocrinol Metab. 2018;29(2):86–98.
Wen YA, Xing X, Harris JW, Zaytseva YY, Mitov MI, Napier DL, et al. Adipocytes activate mitochondrial fatty acid oxidation and autophagy to promote tumor growth in colon cancer. Cell Death Dis. 2017;8(2):e2593.
Rossin D, Calfapietra S, Sottero B, Poli G, Biasi F. HNE and cholesterol oxidation products in colorectal inflammation and carcinogenesis. Free Radic Biol Med. 2017. https://doi.org/10.1016/j.freeradbiomed.2017.01.017.
Hussain A, Qazi AK, Mupparapu N, Guru SK, Kumar A, Sharma PR, et al. Modulation of glycolysis and lipogenesis by novel PI3K selective molecule represses tumor angiogenesis and decreases colorectal cancer growth. Cancer Lett. 2016;374(2):250–60.
Li S, Guo B, Song JW, Deng XL, Cong YS, Li PF, et al. Plasma choline-containing phospholipids: potential biomarkers for colorectal cancer progression. Metabolomics. 2013;9(1):202–12.
Zhao Z, Xiao Y, Elson P, Tan H, Plummer SJ, Berk M, et al. Plasma lysophosphatidylcholine levels: potential biomarkers for colorectal cancer. J Clin Oncol. 2007;25(19):2696–701.
Mirnezami R, Spagou K, Vorkas PA, Lewis MR, Kinross J, Want E, et al. Chemical mapping of the colorectal cancer microenvironment via MALDI imaging mass spectrometry (MALDI-MSI) reveals novel cancer-associated field effects. Mol Oncol. 2014;8(1):39–49.
Li F, Qin X, Chen H, Qiu L, Guo Y, Liu H, et al. Lipid profiling for early diagnosis and progression of colorectal cancer using direct-infusion electrospray ionization Fourier transform ion cyclotron resonance mass spectrometry. Rapid Commun Mass Spectrom. 2013;27(1):24–34.
Crotti S, Agnoletto E, Cancemi G, Di Marco V, Traldi P, Pucciarelli S, et al. Altered plasma levels of decanoic acid in colorectal cancer as a new diagnostic biomarker. Anal Bioanal Chem. 2016;408(23):6321–8.
Triebl A, Trötzmüller M, Hartler J, Stojakovic T, Köfeler HC. Lipidomics by ultrahigh performance liquid chromatography-high resolution mass spectrometry and its application to complex biological samples. J Chromatogr B. 2017;1053:72–80.
Zhang Q, Xu H, Liu R, Gao P, Yang X, Jin W, et al. A Novel Strategy for Targeted Lipidomics Based on LC-tandem-MS parameters prediction, quantification, and multiple statistical data mining: evaluation of lysophosphatidylcholines as potential cancer biomarkers. Anal Chem. 2019;91(5):3389–96.
Bligh EG, Dyer WJ. A rapid method of total lipid extraction and purification. Can J Biochem Physiol. 1959;37(8):91–7.
Tsugawa H, Cajka T, Kind T, Ma Y, Higgins B, Ikeda K, et al. MS-DIAL: data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat Methods. 2015;12(6):523–6.
Gika HG, Macpherson E, Theodoridis GA, Wilson ID. Evaluation of the repeatability of ultra-performance liquid chromatography-TOF-MS for global metabolic profiling of human urine samples. J Chromatogr B. 2008;871(2):299–305.
Want EJ, Wilson ID, Gika H, Theodoridis G, Plumb RS, Shockcor J, et al. Global metabolic profiling procedures for urine using UPLC-MS. Nat Protoc. 2010;5(6):1005–18.
Zhang J, Zhang L, Ye X, Chen L, Zhang L, Gao Y, et al. Characteristics of fatty acid distribution is associated with colorectal cancer prognosis. Prostaglandins Leukot Essent Fatty Acids. 2013;88(5):355–60.
Zhao Z, Xiao Y, Elson P, Tan H, Plummer SJ, Berk M, et al. Plasma lysophosphatidylcholine levels: potential biomarkers for colorectal cancer. J Clin Oncol. 2007;25(19):2696–701.
