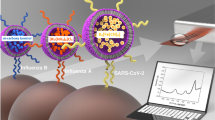
Abstract
A multi-analyte biosensor based on nucleic acid hybridization and liposome signal amplification was developed for the rapid serotype-specific detection of Dengue virus. After RNA amplification, detection of Dengue virus specific serotypes can be accomplished using a single analysis within 25 min. The multi-analyte biosensor is based on single-analyte assays (see Baeumner et al (2002) Anal Chem 74:1442–1448) developed earlier in which four analyses were required for specific serotype identification of Dengue virus samples. The multi-analyte biosensor employs generic and serotype-specific DNA probes, which hybridize with Dengue RNA that is amplified by the isothermal nucleic acid sequence based amplification (NASBA) reaction. The generic probe (reporter probe) is coupled to dye-entrapping liposomes and can hybridize to all four Dengue serotypes, while the serotype-specific probes (capture probes) are immobilized through biotin–streptavidin interaction on the surface of a polyethersulfone membrane strip in separate locations. A mixture of amplified Dengue virus RNA sequences and liposomes is applied to the membrane and allowed to migrate up along the test strip. After the liposome-target sequence complexes hybridize to the specific probes immobilized in the capture zones of the membrane strip, the Dengue serotype present in the sample can be determined. The amount of liposomes immobilized in the various capture zones directly correlates to the amount of viral RNA in the sample and can be quantified by a portable reflectometer. The specific arrangement of the capture zones and the use of unlabeled oligonucleotides (cold probes) enabled us to dramatically reduce the cross-reactivity of Dengue virus serotypes. Therefore, a single biosensor can be used to detect the exact Dengue serotype present in the sample. In addition, the biosensor can simultaneously detect two serotypes and so it is useful for the identification of possible concurrent infections found in clinical samples. The various biosensor components have been optimized with respect to specificity and sensitivity, and the system has been ultimately tested using blind coded samples. The biosensor demonstrated 92% reliability in Dengue serotype determination. Following isothermal amplification of the target sequences, the biosensor had a detection limit of 50 RNA molecules for serotype 2, 500 RNA molecules for serotypes 3 and 4, and 50,000 molecules for serotype 1. The multi-analyte biosensor is portable, inexpensive, and very easy to use and represents an alternative to current detection methods coupled with nucleic acid amplification reactions such as electrochemiluminescence, or those based on more expensive and time consuming methods such as ELISA or tissue culture.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dengue virus, a single stranded RNA virus of the family Flaviviridae, is the most common cause of arboviral disease in the world. There are four closely related but antigenically distinct serotypes (Dengue 1–4). All four serotypes produce a similar illness (Dengue fever, DF) characterized by mild, flu-like symptoms and induce life-long immunity that is specific to the infecting serotype. However, more severe and potentially fatal diseases, Dengue hemorrhagic fever (DHF) and Dengue shock syndrome, marked by abnormal vascular permeability, are developed after secondary infection with a different serotype. Based on recent studies [1], T-cells activated during the second infection have less affinity for the infecting serotype than the previously encountered serotype, causing a delay in clearing the virus and contributing to immunopathology. Dengue virus is a major public health concern in tropical and subtropical areas. It causes an estimated 50 million illnesses annually, including 250,000–500,000 cases of DHF with 5–10% mortality. A total of 2.5 billion people, representing more than two-fifths of the world’s population, live in areas potentially at risk from Dengue virus infection [2, 3].
No effective vaccine or specific therapeutic agents exist to prevent or cure the disease caused by Dengue virus, and only a few control programs have been shown to be effective against the mosquito vectors. Over the past 60 years, the incidence, distribution, and clinical severity of DF/DHF has increased dramatically. Therefore, reliable and fast diagnostic methods are necessary for the surveillance and proper treatment of Dengue virus infection. Serological tests (MAC-ELISA, ELISA, and so on) are currently available to measure IgM and IgG antibodies to the Dengue virus [4, 5]. However, they cross-react to a variable extent with other flaviviruses and generally have to be performed at least five days after the onset of illness since detectable levels of specific antibodies are not present prior to that. Presently, isolation of the virus is the most accurate method of determining the identity of the specific Dengue serotype responsible for a particular infection. It is frequently possible to isolate the virus from clinical specimens during the early acute phase of DF [6]. Nevertheless, such tissue culture-based methods for the isolation and identification of Dengue virus require a week or longer of labor-intensive work. Molecular biological diagnosis systems, such as RT-PCR and TaqMan RT-PCR also can be used to detect viral RNA rapidly and specifically from the patient’s serum in the viremia phase [7, 8]. But they often suffer from sample contamination and require expensive and unwieldy instrumentation unsuitable for field applications, especially when analysis costs are a major concern. Developing a nucleic acid sequence-based amplification reaction for the identification of Dengue viruses [9] circumvents thermocycling during the amplification reaction and is therefore an ideal reaction for miniaturization and eventually inexpensive field application. However, this technology must be paired with a simple detection device in order to prove field-usable.
