Abstract
Analytical techniques for detection of ochratoxin A (OTA) in food products and blood serum are of great significance. In this study, a fluorescent aptasensor was developed for sensitive and specific detection of OTA, based on a DNA pyramid nanostructure (DPN) and PicoGreen (PG) dye. The designed aptasensor inherits characteristics of DPN, such as high stability and capacity for PG loading. PG, as a fluorescent dye, could bind to double-stranded DNA (dsDNA). In the absence of OTA, the pyramid structure of DPN remains intact, leading to a very strong fluorescence emission. Because of higher affinity of aptamer for its target relative to its complementary strand, upon addition of target, the pyramid structure of DPN is disassembled, leading to a weak fluorescence emission. The presented aptasensor showed high specificity toward OTA with a limit of detection (LOD) as low as 0.135 nM. Besides, the designed sensing strategy was successfully utilized to recognize OTA in serum and grape juice with LODs of 0.184 and 0.149 nM, respectively.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ochratoxin A (OTA), a prevalent toxic mycotoxins, is generated by different specious of fungi, such as Penicillium and Aspergillus [1, 2]. OTA can occur in a broad range of food matrices, such as grape juice, cereals, wine, barely, milk, and meat products [3–5]. OTA is hepatotoxic, nephrotoxic, teratogenic, immunotoxic, and mutagenic to humans and animals [6, 7]. Considering these adverse effects, the design of sensitive approaches for the recognition of OTA in food and serum are in great demand.
So far, different analytical approaches have been utilized for the detection of OTA, including mass spectrometry (MS), enzyme-linked immunosorbent assay (ELISA), and high-performance liquid chromatography (HPLC). Generally, these detection procedures are laborious, expensive, and need skilled personnel and sophisticated instruments [1, 2, 6, 8].
Application of aptasensors in analytical techniques is growing fast. Aptamers are single-stranded DNA or RNA nucleic acids artificially produced by an in vitro procedure named SELEX (systematic evolution of ligands by exponential enrichment) [9, 10]. Aptamers interact with a variety of targets, ranging from metal ions to amino acids and entire cells with high affinity and specificity [11, 12]. Aptamers possess significant advantages over traditional antibodies, e.g., small size, no or low toxicity and immunogenicity, high reproducibility, low cost, high thermal stability, and ease of preparation and modification [12–15]. Owing to these outstanding features, aptamers are promising molecular sensing elements for the construction of diverse sensing strategies [16–18].
Fluorescence has been largely utilized for analytical approaches, because of its ease of detection and high sensitivity [19, 20].
So, in this work a novel fluorescent aptasensor was presented for OTA detection based on a DNA pyramid nanostructure (DPN) and PicoGreen (PG) dye. The developed DPN is a DNA nanostructure which has short double-stranded DNA (dsDNA) building blocks. The OTA aptamer (Apt) was placed in the structure of DPN. The fabricated DPN has outstanding features, such as high stability, monodispersity, and high capacity for PG loading. These unique characteristics make the developed DPN an ideal sensing platform for detection of OTA. A 36-base ssDNA, which interacts with OTA selectively [21, 22], was utilized as a molecular recognition element.
Materials and methods
Materials
A Polygen DNA synthesizer (Polygen, Germany) was used to synthesize all the sequences (Table 1). PicoGreen (PG, 200×) was purchased from Invitrogen (USA). Ochratoxin A (OTA), warfarin, plasma from rat, acetamiprid, salmon sperm DNA, ricin A, and aflatoxin B1 and G1 were provided by Sigma-Aldrich (USA).
Preparation of DPN
Final concentrations of 10 μM from A, B, C, and D ssDNAs were mixed in TM buffer containing 10 mM Tris-HCl, pH 7.4 and 20 mM MgCl2. The mixture was heated for 5 min at 90 °C and then cooled to 4 °C on ice. Formation of DPN was evaluated by 2 % agarose gel electrophoresis. The size of DPN was assessed using a particle size analyzer (Malvern, UK).
Influence of concentration of PG on fluorescence intensity of aptasensor
Increasing amounts of PG (0–20 μl 20×) were added to 100 nM DPN (100 μl final volume) and incubated for 7 min at room temperature. Then, fluorescence spectra, λEx = 480 nm, were recorded on a Synergy H4 microplate reader (BioTek, USA).
Influence of incubation time of PG on function of aptasensor
Ten microliters of 20× PG was added to 90 μl DPN (final concentration of 100 nM). The solutions were incubated at room temperature from 0 to 10 min. Then, fluorescence spectra were measured.
