Abstract
We established an optimized biofilm observation method using a hydrophilic ionic liquid (IL), 1-butyl-3-methylimidazolium tetrafluoroborate ([BMIM][BF4]). In the present study, a biofilm was formed by Staphylococcus epidermidis. Using field emission (FE) scanning electron microscopy (SEM) and transmission electron microscopy (TEM), the colonization of assemblages formed by microbial cells was observed as a function of the cultivation time. FE-TEM analysis revealed that the fibril comprises three types of protein. In addition, the ultrastructure of each protein monomer was visualized. It was expected that the curly-structured protein plays an important role in extension during fibril formation. Compared to the conventional sample preparation method for electron microscopy, a fine structure was easily obtained by the present method using IL. This observation technique can provide valuable information to characterize the ultrastructure of the fibril and biofilm that has not been revealed till date. Furthermore, these findings of the molecular architecture of the fibril and the colonization behavior of microbial cells during biofilm formation are useful for the development of antibacterial drugs and microbial utilization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteria are well known for their adhesive behavior on solid surfaces to form an optimal environment for growth. A biofilm is an assemblage of surface-associated microbial cells that is enclosed in an extracellular polymeric substance (EPS) matrix [1, 2]. They can cause infections and contaminations because it is irreversibly associated with the surface and enclosed in a matrix of primarily polysaccharide material [3, 4]. Hence, many researchers focused on microbial adhesion [5, 6]. It has been reported that EPS and filamentous cell appendages facilitate bacterial adhesion between a cell and its surface [7–11]. Therefore, we need to understand the roles of filamentous cell appendages and EPS to reveal the mechanism of biofilm formation. Especially, Staphylococcus epidermidis is well known as a commensal microorganism of human skin, nares, and mucous membranes [12, 13]. Although fibrillar structures of S. epidermidis have been observed [11, 14, 15], observations of the ultrastructure of fibrils and detailed information on biofilm formation have not been reported till date.
Electron microscopes are widely used as important tools of material characterization because of technological advances, particularly in high-resolution image, analysis equipment, and 3D observation abilities [16, 17]. Especially in the inorganic materials research field, the spherical-aberration-correction (Cs + correction) technique of the objective lens in a transmission electron microscope (TEM) has opened up a new field in high-resolution imaging [18, 19]. Even single atom images were obtained using the aberration-corrected TEM and STEM [20]. In the biological materials research field, only some researchers reported high-resolution image using TEM [21–23]. However, the preparation method of biological samples requires special techniques and is time consuming. Furthermore, it has been reported that glutaraldehyde fixation produces artifacts. Therefore, to observe the actual morphologies of hydrous samples using a simple technique, new methodologies have been extensively searched.
Room-temperature (RT) ionic liquids (ILs) have attracted significant attention as novel, eco-friendly materials. These ILs comprise cations and anions, which exist in a liquid state at RT [24]. Because of their negligible vapor pressure and high conductivity, various studies have also revealed that combinations of ILs with samples can be directly observed by electron microscopy without the addition of a conductive coating [25, 26]. On the other hand, the observation of biological materials by electron microscopy without structural deformation is a real challenge. In this regard, combining ILs with biological materials can be the most simple and attractive approach to reveal the morphological structure by electron microscopy [27, 28]. We found that the osmotic pressure of biological materials needs to be maintained to observe the fine morphology [29]. However, TEM observations of biological materials using ILs have not been reported till date. Therefore, it is necessary to develop an optimized method for biological materials, which is applicable to both scanning electron microscopy (SEM) and TEM observations.
In the present study, we investigated the simple observation method for S. epidermidis with the aid of IL using field emission (FE)-TEM and FE-SEM. In addition, we visualized the protein structure of a fibril using this optimized observation technique.
