Abstract
Lactoferrin (LF) is an important multifunctional protein that comprises a large fraction of the protein mass in certain human fluids and tissues, and its concentration is often used to assess health and disease. LF can be nitrated by multiple routes, leading to changes in protein structure, and nitrated proteins can negatively impact physiological health via nitrosative stress. Despite an awareness of the detrimental effects of nitrated proteins and the importance of LF within the body, cost-effective methods for detecting and quantifying nitrated lactoferrin (NLF) are lacking. We developed a procedure to selectively quantify NLF using sandwich enzyme-linked immunosorbent assay (ELISA), utilizing a polyclonal anti-LF capture antibody paired with a monoclonal anti-nitrotyrosine detector antibody. The assay was applied to quantify NLF in samples of pure LF nitrated via two separate reactions at molar ratios of excess nitrating agent to the total number of tyrosine residues between 10/1 and 100/1. Tetranitromethane (TNM) was used as a laboratory surrogate for an environmental pathway selective for production of 3-nitrotyrosine, and sodium peroxynitrite (ONOO−) was used as a surrogate for an endogenous nitration pathway. UV-vis spectroscopy (increased absorbance at 350 nm) and fluorescence spectroscopy (emission decreased by > 96%) for each reaction indicate the production of NLF. A lower limit of NLF detection using the ELISA method introduced here was calculated to be 0.065 μg mL−1, which will enable the detection of human-physiologically relevant concentrations of NLF. Our approach provides a relatively inexpensive and practical way to assess NLF in a variety of systems.

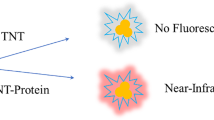
We developed a procedure to selectively quantify nitrated lactoferrin (NLF) protein using a sandwich enzyme-linked immunosorbent assay (ELISA) and verified results against several spectroscopic techniques. Our approach provides an inexpensive and practical way to assess NLF in a variety of systems.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lactoferrin (LF) is an iron-binding protein that is present in many tissues and fluids of the human body and is critical to diverse physiological functions [1]. LF exists in various exocrine fluids such as tears, milk, and saliva [2, 3] and possesses significant antifungal, antiviral, and antibacterial functions within the body [4]. Lactoferrin has a molecular weight of ca. 80 kDa and contains 703 amino acids, including 20 tyrosine residues and two globular lobes linked by an α-helix [5]. Each lobe contains two tyrosine residues at the center of its structure that participate with other amino acids as one iron-binding site [2]. The concentration of LF in human tissues has not been well-studied but is critical for physiological function [6].
Chemical modification of proteins like LF can also play important roles in controlling protein function. Free radicals such as reactive oxygen species (ROS) and reactive nitrogen species (RNS) are broadly important within the human body and as exogenous agents because they can lead to modifications that can affect protein function and harm the living system [7]. Specifically, the presence of nitrotyrosine-modified proteins is associated with many neurodegenerative, cardiovascular, respiratory, and ocular surface diseases [8]. One mechanism by which tyrosine nitration may affect physiology is through protein denaturation leading to an irreversible loss of function [9]. Damage to tissue can also induce an increase in NO synthesis, releasing more peroxynitrite and leading to further increases in protein nitration [10]. Thus, 3-nitrotyrosine (NTyr) can be used as a biomarker for nitrosative stress diseases in body fluids such as blood, urine, and tears [11]. Many ROS and RNS exist within the human body and contribute to the pathology of a variety of medical conditions. Peroxynitrite, in particular, has been shown to induce nitration of tyrosine (Tyr) residues as well as those of several other amino acids [12]. One important pathway by which ONOO− induces tyrosine nitration involves an electron oxidation reaction of the phenolic ring, which forms an intermediate radical at the 3-position of the ring, followed by combination with NO2 to generate NTyr, as summarized in Scheme 1 [13].
Significant effort has gone into the investigation of ROS in the atmosphere and its effects on human health. By comparison, relatively little study has been invested in what parallel role RNS may play in the same environmental systems [14]. One source of exogenous RNS is free radicals and reactive gases in the air that are emitted, directly or indirectly, by combustion sources and are higher in polluted, urban areas [15]. Contact between adjuvants such as pollutant gases and proteins on the surfaces or fluids of the skin, eyes, mucous membranes, and lungs can provide an important pathway of chemical reactivity [16], and exposure to nitrogen oxide gases influences many cardiovascular, respiratory, and ocular surface diseases [17]. Investigating the amount of nitrated proteins in urban road dust indicates that proteins on the surface of pollen and other biological particles can be nitrated in urban air and thus could influence human health [18].