Dobrzyńska I, Szachowicz-Petelska B, Sulkowski S, Figaszewski Z. Changes in electric charge and phospholipids composition in human colorectal cancer cells. Mol Cell Biochem. 2005;276(1-2):113–9.
Hao B, Yu M, Sang C, Bi B, Chen J. Dyslipidemia and non-small cell lung cancer risk in Chinese population: a case-control study. Lipids Health Dis. 2018;17(1):278.
Li Z, Guan M, Lin Y, Cui X, Zhang Y, Zhao Z, et al. Aberrant lipid metabolism in hepatocellular carcinoma revealed by liver lipidomics. Int J Mol Sci. 2017;18(12):2550. https://doi.org/10.3390/ijms18122550.
Di Leo L, Vegliante R, Ciccarone F, Salvatori I, Scimeca M, Bonanno E, et al. Forcing ATGL expression in hepatocarcinoma cells imposes glycolytic rewiring through PPAR-α/p300-mediated acetylation of p53. Oncogene. 2018. https://doi.org/10.1038/s41388-018-0545-0.
Petan T, Jarc E, Jusović M. Lipid droplets in cancer: guardians of fat in a stressful world. Molecules. 2018;23(8):1941. https://doi.org/10.3390/molecules23081941.
Cotte AK, Aires V, Fredon M, Limagne E, Derangère V, Thibaudin M, et al. Lysophosphatidylcholine acyltransferase 2-mediated lipid droplet production supports colorectal cancer chemoresistance. Nat Commun. 2018;9(1):322.
Mitra R, Le TT, Gorjala P, Goodman OB Jr. Positive regulation of prostate cancer cell growth by lipid droplet forming and processing enzymes DGAT1 and ABHD5. BMC Cancer. 2017;17(1):631.
Zhang X, Saarinen AM, Hitosugi T, Wang Z, Wang L, Ho TH, et al. Inhibition of intracellular lipolysis promotes human cancer cell adaptation to hypoxia. Elife. 2017:001. https://doi.org/10.7554/eLife.31132.
Yang D, Li Y, Xing L, Tan Y, Sun J, Zeng B, et al. Utilization of adipocyte-derived lipids and enhanced intracellular trafficking of fatty acids contribute to breast cancer progression. Cell Commun Signal. 2018;16(1):32.
Chen G, Zhou G, Aras S, He Z, Lucas S, Podgorski I, et al. Loss of ABHD5 promotes the aggressiveness of prostate cancer cells. Sci Rep. 2017;7(1):13021.
Ye K, Wu Y, Sun Y, Lin J, Xu J. TLR4 siRNA inhibits proliferation and invasion in colorectal cancer cells by downregulating ACAT1 expression. Life Sci. 2016. https://doi.org/10.1016/j.lfs.2016.05.012.
Yue S, Li J, Lee SY, Lee HJ, Shao T, Song B, et al. Cholesteryl ester accumulation induced by PTEN loss and PI3K/AKT activation underlies human prostate cancer aggressiveness. Cell Metab. 2014;19(3):393–406.
Yore MM, Syed I, Moraes-Vieira PM, Zhang T, Herman MA, Homan EA, et al. Discovery of a class of endogenous mammalian lipids with anti-diabetic and anti-inflammatory effects. Cell. 2014;159(2):318–32.
Zhu QF, Yan JW, Gao Y, Zhang JW, Yuan BF, Feng YQ. Highly sensitive determination of fatty acid esters of hydroxyl fatty acids by liquid chromatography-mass spectrometry. J Chromatogr B. 2017. https://doi.org/10.1016/j.jchromb.2017.06.045.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (no. 81403086), the Fundamental Research Funds for the Central Universities of Central South University (no. 2018zzts901), and the Natural Science Foundation of Hunan Province, China (no. 2018JJ4020).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 417 kb)
Rights and permissions
About this article
Cite this article
Liu, T., Peng, F., Yu, J. et al. LC-MS-based lipid profile in colorectal cancer patients: TAGs are the main disturbed lipid markers of colorectal cancer progression. Anal Bioanal Chem 411, 5079–5088 (2019). https://doi.org/10.1007/s00216-019-01872-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-01872-5