Abstract
Using spike-in controls designed to mimic mammalian mRNA species, we used the quantitative reverse transcription polymerase chain reaction (RT-qPCR) to assess the performance of in vitro transcription (IVT) amplification process of small samples. We focused especially on the confidence of the transcript level measurement, which is essential for differential gene expression analyses. IVT reproduced gene expression profiles down to approximately 100 absolute input copies. However, a RT-qPCR analysis of the antisense RNA showed a systematic bias against low copy number transcripts, regardless of sequence. Experiments also showed that noise increases with decreasing copy number. First-round IVT preserved the gene expression information within a sample down to the 100 copy level, regardless of total input sample amount. However, the amplification was nonlinear under low total RNA input/long IVT conditions. Variability of the amplification increased predictably with decreasing input copy number. For the small enrichments of interest in typical differential gene expression studies (e.g., twofold changes), the bias from IVT reactions is unlikely to affect the results. In limited cases, some transcript-specific differential gene expression values will need adjustment to reflect this bias. Proper experimental design with reasonable detection limits will yield differential gene expression capability even between low copy number transcripts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Assaying global gene expression profiles from small samples, especially those below 10 ng total RNA down to the single-cell level (approximately 10 pg total RNA), is desirable for a variety of biological analyses [1]. It was previously reported that extraction-to-microarray processing produces samples that can only be quantified at the approximately 100,000 copy level [2]. We sought to characterize bias and variability in sample in vitro transcription (IVT) amplification, establish the linearity of amplification, and estimate confidence of the differential gene expression analyses of low copy number genes.
Amplification of most samples is necessary for gene expression profiling, whether the downstream analysis consists of quantitative reverse transcription polymerase chain reaction (RT-qPCR), high-throughput sequencing [3, 4], or microarrays [5]. Many amplification methods are available, including some PCR-based methods that give good to excellent sensitivity for small samples [5], but the most widely utilized method continues to be the in vitro transcription (“linear amplification”) protocol [6]. This protocol was rapidly adopted to amplify mRNA samples for microarrays without systematic study of potential biases on samples until Wang et al. and Baugh et al. [7, 8], who both found IVT produced acceptable qualitative and quantitative representation of gene expression levels despite some artifacts limiting low-abundance transcripts from being amplified. Characterizing amplification can be achieved using a sample with a known mRNA profile, allowing one to map pre- and post-amplification expression levels. The External RNA Control Consortium (ERCC) developed pools of synthetic mRNA with known profiles and copy numbers for spiking into samples to quantitatively assess performance of gene expression measurements [9, 10] and have shown utility in RT-qPCR comparisons between different systems [11]. The ERCC RNA spike-ins were employed here to investigate the degree to which the amplification is biasing sample profiles and how variability of observed amplification depends on the number of input copies (for both individual RNA species and small total RNA input).
Neither previous study characterizing IVT amplification found major problems or biases with the method, though modifications towards optimization were introduced [7, 8]. Later, Nygaard et al. discussed the limitations for small sample amplification [12]. What is noteworthy about this work was the shift in discussion from absolute sample size towards copy number detection. Briefly, reliable detection was limited in smaller sample sizes to the most abundant transcripts, and the overall amplification was nearly constant (10,000-fold) across approximately two orders of magnitude input total RNA concentration range (3 ng up to 115 ng). It was stated that there was a decrease in the probability of maintaining abundance profiles from small numbers of template in amplification, as determined by array experiments. From a global processing viewpoint for small samples and considering all inefficiencies, the two-round amplification and subsequent hybridization to the array surface appear to have a practical single transcript detection limit of approximately 100,000 input copies. Left unclear was which step(s) require(s) further optimization.
In this work, we examined the profiles of ERCC RNA spike-ins after one round of IVT amplification. In doing so, we determined the degree of post-IVT bias and/or uncertainty using a priori knowledge of the number of input copies for all species of interest. Also note that the second round of amplification was specifically excluded because there are additional effects which likely confound the species profiles generated from the first round, namely, biases that arise from random hexamer priming of the antisense RNA (aRNA) [13, 14]. RT-qPCR using hydrolysis (TaqMan®) probes enabled characterization of the aRNA and revealed nonlinearity (i.e., all transcripts were not equally amplified) in the IVT amplification from at least 107 input copies down to approximately 100 input copies, depending on the level of differential expression sought. We characterized the variability of aRNA levels as a function of the number of input copies. The results suggest that IVT provides accurate amplification for practically all sample sizes of interest, despite bias from the IVT nonlinearity. The variability in the data increases with decreasing input copy number for a given species in a way that appears predictable, following a “Horwitz Trumpet”-like profile [15]. This knowledge enables accurate estimation of confidence in low copy number differential gene expression analyses when appropriate numbers of replicates are incorporated into the experimental design.