In the atmosphere, tyrosine nitration is thought to proceed through a mechanism by which O3 and NO2 gases selectively produce a 3-NTyr product [19,20,21], whereas reaction with ONOO− can modify a wider set of amino acid residues, as discussed. Tetranitromethane (TNM) is frequently used in the laboratory as a surrogate for the heterogeneous O3 and NO2 mechanism, because it also selectively produces the 3-NTyr product [18, 22]. Sokolovsky et al. illustrated the TNM mechanism that inserts a nitro group into the aromatic ring specifically at the ortho position to produce 3-NTyr as in Scheme 2 [23]. It is important to note that both Schemes 1 and 2 lead to nitrated tyrosine residues but that these can be at different sites within the protein.
A number of laboratory techniques have been utilized to detect and quantify NTyr, especially for medically relevant research endeavors [24]. Among those, tandem mass spectrometric (MS/MS) analysis is the most sensitive for detecting low concentrations of NTyr and has been used to investigate the relationship of nitrated proteins to many diseases, but requires very high up-front (ca. $300k to $1000k) and maintenance costs [8, 20]. To overcome some of the limitations presented by MS analysis, many studies now show that enzyme-linked immunosorbent assay (ELISA) can be applied to selectively detect and quantify low concentrations of nitrotyrosine within a sample (e.g., [25]). ELISA has been used to quantify LF concentration in milk, plasma, and tear fluid [3] as well as NTyr in plasma and urine [26]. Only once has the detection of lactoferrin nitration been reported directly, however. Teuwissen et al. used a combination of iron-binding capacity and optical rotator dispersion to detect changing tyrosine properties upon nitration of LF [27].
Despite the importance of LF and nitrated proteins within the human system and as biomarkers for many diseases, very little has been reported on chemical modifications of LF. This is due in part to the challenges to inexpensively and selectively quantify nitrated lactoferrin (NLF). We summarize two independent methodologies to nitrate LF via aqueous reaction, utilizing TNM as a surrogate for exogenous atmospheric nitration and ONOO− as a model of endogenous physiological nitration pathway. Spectrophotometry was utilized as a method to show modifications to the protein. Further, we describe herein the development and optimization of a novel sandwich ELISA technique to selectively quantify NTyr in lactoferrin after nitration by each pathway. Using the reported ELISA protocol, we anticipate the ability to selectively quantify NLF at physiologically relevant concentrations, nitrated by either endogenous or exogenous pathways in human tissue and fluid samples, thus providing researchers without access to expensive instrumentation an accurate and effective way to measure NLF.
Experimental
Endogenous nitration of LF using sodium peroxynitrite
The peroxynitrite reaction was prepared following the method reported by Selzle et al. [28], replacing Bet v 1 with LF and adjusting for differences in the number of tyrosine residues between the two proteins. According to this procedure, 200-μL aliquots of 5.0 mg mL−1 LF in aqueous solution were pipetted into 2.5-mL glass vials. PBS buffer, followed by NaONOO, was then added, and reaction solutions were stirred on ice for 2 h. NaONOO was added in different amounts corresponding to peroxynitrite to tyrosine (ONOO−/Tyr) molar ratios of 10/1, 20/1, 30/1, 40/1, 60/1, 80/1, and 100/1. PBS buffer was added to a total volume of 1000 μL for each solution at a NLF concentration of 1.0 mg mL−1. Previous work showed that concentrations of LF higher than this produced lower nitration reaction efficiency [29]. Details regarding materials and reagents are listed in the Electronic Supplementary Material (ESM).
Exogenous nitration of LF using tetranitromethane
The tetranitromethane nitration reaction was prepared following the method reported by Teuwissen et al. [27]. Similar to the above procedure, 200-μL aliquots of 5.0 mg mL−1 LF in aqueous solution were combined with Tris buffer, followed by 200 μL of ethanol and TNM, in 2.5-mL glass vials. TNM was added in amounts corresponding to tetranitromethane to tyrosine (TNM/Tyr) molar ratios between 10/1 and 100/1. The mixtures were stirred at room temperature (RT) for 4 h. Tris buffer was added to a total volume of 1000 μL at a NLF concentration of 1.0 mg mL−1.