Function study of fabricated fluorescent aptasensor
Fluorescent intensity was utilized to evaluate the interaction of OTA and the developed aptasensor. Four microliters of OTA (10 nM) was added to 86 μl DPN (100 nM). After the mixture was incubated for 45 min at room temperature, 10 μl 20× PG was added and the fluorescence emission spectra were recorded after 7 min.
Quantitative detection of OTA using fluorescence measurement
Different amounts of OTA (0–20 nM) were mixed with DPN (100 nM) and incubated at room temperature for 45 min, followed by incubation with 10 μl 20× PG for 7 min. Then, fluorescence intensities, λEx = 480 nm and λEm = 530 nm, were measured.
Selectivity evaluation of the aptasensor
The selectivity was investigated both in the buffer and grape juice in the presence of 10 and 50 nM aflatoxin B1, warfarin, and aflatoxin G1 as structural analogues of OTA and other toxic substances, including acetamiprid and ricin A.
Stability of DPN in serum
Serum stability of DPN was investigated by agarose gel electrophoresis. Thus, 2 μM DPN was treated with rat serum for 3 h. At specific times, the samples were extracted with phenol/chloroform and 2 % agarose gel electrophoresis was performed.
Detection of OTA in serum and grape juice
To appraise the operation of the designed aptasensor in grape juice, increasing amounts of OTA (0–20 nM) and 0.1 mg/mL salmon sperm DNA were spiked into grape juice, and the fluorescent intensity was used to determine the concentrations of OTA.
The same experiment was repeated to measure the concentration of OTA in serum.
Results and discussion
Detection mechanism of the fluorescent aptasensor
The fabricated fluorescent aptasensor was based on the pyramid nanostructure of DPN, target-induced disassembly of the DPN structure, and the ability of PG to act as a signal probe. In this study, PG was utilized as a fluorescent probe owing to its high specificity and sensitivity for binding to dsDNA. Upon binding to dsDNA, fluorescence of PG increases more than 1000-fold, while the free PG exhibits no fluorescence [23, 24].
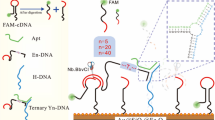
The principle of the developed fluorescent aptasensor is illustrated in Scheme 1. In the absence of OTA, the pyramid structure of DPN remains intact. So, a very strong fluorescence emission is observed following the addition of PG to the DPN, because PG could intercalate into dsDNA blocks of DPN.
OTA detection based on fluorescent assay. a In the presence of OTA, the structure of DPN is disassembled and DPN has less dsDNA building blocks in its structure. Thus, weak fluorescence emission is observed upon the addition of PG to the disassembled DPN. b In the absence of target, the DPN structure remains intact and contains more dsDNA building blocks. So, addition of PG to the DPN leads to very strong fluorescence emission
In the presence of OTA, Apt leaves its complementary strand, the Apt–OTA conjugate forms, and the structure of DPN is disassembled. It has been demonstrated that the aptamer interacts with its corresponding target with a greater binding constant compared to its complementary strand [23, 25, 26]. Upon addition of PG, a weak fluorescence emission is obtained, because of the presence of less dsDNA building blocks in the structure of disassembled DPN.
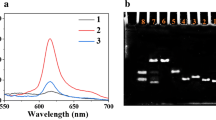
A fluorescence assay was used to evaluate the function of the aptasensor. As shown in Fig. 1, the PG alone had the minimum fluorescence intensity (green curve), owing to its fluorescence characteristic. In the absence of OTA, upon the addition of PG to DPN, the fluorescence intensity was significantly enhanced (red curve), verifying that the designed DPN remained intact and high amounts of PG could be bound to the dsDNA building blocks of DPN. In the presence of target following the addition of PG to DPN, the fluorescence intensity was significantly decreased (blue curve), confirming the formation of Apt–OTA conjugate and disassembly of DPN structure. So, less dsDNA building blocks were available in the structure of disassembled DPN for PG loading.