Materials and methods
Materials
The bacterial strain used in this study was S. epidermidis PAGU283 (=ATCC14990T). This strain was stored at −80 °C and routinely grown at 37 °C for 24 h in 0.5 % CO2 in tryptone soy broth supplemented with 0.25 % glucose (Becton, Dickinson and Company Co., USA). Subsequently, the absorbance of the S. epidermidis medium was adjusted to 0.01 or 0.2 using a UV/visible spectrophotometer at a wavelength of 520 nm (Ultrospec 2100 pro, GE Healthcare Life Sciences Co., UK). It should be noted that the absorbance was adjusted to 0.2 for formation of the biofilm shown in Figs. 2e, f; 3d; and 4a, b, and to 0.01 for all other biofilms. S. epidermidis was grown in 24-well plates and incubated for 12, 16, and 24 h at 37 °C. IL used for electron microscopic observation was 1-butyl-3-methylimidazolium tetrafluoroborate ([BMIM][BF4]), which was purchased from Kanto Chemical Co., Japan, and dried in a vacuum desiccator for 1 day. The water content of IL was 128 ppm.
Structural observations of biofilms using FE-SEM and TEM
The surface morphology of biofilms was observed using FE-SEM (JEOL Co., JXA-8530FA, Japan) with an accelerating voltage of 10.0 kV. The internal structure of biofilms was observed using FE-TEM (JEOL Co., JEM 2100F, Japan) with an accelerating voltage of 120.0 kV. The S. epidermidis biofilm, prepared as described in “Materials,” was characterized using FE-SEM and FE-TEM.
The biofilm formed in a well was washed with purified water. Subsequently, the biofilm was treated with phosphate-buffered saline (PBS) buffer solution for 2 h. The biofilm was washed with water and treated with IL solution for 5 min. Following this, it was scraped using a cell scraper. The IL solution was prepared at an IL to water weight ratio of 1:200. The biofilm for FE-SEM and FE-TEM observations was skimmed off by a copper mesh with carbon-coated plastic microholes for organic samples (HRC-C10, Okenshoji Co., Japan). The biofilm sample was then placed in a centrifuge at 10,000 rpm and maintained under vacuum conditions for 30 min. The detailed methodology is illustrated in Fig. 1. For comparison, the conventional preparation method for FE-TEM observation was performed. The details of the preparation method have been described in the Electronic Supplementary Material (ESM)
Results and discussion
Morphological observation of biofilm formation
We studied the S. epidermidis biofilms prepared at different incubation times. These biofilms were prepared combing with IL following pretreatment using the developed method, as shown in Fig. 1. The morphologies of the biofilms incubated at 37 °C for 12, 16, and 24 h are shown in Fig. 2a–f. After 12 h of cultivation, the biofilm comprised bacteria with sizes around 1 μm. Each bacterium could be distinguished although the cells were adhered to each other (Fig. 2b). The spherical shapes of the S. epidermidis cells changed to angular shapes during the biofilm formation process. Some holes can be observed in the biofilm morphology. In this process, evolution into the 3D structure was not drastically proceeded with. It is well known that the EPS is essential for the cell, device, implant, etc. to adhere. Moreover, polysaccharide intercellular adhesion (PIA) synthesized by ica gene products has important roles as the major functional component and in the accumulation of S. epidermidis biofilms [30, 31]. Another exopolymer related to the accumulation is protein. We will discuss about such typed proteins in “Structural observation of the molecular architecture of the fibril.” Figure 2c, d shows FE-SEM images of the biofilms after 16 h of cultivation. It can be clearly observed that primary polysaccharide materials were formed on the surface of the bacteria and the boundary between a cell and another cell disappeared. We also observed the biofilm morphology after 24 h of cultivation (Fig. 2e). In contrast to the morphologies shown in Fig. 2a–d, an agglomerated morphology was observed. In this process, a 3D structure was formed during biofilm maturation. A thick film, which comprised EPS-enclosed polysaccharide, covered the surface of the bacteria. Thus, 1-μm-sized cells were not observed on the surface of the biofilm incubated for 24 h (Fig. 2f).
