Abstract
Bacterial quorum-sensing regulatory systems can be summarized in a simple model wherein an autoinducer molecule accumulates in cultures and stimulates regulatory changes in gene expression upon reaching a critical threshold concentration. Although quorum sensing was originally thought to be an isolated phenomenon governing the regulation of a handful of processes in only a few bacteria, it is now considered to be a widespread mechanism for coordinating bacterial gene expression. Over decades of research, investigations of autoinducer-mediated regulation have revealed that these systems are far more complicated than originally appreciated, and such discoveries have accelerated recently with the application of molecular and genomic tools. The focus of this review is to highlight recent advances describing complexities that go beyond the simple model of quorum sensing. These complexities include the regulation of autoinducer production and degradation, the presence of multiple quorum-sensing systems in individual bacteria that regulate diverse genes, often in coordination with other regulatory elements, and the influence of interorganismal interactions on quorum sensing.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Studies of autoinduction or quorum sensing were initiated over forty years ago by curious investigators attempting to explain the pattern of light production in cultures of bioluminescent marine bacteria [1–3]. They found that these bacteria exude a bacterial pheromone or “autoinducer” that accumulates in growing cultures and stimulates bioluminescence only when it has reached a critical threshold concentration [4, 5]. The model that emerged from these studies is illustrated simplistically in Fig. 1a. Importantly, this model explained why some bacteria, such as Vibrio fischeri, did not produce bioluminescence when cells were dilute but glowed brightly when populations became more densely packed [5, 6]. This bioluminescence-stimulating compound of V. fischeri was identified over a decade later as a N-acylhomoserine lactone (AHL), specifically N-3-oxo-hexanoyl homoserine lactone (Fig. 1b) [4].
A “simple” model of autoinduction or quorum sensing in V. fischeri. a The AHL autoinducer (AI, triangle) is constitutively produced by the bacterial cells (shaded capsule). At low cell densities very little AI is present. As cell density increases, AI accumulates until a threshold level is reached that then activates the production of light. b Structure of the AHL autoinducer molecule produced by V. fischeri LuxI, N-3-oxo-hexanoyl homoserine lactone, also known as 3-oxo-C6-homoserine lactone. Different AHL synthases produce AHL signal molecules that vary in the length of, and substitutions on, the acyl side chain (the variable region is denoted by parentheses)
Although the autoinduction model was initially met with skepticism, it became widely accepted after the isolation and characterization of the genes necessary for light production and regulation of luminescence expression [7, 8]. Through these studies it was discovered that the regulatory functions and enzymatic activities necessary for light production in V. fischeri were encoded on an 8-kb DNA fragment, consisting of the luxICDABEG operon and the divergently transcribed luxR gene (Fig. 2). The luxCDABEG genes encode the enzymes necessary for light production, while luxI encodes the AHL synthase and luxR the AHL receptor. From this information, a more detailed understanding of autoinduction of bioluminescence emerged, where LuxI produces the AHL molecule, which is freely diffusible across the cell membrane [9] and accumulates in the culture medium, and where AHL bound to LuxR activates transcription of the luxICDABEG operon, resulting in the production of light (Fig. 2). Presumably, the threshold concentration of autoinducer required to stimulate luminescence is defined by the binding affinity of LuxR for its cognate AHL.
Revised model of autoinduction or quorum sensing in V. fischeri, reflecting the identification of the genes encoding the regulatory functions and enzymatic activities necessary for light production. a Under low cell densities, the AHL synthase LuxI is constitutively expressed. LuxI produces autoinducer (AI or AHL), which begins to accumulate but does not reach a threshold level, and light is not produced. b Under high cell densities, AI accumulates to the threshold level and binds the regulator LuxR, which activates transcription of the luxICDABEG operon. The products of the luxCDABEG genes are responsible for light production. The synthesis of AI is regulated in a feedforward manner, where the activity of LuxI stimulates its expression
The general consensus at the time was that autoinduction was an obscure phenomenon present in bioluminescent marine bacteria [3], and that it was related to their dual lifestyles. Bioluminescent bacteria can be found as dilute free-living planktonic cells, but some also inhabit the light organs of squid and fish [10], where they are maintained at high densities of 109 to 1010 cells per ml. During such dense symbiotic growth in their host, autoinducer accumulates to sufficient levels to trigger bacterial light production that is used in animal behaviors such as camouflaging by counter-illumination [11]. The density-dependent induction of bioluminescence ensures that the energetically expensive process of light production [12] is not triggered in the bacteria during free-living growth as planktonic cells in the ocean.