In all cases described below, NLF used as standard reference material for ELISA protocols was produced using the 40/1 M ratio of TNM/Tyr reaction described here. Also, in all cases, LF and NLF concentrations are reported as ± 5%, which was limited by the precision of the LF mass amount in the originally purchased vial.
Spectroscopic analysis
For all spectroscopic analyses, quartz cuvettes were used, and LF and NLF samples were diluted in PBS buffer. Dilution calculations were performed by taking the molecular mass of LF, irrespective of mass changes due to nitration.
UV-vis absorption spectroscopy was performed using a Cary BIO 100 spectrophotometer. Absorption spectra were acquired in 2-nm steps from 250 to 500 nm.
Fluorescence spectroscopy was conducted using a Varian Cary Eclipse fluorescence G9800A spectrophotometer equipped with a xenon lamp as excitation source. The excitation wavelength was stepped every 2 nm, and fluorescence emission spectra were acquired from 200 to 500 nm at 2 nm resolution.
Direct ELISA for characterization of nitrotyrosine antibody
Direct ELISA was used to determine the optimal dilution factor of the detector antibody that was later used in the sandwich ELISA procedure. Nitrated bovine serum albumin (NBSA, 6/1 TNM/Tyr) was serially diluted into carbonate buffer producing five concentrations of NBSA. Microwells were coated in triplicate with each NBSA solution, and pure carbonate buffer was used as a blank. The plate was covered, sealed with sterile tape, and incubated at 4 °C for 17 h overnight. After incubation, the wells were washed in duplicate by adding 200 μL of PBST (PBS with 0.05% Tween). Each subsequent incubation step also followed a similar process of sealing, continuous shaking during the incubation period using a rocking platform (UltraRocker, BIO-RAD), and then duplicate washing with PBST. After washing, the wells were blocked with 200 μL of blocking buffer (5% BSA in PBS) and incubated at RT for 2 h. After washing, mouse monoclonal to nitrotyrosine biotinylated antibody (anti-NTyr) was diluted in blocking buffer in four different dilution ratios between 1:100 and 1:1000. Fifty microliters of each dilution was added in triplicate and incubated at RT for 1 h, thus utilizing 12 wells (four dilution ratios × triplicate measurements) for each NBSA concentration. Next, 50 μL of streptavidin-HRP (diluted 1:10,000 in blocking buffer) was added before incubation at RT for 1 h. Each plate was developed by adding 100 μL of TMB substrate to each well and incubating at RT until the color turned blue (ca. 10 min). The substrate reaction was stopped after 10 min by adding 100 μL of 0.5 N aqueous sulfuric acid, prompting the solution to turn yellow. The optical absorbance of each well was measured using a microwell plate reader (Tecan Infinite, M1000 PRO). Following standard procedures [30], the absorbance of the substrate background at 620 nm (blue) was subtracted from the absorbance at 450 nm (yellow) to produce the absorbance values reported on all plots.
Sandwich ELISA for quantification of NLF
For the sandwich ELISA procedure introduced here, goat polyclonal to lactoferrin non-conjugated antibody (anti-LF) was used as the capture antibody, and mouse monoclonal to nitrotyrosine biotinylated antibody (anti-NTyr) was used as the detector antibody (clone number CC.22.8C7.3). Details regarding the choice and dilution ratio of antibodies are outlined in the ESM (Section S.6). Microwells were coated with 50 μL of diluted capture antibody and incubated at 4 °C overnight. The plate was then washed and blocked as described above. NLF was serially diluted into blocking buffer producing NLF solutions at eight concentrations. After a second washing step, 50 μL of each of the eight NLF solutions was added in triplicate to the plate and incubated at RT for 1 h. Blocking buffer was used for blank measurements. Fifty microliters of the detector antibody, diluted in blocking buffer to a ratio of 1:100, was then added, and the plate was incubated with shaking at RT for 1 h. Streptavidin-HRP and TMB were added following the ELISA protocol, as described above.