FE-SEM images of biofilm of S. epidermidis prepared after different incubation times following pretreatment, as shown in Fig. 1. The biofilms were incubated at 37 °C for 12 h (a, b), 16 h (c, d), and 24 h (e, f)
Figure 3 shows the fine morphology of the biofilms from selected areas (designated as i–iv in Fig. 2). It should be noted here that some morphological changes occurred when we observed the samples at a high magnification for a long time. It was caused by the radiation damage of the electron beam. However, we can reduce the damage using osmium coating. In the present study, we could observe the fine morphology without any conductive coating. The process of biofilm formation according to the cultivation time can be clearly observed. In Fig. 3a, rod-like structures with a diameter of <100 nm were observed. Franson et al. reported the presence of “foot” processes sized 100 nm between the cell and the substrate [1]. In addition, Dubey et.al visualized nanotubes in neighboring cells [32]. The structures of these “foot” processes and nanotubes were similar to the rod-like structure as shown in Fig. 3a. The nanotubes associated with cytoskeletal and motor proteins play like an active transport [33]. In Fig. 3b, the voids between bacteria cells were filled primarily by polysaccharide materials, and the surface roughness of the biofilm was drastically increased. These phenomena proceed further as shown in Fig. 3c. In Fig. 3c, a fine-sized EPS-enclosed polysaccharide was observed on the surface of the biofilm. Characklis et al. have reported that the surface roughness of the biofilm increases as microbial colonization proceeds [34]. Our results are in good agreement with these findings. In Fig. 3d, some holes <500 nm in size still remained on the biofilm. These FE-SEM results indicate that fine morphologies can be clearly observed using hydrophilic IL without any conductive coating. Ishizaki et al. have reported that a simple technique using hydrophilic IL enables the observation of the fine thread-like morphologies of cultured human cells [28]. In good agreement with this report, we observed fine morphologies of filamentous fibrils using an IL. It can be considered that the simple observation method using hydrophilic IL will be useful for observing the filamentous morphology of biological materials. It should be noted here that many fibrils were observed on the surface of bacteria (ESM Fig. S1). It can be seen that the fibrils anchored to the cell grow toward another cell. The fibrils function as a ladder from one cell to another, linking the cells. To further characterize the ultrastructure of fibrils derived from S. epidermidis, we will discuss the details in “Structural observation of the molecular architecture of the fibril.”
FE-SEM images of the biofilms from selected areas (designated i–iv) showed in Fig. 2. Areas i, ii, iii, and iv correspond to a, b, c, and d, respectively
In order to understand the inner part of the biofilm, we performed cross-sectional observation. Figure 4 shows FE-SEM images of the biofilm formed after 24 h of cultivation. Bacterial cells anchored on the surface of the biofilm and buried under EPS-enclosed polysaccharide were observed. The adhered bacteria were linked to the biofilm and existed as an agglomerate (Fig. 4a). The shapes of the cells transformed from a spherical form to a flat form. Figure 4b shows the cross-section image of Fig. 4a. The EPS product wall measuring several hundred nanometers was clearly observed on the surface of the biofilm, and collapsed cells surrounded by EPS-enclosed polysaccharide were observed in the inner area of the biofilm agglomerate. Interstitial voids were present in the inner area of the biofilm covered by the EPS product wall. It can be considered that water channels exist in the EPS matrix and cells. These water channels are well known to play important roles in liquid flow, allowing the diffusion of nutrients and oxygen. However, the nanostructure of the fibril was not clearly observed using FE-SEM. Therefore, we attempted to reveal the fibril ultrastructure using TEM. Furthermore, we tried to observe the internal structure of the biofilm to understand the biofilm formation process.
a Conventional FE-SEM image and b cross-sectional FE-SEM image of biofilm after 24 h of cultivation pretreated with new method according to Fig. 1
Structural observation of biofilm formation
Figure 5 shows FE-TEM images of the biofilm that was formed after 12 and 16 h of cultivation with IL treatment and kept under vacuum conditions for 30 min. The details of the preparation method are described previously in Fig. 1. After 12 h of cultivation, cells with a size of 1 μm formed a microcolony structure (Fig. 5a). After 16 h of cultivation, a slime-like structure was observed around the cells. It can be considered that the agglomerated slime-like structure is derived from EPS-enclosed polysaccharide. Moreover, fine EPS products were observed around S. epidermidis cells (indicated by arrows). The EPS products were <200 nm in size, and they were partially observed. The size of the EPS products was very similar to that shown in Fig. 4a, b. As the cultivation time increased, EPS production drastically increased to cover the bacterial cells. In Fig. S1 in the ESM, the morphology of the fibril was observed using FE-SEM. To understand further the fibril structure and role, TEM study was also performed. Figure 5c–f shows the process of fibril adhesion to a cell. After 12 h of cultivation, the biofilm was observed in four selected areas (Fig. 5c–f). Figure 5c shows some fibrils that were <200 nm in length. The shaft was approximately 50 nm in width. On the other hand, a 2.5-μm-long fibril with a 30-nm-wide shaft was also observed (Fig. 5d). Figure 5e shows S. epidermidis fibrils adhered to another cell via thin adhesion threads. These fibrils extended straight from one cell to another without branching. Figure 5f shows the fibrils attached to the biofilm formed in another area. Interestingly, these fibrils were formed by several threads with diameters <30 nm (indicated by arrows). The cells bound via several threads formed a wrinkled structure. It is assumed that this wrinkled structure is caused by the formation of several fibrils. EPS-enclosed polysaccharide within cells may be used to form the bundle structure of the fibrils. It is a similar phenomenon to that observed in Fig. 2d.