Although the study of autoinduction was initiated to explain a curious phenomenon in what were considered to be “unimportant” marine bacteria [3], over the years similar systems have been found in a variety of bacteria [13, 14]. AHL signals with varying acyl side chains are commonly used as autoinducers by Gram-negative bacteria [15, 16], but other autoinducer molecules such as peptides [17], quinolones [18], and 4,5-dihydroxy-2,3-petanedione derivatives collectively known as “AI-2” [19, 20] have also been discovered. The behaviors regulated in these systems are usually those that are more productive when undertaken as groups of cells rather than as individuals, and/or behaviors that would be more profitable for cells in a diffusion-limited environment, such as the secretion of extracellular enzymes. Examples of autoinducer-triggered behaviors include secretion of proteases or virulence factors, antibiotic production, sporulation, biofilm formation, genetic competence, conjugative DNA transfer, and, as mentioned above, bioluminescence.
With the discovery of many new autoinducers and population-based regulatory systems, the term “quorum sensing” (QS) was coined in the mid-1990s [21] to convey the idea that bacteria used autoinducers to census their culture density and to make certain decisions only when a critical density, a “quorum,” was present. By making the analogy to the practice of many organizations requiring a quorum of members present to conduct business, the term was readily understandable by those outside the field and it highlighted the social nature of bacteria. However, in human endeavors the rules of order are typically complex. For example, in the United States House of Representatives a quorum could be two members, one hundred members, or two hundred and eighteen members, depending on the action being taken. Moreover, a quorum is a starting point required to do business, but another tally is still necessary to make decisions, and for some group decisions multiple votes may be required, different groups may have to concur, and external forces may try to subvert the process.
It should come as no surprise that systems regulating group decisions by bacteria are at least as complex as those governing societies. Over the years it has become clear that bacterial QS systems do not respond purely to the number of cells present, and recent applications of molecular and genomic tools have accelerated the discovery of such complexities. These studies have revealed that the processes of autoinducer-mediated gene regulation are often far more complex than the simple model presented in Figs. 1a or 2. This complexity is reflected in the regulation of autoinducer production, the presence and regulation of endogenous systems for degrading autoinducers, the multiple QS systems found in individual bacteria, the coordination of QS systems with other regulatory elements, and the influence of other organisms on QS in bacteria. In this review we highlight recent advances in the QS field that have described these complex QS pathways.
Autoinducer accumulation: multiple levels of control
In the simplest model of QS, autoinducer is produced at a constant rate and accumulates at a constant rate. If this were true, autoinducer concentration would be a relatively straightforward reflection of bacterial numbers and autoinducer diffusion rates over time; however, quorum signaling is considerably more complicated, even in clonal cultures. Many bacteria regulate autoinducer production and some have the capacity to destroy their own autoinducers. Thus, the concentration of autoinducer actually reflects a complicated set of parameters including environmental influences.
It has been known for a long time that bacteria can regulate the production of autoinducer. Perhaps the most obvious example of this is the AHL synthase LuxI in V. fischeri, whose activity stimulates its own expression (see above and Fig. 2). Given the ubiquity and obvious functional logic of feedback negative regulation in biological systems, such feedforward positive regulation is striking, although not unprecedented. Interestingly, this phenomenon is not limited to V. fischeri, and autoinducer molecules stimulate autoinducer synthases in several bacteria, including the opportunistic pathogen Pseudomonas aeruginosa [22], the tumor-generating plant pathogen Agrobacterium tumefaciens [23], and the human pathogen Streptococcus pneumoniae [24]. Although such feedforward regulation means that autoinducers do not accumulate at a steady rate, it does preserve a model of QS with internal control.
Perhaps more importantly, from an ecological perspective, regulation of autoinducer production can also respond to inputs from outside the QS systems themselves, thereby embedding QS systems in a larger, environmentally responsive transcriptional network. Combined with the feedforward autostimulatory nature of some QS systems mentioned above, these environmental inputs can be greatly amplified. Moreover, the regulatory networks controlling QS can be quite complex. For example, the QS systems of P. aeruginosa are modulated by several other regulators, including VsqR, RsaL, QscR, GacA/RsmAZ, Vfr, RelA, and RpoS [25]. The precise environmental signals detected by these systems are not always clear; however, the latter two regulators respond to the nutritional/growth status of the cell. rpoS may also be regulated by QS, although this finding is controversial [26, 27]. This connection of QS to the availability of growth substrates represents a common, though not universal, theme in the environmental regulation of QS. For example, catabolite repressor (Crp) stimulates autoinducer synthesis in V. fischeri in response to growth substrates [28, 29]. By embedding quorum signaling with such regulatory systems, bacteria are able to modulate the production of autoinducers such that their concentration reflects not only cell density but also specific parameters of their environment.
