Abstract
It is well known that saccharides and their glycoconjugates can have an important influence on various serious pathologic stages such as cancer. They can regulate tumor proliferation, invasion, hematogenous metastasis, and angiogenesis. These facts clearly show the importance of cancer saccharide recognition. In medicine, sensor analysis is one of the best methods for recognition and determination of biologically important analytes. The development and study of sensors for saccharide tumor markers can open a new way for their detection. Therefore, this review is focused on recognition of saccharide-based cancer markers by natural or synthetic selective ligands working as bio- and chemosensors. The design and application of these ligands for cancer diagnosis is a useful direction of research. Moreover, it also opens the possibility of using these agents for the targeted drug transport required for advanced anticancer therapy.

Boronic acid sensor [56]
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Recently, a slight decline in the incidence of cancer has been achieved worldwide, but long-term mortality rates still remain high [1]. For successful therapy, early diagnosis of cancer plays the key role. Implementation of early detection in traditionally used clinical methods is necessary for significant reduction of the morbidity and mortality caused by cancer [2].
For decades, microscopy of biopsy samples was the principal diagnostic method. However, this method suffers from subjectivity and a limited ability to detect the early events of cancer [3]. To fulfill the demand for the earliest possible diagnosis, new tools have to be found and applied. It is well known that when a tumor is detected, certain changes at the molecular level have already occurred. The main goal of the new diagnostic approaches is to recognize these changes as early as possible. This recognition can be based on a specific interaction of diagnostic agents with suitable molecular partners, i.e., cancer biomarkers [4]. Biomarkers, divided by several structural motifs (e.g., proteins, saccharides, metabolites, and nucleic acids), represent molecular signatures of the cell phenotype. They can be used for specific detection and recognition of particular cell types, such as cancer cells. In addition, biomarkers can be used for prediction of disease progress for chosen and optimized therapy.
The development of molecular tools for cancer diagnosis and prognosis is already in progress and it is still evolving [5]. Many biomarkers of cancers have been identified [4, 5]. An ideal recognition preferably employs biomarkers (targets) which are overexpressed on all tumor cells but not on the normal cells, and at the same time are required for cell survival, proliferation, or other critical functions [6]. The recognition component of a diagnostic agent can also be used in the drug delivery system to enhance its efficacy of medication [7].
Methodological approach
In research and clinical practice genomic, proteomic, and metabolic methods are usually used [8]. The genomic method [9] (microarays, serial analysis of gene expression, or PCR) provides information about the expression profile and mutation of the genes. Such information can be used for the prognosis of the patient (level of messenger RNA) or to pursue the effect of targeted therapies (somatic change of DNA). Proteomic methods [4] (ELISA, radioimmunoassay, MudPIT, surface-enhanced laser desorption/ionization time-of-flight analysis) can be used for the diagnosis of blood malignancies and for the determination of protein translation modification (phosphorylation). Metabolomics (analysis of metabolic pattern) reflects a global change of cancer cells and therefore metabolic analysis can be used for cancer recognition (low intracellular and extracellular pH) [10] and higher levels of some bioanalytes [11] (pyruvate [12], lactate [12, 13], metals [14, 15], and others). Effective diagnostic methodology requires determination of more biomarkers by a different diagnostic method [16]. Therefore, the identification of new cancer markers and the development of methods for their selective recognition and determination are intensively sought. Besides the methods mentioned above, there is a new emerging field of glycomic research, which can be a useful tool for cancer diagnosis and a good starting point for the development of a controlled and targeted drug delivery system [7].
In glycomics, cancer recognition is focused on saccharide patterns of cancer cells [17, 18]. It is well known that the saccharide substitution pattern of cell receptors is significantly changed during oncogenic transformation [18]. Such a change was observed in various stages of many cancer types and provides important information about cancer progress, immune response, drug resistance, metastatic capacity, and malignancy. It includes overexpression of the cell-surface polysaccharides (glycosaminoglycans [19, 20], polysialic acid [21]) and oligosaccharides (Lewis antigene [22], saccharide part of gangliosides [23]), and modification of the surface receptors (e.g., sialylation of glycolipids) [24].