Data analysis
Data points are shown here as mean ± sample standard deviation (s). Statistical analysis was performed using Igor Pro 7 (Wavemetrics, Inc., OR, USA). The experiments were carried out in triplicate, and the results were subjected to t tests to provide a measure of significance.
Results and discussion
Detection of NLF using absorption spectroscopy
After nitration of LF via the two pathways (TNM and ONOO−), nitration products were detected using UV-vis absorption spectroscopy. Figure 1 shows absorption spectra of LF and NLF nitrated separately by ONOO− (\( {\mathrm{NLF}}_{ONOO^{-}} \)) and TNM (NLF TNM ) reactions. Absorption measurements at 280 nm are commonly used as a general tool for the detection of proteins that contain at least one of the three aromatic amino acids Phe, Tyr, or Trp. The spectrum of LF (Fig. 1, black trace) shows one peak at 280 nm, which represents absorption by a combination of these aromatic amino acids. The absorption spectrum of NLF TNM (Fig. 1, red trace) shows a peak at 350 nm, consistent with the peak location of nitrated tyrosine [31]. An additional product of the reaction is nitroformate, which also absorbs at 350 nm [32]. As a result, the reaction mixture was purified (see ESM Section S.3), and the absorption spectra were compared before and after purification to verify the presence of NLF without interference. Both spectra show the peak at 350 nm, qualitatively verifying the presence of the product. Quantitative comparison is more complicated due to uncertainties in concentration introduced by dilution during purification.
The absorption spectrum of \( {\mathrm{NLF}}_{ONOO^{-}} \) (Fig. 1, blue trace) shows a NLF peak at 350 nm as well as a double-humped peak at 280 nm from unreacted Phe, Tyr, and Trp in the protein. The \( {\mathrm{NLF}}_{ONOO^{-}} \) product was not purified before spectroscopic analysis, because secondary reaction products and degradation by-products of ONOO− do not overlap with the 350-nm peak [33]. As a result, the quantity of the \( {\mathrm{NLF}}_{ONOO^{-}} \) product is unambiguous here. In the case of the ONOO− reaction, modification of the protein is not limited to tyrosine, and the absorption spectrum of the nitrated product can be broadened by nitration of Phe, Tyr, and Trp residues [34]. Further, the relative tyrosine nitration yield from ONOO− is reduced compared to the TNM reaction, because the ratio of ONOO− to the sum of all nitrate-able amino acids is lower than the ratio of TNM to Tyr [35].
Detection of NLF using fluorescence spectroscopy
Fluorescence spectroscopy has also been frequently used to detect protein modifications, due to changes that can take place in protein molecular conformation [36]. Tyr exhibits among the highest fluorescence quantum yields of all amino acids, and it is the primary source of fluorescence in proteins without Trp. When situated sufficiently close (ca. < 5 nm) to Trp within a protein structure, however, Tyr experiences efficient resonance energy transfer to Trp and so is often not responsible for observed fluorescence [37, 38]. LF exhibits relatively strong fluorescence emission because it contains a large number of aromatic amino acids, including Phe (32), Tyr (20), and Trp (10) [39]. Fluorescence from nitrated proteins has previously been shown to reduce total intensity from native proteins and can thus be used as a quantitative measure of nitration [40, 41].
Figure 2 shows fluorescence excitation emission matrices (EEM) for native LF as well as for NLF TNM and \( {\mathrm{NLF}}_{ONOO^{-}} \). In the presence of nitrating agents, the addition of a nitro group (NO2) onto Tyr decreases the pKa of the hydroxyl group in the phenolic ring. As a result, the substitution of a large hydrophobic NO2 group sterically hinders the tyrosine structure and can cause conformational changes to the protein [13]. This reduces the absorbance intensity due to the unmodified protein and also the intensity of the fluorescence emission.
Fluorescence EEMs for native LF (a), NLF TNM (b), and \( {\mathrm{NLF}}_{ONOO^{-}} \) (c). Diagonal lines indicate elastic scattering from excitation source. EEM color scale indicates emission intensity, in logarithmically scaled arbitrary units (arb.). Peak emission intensity at λ EX 280 nm for each panel summarized in panel (d)
LF shows strong emission centered at λ Em 330 nm for two excitation bands at λ Ex 224 and 280 nm due to a large number of Tyr and Trp residues, consistent with previous studies [42]. Both nitrated products show dramatically decreased emission intensity (Fig. 2d; NLF TNM , 98% decrease and \( {\mathrm{NLF}}_{ONOO^{-}} \), 96% decrease) due to the addition of NO2 groups to LF, qualitatively matching observations for nitrotyrosine reported by Crow et al. [40]. The emission intensity of NLF TNM is reduced by a greater factor than \( {\mathrm{NLF}}_{ONOO^{-}} \), again because ONOO− nitrates more broadly than TNM.