Interestingly, bacteria can also control the local concentration of autoinducer by degrading signals that they have already generated. Investigation of bacteria consuming their own autoinducers has recently gained momentum, but relatively few examples have been examined and the specifics of the systems are not yet as well-understood as the synthesis of autoinducers. Both the prevalence of QS systems and the results of bioinformatic searches for homologs of known quorum-signal degraders [30] suggest that these pathways may be more widespread than has been appreciated, although their prevalence and ecological importance are still uncertain.
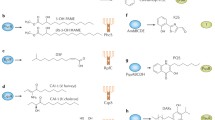
What is clear is that diverse systems exist for autoinducer degradation in the bacteria that produce these signals and that these autoinducer-degrading systems are themselves regulated. For example, both A. tumefaciens and P. aeruginosa can degrade the AHL molecules that they produce, but they do so in biochemically distinct ways. A. tumefaciens produces a lactonase, AttM, that hydrolyzes AHLs to produce N-acylhomoserines [31], whereas P. aeruginosa encodes multiple AHL acylases, including PvdQ and QuiP, capable of cleaving the acyl side chain from homoserine lactones (Fig. 3a) [30, 32, 33]. As might be expected given the variability of AHL acyl groups, these AHL acylases display some substrate specificity; however, they do act on the signal molecules produced by P. aeruginosa.
Mechanisms for enzymatic degradation of signal molecules in bacteria. a AHL acylases and lactonases can degrade AHL molecules with various acyl side chains. A generic AHL molecule is shown to illustrate the enzymatic processes. b Proposed degradation pathway for AI-2 in Salmonella or E. coli (modified from [43]). The AI-2 molecule shown is (2R, 4S)-2-methyl-2,3,3,4-tetrahydroxytetrahydrofuran or R-THMF, which is phosphorylated by LsrK. The phosphorylated AI-2 molecule is ultimately degraded by LsrF and LsrG. The method by which LsrF and LsrG degrade AI-2 is currently under investigation
Just as autoinducer synthesis is regulated, and thereby responsive to environmental inputs, so too are autoinducer degradation pathways modulated and embedded in regulatory networks. For example, the P. aeruginosa AHL acylase PvdQ is upregulated elevenfold in response to iron limitation [34], and expression of the A. tumefaciens lactonase AttM is induced through a RelA-dependent mechanism when cells are starved of carbon or nitrogen [35]. At least superficially, theses examples of autoinducer degradation being tied to nutrient availability are reminiscent of the regulation of autoinducer production. Most strikingly, autoinducer production by P. aeruginosa and autoinducer degradation by A. tumefaciens are each regulated by the starvation response regulator RelA. However, RelA activates both of these systems, so there is not a consistent pattern toward increasing or decreasing autoinducer accumulation based on starvation.
Controlled degradation of autoinducers could make for a more dynamic and responsive signaling system; however, an alternative model would be that autoinducers are degraded simply for nutrition, as sources of carbon, nitrogen, or energy. One argument against the latter model is that autoinducers generally function at very low concentrations and would not appear to be attractive growth substrates. Importantly, the autoinducer degradation systems described above catalyze AHL turnover at these low, physiologically relevant concentrations. Moreover, although P. aeruginosa is able to utilize AHL as a growth substrate, A. tumefaciens is not. Similarly, although Salmonella degrade their AI-2 signal, as described below, they do not appear to use it as a carbon source [36]. When all of these factors are considered, it seems most likely that the regulation of autoinducer degradation allows bacteria to “take back” an existing signal should conditions change.