Experimental
Certain commercial equipment, instruments, or materials are identified in this paper to foster understanding. Such identification does not imply recommendation or endorsement by the National Institute of Standards and Technology, nor does it imply that the materials or equipment identified are necessarily the best available for the purpose.
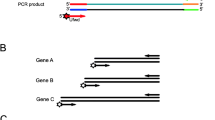
Messenger RNA was amplified using in vitro transcription then assayed using RT-qPCR. Purified total RNA from Jurkat T cells (Strategene) at 1 μg/μL (25 μg total) was used as the stock for the background of the sample. To this, 62.5 ng of mixed ERCC spike-in RNA controls [9, 10] (Electronic supplementary material) was added. Assuming mRNA levels are 2 % of the total RNA, the ERCC RNA amounts to approximately 12.5 % of the mRNA. The dynamic range of the studied ERCC RNA species was 213 (approximately 8,200) (Table S1, Electronic supplementary material). Sample sizes ranged from 10 ng to 1 pg. In vitro transcription of the total + ERCC RNA was achieved following the Ambion MessageAmp II® protocol (Life Technologies), with a 14-h incubation at 37 °C on a thermocycler (PTC-200, MJ Research), volumes scaled down 4× for the RT and second strand synthesis steps, with 5 to 10 μL IVT reactions. Post-IVT aRNA was purified using the kit protocols and concentrated using a SpeedVac (Thermo Scientific) to less than 25 μL, then diluted to 25 μL using nuclease-free water (Life Technologies).
The aRNA was characterized by TaqMan® RT-qPCR (Life Technologies, TaqMan® One-Step RT-PCR master mix reagents kit) with 10- or 20-μL reactions on an ABI 7900 (Life Technologies) (Electronic supplementary material). Primer pairs and hydrolysis probe designs were developed by Genomic Health (CA) and synthesized by IDT (Coralville, IA) and Biosearch Technologies (Novato, CA) (Table S1), respectively. For these assays, concentrations of 900 nmol/L for primers and 200 nmol/L of probes were used. A 1-ng total Jurkat T cell RNA + ERCC spike in RNA controls was used as the control for the reactions on all plates to monitor assay performance. In addition, we prepared and analyzed 10-fold dilutions of the Jurkat RNA + ERCC spike in RNA from 9 ng down to 0.0009 ng for every ERCC RNA transcript to confirm the linearity of the RT-qPCR assay. Every RT-qPCR assay was run in triplicate, and data were only included when all three replicates reported a threshold cycle value. Non-specific amplification was not observed from the TaqMan® assays using no-template controls. The quantitative cycle number (C q) values for the ERCC RNA showed no significant change over the course of study. Note that plotted data use log2 values of the input copy numbers rather than log10. Plots of C q vs. log2 (input copy number) should ideally give slopes of (−1). To convert the slope back to the traditional log10 value, multiply by 3.33.
The sensitivity of the RT-qPCR assay meets or exceeds the requirements for this study. We first assume single copy detection of DNA at C q values of approximately 36 from our experience with the ABI 7900. For typical IVT, a 103- to 104-fold amplification was expected, which would decrease C q by approximately (10 to 13) cycles. The aRNA were aliquoted into twenty-five 1-μL samples, increasing C q by approximately 4.6 cycles for each assay. Any efficiency reduction due to species-specific priming and reverse transcription also result in approximately one to four cycles higher C qs. Hence, the predicted required detection limit was at C q values between 30 and 33 cycles, which was within the capabilities of this RT-qPCR system.
Results and discussion
Linearity of IVT in the low copy number regime
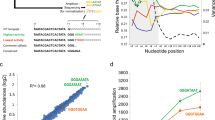
We analyzed C q values vs. the input copy number for each transcript to measure the linearity of the IVT reaction compared with non-IVT-amplified controls (Fig. 1 and Table 1). The data represent input copies from approximately 100 to 107 input copies, and input RNA amounts from approximately 0.1 pg to 10 ng of total Jurkat RNA with ERCC spike-ins. We found that each RT-qPCR assay had slightly different RT behavior (Table 1, intercepts) due to the different primers used with each RNA species; thus, all species cannot be plotted on a single line.