Glycan cancer markers
Glycans are covalent assemblies of sugars (oligosaccharides and polysaccharides) that exist in either free form or in covalent complexes with proteins or lipids. These glycans might be regarded as part of a larger array of “metastatic codes” that a tumor’s glycan profile (or “glycotype”) might represent. For example, the serological markers CA125, CA19-9, and CA15-3 are mucin glycoconjugates that are commonly overexpressed by ovarian, pancreatic, and breast adenocarcinomas, respectively, and their serum levels correlate with tumor burden and prognosis [18, 25, 26]. Mass spectrometry [25, 26], chromatography [25], or sophisticated methods such as lectin/antibody microarrays [26, 27] are used for recognition and determination of glycan markers. Important saccharide markers include glycosaminoglycans, polysialic acid, gangliosides, and Lewis antigens.
Glycosaminoglycans are one of the most important cancer saccharides. They are functional linear heteropolysaccharides which participate in and regulate a number of cellular events and physiological/pathological processes [28]. Glycosaminoglycans can undoubtedly influence tumor cell proliferation, metastasis, and cancer progression. Polysulfated glycosaminoglycans (heparan sulfate, chondroitin sulfate, dermatan sulfate, and keratin sulfate) [19, 29, 30] are polysaccharides with high structural variability and negative charge. Inhibition of the expression of their receptors [31] or their glycosylation [32, 33] can be useful way for reduction of tumorigenic phenotype and angiogenesis. They can be used for recognition of pancreatic carcinoma [34], breast cancer [35], prostate cancer [36], thyroid cancer [30], astrocyte tumor [37], and glycoblastoma [38]. Hyaluronic acid is a nonsulfated glycosaminoglycan composed of repeating units of alternating uronic acids and N -acetylglucosamine [20, 39]. The overexpresion of hyaluronic acid was found in cancer of the neck, head, thyroid gland, liver, lung, prostate gland, and ovary among other organs and structures [40] and to be involved in stimulation of metastatic activity, angiogenesis, and tumor cell resistance.
Polysialic acid [41] is a large negatively charged linear homopolymer of a 2,8-sialic acid residue mainly associated with neural cell adhesion molecule. Medulloblastoma, neuroblastoma, and alveolar rhabdomyosarcoma are characterized by a high level of polysialic acid [21]. The elevated expression can cause higher migration ability of cancer cells [42]. In brain tumors, high expression of polysialic acid correlates with high tumor invasiveness and high risk of metastasis.
Gangliosides are sialic acid containing glycosphingolipids that are primarily expressed in the plasma membrane, and play an important role in cell growth and differentiation [43]. Overexpression of some gangliosides was observed in many tumors of neuroectodermal or epithelial origin, such as glioma, medulloblastoma, neuroblastoma, melanoma, head and neck tumors, breast cancer, and teratomas [23]. GD1a expression is used as marker for ovarian cancers [44]. In tumor biological processes, glycosylated molecules play key roles in protection of cancer cells from immune system regulation, in cell adhesion/motility, and thus in the initiation of tumor metastasis. Inhibition of cancer ganglioside function is a useful method for the reduction of tumor aggressiveness, immumoprotection, and angiogenesis [45].
Lewis blood group antigens are also common for various types of malignancies [22]. Lea and Leb antigens are important tissue blood groups, whereas Lex and Ley antigens in healthy individuals are only expressed, at relatively low levels, by a few tissues (epithelial cells). Lewis antigens can be synthesized de novo or overexpressed in the majority of human carcinomas of the colon, bladder, breast, and lungs, and are often associated with advanced forms of malignancies [46]. The presence of sialyl-Lea and sialyl-Lex is associated with bad prognosis in various types of cancers [22].
Selective ligands for recognition of glycan tumor markers
The development and study of selective ligands (biosensor and chemosensors) for glycan tumor markers can open a new way of their detection. A biosensor for cancer saccharide recognition can be represented by fluorescence-labeled monoclonal antibody directed against cancer saccharide receptors or by certain lectins with specificity for cancer saccharide markers [47].