Design of sandwich ELISA assay for quantification of NLF
The sandwich ELISA developed here was designed to detect NTyr bound to LF protein in NLF samples. The assay first utilizes polyclonal anti-LF capture antibodies to bind LF proteins. This antibody recognizes a specific sequence of 13 amino acids in the LF protein (see ESM Section S.4) [43]. Second, monoclonal anti-NTyr detector antibodies are used to attach to specific NTyr residues bound on captured NLF proteins. For the capture process, polyclonal antibodies were used to enhance the probability of binding, given the many sites at which nitration could affect LF binding, whereas monoclonal antibodies were used as detector antibodies to selectively enhance the assay’s sensitivity (ESM Fig. S2). The scheme is conceptually similar to the sandwich ELISA protocol introduced by Franze et al., which detects NTyr in nitrated birch pollen protein (Nitro-(3)-Bet v 1) and used rabbit polyclonal anti-Bet v 1 protein as capture and mouse monoclonal anti-NTyr as detector antibodies [19]. The detector antibody chosen here has been used in many immunoassay applications [44] because it has low cross-reactivity [45] and high sensitivity [46]. See ESM Section S.6 and Figs. S1–S2 for additional details about assay development.
Development of sandwich ELISA
In order to develop the modified sandwich ELISA for detection of NLF, each of the two antibodies was tested separately using direct ELISA and combined into a single sandwich ELISA. As a first step, direct ELISA was used to determine the optimal buffer dilution ratio of NBSA antigen to the nitrotyrosine biotinylated antibody (analogous to ESM Fig. S3 without the initial capture antibody step). Four dilution ratios from 1:100 to 1:1000 were investigated. The 1:100 dilution exhibited the highest binding sensitivity, shown as the steepest slope in Fig. 3.
As a second step, the full sandwich ELISA was used to optimize the dilution of capture antibody (anti-LF). The 1:1000 (v/v) dilution of capture antibody was selected because it exhibited binding sensitivity similar to that of the 1:250 (v/v) dilution, but requires less antibody volume (ESM Fig. S4). Using the higher dilution thus achieves sensitive detection at a lower cost.
A calibration curve of optical absorption versus NLF concentration was developed as the final detection step of the sandwich ELISA (Fig. 4). A range of molar ratios between 10/1 and 100/1 was investigated (ESM Fig. S5–S6), and the 40/1 M ratio of TNM/Tyr was chosen as the protein standard. Sandwich ELISA results show a decrease in detected NLF concentration produced at nitration ratios higher than 40/1. The reason for this could be due to a reduction in antibody binding due to LF changes at capture antibody binding sites or as a result of denaturation of LF protein. Results published by Teuwissen et al. also show NLF TNM properties peaking at a 40/1 M ratio [27].
The limit of detection (LOD) for the newly developed assay was calculated by two separate methods by measuring the mean + three times the sample standard deviation of a set of three absorbance measurements following sandwich ELISA. LOD LF was calculated as 0.032 absorbance units (AU) by replacing native LF for NLF as the antigen in the assay, and LOD blank was calculated as 0.022 AU by withholding any antigen from the assay. A minimum detectable NLF concentration of 0.065 μg mL−1 was calculated as 3 s LF /m, where s LF is the standard deviation of the blank measurement and m is the slope of the NLF calibration curve. Using the responses from the sandwich ELISA (Fig. 4), t tests showed the NLF measurement to be statistically different (p value < 0.01) from the LF measurement at all antigen concentrations measured (minimum 0.18 μg mL−1; see ESM Table S1).
To confirm the results from the new ELISA protocol, Fig. 5 shows that the molecular absorbance of NLF (black traces) and responses from the sandwich ELISA (red bars) each increase with increasing molar ratio of TNM or ONOO− to Tyr. ESM Fig. S5 shows an extension of the ELISA response to higher ratios of nitrating agent.