Biosensors based on nucleic acid hybridization and liposome signal amplification have been shown to be very successful for the development of inexpensive, rapid, and easy to handle systems for the detection and quantification of RNA molecules [10, 11, 12, 13, 14]. Our laboratory has previously reported the use of RNA biosensors for the rapid, sensitive and serotype-specific detection of Dengue virus [15]. However, in a few cases cross-reactivity between serotype-specific biosensors was encountered. Moreover, the performance of four separate assays was required to discriminate one serotype from the others. Each individual assay employed a single membrane that had a single-capture zone. In the present work, we describe a novel biosensor, which permits the simultaneous identification of all four Dengue virus serotypes by immobilization of all four serotype-specific DNA capture probes in discrete zones on a single-membrane strip. The biosensor principle and format are shown in Fig. 1.
Principle of the biosensor assay. A DNA reporter probe is coupled to a liposome while a DNA capture probe is immobilized on a polyethersulfone membrane by means of a biotin-streptavidin linkage. When a specific Dengue virus RNA is present (a), a sandwich is formed between the reporter probe, the RNA molecule and the capture probe. The amount of liposomes captured in the detection zone is proportional to the amount of Dengue virus RNA. No signal will be observed when nonspecific RNA is present (b)
The biosensor is based on nucleic acid hybridization; serotype-specific and generic DNA probes hybridize with Dengue viral RNA. The specificity of the probes was previously described [9]. The generic probe (reporter probe), coupled to liposomes with encapsulated fluorescent dye, was designed to hybridize to sequences found in all four Dengue serotypes. The serotype-specific probes (capture probes) were immobilized on the surface of a polyethersulfone membrane strip via biotin–streptavidin interaction. Viral RNA was amplified using the nucleic acid sequence based amplification (NASBA) reaction, which exclusively amplifies single-stranded RNA molecules under isothermal conditions and results in more than a 1010 -fold increase in RNA concentration within a 90-min reaction [16]. In the biosensor assay, liposomes were mixed with the amplified Dengue virus RNA and the mixture was applied to a polyethersulfone membrane. By capillary action, the mixture migrated along the membrane strip and upon passing the capture zone a sandwich was formed by hybridization of probe-tagged liposomes, RNA target molecules, and the immobilized capture probe. Unbound liposomes accumulated at the end of the membrane strip. Therefore, the amount of liposomes present in the immobilized complexes correlated directly with the concentration of viral RNA present in the sample. The new biosensor assay format utilized only two cartridges with multi-zone membrane strips: one for sample analysis and one for a negative control. By judicious arrangement of the specific order of the immobilized capture probes and the use of unlabeled oligonucleotides (cold probes) the cross-reactivity between Dengue virus serotypes could be dramatically reduced. The multi-analyte biosensor was successfully applied to the determination of concurrent infections and to the identification of coded samples.
Materials and methods
Reagents
All general chemicals and buffer reagents were obtained from Sigma Company, St. Louis, MO, USA. Organic solvents were purchased from Aldrich Chemical Company, Milwaukee, WI, USA. Membranes were obtained from Pall/Gelman Company, Port Washington, NY, USA. Lipids were purchased from Avanti Polar Lipids, Alabaster, AL, USA. Sulforhodamine B and streptavidin were acquired from Molecular Probes Company, Eugene, OR, USA. All oligonucleotides were purchased from Qiagen, Valencia, CA, USA.
DNA probes
DNA oligonucleotides (probes) had been designed earlier [9] and their sequences were given in [15]. A generic sequence was introduced into Dengue virus RNA during the NASBA reaction and, regardless of the specific serotype, was therefore present in each amplified Dengue RNA. A generic probe [15] complementary to this region was used as reporter probe and coupled to liposomes via an amine group at the 3′ end. A conserved capture probe [15] was used that could hybridize to a region contained within the RNA found in all four Dengue serotypes. The conserved and four serotype-specific capture probes were modified with a biotin at the 5′ end and immobilized on the streptavidin coated membrane surface. The sequence layout is shown in Fig. 2.