A similar principle utilizing antibody or lectin conjugates [48] has been applied in various drug delivery systems for anticancer drugs. The development of lectin supramolecular complexes with metal porphyrins for targeted photodynamic therapy [49, 50], of lectin with galactose specificity, or of galactose–cyclodextrin for targeted transport of doxorubicin [51] represent other options for lectin-mediated drug delivery systems. Lectin/antibody carriers with suitable selectivity recognize and bind target cancer markers and thereby they provide a high level of slowly liberated anticancer agents in the cancer cells and a low level in normal cells. In addition, some lectins exhibit their anticancer effect by activation of cell death receptors [52].
Design of optical chemosensors is based on CH/π saccharide interaction [53]. Construction of these sensors can be based on smaller aromatic systems, a hydrophobic cavity, or metallocomplexes. Their function can be further improved by introduction of cationic charge and saccharide binding groups (e.g., boronic acid).
The ligands enabling cationic recognition are suitable for recognition of anionic polymers as are some saccharide cancer markers (glycosaminoglycans and polysialic acid). Good inspiration in this field might come from known sensors recognizing heparin (e.g., cyanine bases [54], cationic polymer [55]) for anionic saccharide polymers, mainly heparin (Fig. 1).
Cyanine dye sensor [54] for heparin
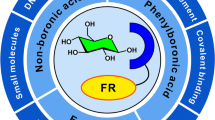
Some of those structural motifs such as cyanine bases have potential in anticancer treatment (photodynamic therapy and chemotherapy). For saccharide recognition, simple sensors based on boronic acid are usually applied [56] (Fig. 2). Recently, a colorimetric sensor with high selectivity for sialic acid was discovered.
Boronic acid sensor [56]
The idea of a hydrophobic cavity was inspired by nature’s design of lectins [57]. The hydrophobic cavity reduces the negative influence of water on the stability of the ligand–receptor complex. This phenomenon can be exploited to couple chemosensors with lectins [58]. It is based on reduction of the negative influence of water on the stability of the ligand–receptor complex. This goal can be achieved by coupling chemosensors with lectins. The idea of a hydrophobic cavity can be also applied to chemosensor design. These sensors use polyaromatic or better a heteoroaromatic core as a signal and central unit (porphyrin derivatives [59–65], cryptan system [66], and others) with substitution of hydrophilic and analyte binding groups (binaphthols [59] phosphonates [60] steroids [61], boronic acids [62] bile acid [63], PEG [64], cyclodextrin [64], and others). Similarly, Yang et al [67, 68]. prepared fluorescent diboronic acid probes for specific determination of cancer cells with overexpressed sialyl Lex carbohydrate (Fig. 3).
They observed high selectivity for the target analyte and human hepatocellular carcinoma cells with overexpression of this marker as compared with modified cells without expression of the marker studied.
Diboronic acid probe [67]
A metal–porphyrin dimer could be a possible sensor structural motif for Lex and Lea antigens [69, 70] (Fig. 4).
Promising results were also obtained in our laboratory by using bile acid–porphyrin conjugates [63] (Fig. 5). The conjugates showed high affinity for the cancer saccharide marker (heparin sulfate, hyaluronic acid, and sialic acid) and unique selectivity for transformed tumor cells (murine sarcoma, human colorectal adenocarcinoma, and chick embryo fibroblast sarcoma) in comparison with normal nontransformed cells (Fig. 6).
Such selectivity resulted in high photodynamic efficacy of this structural motif for cancer cells and very low photodynamic efficacy for normal cells.
Bile acid–porphyrin sensor [63]
The affinity of the saccharide motif for some fluorescent metal ions such as lanthanides [71] is a good basic point of sensor design. For example, Alpturk et al [72]. developed a europium complex for effective detection of a sialic acid cancer marker as GD1a (Fig. 7). This promising work showed the strong potential of this sensor type in cancer diagnosis.