NLF samples produced using the TNM reaction were tested in both purified and non-purified forms, following procedures outlined above. Absorbance values and sandwich ELISA results show no statistical differences, and so all NLF TNM data shown here (Figs. 4 and 5) are shown without any purification (see ESM Fig. S7). The ability of the assay to work effectively without purifying the product is important, because it shows that the ELISA protocol developed here can be applied broadly to samples extracted from tissues and fluids without the need to perform additional purifying steps that can increase analysis costs and quantitative uncertainty. In contrast to this, the \( {\mathrm{NLF}}_{ONOO^{-}} \) samples can degrade over time if produced via the reactions outlined above. \( {\mathrm{NLF}}_{ONOO^{-}} \) samples thus require purification before storage to limit agglomeration and protein changes induced by the presence of excess ONOO− [47].
Conclusions
NLF samples were synthesized using two different nitration reagents in varying ratios of molar excess. ONOO− was used as an endogenous model and TNM was used as an exogenous model. Both nitrated products were characterized by UV-vis and fluorescence spectroscopy, showing dramatic changes upon nitration. To quantify the concentration of NLF produced from each reaction, we developed a novel sandwich ELISA for the selective detection of NTyr bound within NLF antigens. We demonstrate a LOD LF of 0.032 AU and a LOD blank of 0.022 AU, translating to a lower limit of detectable NLF concentration of 0.065 μg mL−1. An important feature of this sandwich ELISA protocol is that is does not require sample purification, meaning that quantitative uncertainty introduced from imprecise dilution through washing steps can be avoided. The NLF sandwich ELISA introduced here is a novel method that was demonstrated to be applicable to the quantification of nitrotyrosine in lactoferrin. LF is the most concentrated protein in human tears, representing up to 25% of the protein mass, or approximately 2.2 mg mL−1 [3]. The concentration of ONOO− generated in inflamed corneal tissue has been shown to be in the approximate range of 3–30 μM [48]. NLF has never been quantified in tissue or fluid samples, but if even 30 of every 106 LF molecules (30 ppm) were nitrated by ONOO−, NLF concentrations would be above detection limits reported here. Thus, NLF is likely to be detectable by this procedure at physiologically relevant concentrations. As a result, this assay has important application to the investigation of NLF produced through inflammatory pathways in human tear, milk, and ocular tissue samples, and also through inflammation linked to traffic-borne air pollution.
References
Kanyshkova TG, Buneva VN, Nevinsky GA. Lactoferrin and its biological functions. Biochem Mosc. 2001;66(1):1–7.
Vorland LH. Lactoferrin: a multifunctional glycoprotein. APMIS. 1999;107(7–12):971–81.
Kijlstra A, Jeurissen SH, Koning KM. Lactoferrin levels in normal human tears. Br J Ophthalmol. 1983;67(3):199–202.
Jenssen H, Hancock REW. Antimicrobial properties of lactoferrin. Biochimie. 2009;91(1):19–29.
Lönnerdal B, Iyer S. Lactoferrin: molecular structure and biological function. Annu Rev Nutr. 1995;15(1):93–110.
González-Chávez SA, Arévalo-Gallegos S, Rascón-Cruz Q. Lactoferrin: structure, function and applications. Int J Antimicrob Agents. 2009;33(4):301–e1.
Valko M, Leibfritz D, Moncol J, Cronin MTD, Mazur M, Telser J. Free radicals and antioxidants in normal physiological functions and human disease. Int J Biochem Cell Biol. 2007;39(1):44–84.
Yeo W-S, Lee S-J, Lee J-R, Kim K-P. Nitrosative protein tyrosine modifications: biochemistry and functional significance. BMB Rep. 2008;41(3):194–203.
Gow AJ, Duran D, Malcolm S, Ischiropoulos H. Effects of peroxynitrite-induced protein modifications on tyrosine phosphorylation and degradation. FEBS Lett. 1996;385(1–2):63–6.
Pfeiffer S, Lass A, Schmidt K, Mayer B. Protein tyrosine nitration in cytokine-activated murine macrophages involvement of a peroxidase/nitrite pathway rather than peroxynitrite. J Biol Chem. 2001;276(36):34051–8.