DPN characterization
The production of DPN was confirmed using a gel retardation assay. As shown in Fig. 2a, the band of DPN was retarded compared to the bands of other DNA combinations lacking one or more strands. The size of DPN was 14 ± 1.2 nm as determined by the particle size analyzer. A stability assay indicated that DPN was stable over 3 h when incubated in serum (Fig. 2b).
a Agarose gel electrophoresis of DPN formation. Lane 1 A sequence, lane 2 A + B sequences, lane 3 A + B + C sequences, lane 4 A + B + C + D sequences (DPN). b 2 μM DPN was incubated in rat serum and its stability was analyzed by gel electrophoresis at the different times. Lane 1, 0 h; lane 2, 1.5 h; lane 3, 3 h
Optimization of detection conditions
Optimum concentration of PG
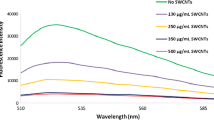
To obtain the optimum concentration of PG for complete reaction with DPN, different amounts of PG were added to a fixed concentration of DPN. As indicated in Fig. 3a, 10 μL 20× PG afforded the maximum fluorescence intensity, indicating that the maximum conjugation of PG with DPN happened at this concentration.
Parameters involved in the fluorescence intensity of the aptasensor. a Fluorescence spectra of DPN in the presence of various concentrations of PG (2, 5, 10, and 20 μL 20× from bottom to top). b Fluorescence spectra of DPN as a function of PG incubation time (1, 2, 5, 7, and 10 min from bottom to top)
Optimum incubation time of PG
Upon the addition of PG to DPN, the fluorescence intensity was monitored up to 10 min. The fluorescence intensity enhanced rapidly as the incubation time increased to 7 min (Fig. 3b), which indicated the required time for loading of PG onto DPN.
Quantification of OTA
Figure 4a shows the relative fluorescence intensity of the aptasensor at various concentrations of OTA. As the concentration of OTA increased, the relative fluorescence intensity increased until a maximum value was obtained at 10 nM. The analytical approach displayed a linear range (0.3–10 nM) toward target (Fig. 4b). The limit of detection (LOD) was calculated to be 0.135 nM (54.5 pg/mL), determined as three times the standard deviation of the blank/slope.
a Relative fluorescence intensity of the aptasensor as a function of OTA concentration. b OTA standard curve. F 0 and F are the fluorescence intensities at 530 nm before and after addition of OTA, respectively. c Relative fluorescence intensity of the fabricated aptasensor in the presence of different substances (10 nM) in the buffer. d Relative fluorescence intensity of the fabricated aptasensor in the presence of different substances in the grape juice. e Relative fluorescence intensity of the fabricated aptasensor in the presence of different substances (50 nM) in the buffer. F 0 and F are the fluorescence intensities at 530 nm before and after addition of each substance, respectively
Reported detection limits of OTA in other articles are shown in Table 2. Relative to the designed fluorescent aptasensor, most of these analytical techniques are expensive, laborious, and have higher LODs.
Specificity of the aptasensor
In addition to sensitivity, selectivity is an important factor for sensing applications. The relative fluorescence intensity of the assay toward OTA was significantly more than other materials (Fig. 4c–e). These results proved the high selectivity of the analytical assay toward OTA.
Detection of OTA in real samples
The aptasensor was utilized to detect OTA in grape juice and serum as a multiplex biological fluid with a blend of proteins and other interfering matrices. Rising amounts of OTA were spiked into serum and grape juice and LODs were determined to be 0.184 nM (74.3 pg/mL) and 0.149 nM (60.2 pg/mL), respectively (Fig. 5).
a Relative fluorescence intensity of the designed aptasensor upon the addition of various concentrations of OTA in serum. b OTA standard curve in serum. c Relative fluorescence intensity of the developed aptasensor upon the addition of various amounts of OTA in grape juice. d OTA standard curve in grape juice. F 0 and F are the fluorescence intensities at 530 nm before and after addition of OTA, respectively
The calculated LOD for grape juice was lower than the maximum permitted level of OTA (7.5 nM) stipulated by the European Commission [31]. These results confirmed the successful application of the designed fluorescent aptasensor for recognition of OTA in grape juice and serum.
Conclusion
A fluorescent aptasensor was developed for ultrasensitive and specific detection of OTA, based on the pyramid nanostructure of DPN, PG, and target-induced disassembly of the DPN structure. The measured LOD for OTA was as low as 0.135 nM. Furthermore, the designed aptasensor could recognize OTA in serum and grape juice with LODs of 0.184 and 0.149 nM, respectively.
References
Wang C, Qian J, Wang K, Liu Q, Dong X, Huang X. Magnetic-fluorescent-targeting multifunctional aptasensorfor highly sensitive and one-step rapid detection of ochratoxin A. Biosens Bioelectron. 2015;68:783–90. doi:10.1016/j.bios.2015.02.008.