Threshold cycle vs. input copy number for six ERCC RNA species. Unamplified ERCC transcripts (blue plus signs with dashed blue line fit) were compared with IVT-amplified transcripts (various black shapes, with solid black line fit). Each amplified transcript follows the same slope with the input copy number and is significantly different than the unamplified control (see Table 1). This indicates that the IVT and PCR amplify without any statistically significant bias towards a particular species, but that IVT is significantly biased against low input copy number. The differences in intercepts of the linear fits indicate a bias in the reverse transcription step of the RT-qPCR due to the species-specific primer
First, the slopes of C q vs. log2(input copy number) of the non-IVT-amplified controls were −1, confirming that the RT-qPCR assay is not a source of bias. The slopes from the regressions of threshold cycle vs. log2(input copy number) for all IVT-amplified transcripts were statistically indistinguishable, supporting the hypothesis that IVT amplifies all cDNA species (i.e., sequences) the same. However, the amplification does not appear to be linear in this copy number regime with respect to input copy number. The average slope across all species of C q vs. log2(input copy number) was calculated using Graybill–Deal and DerSimonian–Laird, and was observed to be −1.24 ± 0.06 (average ± 95 % confidence interval) instead of the expected −1 for a linear relationship as observed for the controls (−1.04 ± 0.03).
The deviation from −1 implies a bias in amplification. The bias is small and not enough to drastically change the apparent expression profiles for most applications where small differences in expression profiles are studied (e.g., an observed twofold difference is actually only 1.7-fold), as other sources of error are likely to be more significant. The bias generates the most significant alterations of the measured transcript profiles over wide differences (more than an order of magnitude) of input copy numbers and especially between low and moderate-to-high copy number differences. This was in line with previous results [8] but not quantified.
Experimental evidence suggests that the low-abundance transcripts are inefficiently amplified by IVT compared with higher-abundance transcripts. The reason for this is unclear. The initial reverse transcription of mRNA using the T7-oligo(dT)24VN primer may be relatively inefficient, reducing the copy numbers of transcripts available for subsequent reactions. Because the priming is template independent, one would not expect sequence-specific selection of transcripts, and our results do not suggest any transcript enriched or suppressed relative to the others. The RNase H and DNA polymerase reactions are not expected to significantly contribute to variability, as the cDNA is relatively stable and those enzymes are efficient compared with reverse transcriptases. The T7 RNA polymerase is known to have a relatively short (50 min) half-life under reaction conditions [16]. But, low copy number transcripts would be equally likely to experience deactivation rates as high-copy-number transcripts, assuming length and sequence (i.e., terminators, base repeats) were not significant factors, which, for our set of transcripts, they are not (see Electronic supplementary material).
One reviewer suggested that conversion of aRNA back to cDNA could contribute additional uncertainty in the PCR assay. As previously mentioned, reverse transcription efficiencies are typically low for in vitro analyses, including gene-specific and non-specific (e.g., poly(T), random hexamer) priming. However, for RT-qPCR of the same transcript at varying concentration, this RT efficiency was constant (Fig. 1, Table 1), and differences in IVT amplification from transcript concentration became apparent. Note that all six transcripts showed the same amplification efficiencies.
As a result, IVT amplification appears to result in a slight decrease in the ratio of relative expression levels of low abundant to highly abundant transcript species. Expected yield for an IVT amplification of a typical input amount of material (e.g., 104 copies) is approximately 1,000×, yet our observed nonlinearity of amplification suggests that low copy number transcripts (e.g., 100 copies) may yield only 200×. Multiple rounds of amplification for small (less than 10 ng) samples may even compound this problem, though that remains to be studied. This amplification bias could potentially lead to misidentifying genes that appear differentially expressed above some predetermined threshold, but were in fact significantly less than what was measured.
We also examined reproducibility by plotting the C q values for repeat experiments (Fig. 2). This plot shows all species plotted simultaneously and eliminates any transcript-specific biases related to the qPCR step. There was not a significant contribution of data above 30 cycles, which agrees with our prediction for IVT followed by RT-qPCR.