Dalle-Donne I, Scaloni A, Giustarini D, Cavarra E, Tell G, Lungarella G, et al. Proteins as biomarkers of oxidative/nitrosative stress in diseases: the contribution of redox proteomics. Mass Spectrom Rev. 2005;24(1):55–99.
Beckman JS. Oxidative damage and tyrosine nitration from peroxynitrite. Chem Res Toxicol. 1996;9(5):836–44.
Radi R. Protein tyrosine nitration: biochemical mechanisms and structural basis of its functional effects. Acc Chem Res. 2013;46(2):550–9.
Wiseman H, Halliwell B. Damage to DNA by reactive oxygen and nitrogen species: role in inflammatory disease and progression to cancer. Biochem J. 1996;313(Pt 1):17-29.
Reinmuth-Selzle K, Kampf CJ, Lucas K, Lang-Yona N, Fröhlich-Nowoisky J, Shiraiwa M, et al. Air pollution and climate change effects on allergies in the anthropocene: abundance, interaction, and modification of allergens and adjuvants. Environ Sci Technol. 2017;51(8):4119–41.
Lakey PSJ, Berkemeier T, Tong H, Arangio AM, Lucas K, Pöschl U, et al. Chemical exposure-response relationship between air pollutants and reactive oxygen species in the human respiratory tract. Sci Rep. 2016;6:32916.
Patel RP, McAndrew J, Sellak H, White CR, Jo H, Freeman BA, et al. Biological aspects of reactive nitrogen species. Biochim Biophys Acta Bioenerg. 1999;1411(2–3):385–400.
Franze T, Weller MG, Niessner R, Pöschl U. Protein nitration by polluted air. Environ Sci Technol. 2005;39(6):1673–8.
Franze T, Weller MG, Niessner R, Pöschl U. Enzyme immunoassays for the investigation of protein nitration by air pollutants. Analyst. 2003;128(7):824–31.
Reinmuth-Selzle K, Ackaert C, Kampf CJ, Samonig M, Shiraiwa M, Kofler S, et al. Nitration of the birch pollen allergen Bet v 1.0101: efficiency and site-selectivity of liquid and gaseous nitrating agents. J Proteome Res. 2014;13(3):1570–7.
Kampf CJ, Liu F, Reinmuth-Selzle K, Berkemeier T, Meusel H, Shiraiwa M, et al. Protein cross-linking and oligomerization through dityrosine formation upon exposure to ozone. Environ Sci Technol. 2015;49(18):10859–66.
Shuker DE, Prevost V, Friesen MD, Lin D, Ohshima H, Bartsch H. Urinary markers for measuring exposure to endogenous and exogenous alkylating agents and precursors. Environ Health Perspect. 1993;99:33–7.
Sokolovsky M, Riordan JF, Vallee BL. Tetranitromethane. A reagent for the nitration of tyrosyl residues in proteins. Biochemistry. 1966;5(11):3582–9.
Crow JP, Beckman JS. Quantitation of protein tyrosine, 3-nitrotyrosine, and 3-aminotyrosine utilizing HPLC and intrinsic ultraviolet absorbance. Methods. 1995;7(1):116–20.
Engvall E. [28] Enzyme immunoassay ELISA and EMIT. Methods Enzymol. 1980;70:419–39.
Jamshad K, Brennan MD, Bradley N, Beirong GAO, Bruckdorfer R, Jacobs M. 3-Nitrotyrosine in the proteins of human plasma determined by an ELISA method. Biochem J. 1998;330(2):795–801.
Teuwissen B, Masson PL, Osinski P, Heremans JF. Metal-combining properties of human lactoferrin. FEBS J. 1973;35(2):366–71.
Selzle K, Ackaert C, Kampf CJ, Kunert AT, Duschl A, Oostingh GJ, et al. Determination of nitration degrees for the birch pollen allergen Bet v 1. Anal Bioanal Chem. 2013;405(27):8945–9.
Vincent JP, Lazdunski M, Delaage M. On the use of tetranitromethane as a nitration reagent. The reaction of phenol side-chains in bovine and porcine trypsinogens and trypsins. FEBS J. 1970;12(2):250–7.
Rajasekariah GH, Kay GE, Russell NV, Smithyman AM. Assessment of assay sensitivity and precision in a malaria antibody ELISA. J Immunoassay Immunochem. 2003;24(1):89–112.