Europium complex [72]
In the branch of target transport, these ligands can be used for improving drug delivery selectivity as the recognition part of these systems, or as a conjugate with anticancer agents. For example, our study focusing on bile–porphyrin conjugates clearly showed high potential of targeted anticancer therapy based on cancer saccharide recognition [63]. In the field advanced drug delivery, Lee et al [7]. studied the influence of a conjugated specific chondroitin sulfate ligand on drug delivery selectivity for the human renal adenocarcinoma cell line and its anticancer effectiveness. They observed a significant improvement of the drug delivery property for cancer cells with overexpressed target marker and an inconsiderable change for cancer cells with a reduced marker level.
Conclusion
Knowledge of tumor saccharide pattern can be also exploited for the development of diagnostic methods and targeted anticancer systems. We presented in this review the use of glycomics, mainly optical biochemical and chemical sensors, for recognition of cancer saccharide markers. The examples discussed help to highlight the design principle of natural or synthetic selective ligands as optical sensors for cancer diagnosis. The sensors described clearly demonstrate the high potential of this approach for cancer diagnosis. This review also focused on the exploitation of cancer saccharide ligands as targeted anticancer therapy. Their in vivo and in vitro study demonstrated the strong potential of this approach in targeted anticancer treatment.
References
Warshawsky D, Landolph JR (2005) Molecular carcinogenesis and the human biology of human cancer. Taylor & Francis, London
Negm RS, Verma M, Srivastava S (2002) 2002. Trends Mod Med 8:288–229
Saffroy R, Pham P, Reffas M, Takka M, Lemoine A, Debuire B (2007) Clin Chem Lab Med 45:1169–1179
Hamdan MH (2007) Cancer biomarkers: analytical techniques for discovery. Wiley, Totowa
Ransohoff DF (2008) J Natl Cancer Inst 100:1419–1420
Abou-Jawde R, Choueiri T, Alemany C, Mekhail T (2003) Clin Ther 8:2121–2137
Lee CM, Tanaka T, Murai T, Kondo M, Kimura J, Su W, Kitagawa T, Ito T, Matsuda H, Miyasaka M (2002) Cancer Res 62:4282–4288
Griffin JL, Kauppinen RA (2007) J Proteome Res 6:498–505
Kawakami Y, Fujita T, Matsuzaki Y, Sakurai T, Tsukamoto M, Toda M, Sumimoto H (2004) Cancer Sci 95:784–791
Adams DJ (2005) Dev Curr Med Chem Anticancer Agents 5:1–13
Gao H, Dong B, Liua X, Xuan H, Huang Y, Lina D (2008) Anal Chim Acta 7:269–277
Sattler UGA, Walenta S, Mueller-Klieser W (2007) Anaesthesist 56:466–469
Sattler UGA, Hirschhaeuser F, Mueller-Klieser WF (2010) Curr Med Chem 17:96–108
Pasha Q, Malik SA, Iqbal J, Shaheen N, Shah MH (2008) Environ Monit Assess 147:377–388
Geraki K, Farquharson MJ, Bradley DA (2002) Phys Med Biol 47:2327–2339
Celis JE, Moreira JM, Gromova I, Cabezon T, Ralfkiaer U, Guldberg P, Straten P, Mouridsen H, Friis E, Holm D, Rank F, Gromov P (2005) FEBS J 272:2–15
Wuhrer M (2007) Expert Rev Proteomics 4:135–136
Fuster MM, Esko JD (2005) Nat Rev Cancer 5:526–542