The location of the reporter and capture probes in Dengue virus RNA. The generic reporter probe coupled to dye-entrapping liposomes hybridizes to a generic sequence present in every amplified Dengue virus RNA. The four serotype-specific capture probes and the conserved capture probe are immobilized onto the surface of a membrane strip
In addition, DNA “cold probes” were used to prevent the cross-reactivity of different serotypes (see “Results and discussion”). They are unlabeled oligonucleotides with sequences identical to serotype-specific capture probes [15]. Additional cold probe 3 was designed with 5′-Agg gAA gCT gTA CCT CCT TgC AAA g-3′ sequence. All probes were obtained desalted and lyophilized.
Liposome preparation
The liposomes were prepared using a modified version [13] of the reversed phase evaporation method described by Siebert et al [17].
Conjugating reporter probe to liposome surface
The reporter probe modified with an amine group at the 3′ end was coupled to acetylthioacetate-tagged liposomes following the usual protocol [13].
Multi-zone membrane preparation
The membranes were cut into 4.5×100 mm strips. Five mixtures each containing 20 pmol streptavidin and 60 pmol capture probe (one conserved and four serotype-specific) in 1 μl of 0.4 M sodium carbonate buffer with 5% methanol were incubated for at least 15 min at room temperature. Capture zone 2 was created by pipetting 1 μl of the capture probe–streptavidin solution directly onto the membrane approximately 1.5 cm from the bottom (origin) of the membrane strip. Then in a similar manner the other capture zones were created at 1 cm intervals along the same membrane strip in the final order of 2, 4, 4, 3, 1, and C (conserved) (the order will be discussed later in the “Discussion”). Therefore, each membrane strip had six capture zones.
The subsequent procedures of oligonucleotide immobilization, membrane blocking and their storage were the same as described previously [13].
Biosensor assay format
A vertical flow assay format was developed. In disposable culture tubes (10×75 mm, VWR) 2 μl of liposomes, 2 μl of a hybridization solution (master mix) (for composition, see below), and 2 μl of target sequence (amplified Dengue virus RNA) or water (a negative control) were incubated for 15 min at room temperature. A volume of 1 μl of 0.4 mM cold probe 4 was applied directly onto the membrane strip between the two Dengue 4 capture zones and the membrane was then immediately inserted into the mixture solution. As soon as the hybridization mixture completely soaked into the strip, 40 μl of running buffer (for composition, see below) were added at the bottom of the glass tube and allowed to migrate up the strip. After the running buffer reached the upper end of the strip (~10 min), membranes were removed from tubes and dried in the air for 20–30 min (drying can be omitted to speed up the assay time). The capture zones were analyzed with the BR-10 reflectometer (ESECO; Cushing, OK, USA) by placing the capture zone directly under the reflectometer opening. The reflectometer measures the reflectance of light at a wavelength of 560 nm, which is close to the absorbance maximum of sulforhodamine B encapsulated within the liposomes. The reflectometer was calibrated each time before use with white and magenta strips to a minimum value of zero and a maximum of 155 arbitrary units (AU), respectively. Both the master mix and running buffer solutions contained formamide, SSC (1 x SSC contains 15 mM sodium citrate and 150 mM NaCl, pH 7.0), Ficoll type 400, sucrose and Triton X-100. The concentrations of each component were optimized for the biosensor assay. Additionally, different liposome and target sequence concentrations were also investigated. The optimized hybridization mixture had the following composition for all assays: 2 μl of master mix (60% formamide, 6xSSC, 0.15 M sucrose, 0.8% Ficoll, 0.01% Triton X-100), 2 μl of liposomes and 2 μl of target sequence (dilution 1:5 or 1:10). The optimal running buffer concentrations were 10% formamide, 3xSSC, 0.2 M sucrose, 0.2% Ficoll, 0.01% Triton X-100.
Dengue virus samples
In the assays four different sets of Dengue RNA samples were used (Table 1).