The dynamic nature of autoinducer concentration, mediated by controlled synthesis and degradation of the signal, is well-illustrated in the accumulation of AI-2 in culture. AI-2 production can be considered an offshoot, or even a by-product, of the activated methyl cycle, and not surprisingly AI-2 accumulation is connected to central metabolism [37]. The role of AI-2 as a true QS autoinducer is best understood in Vibrio harveyi, where recognition of AI-2 by LuxP and LuxQ initiates a regulatory phosphorylation cascade [38, 39]; however, the regulation of AI-2 accumulation is more thoroughly studied in Escherichia coli and Salmonella. In E. coli, the autoinducer synthase LuxS appears to be modulated by CRP based on carbon-source utilization of the cell, and in both bacteria the Pfs enzyme that presumably supplies LuxS with substrate is also transcriptionally regulated in response to the carbon source available [40, 41]. In Salmonella, AI-2 production correlates well with pfs expression [40]. Moreover, these bacteria possess an elaborate system for sequestering and destroying the AI-2 already released in culture. The current model (Fig. 3b) proposes that LsrB binds AI-2 and shuttles it back into the cell via a transporter composed of LsrA, LsrC, and LsrD, and that AI-2 is then phosphorylated by LsrK and ultimately degraded by LsrF and LsrG [41, 42]. Like pfs, the lsr genes are themselves regulated, in part by CRP [41, 43] and also by LsrR, which derepresses most lsr genes in response to internalized and phosphorylated AI-2. These systems, and others regulators that are not yet fully described, function such that AI-2 accumulation is influenced by carbon source, osmolarity, pH, iron, oxygenation, redox state, various stresses, and AI-2 itself [44–46]. In contrast to the simple model of autoinducer reflecting cell numbers, AI-2 levels in batch cultures of E. coli and Salmonella usually peak and then decline.
In addition to being consumed, autoinducers may be removed from a system simply by diffusing away. Under many laboratory conditions where QS is studied, for example with cultures grown in flasks or test tubes, the concentration of autoinducer is generally not affected by local diffusion, because samples are well-mixed and the system is closed. However, in different real-world environments, the same number of cells producing and consuming autoinducer at the same rates may experience different autoinducer concentrations based on local autoinducer diffusion rates. Recently, an alternative view for the function of QS has emerged where autoinducer functions as an environmental probe to allow a bacterial cell to determine the extent of mixing and diffusion in its immediate environment [47]. It is argued that autoinducer-triggered behaviors can benefit single cells in a diffusion-limited environment, in contrast to the idea that bacteria utilize QS to act cooperatively. This raises an interesting issue as to the evolution of signaling pathways and the relation of the control of synthesis and degradation of signals to the particular environmental niche occupied by a bacterium, an area that warrants further exploration.
It is now clear that autoinducers do not necessarily accumulate at a constant rate and are therefore not purely “census-taking” molecules. Even for clonal cultures grown in shake flasks in the laboratory, the concentration of autoinducer reflects not only cell density but also the growth conditions. Environmental signals are transduced through regulatory networks that control both autoinducer synthesis and, at least in some instances, autoinducer degradation as well. Moreover, the “feedforward” nature of many QS systems can greatly amplify these environmental inputs, and the net effects on the autoinducer-to-cell ratio can be dramatic. Finally, environment-specific barriers to autoinducer diffusion may influence the local accumulation of autoinducer and QS behaviors. Thus, autoinducer concentrations are dynamic and very much context-dependent.
Another layer of complexity: multiple QS systems in bacteria
When autoinduction or QS was first described in V. fischeri, it was thought that this bacterium produced a single AHL autoinducer that was produced by LuxI and detected by a single receptor protein, LuxR [7, 8]. However, it has since been found that V. fischeri possesses another AHL synthase, AinS, which produces a second structurally distinct AHL signal, octanoyl homoserine lactone, recognized by both LuxR and AinR, which is also called LuxN [48]. V. fischeri also produces an AI-2 type signaling molecule [49]. These findings actually followed the first discovery of multiple QS networks within an individual bacterium, which was described in another bioluminescent marine bacterium, V. harveyi [50]. The presence of multiple QS systems is now known to be a common trait in bacteria, leading to questions on the identities of the genes regulated by each system and the role(s) of multiple systems in bacterial biology.
Molecular biological tools and whole-genome analyses have been powerful tools in identifying genes belonging to QS regulons in bacteria, most notably in the opportunistic pathogen, P. aeruginosa. In particular, P. aeruginosa DNA microarray analyses have provided new insight into the complexities inherent in QS-controlled gene expression and have provided a framework for comparative studies in other organisms (reviewed in [25]). Through these analyses, it has become apparent that the multiple QS systems in P. aeruginosa influence the expression of a wide range of genes, and, as mentioned above, QS gene regulation is embedded in a network of regulatory pathways that respond to environmental signals.