Liu D (2005) In: Garg HG, Linhardt RJ, Hales CA (eds) Elsevier, Amsterdam
Lokeshwar VB, Rubinowicz D, Schroeder GL, Forgacsi E, Minnai JD, Block NL, Nadji M, Lokeshwar BL (2001) J Biol Chem 276:11922–11932
Gurlek A, Karavitaki N, Ansorge O, Wass JHV (2007) Eur J Endocrinol 156:143–157
Dall’Olio F, Chiricolo MG (2001) Glycoconj J 18:841–850
Birklé S, Zeng G, Gao L, Yu RK, Aubr J (2003) Biochimie 85:455–463
Cylwik B, Chrostek L, Szmitkowski M (2005) Pol Merkur Lekarski 19:237–241
An HJ, Kronewitter SR, Leoz ML, Lebrill CB (2009) Curr Opin Chem Biol 13:601–607
Abbott KL, Lim JM, Wells L, Benigno BB, McDonald JF, Pierce M (2010) Proteomics 10:470–481
Hu D, Wong DT (2009) Proteomics Clin Appl 3:148–154
Yip GW, Smollich M, Gotte M (2006) Mol Cancer Ther 5:2139–2138
Malavaki C, Mizumoto S, Karamanos N, Sugahara K (2008) Connect Tissue Res 49:3–4
Magro G, Perissinotto D, Schiappacassi M, Goletz S, Otto A, Muller EC, Bisceglia M, Brown G, Ellis T, Grasso S, Colombatti A, Perris R (2003) AJP 163:183–193
Jiang X, Couchman JR (2003) J Histochem Cytochem 51:1393–1410
Zhou Z, Wang J, Cao R, Morita H, Soininen R, Chan KM, Liu B, Cao Y, Tryggvason K (2004) Cancer Res 64:4699–4702
Li F, Dam GB, Murugan S, Yamada S, Hashiguchi T, Mizumoto S, Oguri K, Okayama M, Kuppevelt TH, Sugahar K (2008) J Biol Chem 283:34294–34304
Theocharis AD, Tsara MA, Papageorgacopoulou N, Karavias DD, Theocharis DA (2000) Biochim Biophys Acta 1502:201–206
Suwiwat S, Ricciardelli C, Tammi M, Tammi M, Auvinen P, Kosma VM, LeBaron RG, Raymond WA, Tilley WD, Horsfall DJ (2004) Clin Cancer Res 10:2491–2498
Sakko AJ, Butler MS, Byers S, Reinboth BJ, Stahl J, Kench JG, Horvath LG, Sutherland RB, Stricker PD, Henshall SM, Marshall VR, Tilley WD, Horsfall DJ, Ricciardell C (2008) Cancer Epidemiol Biomarkers 17:2488–2497
Kato Y, Hayatsu N, Kaneko MK, Ogasawara S, Hamano T, Takahashi S, Nishikawa R, Matsutani M, Mishima K, Narimatsu N (2008) Biochem Biophys Res Commun 369:1041–1046
Hayatsu N, Ogasawara S, Kaneko MK, Kato Y, Narimatsu H (2008) Biochem Biophys Res Commun 368:217–222
Itano N, Zhuo L, Kimata K (2008) Cancer Sci 99:1720–1725
Garg HG, Hales CA (2004) In: Patel S, Page MJ (eds) Elsevier, Amsterdam
Gascon E, Vutskitsb L, Kiss JZ (2007) Brain Res Rev 56:101–118
Suzuki M, Suzuki M, Nakayama J, Suzuki A, Angata K, Chen S, Sakai K, Hagihara K, Yamaguchi Y, Fukuda M (2005) Glycobiology 15:887–894
Lahiri S, Futerman AH (2007) Cell Mol Life Sci 64:2270–2284
Prinetti A, Aureli M, Illuzzi G, Prioni S, Nocco V, Scandroglio F, Gagliano N, Tredici G, Rodriguez-Menendez V, Chigorno V, Sonnino S (2010) Glycobiology 20:62–77
Fredman P, Hedberg K, Brezicka T (2003) Biodrugs 17:155–167
Marionneau S, Cailleau-Thomas A, Rocher J, Le Moullac-Vaidye B, Ruvoën N, Clément M, Le Pendu J (2001) Biochimie 83:565–573
Jelinek R, Kolusheva S (2004) Chem Rev 104:5987–6015
Gabor F, Bogner F, Weissenboeck A, Wirth M (2004) Adv Drug Deliv Rev 56:459–480
Komath SS, Kavithab M, Swamy MJ (2006) Org Biomol Chem 4:973–988
Komath SS, Bhanu K, Maiya BG, Swamy MJ (2008) Biosci Rep 20:265–276