Since we did not use live virus preparations in our laboratories at Cornell University, amplified RNA sequences were shipped to us by Advanced BioScience Laboratories, Inc. Briefly, seed stocks of all four serotypes of Dengue virus of four sets were prepared in Vero cells, and virus titers were determined by plaque assays [18]. Dengue viral RNA was extracted using the method of Boom et al [19] resulting in a final volume of 50 μl of elution buffer. A 5 μl portion of the extract was then amplified using NASBA [9]. For reamplification, Dengue amplicons of set 1 were diluted 1:10 for Dengue 1 and 1:10,000 for Dengue 2–4 and NASBA was performed as usual, using 5 μl of the diluted template. Before shipment to our laboratory, the amplification reaction products were characterized by electrochemiluminescence (ECL) analysis [9] (Table 2).
Results and discussion
The hybridization buffers (master mix and running buffer) were initially optimized with respect to signal-to-noise ratio and cross-reactivity of the different serotype RNA sequences to the various capture probes. This was especially important since, unlike the previously described single-analyte biosensors [15] or ECL methods [9], only limited amounts, or none at all, of cold probes could be added to the multi-analyte assay. It was observed that the use of either cold probe 1 [15] or cold probe 3 was not feasible in the multi-analyte biosensor since their presence diminished the intensity of both specific as well as nonspecific signals. The concentrations of SSC (0–20x) and formamide (0–60%) were varied in the master mix and running buffer in order to optimize the stringency conditions. In addition, the amount of capture probes (10–130 pmoles) in each zone and the amount of liposomes (2–6 μl) were optimized.
Design of multi-analyte biosensor
The multi-analyte biosensor design was based on the cross-reactivity investigation of one of the amplicon sets that was available in the beginning of this study (set 2, Table 1). For the cross-reactivity analysis, biosensors with single capture zone design were used [15]. Each Dengue virus serotype was analyzed with all four serotype-specific single-analyte biosensors. As an example, the results for Dengue virus serotype 4 are shown in Fig. 3.
Reflectometer readings for the specific and nonspecific signals are given in AU in Fig. 4. As can be seen, biosensors for Dengue serotypes 2 and 4 were highly specific, but biosensors 1 and 3 had cross-reactivities with Dengue serotype 4. Derived from these results, an initial design of a multi-analyte biosensor was suggested (Fig. 5a).
Cross-reactivity of Dengue serotype-specific biosensors. Amplicons of all four serotypes were investigated with the biosensor designed for the detection of serotype 1, then with the serotype-specific biosensor 2, and so on. The individual serotype 1 and 3 biosensors also gave positive signals when Dengue 4 was analyzed. The background signal (4–7 AU) obtained from assays without Dengue RNA was subtracted from all raw reflectometer values. No cold probes were used in these assays
a Initially proposed capture zone order. Biotinylated capture probes (60 pmoles) complementary to Dengue serotypes 2, 4, 3, 1 and conserved regions were immobilized on membrane strips in the order shown. b Final design of the multi-analyte biosensor for the detection of four Dengue virus serotypes. Each capture zone contained 60 pmoles of specific probes, and the concentration of cold probe 4 was 0.4 mM
The position for capture zone 2 was selected near the origin of the membrane test strip based on the previous results that biosensor 2 was always highly specific [15]. Capture zone 4 was placed before 3 and 1 assuming that if all Dengue 4 could be captured, false positive signals would not be observed in capture zones 1 and 3. In fact, we did successfully eliminate the cross-reactivity between Dengue serotypes 4 and 3. However, strong nonspecific signal was still detected on capture zone 1 when Dengue 4 was analyzed. In order to bind all Dengue 4 RNA on one capture site we initially varied the amount of specific capture probe 4 immobilized on the membrane strip in the range from 10 to 130 pmoles. It was observed that the optimum was 60 pmoles. Higher quantities of capture probes, however, resulted in decreased signal intensity, presumably due to steric hindrance in the limited area of the capture zone. Furthermore, nonspecific interactions between oligonucleotides at very high quantity could interfere with the specific hybridization to the target RNA. Therefore, a second capture zone 4 was introduced immediately adjacent to the first one, thus ultimately increasing the amount of capture probe available for binding to the target RNA. However, the use of two successive capture zones for Dengue 4 did not entirely solve the cross-reactivity problem: Dengue virus serotype cross-reactivity to capture probe zone 1 was still observed. Finally, a solution of cold probe 4 [15] was added between the two capture zones 4. The concentration of the cold probe was optimized to be 0.4 mM. At this concentration when Dengue 4 was analyzed, positive signals were only observed within the two capture 4 zones and the conserved zone. Therefore, the increased amount of immobilized capture probe and the presence of free cold probe 4 were sufficient to bind to all of Dengue virus serotype 4 RNA present in a sample and prevent subsequent nonspecific binding to capture probes 3 and 1. The final design of the multi-analyte biosensor is shown in Fig. 5b. Subsequently, the multi-analyte biosensor was tested with all four serotypes (Fig. 6).