FE-TEM images of the biofilm formed after a 12 h and b 16 h of cultivation following pretreatment using the new method, as shown in Fig. 1. FE-TEM images of fibrils attached to cells formed after 12 h of cultivation. c Initial stage of the adhesion process, d attachment of a long fibril, e attachment of variable numbers of short fibril, and f fibril formed by several threads. Scale bar 50 nm
To further understand the biofilm formation process on the basis of the bundle structure, the attachment region between a cell and another cell was observed carefully. Figure 6 shows FE-TEM images of the biofilm after 12 h of cultivation. The bundled bound substances between cells were clearly observed. The shafts were 100–200 nm in width (indicated by arrows). It should be noted that there is a mesh grid at the back of the cells in the upper right area of Fig. 6a (indicated by lines). Figure 6b shows a magnified image of area i in Fig. 6a. Some rod-like substances were observed between the cells. It can be considered that the fibril, which existed as a rod-like structure involving the EPS-enclosed polysaccharide, was bundled. The contrast of the rod-like structure was not constant. Thereby, it was expected that a rod-like structure is not solid. The surface morphology of the cell and fibril could be observed using FE-SEM (Fig. 3). In addition, TEM analysis determined the internal structure of the fibril. From these observations, it seems that the rod-like structure has a tubular structure. It is considered that the rod-like structure can work as an active transport. The correct structure was elucidated using a combination of both types of electron microscopy. In the present study, we tried to observe the fibrils using the conventional method for TEM observation in order to compare the new method using a hydrophilic IL. TEM images derived from the conventional method showed clearly inner structures of cells (ESM Fig. S2). However, the structures of fibrils were not clearly observed, although some fibrils were found between the cells.
a FE-TEM image of substances bound between cells after 12 h of cultivation following pretreatment using the new method, as shown in Fig. 1. b High-magnification image of selected area i
Structural observation of the molecular architecture of the fibril
To obtain further structural information and determine the molecular architecture of S. epidermidis, we observed the ultrastructure of the fibril by using FE-TEM. Figure 7a shows a TEM image of the fibril. It was confirmed that the length of the fibril was 340 nm and the width of the shaft was approximately 20 nm. The fibril had bamboo-like structures in the middle region of the shaft. However, the part that adhered to the cell wall and the tip of the fibril lacked ring-shaped structures. The high-magnification images show the ultrastructure of the fibril. It was clearly observed that the fibril was anchored to the cell wall (Fig. 7b). The structure was changed 80 nm from the bottom of the fibril (Fig. 7c). From 80 nm toward the tip side, ring-shaped structures were observed (Fig. 7c, d). Vesicle structures that were 10–20 nm in diameter were observed inside the fibril structure. This is the first observation of protein monomers at such high magnification. It can be considered that each protein was tightly bound to the next protein. The tip side was formed by a circle-shaped protein with a diameter of 20 nm. The orientation of the protein at the tip end was different from that of the shaft proteins at the bottom and middle regions of the fibril. It can be considered that this protein is responsible for adhesion. The fibril just beginning to be formed was also observed (Fig. 7f). The length of the fibril in Fig. 7f was approximately 50 nm, and the width of the shaft was approximately 20 nm. This can be because the bottom of the fibril (approximately 80 nm in diameter) has a different protein arrangement than the middle region of the fibril, as described above. The tip of the fibril was formed by structures that appeared curled. It is expected that these structures would extend during fibril formation.