Riordan JF, Sokolovsky M, Vallee BL. Tetranitromethane. A reagent for the nitration of tyrosine and tyrosyl residues of proteins1. J Am Chem Soc. 1966;88(17):4104–5.
Sokolovsky M, Fuchs M, Riordan JF. Reaction of tetranitromethane with tryptophan and related compounds. FEBS Lett. 1970;7(2):167–70.
Jankowski JJ, Kieber DJ, Mopper K. Nitrate and nitrite ultraviolet actinometers. Photochem Photobiol. 1999;70(3):319–28.
Ter Steege JCA, Koster-Kamphuis L, van Straaten EA, Forget PP, Buurman WA. Nitrotyrosine in plasma of celiac disease patients as detected by a new sandwich ELISA. Free Radic Biol Med. 1998;25(8):953–63.
Ischiropoulos H, Zhu L, Beckman JS. Peroxynitrite formation from macrophage-derived nitric oxide. Arch Biochem Biophys. 1992;298(2):446–51.
Kronman MJ, Holmes LG. The fluorescence of native, denatured and reduced-denatured proteins. Photochem Photobiol. 1971;14(2):113–34.
Lakowicz JR. Principles of fluorescence spectroscopy: Springer Science & Business Media; 2013.
Pöhlker C, Huffman JA, Pöschl U. Autofluorescence of atmospheric bioaerosols—fluorescent biomolecules and potential interferences. Atmos Meas Tech. 2012;5(1):37–71.
Bläckberg L, Hernell O. Isolation of lactoferrin from human whey by a single chromatographic step. FEBS Lett. 1980;109(2):180–4.
Crow JP, Ischiropoulos H. [17] Detection and quantitation of nitrotyrosine residues in proteins: in vivo marker of peroxynitrite. Methods Enzymol. 1996;269:185–94.
De Filippis V, Frasson R, Fontana A. 3-Nitrotyrosine as a spectroscopic probe for investigating protein protein interactions. Protein Sci. 2006;15(5):976–86.
Meyer A, Betzel C, Pusey M. Latest methods of fluorescence-based protein crystal identification. Acta Cryst. 2015;71(2):121–31.
abcam. Anti-Lactoferrin antibody (ab77548). 2017. Accessed 26 Jun 2017.
Ferrante RJ, Shinobu LA, Schulz JB, Matthews RT, Thomas CE, Kowall NW, et al. Increased 3-nitrotyrosine and oxidative damage in mice with a human copper/zinc superoxide dismutase mutation. Ann Neurol. 1997;42(3):326–34.
Chou SM, Wang HS, Komai K. Colocalization of NOS and SOD1 in neurofilament accumulation within motor neurons of amyotrophic lateral sclerosis: an immunohistochemical study. J Chem Neuroanat. 1996;10(3):249–58.
Aggarwal S, Gross CM, Kumar S, Datar S, Oishi P, Kalkan G, et al. Attenuated vasodilatation in lambs with endogenous and exogenous activation of cGMP signaling: role of protein kinase G nitration. J Cell Physiol. 2011;226(12):3104–13.
Estillore AD, Trueblood JV, Grassian VH. Atmospheric chemistry of bioaerosols: heterogeneous and multiphase reactions with atmospheric oxidants and other trace gases. Chem Sci. 2016;7(11):6604–16.
Ashki N, Chan AM, Qin Y, Wang W, Kiyohara M, Lin L, et al. Peroxynitrite upregulates angiogenic factors VEGF-A, BFGF, and HIF-1α in human corneal limbal epithelial cells. Invest Ophthalmol Vis Sci. 2014;55(3):1637–46.
Acknowledgements
Amani Alhalwani acknowledges scholarship support from King Saud bin Abdulaziz University for Health Sciences (KSAU-HS).
Funding
This study received funding support through a grant from the University of Denver Knoebel Institute for Healthy Aging (KIHA).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 250 kb)
Rights and permissions
About this article
Cite this article
Alhalwani, A.Y., Repine, J.E., Knowles, M.K. et al. Development of a sandwich ELISA with potential for selective quantification of human lactoferrin protein nitrated through disease or environmental exposure. Anal Bioanal Chem 410, 1389–1396 (2018). https://doi.org/10.1007/s00216-017-0779-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-017-0779-7