Oda Y, Yanagisawa M, Maruyama M, Hattori K, Yamanoi T (2008) Bioorg Med Chem 16:8830–8840
Liu B, Bian HL, Bao JK (2010) Cancer Lett 287:1–12
Stanca-Kaposta EC, Gamblin DP, Screen J, Liu B, Snoek LC, Davis BG, Simons JP (2007) Phys Chem 9:4444–4451
Bříza T, Kejík Z, Císařová I, Králová J, Martásek P, Král V (2008) Chem Commun 1901–1903
Sun W, Bandmann H, Schrader T (2007) Chem Eur J 13:7701–7707
Yang Y, Lewis PT, Escobedo JO, St. Luce NN, Treleaven WD, Cook RL, Strongin RM (2004) Collect Czech Chem Commun 69:1282–1291
Muraki M, Ishimura M, Harata K (2002) Biochim Biophys Acta 1569:10–20
Rusin O, Král V, Escobedo JO, Strongin RM (2004) Org Lett 6:1373–1376
Rusin O, Lang K, Král V (2002) Chem Eur J 8:655–663
Král V, Rusin O, Charvátová J, Anzenbacher P, Fogl J (2000) Tetrahedron Lett 41:10147
Dukh M, Šaman D, Lang K, Pouzar V, Černy I, Drašar P, Král V (2003) Org Biomol Chem 1:3458–3463
Jiang S, Escobedo JO, Kim KK, Alpturk O, Samoei GK, Fakayode SO, Warner IM, Rusin O, Strongin RM (2006) J Am Chem Soc 128:12221–12228
Králová J, Koivukorpi J, Kejík Z, Poučková P, Sievanen E, Kolehmainen E, Král V (2008) Org Biomol Chem 6:1548–1552
Králová J, Bříza T, Moserová I, Dolenský B, Vašek P, Poučková P, Kejík Z, Kaplánek R, Martásek P, Dvořák M, Král V (2008) J Med Chem 51:5964–5973
Králová J, Kejík Z, Bříza T, Poučková P, Král A, Martásek P, Král V (2010) J Med Chem 53:128–138
Rusin O, Kral V, Schmidtchen FP (2001) Org Lett 3:873–876
Yang W, Gao S, Gao X, Karnati VV, Ni W, Wang B, Hooks WB, Carson J, Weston B (2002) Bioorg Med Chem Lett 12:2175–2177
Yang W, Fan H, Gao X, Gao S, Karnati VV, Ni W, Hooks WB, Carson J, Weston B, Wang B (2004) Chem Biol 11:439–448
Hirata O, Kubo Y, Takeuchi M, Shinkai S (2004) Tetrahedron 60:11211–11218
Sugasaki A, Sugiyasu K, Ikeda M, Takeuchi M, Shinkai S (2001) J Am Chem Soc 123:10239–10244
Harte SCMG, AJ QSJ, Gunnlaugsson T (2008) Coord Chem Rev 252:2512–2527
Alpturk O, Rusin O, Fakayode SO, Wang W, Escobedo JO, Warner IM, Crowe VE, Kral V, Pruet JM, Strongin RM (2006) Proc Natl Acad Sci 103:9756–9760
Acknowledgements
This work was supported by grants from the Ministry of Education of the Czech Republic (grants MSMT 1M 6837805002, MSM6036137307, MSM0021620857, AV0Z50520514; projects LC512, LC06077, and MSM6036137307) and from the Grant Agency of the Czech Republic (grant 203/09/1311) and, in part, by project AV0Z50520514 and grant KAN200200651 awarded by the Grant Agency of the Academy of Sciences of the Czech Republic.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in the special issue Optical Biochemical and Chemical Sensors (Europtrode X) with guest editor Jiri Homola.
Rights and permissions
About this article
Cite this article
Kejík, Z., Bříza, T., Králová, J. et al. Selective recognition of a saccharide-type tumor marker with natural and synthetic ligands: a new trend in cancer diagnosis. Anal Bioanal Chem 398, 1865–1870 (2010). https://doi.org/10.1007/s00216-010-4124-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-010-4124-7