Molecular architecture of fibril after 12 h of cultivation following pretreatment using the new method, as shown in Fig. 1. a Combined FE-TEM image of a fibril (approximately 18∼22 proteins), b high-magnification image of a protein subunit anchored to the cell wall surface, c protein subunit anchored to the middle region of a fibril, d protein subunit at the middle region of a fibril, e protein subunit at the tip of a fibril, and f another fibril anchored to the cell wall
The fibrillar structures of S. epidermidis were reported by some researchers [11, 15]. It was found that Aap is the essential component of the fibril and accumulation-associated protein [11, 15, 35]. Furthermore, Fbe, AtlE, Bap, Embp, and Bhp proteins have been reported as proteinaceous intercellular adhesins in biofilm accumulation [36–39]. Aap is a cell wall-anchored protein with an N-terminal signal sequence and a C-terminal sorting signal including an LPXTG motif followed by a hydrophobic transmembrane region and a positively charged cytoplasmic tail. Moreover, it was reported that Aap is fibrillar adhesion [15]. Nelson et.al reported that fibrinogen-binding protein Fbe and accumulation-associated protein Aap play major roles in the adherence of cells to accumulative growth on the polymer surface [14].
In the present study, the three types of protein structure regions of the fibril were revealed with the aid of a novel technique using IL. It can be expected that the curled protein was derived from Aap protein and mainly fibrillar structure comprise Aap and Fbe proteins. Moreover, Embp protein has extracellular matrix-binding properties [38]. Thus, the slime-like layer can be formed related to Embp protein (Fig. 5).
We successfully observed the ultrastructure of the fibril under high magnification using IL. Compared to the conventional method, the protein formation of the fibril was clearly observed. Although field emission-type TEM was used for characterization, electron beam damage was not observed. On the other hand, the individual protein structure was not found using the conventional method (ESM Fig. S2c). It should be noted here that excess IL around the biofilm was removed using a centrifuge. In addition, PBS buffer was used for sample preparation. When the sample was prepared without treatment of the PBS buffer, the biofilm was partially broken during the sample preparation. Normally, conventional processing of biological materials for TEM observation takes more than a few days and consists of fixation with glutaraldehyde, dehydration, resin embedding, slicing of sections, and counterstaining. The new method enables to reduce preparation time. The new method using IL takes only 3 h because processes such as fixation, dehydration, resin embedding, slicing of sections, and counterstaining are skipped. Therefore, we can observe the ultrastructure of bacteria without any technique and specific apparatus. The information on fibril structure obtained in the present study will help in the understanding of the development of biofilms by aggregation or coaggregation.
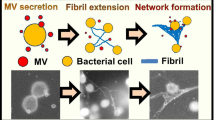
Conclusions
We have successfully observed the ultrastructure of S. epidermidis with the help of electron microcopy techniques using a hydrophilic IL [BMIM][BF4]. Furthermore, the complex process of biofilm formation was elucidated by visualizing biofilms after different cultivation times. In particular, the fibril, which plays an important role in cell adhesion, was characterized using FE-SEM and FE-TEM. Our TEM study confirmed that the fibril has a protein with a bamboo-like-structured protein at the middle region. We can summarize the mechanism of biofilm formation as follows: (1) a fibril derived from a cell extends toward another cell and subsequently adheres to the surface of the other cell; (2) the fibril forms a bridge between cells; (3) bundle structures consisting of fibril and EPS-enclosed polysaccharide are formed; (4) EPS product sized 200 nm accumulates on the surface of the cell and between cells; and (5) film-like structured EPS product covers the cell agglomerates. Consequently, this electron microscopic observation using an optimization method for sample preparation enables the observation of the ultrastructure of biological materials. It is concluded that compared with the conventional technique, the time required for sample preparation can be drastically reduced using the optimization method. The structural observation of the fibril in this study will help reveal the model of the fibril of S. epidermidis. Moreover, the findings related to the mechanism of biofilm formation and the fibril structure can be very significant for the development of a vaccine and microbial utilization. The simple optimization method using IL developed in the present study can be widely applied to characterize biological materials such as microorganisms, plants, and animals without any special apparatus.