Analysis of four Dengue virus serotypes with the multi-analyte biosensor. For a negative control water was used instead of Dengue amplicon. Reflectometer readings for serotype-specific biosensors are shown in Table 3, set 2
In this way a functional multi-analyte biosensor was successfully developed in which serotype-specific signals with no cross-reactivities were obtained, as well as a generic signal through the generic capture site C at the end of the strip.
Testing the multi-analyte biosensor with samples of Dengue virus from different strains
Dengue virus amplicons from other virus strains (sets 2–4) were tested with the optimized multi-analyte biosensor.
Despite diverse cross-reactivity of different strains (their cross-reactivity pattern was investigated as described before using single-analyte biosensor design (Figs. 3 and 4)), the multi-analyte biosensor worked very well with all Dengue virus samples and all four serotypes were identified correctly. The results were similar to those shown in Fig. 6 and are summarized in Table 3. No visible pink color was noticed for the background, with reflectometer values equal to 4–7 AU. Only a few pale pink spots were observed on the multi-analyte test strips that corresponded to nonspecific interactions between sample and specific probes and had reflectometer values of 12±2. Therefore, a biosensor signal above 14 was considered positive. The values for the positive signals varied between 16 and 44, which corresponded to the ECL values of the signals given in Table 2. Due to the manner in which NASBA amplification was performed on these samples, quantitative results could not be obtained, so the RNA concentrations in the amplicons do not correlate with the number of virus particles in the sample. Therefore, the biosensor could only be used for a qualitative analysis as well. However, if a quantitative NASBA reaction was performed [20], the multi-analyte biosensor could also provide quantitative data.
Simultaneous detection of two serotypes
Outbreaks involving more than one serotype of Dengue virus have been reported in several countries in Southeast Asia as well as in Central and South America [21]. Concurrent infections by two serotypes have been observed in the same individual [22, 23]. Therefore, the developed multi-analyte biosensor was tested with different mixtures of two Dengue virus samples. All possible combinations of Dengue virus serotypes (D1-D2, D1-D3, D1-D4, D2-D3, D2-D4, D3-D4) were investigated. The results are shown in Fig. 7.
Simultaneous detection of two Dengue virus serotypes. Amplicons from set 4 were used in the experiment. Reflectometer readings were similar to those shown in Table 3 (set 4)
The specific signals for each of two serotypes can be observed clearly. Therefore, the multi-analyte biosensor can be successfully applied to the identification of concurrent infections in clinical samples.
Sensitivity
A known number of RNA molecules of each Dengue virus serotype was amplified using NASBA and subsequently analyzed using ECL (Table 4) and the multi-analyte biosensor. Using these sets of data, it was determined that the multi-analyte biosensor could detect samples with ECL values of 100,000 or higher. Therefore, the multi-analyte biosensor could detect 50,000 RNA molecules of Dengue 1, 50 RNA molecules of Dengue 2 and 500 RNA molecules of Dengue 3 and 4. Since naturally-occurring Dengue samples would be expected to have ECL values above 100,000, our multi-analyte biosensor will provide excellent analytical sensitivity.
Coded sample identification
A set of 20 coded samples was provided for our laboratory by our collaborators at ABL and IBI. A very low number of RNA molecules (500 copies) was amplified using NASBA resulting in ECL values of 14,000–31,300 for Dengue 1, 336,700–835,700 for Dengue 2, 72,300–300,100 for Dengue 3, and 124,100–502,300 for Dengue 4 (Table 5). Since the ECL values for Dengue 1 and partly also for Dengue virus 3 were below the detection limit of the multi-analyte biosensor, samples containing these serotypes were not identified correctly. However, eliminating out of consideration the samples of serotype 1 and 3 with ECL values below the detection threshold, it was possible to identify the correct serotype present in 92% of the samples, including samples containing only water.
In the future, we will continue with a second coded-sample experiment, using higher amounts of RNA molecules (at least 50,000) than those tested here.