Using microarray analysis of specific P. aeruginosa QS regulatory mutants, three independent groups identified hundreds of genes belonging to the two QS regulons [25, 51–53]. While many predicted suites of genes (e.g., those encoding virulence factors) were QS-regulated, these analyses also identified genes whose products are involved in “housekeeping” cell processes such as central metabolism [25], demonstrating that QS signaling can have global effects on gene expression in bacteria and can influence cell physiology. However, when comparing the three independent studies, most of the “regulated” genes were identified by only one or two of the groups, and only 102 QS-induced genes were common to all three data sets [25]. Considering the three experiments were performed in separate laboratories with differences in experimental conditions, this interesting result suggests that environmental conditions can have profound influences on QS gene regulation.
One surprising result of these studies is that there does not appear to be a single cell density at which all QS-regulated genes are induced or repressed, and the “quorum” required for expression depends on the gene and environmental conditions. The expression of individual QS genes is influenced by the presence and/or absence of each of the two P. aeruginosa AHL signals, so the cells contain distinct regulons responsive to one or both signals, each of which has its own critical threshold concentration. In addition, many QS-controlled genes do not respond to exogenously provided AHL signals until the cells reach the stationary phase of growth [25, 52], which can be explained by the requirement for the stationary-phase σ factor RpoS for activation of QS [26, 27, 54], or the presence of inhibitors in the culture medium [55]. These results show that QS gene regulation is not solely influenced by signal accumulation, but requires the presence of additional factors and is responsive to environmental conditions (reviewed in [25, 56]). The fact that QS gene regulation is entangled in a network of other regulatory pathways suggests that bacteria tightly control this process so that particular QS regulon genes are only expressed under the appropriate conditions. This leads to the question of the role of these complex multiple QS systems in bacterial growth in a “natural” environment, for example during host colonization.
The use of model systems such as the symbiosis between V. fischeri and the Hawaiian bobtail squid Euprymna scolopes are beginning to reveal the importance of multiple QS systems for bacteria inhabiting a natural environment. This system has been studied for over 15 years as a model for the extracellular colonization of host tissue by cooperative and pathogenic bacteria [57, 58]. As mentioned above, V. fischeri contains two AHL-based QS systems, LuxI/LuxR and AinS/AinR, as well as a LuxS/AI-2-based QS system, and these each influence the expression of bioluminescence [7, 8, 48]. Since luminescence is an integral part of the symbiotic relationship, the role of each system in light production and host colonization were investigated.
These studies revealed a complex picture of the involvement of each system in the regulation of light production and host colonization (reviewed in [59]). Analysis of luxI, luxS and ainS mutants during growth in liquid culture and during host colonization revealed a stepwise activation of the QS systems with increasing cell density [60–62]. ainS and luxS influence light production in broth cultures and are important for initiation of the symbiosis, conditions that correspond to lower cell densities. The AHL signal produced by AinS appears to be the dominant signal under these circumstances, with AI-2 playing a lesser role [49]. In contrast, the presence of luxI has little if any influence on luminescence in liquid culture, but is necessary for boosting luminescence during later stages of the colonization process, corresponding to the higher cell densities present in the squid light organ as compared to laboratory culture conditions. It is important to note that in certain strains of V. fischeri, including the very brightly bioluminescent strains initially characterized, luxI does play a role in stimulating bioluminescence in culture, which explains why it was discovered early on. For reasons that are not yet clear, low expression of luxI in culture appears to be a hallmark of the V. fischeri strains isolated from E. scolopes [63, 64].
The involvement of ainS in initiation of colonization suggested that this QS system influences the expression of genes other than those responsible for light production. A V. fischeri DNA microarray was used to identify genes whose expression was altered in the presence of ainS, and this approach revealed 30 differentially regulated genes potentially involved in motility, transcriptional regulation, metabolism and biosynthesis of extracellular polysaccharides [60]. Although such detailed microarray analyses of the roles of luxI and luxS are not yet published, a proteomic approach suggested, not surprisingly, that the LuxI/LuxR system also controls several target genes involved in a variety of processes, including genes whose function are important during host colonization [65]. Thus, although the LuxI/LuxR, AinS/AinR, and AI-2 type QS systems were initially implicated in the control of bioluminescence, in-depth studies are revealing specific and distinct functions for each system in the biology of V. fischeri and its interactions with the squid host, and the function of autoinducer-mediated gene regulation has proven far more complex than the original model presented in Figs. 1 and 2.