Wei Y, Zhang J, Wang X, Duan Y. Amplified fluorescent aptasensor through catalytic recycling for highly sensitive detection of ochratoxin A. Biosens Bioelectron. 2015;65:16–22. doi:10.1016/j.bios.2014.09.100.
Sanzani SM, Reverberi M, Fanelli C, Ippolito A. Detection of ochratoxin a using molecular beacons and real-time PCR thermal cycler. Toxins. 2015;7(3):812–20. doi:10.3390/toxins7030812.
Yang M, Jiang B, Xie J, Xiang Y, Yuan R, Chai Y. Electrochemiluminescence recovery-based aptasensor for sensitive ochratoxin A detection via exonuclease-catalyzed target recycling amplification. Talanta. 2014;125:45–50. doi:10.1016/j.talanta.2014.02.061.
Zhu Z, Feng M, Zuo L, Wang F, Chen L, Li J, et al. An aptamer based surface plasmon resonance biosensor for the detection of ochratoxin A in wine and peanut oil. Biosens Bioelectron. 2015;65:320–6. doi:10.1016/j.bios.2014.10.059.
Chen J, Zhang X, Cai S, Wu D, Chen M, Wang S, et al. A fluorescent aptasensor based on DNA-scaffolded silver-nanocluster for ochratoxin A detection. Biosens Bioelectron. 2014;57:226–31. doi:10.1016/j.bios.2014.02.001.
Lv L, Cui C, Liang C, Quan W, Wang S, Guo Z. Aptamer-based single-walled carbon nanohorn sensors for ochratoxin A detection. Food Control. 2016;60:296–301. doi:10.1016/j.foodcont.2015.08.002.
Wang C, Dong X, Liu Q, Wang K. Label-free colorimetric aptasensor for sensitive detection of ochratoxin A utilizing hybridization chain reaction. Anal Chim Acta. 2015;860:83–8. doi:10.1016/j.aca.2014.12.031.
Du F, Guo L, Qin Q, Zheng X, Ruan G, Li J, et al. Recent advances in aptamer-functionalized materials in sample preparation. TrAC Trends Anal Chem. 2015;67:134–46. doi:10.1016/j.trac.2015.01.007.
Xiang D, Shigdar S, Qiao G, Wang T, Kouzani AZ, Zhou SF, et al. Nucleic acid aptamer-guided cancer therapeutics and diagnostics: the next generation of cancer medicine. Theranostics. 2015;5(1):23–42. doi:10.7150/thno.10202.
Dong Y, Xu Y, Yong W, Chu X, Wang D. Aptamer and its potential applications for food safety. Crit Rev Food Sci Nutr. 2014;54(12):1548–61. doi:10.1080/10408398.2011.642905.
Zhou W, Jimmy Huang PJ, Ding J, Liu J. Aptamer-based biosensors for biomedical diagnostics. Analyst. 2014;139(11):2627–40. doi:10.1039/c4an00132j.
Eissa S, Siaj M, Zourob M. Aptamer-based competitive electrochemical biosensor for brevetoxin-2. Biosens Bioelectron. 2015;69:148–54. doi:10.1016/j.bios.2015.01.055.
Lian Y, He F, Wang H, Tong F. A new aptamer/graphene interdigitated gold electrode piezoelectric sensor for rapid and specific detection of Staphylococcus aureus. Biosens Bioelectron. 2015;65:314–9.
Zhao B, Wu P, Zhang H, Cai C. Designing activatable aptamer probes for simultaneous detection of multiple tumor-related proteins in living cancer cells. Biosens Bioelectron. 2015;68:763–70. doi:10.1016/j.bios.2015.02.004.
Abnous K, Danesh NM, Ramezani M, Emrani AS, Taghdisi SM. A novel colorimetric sandwich aptasensor based on an indirect competitive enzyme-free method for ultrasensitive detection of chloramphenicol. Biosens Bioelectron. 2016;78:80–6. doi:10.1016/j.bios.2015.11.028.
Mohammad Danesh N, Ramezani M, Sarreshtehdar Emrani A, Abnous K, Taghdisi SM. A novel electrochemical aptasensor based on arch-shape structure of aptamer-complimentary strand conjugate and exonuclease I for sensitive detection of streptomycin. Biosens Bioelectron. 2016;75:123–8. doi:10.1016/j.bios.2015.08.017.