Conclusion
We have developed a serotype-specific multi-analyte biosensor for the rapid detection of Dengue virus. Dengue viruses of different strains can be detected very specifically by carrying out a single analysis. The biosensor demonstrated 92% reliability in serotype identification and is able to detect as few as 50 RNA molecules of serotype 4, 500 molecules of serotype 2 and 3, and 50,000 molecules of serotype 1 if an amplification reaction has been carried out prior to the analysis. In addition, the biosensor permits the simultaneous determination of any two serotypes and can therefore be applied to the analysis of concurrent infections. In the future, as clinical samples become available, we will analyze them using the multi-analyte biosensor we have developed.
References
Mongkolsapaya J, Dejnirattisai W, Xu X, Vasanawathana S, Tangthawornchaikul N, Chairunsri A, Sawasdivorn S, Duangchinda T, Dong T, Rowland-Jones S, Yenchitsomanus P, McMichael A, Malasit P, Screaton G (2003) Nat Med 9:921–927
Gubler D (1997) In: Gubler DJ, Kuno G (eds) Dengue and dengue hemorrhagic fever. CAB International, Cambridge, pp 1–22
World Health Organization (2000) Strengthening the implementation of the global strategy for dengue fever/dengue hemorrhagic fever prevention and control. WHO, Geneva
Chakravarti A, Gur R, Berry N, Mathur M (2000) Diagn Microbiol Infect Dis 36:273–274
Balmaseda A, Guzman M, Hammond S, Robleto G, Flores C, Tellez Y, Videa E, Saborio S, Perez L, Sandoval E, Rodriguez Y, Harris E (2003) Clin Diag Lab Immunol 10:317–322
Henchal E, Putnak J (1990) Clin Microbiol Rev 3:376–396
Lanciotti R, Calisher C, Gubler D, Chang G, Vorndam A (1992) J Clin Microbiol 30:545–551
Laue T, Emmerich P, Schmitz H (1999) J Clin Microbiol 37:2543–2547
Wu S, Lee E, Putvatana R, Shurtliff R, Porter K, Suharyono W, Watt D, King C, Murphy G, Hayes C, Romano J (2001) J Clin Microbiol 39:2794–2798
Esch M, Baeumner A, Durst R (2001) Anal Chem 73:3162–3167
Baeumner A, Cohen R, Miksic V, Min J (2003) Biosens Bioelectron 18:405–413
Baeumner A, Hartley H (2003) Anal Bioanal Chem 376:319–327
Baeumner A, Pretz J, Fang S (2004) Anal Chem 76:888–894
Baeumner A, Jones C, Wong C (2004) Anal Bioanal Chem 378:1587–1593
Baeumner A, Schlesinger N, Slutzki N, Romano J, Lee E, Montagna R (2002) Anal Chem 74:1442–1448
Compton J (1991) Nature 360:91–92
Siebert S, Reeves S, Durst R (1993) Anal Chim Acta 282:297–305
Eckels K, Brandt W, Harrison V, McCown J, Russel P (1976) Infect Immunol 14:1221–1227
Boom R, Sol C, Salimans M, Jansen C, Wertheim-van Dillen P, van der Norda J (1990) J Clin Microbiol 20:495–503
Romano J, Shurtliff R, Dobratz E, Gibson A, Hickman K, Markham P, Pal R (2000) J Virol Methods 86:61–70
CDC (1998) Morbidity and mortality weekly report (MMWR). Centers for Disease Control and Prevention, Atlanta, GA, pp 952–956
Wang W, Chao D, Lin S, King C, Chang S (2003) J Microbiol Immunol Infect 36:89–95
Rocco I, Barbosa M, Kanomata E (1998) Rev Inst Med Trop Sao Paulo 40:151–154
Acknowledgements
The authors acknowledge financial support for this project from Innovative Biotechnologies International, Inc., from the National Institute of Allergy and Infectious Diseases (NIAID), Bethesda, MD, USA, from the New York State CAT, Biotechnology Program at Cornell University, and from the Cooperative State Research, Education and Extension Services (NYC-123314). We thank M.G. Sarngadharan and Roxanne N. Shurtliff from Advanced BioScience Laboratories, Inc., for their many discussions and help in moving the project along. We also want to thank Sutee Yoksan from Mahidol University, Thailand and Shuenn-Jue Wu from Naval Medical Research Center, USA, for providing us with Dengue virus samples that were used in this publication.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zaytseva, N.V., Montagna, R.A., Lee, E.M. et al. Multi-analyte single-membrane biosensor for the serotype-specific detection of Dengue virus. Anal Bioanal Chem 380, 46–53 (2004). https://doi.org/10.1007/s00216-004-2724-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-004-2724-9