References
Franson TR, Sheth NK, Rose HD, Sohnle PG (1984) Scanning electron microscopy of bacteria adherent to intravascular catheters. J Clin Microbiol 20:500–505
Kolari M, Schmidt U, Kuismanen E, Salkinoja-Salonen MS (2002) Firm but slippery attachment of Deinococcus geothermalis. J Bacreriol 184:2473–2480
Donlan RM (2002) Biofilms and device-associated infections. Emerg Infect Dis 7:277–281
Clauss M, Trampuz A, Borenz O, Bohner M, Ilchmann T (2010) Biofilm formation on bone grafts and bone graft substitutes: comparison of different materials by a standard in vitro test and microcalorimetry. Acta Biomater 6:3791–3797
Dankert J, Hogt AH, Feijen J (1986) Biomedical polymers: bacterial adhesion, colonization, and infection. Crit Rev Biocompat 2:219–301
An YH, Friedman RJ (1998) Concise review of mechanisms of bacterial adhesion to biomaterial surfaces. J Biomed Mater Res 43:338–348
Ishii S, Koki J, Unno H, Hori K (2004) Two morphological types of cell appendages on a strongly adhesive bacterium, Acinetobacter sp. strain Tol 5. Appl Environ Microbiol 70:5026–5029
Schmid T, Burkhard J, Yeo BS, Zhang W, Zenobi R (2008) Towards chemical analysis of nanostructures in biofilms I: imaging of biological nanostructures. Anal Bioanal Chem 391:1899–1905
Mandlik A, Swierczynski A, Das A, Ton-That H (2008) Pili in gram-positive bacteria: assembly, involvement in colonization and biofilm development. Trends Microbiol 16:33–40
Hilleringmann M, Ringler P, Muller SA, Angelis GD, Rappuoli R et al (2009) Molecular architecture of Streptococcus pneumoniae TIGR4 pili. EMBO J 28:3921–3930
Banner MA, Cunniffe JG, Macintosh RL, Foster TJ, Rohde H et al (2007) Localized tufts of fibrils on Staphylococcus epidermidis NCTC 11047 are comprised of the accumulation-associated protein. J Bacteriol 189:2793–2804
Mack D, Siemssen N, Laufs R (1992) Parallel induction by glucose of adherence and a polysaccharide antigen specific for plastic-adherent Staphylococcus epidermidis: evidence for functional relation to intercellular adhesion. Infect Immun 60:2048–2057
Otto M (2009) Staphylococcus epidermidis—the ‘accidental’ pathogen. Nat Rev Microbiol 7:555–567
Nelson A, Hultenby K, Hell E, Riedel HM, Brismar H et al (2009) Staphylococcus epidermidis isolated from newborn infants express pilus-like structures and are inhibited by the cathelicidin-derived antimicrobial peptide LL37. Pediatr Res 66:174–178
Macintosh RL, Brittan JL, Bhattacharya R, Jenkinson HF, Derrick J et al (2009) The terminal A domain of the fibrillar accumulation-associated protein (Aap) of Staphylococcus epidermidis mediates adhesion to human corneocytes. J Bacteriol 191:7007–7016
Saka H, Mima T, Takeuchi Y, Marukawa T, Arai S et al (2006) Observation of gas-solid and gas-liquid reactions by in-situ environmental holder in TEM. Microsc Microanal 12:764–765
Takahashi C, Yaakob Y, Yusop MZM, Kalita G, Tanemura M (2014) Direct observation of structural change in Au-incorporated carbon nanofibers during field emission process. Carbon 75:277–280
Haider M, Uhlemann S, Schwan E, Rose H, Kabius B, Urban K (1998) Electron microscopy image enhanced. Nat 392:768–769
Yamasaki J, Kawai T, Tanaka N (2004) Direct observation of a stacking fault in Si1-xGex semiconductors by spherical aberration-corrected TEM and conventional ADF-STEM. J Electron Microsc 53:129–135
Mangel S, Aronovitch E, Enyashin AN, Houben L, Bar-Sadan M (2014) Atomic-scale evolution of a growing core-shell nanoparticle. J Am Chem Soc 136:12564–12567
Zakharov VV, Mosevitsky MI (2010) Oligomeric structure of brain abundant proteins GAP-43 and BASP1. J Struct Biol 170:470–483