Ramezani M, Mohammad Danesh N, Lavaee P, Abnous K, Mohammad TS. A novel colorimetric triple-helix molecular switch aptasensor for ultrasensitive detection of tetracycline. Biosens Bioelectron. 2015;70:181–7. doi:10.1016/j.bios.2015.03.040.
Ramezani M, Danesh NM, Lavaee P, Abnous K, Taghdisi SM. A selective and sensitive fluorescent aptasensor for detection of kanamycin based on catalytic recycling activity of exonuclease III and gold nanoparticles. Sens Actuators B Chem. 2016;222:1–7. doi:10.1016/j.snb.2015.08.024.
Pang Y, Rong Z, Wang J, Xiao R, Wang S. A fluorescent aptasensor for H5N1 influenza virus detection based-on the core-shell nanoparticles metal-enhanced fluorescence (MEF). Biosens Bioelectron. 2015;66:527–32. doi:10.1016/j.bios.2014.10.052.
Mishra RK, Hayat A, Catanante G, Istamboulie G, Marty JL. Sensitive quantitation of ochratoxin A in cocoa beans using differential pulse voltammetry based aptasensor. Food Chem. 2016;192:799–804. doi:10.1016/j.foodchem.2015.07.080.
Wang R, Xiang Y, Zhou X, Liu LH, Shi H. A reusable aptamer-based evanescent wave all-fiber biosensor for highly sensitive detection of ochratoxin A. Biosens Bioelectron. 2015;66:11–8. doi:10.1016/j.bios.2014.10.079.
Song Q, Peng M, Wang L, He D, Ouyang J. A fluorescent aptasensor for amplified label-free detection of adenosine triphosphate based on core–shell Ag@SiO2 nanoparticles. Biosens Bioelectron. 2016;77:237–41. doi:10.1016/j.bios.2015.09.008.
Chen Z, Tan L, Hu L, Luan Y. Superior fluorescent probe for detection of potassium ion. Talanta. 2015;144:247–51. doi:10.1016/j.talanta.2015.06.015.
Wu S, Zhang H, Shi Z, Duan N, Fang C, Dai S, et al. Aptamer-based fluorescence biosensor for chloramphenicol determination using upconversion nanoparticles. Food Control. 2015;50:597–604.
Yang C, Wang Q, Xiang Y, Yuan R, Chai Y. Target-induced strand release and thionine-decorated gold nanoparticle amplification labels for sensitive electrochemical aptamer-based sensing of small molecules. Sens Actuators B Chem. 2014;197:149–54.
Liu XP, Deng YJ, Jin XY, Chen LG, Jiang JH, Shen GL, et al. Ultrasensitive electrochemical immunosensor for ochratoxin A using gold colloid-mediated hapten immobilization. Anal Biochem. 2009;389(1):63–8. doi:10.1016/j.ab.2009.03.019.
Yu FY, Chi TF, Liu BH, Su CC. Development of a sensitive enzyme-linked immunosorbent assay for the determination of ochratoxin A. J Agric Food Chem. 2005;53(17):6947–53. doi:10.1021/jf0513922.
Majdinasab M, Sheikh-Zeinoddin M, Soleimanian-Zad S, Li P, Zhang Q, Li X, et al. A reliable and sensitive time-resolved fluorescent immunochromatographic assay (TRFICA) for ochratoxin A in agro-products. Food Control. 2015;47:126–34. doi:10.1016/j.foodcont.2014.06.044.
Taghdisi SM, Danesh NM, Beheshti HR, Ramezani M, Abnous K. A novel fluorescent aptasensor based on gold and silica nanoparticles for the ultrasensitive detection of ochratoxin A. Nanoscale. 2016;8(6):3439–46. doi:10.1039/c5nr08234j.
Yang X, Qian J, Jiang L, Yan Y, Wang K, Liu Q, et al. Ultrasensitive electrochemical aptasensor for ochratoxin A based on two-level cascaded signal amplification strategy. Bioelectrochemistry. 2014;96:7–13.
Acknowledgments
Financial support of this study was provided by Mashhad University of Medical Sciences.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
There is no conflict of interest about this article.
Additional information
Morteza Alinezhad Nameghi and Noor Mohammad Danesh contributed equally to this work.
Rights and permissions
About this article
Cite this article
Nameghi, M.A., Danesh, N.M., Ramezani, M. et al. A fluorescent aptasensor based on a DNA pyramid nanostructure for ultrasensitive detection of ochratoxin A. Anal Bioanal Chem 408, 5811–5818 (2016). https://doi.org/10.1007/s00216-016-9693-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9693-7