Repeat IVT experiments are reproducible. The C qs for the aRNA subjected to RT-qPCR from IVT of equivalent input copies of ERCC RNA are plotted against each other from two different days. The slope of the line is 1.00 ± 0.08, and the intercept is 1.7 ± 1.7 (average ± 95 % confidence interval). This indicates that IVT acts consistently, and a difference between the relative abundances is unlikely due to day-to-day variability. Some transcripts did generate C qs above 32 but did not appear in repeat experiments
Impact of variability on low copy number gene expression analysis experimental design
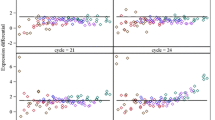
We examined the residuals from the linear regressions (Fig. 3) to measure the global variability one can expect from the IVT process. As the number of input copies decreases, the variability increases. The standard deviation was calculated for each bin of samples (approximately every 10-fold) and tabulated (Table 2). The results indicate that given a large enough ensemble of measurements and appropriate expectations for the expected magnitude of difference in signal, one can quantitatively assess differential gene expression (Δ) even in small samples. Using qPCR for assessing gene expression, the following equation gives the 95 % confidence interval around a mean using Student's t test:
Residuals vs. input copy number of linear fit. The envelope of the residuals (black dots) increases with decreasing input copy number, indicating that as fewer input copies are used, it becomes increasingly difficult to detect differences in actual gene expression vs. the noise of the background. Bins of residuals from similar input copy number samples were established (separated by vertical gray lines) to simplify the analysis. The standard deviation of the binned residuals was fit to a power law with offset (y = 0.51 + 2.4x −0.29). The solid dark gray line indicates the envelope for 1σ, and the dotted gray line shows 2σ. This result indicates the uncertainty one can expect at decreasing levels of input copies and represents a likely “best case” scenario for detecting differential gene expression
where t (0.05, n) is the t test statistic for a 95 % level of confidence given n duplicate experiments, and σ is the observed standard deviation. For example, at the 105 copy level, four replicates should be sufficient for measuring Δ = 2 with 95 % confidence. More stringent requirements at lower levels, e.g., Δ = 1.5 at 100 copies, require 16 replicates. Clearly, sample availability and analysis cost will be limiting factors in the experimental design and should be planned for accordingly.
Conclusions
The IVT reaction appears to be highly effective and reproducible even at the 100 to 1,000 input copy level. Amplification is a significant contributor to the apparent “noise” of gene expression profile analysis of small samples, though not in the strictly stochastic way as previously suspected [2].
The incorporation of the ERCC spike-in control RNA enabled evaluation of IVT performance because the input quantity of each species was known, unlike natural RNA. Single input copy analysis remains a challenge, as detection remains irreproducible and infrequent under the conditions studied here, and not likely to become robust without significantly modifying the IVT and/or RT-qPCR protocols. The variability of the data increased with decreasing copy number, as expected. We provide guidance for measuring differential gene expression under near-optimal conditions using an IVT-qPCR approach and suggest that it may be possible to draw conclusions about expression changes of low copy number genes if the appropriate number of replicates is performed or if gene set enrichment-type analyses can be employed [17].
Amplification appeared to be nonlinear under the conditions of “small” input sample size and 14 h IVT, biased against low copy number transcripts. This should be taken into consideration for differential gene expression studies of low input copy numbers by decreasing the observed fold difference. This reduction appears to follow a power-law fit under the experimental conditions we used, such that the actual fold difference is approximately the measured value to the 0.8 power for this scale of input sample and duration of IVT (i.e., a 10-fold measured difference in expression level is closer to a sixfold difference).
More broadly, IVT amplification's role as a source of variability in gene expression profiling was demonstrated. Combined with second-round amplification, which, by using a random hexamer-primed reverse transcription, is known to bias amplification based on sequence [13] causing misrepresentation of the transcript levels through multiple primings [14], methods will be needed to address these issues. We must address modification of the existing IVT process and/or implement downstream data analysis that takes into account the sequence- and length-based biasing at the individual transcript levels.