Mahmoudi M, Shokrgozar MA, Sardari S, Moghadam MK, Vali H, Laurentc S, Stroeved P (2011) Irreversible changes in protein conformation due to interaction with superparamagnetic iron oxide nanoparticles. Nanoscale 3:1127–1138
Wang Y, Xu J, Wanga Y, Chen H (2013) Emerging chirality in nanoscience. Chem Soc Rev 42:2930–2962
Wilkes JS, Zaworotko MJ (1992) Air and water stable 1-ethyl-3-methylimidazolium based ionic liquids. J Chem Soc Chem Commun 965–967
Torimoto T, Okazaki K, Kiyama T, Hirahara K, Tanaka N et al (2006) Sputter deposition onto ionic liquids: simple and clean synthesis of highly dispersed ultrafine metal nanoparticles. Appl Phys Lett 89:243117
Takahashi C, Shirai T, Fuji M (2012) Observation of interactions between hydrophilic ionic liquid and water on wet agar gels by FE-SEM and its mechanism. Mater Chem Phys 133:565–572
Kuwabata S, Tsuda T, Torimoto T (2010) Room-temperature ionic liquids. A new medium for material production and analyses under vacuum conditions. J Phys Chem Lett 1:3177–3188
Ishigaki Y, Nakamura Y, Takehara T, Nemoto N, Kurihara T et al (2011) Comparative study of hydrophilic and hydrophobic ionic liquids for observing cultured human cells by scanning electron microscopy. Microsc Res Tech 74:415–420
Takahashi C, Shirai T, Fuji M (2013) FE-SEM observation of swelled seaweed using hydrophilic ionic liquid; 1-butyl-3-methylimidazolium tetrafluoroborate. Microsco Res Tech 76:66–71
Heilmann C, Schweitzer O, Gerke C, Vanittanakom N, Mack D et al (1996) Molecular basis of intercellular adhesion in the biofilm-forming Staphylococcus epidermidis. Mol Microbiol 20:1083–1091
O’Gara JP (2007) Ica and beyond: biofilm mechanisms and regulation in Staphylococcus epidermidis and Staphylococcus aureus. FEMS Microbiol Lett 270:179–188
Dubey GP, Yehuda SB (2011) Intercellular nanotubes mediate bacterial communication. Cell 144:590–600
Davis DM, Sowinski S (2008) Membrane nanotubes: dynamic long-distance connections between animal cells. Nat Rev Mol Cell Biol 9:431–436
Characklis WG, McFeters GA, Marshall KC (1990) In: Characklis WG, Marshall KC (eds) Biofilms. Wiley, New York
Hussain M, Herrmann M, von Eiff C, Perdreau-Remington F, Peters G (1997) A 140-kilodalton extracellular protein is essential for the accumulation of Staphylococcus epidermidis strains on surfaces. Infect Immun 65:519–524
Pei L, Flock JI (2001) Functional study of antibodies against a fibrogenin-binding protein in Staphylococcus epidermidis adherence to polyethylene catheters. J Infect Dis 184:52–55
Potter A, Ceotto H, Giambiagi-Demarval M, dos Santos KR, Nes IF et al (2009) The gene bap, involved in biofilm production, is present in Staphylococcus spp. strains from nosocomial infections. J Microbiol 47:319–326
Christner M, Franke GC, Schommer NN, Wendt U, Wegert K et al (2010) The giant extracellular matrix-binding protein of Staphylococcus epidermidis mediates biofilm accumulation and attachment to fibronectin. Mol Microbiol 75:187–207
Mack D, Davies AP, Harris LG, Jeeves R, Pascoe B et al (2013) In: Moriarty F, Zaat SAJ, Busscher HJ (eds) Staphylococcus epidermidis in biomaterial-associated infections. Springer, New York
Acknowledgments
This study was partially supported by the research grant from the Institute of Pharmaceutical Life Sciences, Aichi Gakuin University. The authors are grateful to Dr. Y. Morita of Aichi Gakuin University, Japan, for useful discussions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 258 kb)
Rights and permissions
About this article
Cite this article
Takahashi, C., Kalita, G., Ogawa, N. et al. Electron microscopy of Staphylococcus epidermidis fibril and biofilm formation using image-enhancing ionic liquid. Anal Bioanal Chem 407, 1607–1613 (2015). https://doi.org/10.1007/s00216-014-8391-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-014-8391-6