The study of QS gene regulation in P. aeruginosa and V. fischeri has provided valuable insight into the complex roles of autoinducers as modulators of global regulatory systems that influence many aspects of bacterial physiology and host interactions. Moreover, these two model systems are illustrative of similar discoveries in many other bacteria. Although the initial simple model of quorum sensing (e.g., Fig. 1) requires only a single autoinducer compound to serve as a census-taker and to direct population-based regulatory decisions, it is now clear that several bacteria produce multiple autoinducers, that their cognate regulators are embedded in other regulatory networks, and that the genes they control are not always clearly connected to group behaviors. This surprising complexity and diversity of QS in bacteria is likely a reflection of the evolution of these systems in bacteria that need to respond to fluctuating and diverse conditions, particularly during the transition from life in the environment to living inside a host.
Outside influences on bacterial quorum sensing
Bacteria that regulate gene expression through AHL-based QS are often found in mixed microbial consortia associated with plants or animals. Because QS-related behaviors such as virulence factor production, biofilm formation, and antibiotic production often have detrimental effects on the other members of these dynamic communities, it is not surprising that plants, animals, and other microorganisms have developed methods for interfering with QS signaling processes.
The first indication that compounds from other organisms could influence QS in bacteria came with the discovery that halogenated furanones produced by the marine red alga Delisea pulchra acted as QS mimics that could inhibit bacterial AHL-controlled processes [66]. The mode of action of furanones is to bind to AHL receptor proteins and promote their proteolytic degradation [67, 68]. Intriguingly, D. pulchra is known to be remarkably free of bacterial fouling in marine environments, and the ability to disrupt bacterial QS is thought to be at least partly responsible for this characteristic. This initial report was followed by the discovery that a green alga, Chlamydomonas reinhardtii, and certain higher plants also secrete compounds that affect AHL-based QS regulation in bacteria [69–71]. The chemical structures of these compounds remain to be solved, but interestingly, most appear to stimulate rather than inhibit AHL-controlled gene expression.
The ability to modulate AHL-based bacterial QS is not limited to algae and plants. In addition to the AHL degradation and turnover by autoinducer producers (described above), numerous bacteria that do not produce AHL signals also inactivate AHLs enzymatically through the production of AHL acylases or lactonases (Fig. 3) [72–78]. It may be the case that interfering with QS signaling could give these bacteria a competitive advantage when occupying the same environmental niche as AHL-producing bacteria [79]. In addition, certain bacteria can not only inactivate AHLs, but also utilize them as sole carbon and/or nitrogen sources for growth [75, 78, 80], adding another potential benefit to the degradation of AHL signals. Although each specific AHL signaling molecule may not be very abundant, the total concentration of different AHLs in a microbial community could make them an attractive nutrient source, and AHL degraders may also find less competition for this substrate and thus occupy a specialized niche. Enzymatic degredation of AHL signals has also been described in certain mammalian cell types [81, 82], suggesting that host cells may be able to counteract QS signaling during bacterial infection and colonization. The studies mentioned above provide a just a sample of the ways in which other organisms can influence AHL-based QS systems, and the intense interest in this topic has resulted in several more comprehensive reviews [79, 83–85].
Conclusions
Early studies of autoinducer-mediated gene regulation led to an intuitive and relatively simple model of QS systems as mechanisms of censusing bacterial populations and regulating the expression of genes with roles specific to life as solitary cells or as dense groups. Over the years a much more complex picture has emerged, wherein autoinducers are dynamically controlled, individual cells produce multiple autoinducer molecules, organisms compete by disrupting QS signals, and a wide range of genes are regulated in coordination with other global regulators but not always with direct correlation to population numbers. The discovery that bacteria can use QS to regulate virulence gene expression and other medically and economically important behaviors has led to an explosion of research in this area, and with these applications in mind, future research geared toward unraveling QS in real-world environments will undoubtedly uncover more complexities inherent in these systems.