References
Hartwell LH, Hopfield JJ, Liebler S, Murray AW (1999) From molecular to modular cell biology. Nature 402:C47–C52
Nygaard V, Hovig E (2006) Options available for profiling small samples: a review of sample amplification technology when combined with microarray profiling. Nucleic Acids Res 34:996–1014
Ozsolak F, Ting DT, Wittner BS, Brannigan BW, Paul S, Bardeesy N, Ramaswamy S, Milos PM, Haber DA (2010) Amplification-free digital gene expression profiling from minute cell quantities. Nat Meth 7:619–622
Ozsolak F, Milos PM (2010) RNA sequencing: advances, challenges, and opportunities. Nat Rev Genet 12:87–98
Kurimoto K, Saitou M (2010) Single-cell cDNA microarray profiling of complex biological processes of differentiation. Curr Op Genet Devel 20:470–477
van Gelder RN, von Zastrow ME, Yool A, Dement WC, Barchas JD, Eberwine JH (1990) Amplified RNA synthesized form limited quantities of heterogeneous cDNA. PNAS 87:1663–1667
Wang E, Miller LD, Ohnmacht GA, Liu ET, Marincola FM (2000) High-fidelity mRNA amplification for gene profiling. Nat Biotech 18:457–459
Baugh LR, Hill AA, Brown EL, Hunter CP (2001) Quantitative analysis of mRNA amplification by in vitro transcription. Nucleic Acids Res 29:e29
Reid LH, Lucas AB, Kopf-Sill AR, Chen B, Bromley B, Foy C, Hinkel CS, Boysen C, Liu CM, Ranamukha-arachchi D, Wagar E, Kawasaki ES, Goodsaid FM, Wilmer F, Fischer G, Kiser GL, Causton HC, Fuscoe JC, Brenton JD, Warrington JA, Soriano J, Coller J, Burrill JD, Rhodes K, Kerr KF, Zoon KC, Lee K, Shi LM, Salit M, Satterfield M, Marton M, Cronin M, Conley MP, Williams M, Fero M, Wilson M, Novoradovskaya N, Gilles P, Wolber PK, Ikonomi P, Puri R, Beyer RP, Shippy R, Setterquist R, Elespuru RK, Baker SC, Chervitz SA, Bauer SR, Russell S, Kaysser-Kranich T, Bammler TK, Ryder TB, Sendera TJ, Scherf U, Gao XL, Wu XN, Guo X, Liu ZL (2005) Proposed methods for testing and selecting the ERCC external RNA controls. BMC Genomics 6:150
Baker SC, Bauer SR, Beyer RP, Brenton JD, Bromley B, Burrill J, Causton H, Conley MP, Elespuru R, Fero M, Foy C, Fuscoe J, Gao XL, Gerhold DL, Gilles P, Goodsaid F, Guo X, Hackett J, Hockett RD, Ikonomi P, Irizarry RA, Kawasaki ES, Kaysser-Kranich T, Kerr K, Kiser G, Koch WH, Lee KY, Liu CM, Liu ZL, Lucas A, Manohar CF, Miyada G, Modrusan Z, Parkes H, Puri RK, Reid L, Ryder TB, Salit M, Samaha RR, Scherf U, Sendera TJ, Setterquist RA, Shi LM, Shippy R, Soriano JV, Wagar EA, Warrington JA, Williams M, Wilmer F, Wilson M, Wolber PK, Wu XN, Zadro R (2005) The external RNA controls consortium: a progress report. Nat Methods 2:705–731
Devonshire AS, Elaswarapu R, Foy CA (2011) Applicability of RNA standards for evaluating RT-qPCR assays and platforms. BMC Genomics 12:118
Nygaard V, Holdin M, Løland A, Langaas M, Myklebost O, Hovig E (2005) Limitations of mRNA amplification from small-size cell samples. BMC Genomics 6:147–163
Hansen KD, Brenner SE, Dudoit S (2010) Biases in Illumina transcriptome sequencing caused by random hexamer priming. Nucleic Acids Res 38:e131
Zhang J, Byrne CD (1999) Differential priming of RNA templates during cDNA synthesis markedly affects both accuracy and reproducibility of quantitative competeitive reverse-transcriptase PCR. Biochem J 337:231–241
Thompson M (2004) The amazing Horwitz function. AMC Tech Brief
Arnold S, Siemann M, Scharnweber K, Werner M, Baumann S, Reuss M (2000) Kinetic modeling and simulation of in vitro transcription by phage T7 RNA polymerase. Biotech Bioeng 72:548–561
Submramanian A, Yamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. PNAS 102:15545–15550
Acknowledgments
The authors wish to thank David L. Duewer and Hung-kung Liu for their assistance with the statistical analysis of the data, and summer undergraduate research fellowship (SURF) program participants M. Chris Howell and Alex Peters. We would also like to thank Genomic Health for their generous gift of primer/probe sets for the RT-qPCR assays.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 225 kb)
Rights and permissions
About this article
Cite this article
Kralj, J.G., Salit, M.L. Characterization of in vitro transcription amplification linearity and variability in the low copy number regime using External RNA Control Consortium (ERCC) spike-ins. Anal Bioanal Chem 405, 315–320 (2013). https://doi.org/10.1007/s00216-012-6445-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-012-6445-1