References
Greenberg EP (1997) ASM News 63:371–377
Hastings J, Greenberg E (1999) J Bacteriol 181:2667–2668
Nealson K (1999) In: Dunny, G, Winans S (eds) Cell–cell signaling in bacteria. American Society for Microbiology, Washington, DC, pp 277–289
Eberhard A, Burlingame AL, Eberhard C, Kenyon GL, Nealson KH, Oppenheimer NJ (1981) Biochemistry 20:2444–2449
Nealson KH, Platt T, Hastings JW (1970) J Bacteriol 104:313–322
Farghaly AH (1950) J Cell Physiol 36:165–183
Engebrecht J, Nealson K, Silverman M (1983) Cell 32:773–781
Engebrecht J, Silverman M (1984) Proc Natl Acad Sci USA 81:4154–4158
Kaplan HB, Greenberg EP (1985) J Bacteriol 163:1210–1214
Harvey E (1952) Bioluminescence. Academic, New York
Jones B, Nishiguchi M (2004) Mar Biol 144:1151–1155
Stabb E (2005) ASM News 71:223–229
de Kievit TR, Iglewski BH (2000) Infect Immun 68:4839–4849
Waters CM, Bassler BL (2005) Annu Rev Cell Dev Biol 21:319–346
Fuqua WC, Eberhard A (1999) In: Dunny G, Winans SC (eds) Cell–cell signaling in bacteria. ASM, Washington, DC, pp 211–230
Manefield M, Turner SL (2002) Microbiology 148:3762–3764
Lyon GJ, Novick RP (2004) Peptides 25:1389–1403
Pesci EC, Milbank JB, Pearson JP, McKnight S, Kende AS, Greenberg EP, Iglewski BH (1999) Proc Natl Acad Sci USA 96:11229–11234
Miller ST, Xavier KB, Campagna SR, Taga ME, Semmelhack MF, Bassler BL, Hughson FM (2004) Mol Cell 15:677–687
Schauder S, Shokat K, Surette MG, Bassler BL (2001) Mol Microbiol 41:463–476
Fuqua WC, Winans SC, Greenberg EP (1994) J Bacteriol 176:269–275
Seed PC, Passador L, Iglewski BH (1995) J Bacteriol 177:654–659
Hao G, Burr TJ (2006) J Bacteriol 188:2173–2183
Pestova EV, Havarstein LS, Morrison DA (1996) Mol Microbiol 21:853–862
Schuster M, Greenberg EP (2006) Int J Med Microbiol 296:73–81
Latifi A, Foglino M, Tanaka K, Williams P, Lazdunski A (1996) Mol Microbiol 21:1137–1146
Whiteley M, Parsek MR, Greenberg EP (2000) J Bacteriol 182:4356–4360
Dunlap PV (1989) J Bacteriol 171:1199–1202
Dunlap PV, Greenberg EP (1985) J Bacteriol 164:45–50
Huang JJ, Petersen A, Whiteley M, Leadbetter JR (2006) Appl Environ Microbiol 72:1190–1197
Zhang HB, Wang LH, Zhang LH (2002) Proc Natl Acad Sci USA 99:4638–4643
Huang JJ, Han JI, Zhang LH, Leadbetter JR (2003) Appl Environ Microbiol 69:5941–5949
Sio CF, Otten LG, Cool RH, Diggle SP, Braun PG, Bos R, Daykin M, Camara M, Williams P, Quax WJ (2006) Infect Immun 74:1673–1682
Ochsner UA, Wilderman PJ, Vasil AI, Vasil ML (2002) Mol Microbiol 45:1277–1287
Zhang HB, Wang C, Zhang LH (2004) Mol Microbiol 52:1389–1401
Taga ME, Semmelhack JL, Bassler BL (2001) Mol Microbiol 42:777–793
De Keersmaecker SC, Sonck K, Vanderleyden J (2006) Trends Microbiol 14:114–119
Mok KC, Wingreen NS, Bassler BL (2003) Embo J 22:870–881
Xavier KB, Bassler BL (2003) Curr Opin Microbiol 6:191–197
Beeston AL, Surette MG (2002) J Bacteriol 184:3450–3456
Wang L, Hashimoto Y, Tsao CY, Valdes JJ, Bentley WE (2005) J Bacteriol 187:2066–2076
Taga ME, Miller ST, Bassler BL (2003) Mol Microbiol 50:1411–1427
Xavier KB, Bassler BL (2005) J Bacteriol 187:238–248
DeLisa MP, Valdes JJ, Bentley WE (2001) Biotechnol Bioeng 75:439–450
DeLisa MP, Valdes JJ, Bentley WE (2001) J Bacteriol 183:2918–2928
Surette MG, Bassler BL (1999) Mol Microbiol 31:585–595
Redfield RJ (2002) Trends Microbiol 10:365–370
Gilson L, Kuo A, Dunlap PV (1995) J Bacteriol 177:6946–6951
Lupp C, Ruby EG (2004) J Bacteriol 186:3873–3881
Bassler BL, Wright M, Silverman MR (1994) Mol Microbiol 13:273–286
Hentzer M, Wu H, Andersen JB, Riedel K, Rasmussen TB, Bagge N, Kumar N, Schembri MA, Song Z, Kristoffersen P, Manefield M, Costerton JW, Molin S, Eberl L, Steinberg P, Kjelleberg S, Hoiby N, Givskov M (2003) Embo J 22:3803–3815
Schuster M, Lostroh CP, Ogi T, Greenberg EP (2003) J Bacteriol 185:2066–2079
Wagner VE, Bushnell D, Passador L, Brooks AI, Iglewski BH (2003) J Bacteriol 185:2080–2095
Schuster M, Hawkins AC, Harwood CS, Greenberg EP (2004) Mol Microbiol 51:973–985
Yarwood JM, Volper EM, Greenberg EP (2005) Proc Natl Acad Sci USA 102:9008–9013
Juhas M, Eberl L, Tummler B (2005) Environ Microbiol 7:459–471
Nyholm SV, McFall-Ngai MJ (2004) Nat Rev Microbiol 2:632–642
Ruby EG (1999) J Mol Microbiol Biotechnol 1:13–21
Visick KL (2005) J Bacteriol 187:3603–3606
Lupp C, Ruby EG (2005) J Bacteriol 187:3620–3629
Lupp C, Urbanowski M, Greenberg EP, Ruby EG (2003) Mol Microbiol 50:319–331
Visick KL, Foster J, Doino J, McFall-Ngai M, Ruby EG (2000) J Bacteriol 182:4578–4586
Boettcher KJ, Ruby EG (1990) J Bacteriol 172:3701–3706
Lee KH, Ruby EG (1994) J Bacteriol 176:1985–1991
Callahan SM, Dunlap PV (2000) J Bacteriol 182:2811–2822
Givskov M, de Nys R, Manefield M, Gram L, Maximilien R, Eberl L, Molin S, Steinberg PD, Kjelleberg S (1996) J Bacteriol 178:6618–6622
Manefield M, de Nys R, Kumar N, Read R, Givskov M, Steinberg P, Kjelleberg S (1999) Microbiology 145(Pt 2):283–291
Manefield M, Rasmussen TB, Henzter M, Andersen JB, Steinberg P, Kjelleberg S, Givskov M (2002) Microbiology 148:1119–1127
Gao M, Teplitski M, Robinson JB, Bauer WD (2003) Mol Plant Microbe Interact 16:827–834
Teplitski M, Chen H, Rajamani S, Gao M, Merighi M, Sayre RT, Robinson JB, Rolfe BG, Bauer WD (2004) Plant Physiol 134:137–146
Teplitski M, Robinson JB, Bauer WD (2000) Mol Plant Microbe Interact 13:637–648
Dong YH, Gusti AR, Zhang Q, Xu JL, Zhang LH (2002) Appl Environ Microbiol 68:1754–1759
Dong YH, Xu JL, Li XZ, Zhang LH (2000) Proc Natl Acad Sci USA 97:3526–3531
Kang BR, Lee JH, Ko SJ, Lee YH, Cha JS, Cho BH, Kim YC (2004) Can J Microbiol 50:935–941
Leadbetter JR, Greenberg EP (2000) J Bacteriol 182:6921–6926
Lee SJ, Park SY, Lee JJ, Yum DY, Koo BT, Lee JK (2002) Appl Environ Microbiol 68:3919–3924
Uroz S, Chhabra SR, Camara M, Williams P, Oger P, Dessaux Y (2005) Microbiology 151:3313–3322
Uroz S, D’Angelo-Picard C, Carlier A, Elasri M, Sicot C, Petit A, Oger P, Faure D, Dessaux Y (2003) Microbiology 149:1981–1989
Zhang LH, Dong YH (2004) Mol Microbiol 53:1563–1571
Flagan S, Ching WK, Leadbetter JR (2003) Appl Environ Microbiol 69:909–916
Chun CK, Ozer EA, Welsh MJ, Zabner J, Greenberg EP (2004) Proc Natl Acad Sci USA 101:3587–3590
Ozer EA, Pezzulo A, Shih DM, Chun C, Furlong C, Lusis AJ, Greenberg EP, Zabner J (2005) FEMS Microbiol Lett 253:29–37
Rasmussen TB, Givskov M (2006) Microbiology 152:895–904
Rasmussen TB, Givskov M (2006) Int J Med Microbiol 296:149–161
Shiner EK, Rumbaugh KP, Williams SC (2005) FEMS Microbiol Rev 29:935–947
Acknowledgements
This work was supported by CAREER award MCB-0347317 from the National Science Foundation to EVS. The authors thank J.L. Bose for contributing to Fig. 2.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dunn, A.K., Stabb, E.V. Beyond quorum sensing: the complexities of prokaryotic parliamentary procedures. Anal Bioanal Chem 387, 391–398 (2007). https://doi.org/10.1007/s00216-006-0730-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-006-0730-9