Abstract
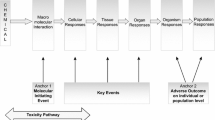
Adverse outcome pathways (AOPs) have been recently introduced as tools to map the mechanisms underlying toxic events relevant for chemical risk assessment. AOPs particularly depict the linkage between a molecular initiating event and an adverse outcome through a number of intermediate key events. An AOP has been previously introduced for cholestatic liver injury. The objective of this study was to test the robustness of this AOP for different types of cholestatic insult and the in vitro to in vivo extrapolation. For this purpose, in vitro samples from human hepatoma HepaRG cell cultures were exposed to cholestatic drugs (i.e. intrahepatic cholestasis), while in vivo samples were obtained from livers of cholestatic mice (i.e. extrahepatic cholestasis). The occurrence of cholestasis in vitro was confirmed through analysis of bile transporter functionality and bile acid analysis. Transcriptomic analysis revealed inflammation and oxidative stress as key events in both types of cholestatic liver injury. Major transcriptional differences between intrahepatic and extrahepatic cholestatic liver insults were observed at the level of cell death and metabolism. Novel key events identified by pathway analysis included endoplasmic reticulum stress in intrahepatic cholestasis, and autophagy and necroptosis in both intrahepatic as extrahepatic cholestasis. This study demonstrates that AOPs constitute dynamic tools that should be frequently updated with new input information.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cholestasis denotes any situation of impaired bile secretion concomitant with an accumulation of bile acids (BAs) in the liver and/or systemic circulation (Nguyen et al. 2014; Noor 2015). Two types of cholestasis are traditionally distinguished, depending on the site of obstruction, namely extrahepatic and intrahepatic cholestasis. The latter is the result of functional defects in bile formation by hepatocytes, while extrahepatic cholestasis is caused by an anatomical blockage outside the liver (Mariotti et al. 2017; Zollner and Trauner 2006). Drug-induced intrahepatic cholestasis constitutes a major subgroup of drug-induced liver injuries (DILI), accounting for as much as 50% of the registered DILI cases (Oorts et al. 2016). DILI is of high clinical importance, deemed responsible for more than half of all cases of acute liver failure (Goldberg et al. 2015). In addition to its clinical relevance, DILI is also one of the major reasons for drug failure during pre-marketing and post-marketing phases of drug development, leading to massive financial losses for pharmaceutical industry (Jones et al. 2018; Lee et al. 2013; Van den Hof et al. 2015). Unfortunately, current pre-clinical animal and in vitro models can only detect about 50–60% of all human clinical DILI cases. This could be explained, at least in part, by significant gaps in the mechanistic understanding of DILI, including drug-induced cholestasis (Bale et al. 2014; Laverty et al. 2010). For this reason, there is an urgent need to map the full mechanistic scenario of drug-induced intrahepatic cholestasis to identify biomarkers that can improve prediction of cholestatic liabilities. An adverse outcome pathway (AOP) seems fit for this purpose, as it provides a conceptual construct that portrays existing knowledge concerning the linkage between a direct molecular initiating event and an adverse outcome at a biological level relevant to risk assessment (Ankley et al. 2010; Villeneuve et al. 2014). An AOP on cholestatic liver injury has been previously introduced by our group, thereby focusing on inhibition of the bile salt export pump (BSEP) as molecular initiating event (Vinken et al. 2013). BSEP plays an essential role in the hepatocyte export of BAs and its inhibition results in intrahepatocyte accumulation of BAs. The latter key event initiates two types of cellular responses, namely a deteriorative response and an adaptive response. The deteriorative response is characterized by the occurrence of inflammation, opening of the mitochondrial permeability pore, oxidative stress, and cell death. The adaptive response reflects a homeostatic reaction to counteract BA accumulation via activation of a number of nuclear receptors (NRs) [i.e. constitutive androstane receptor (CAR), farnesoid X receptor (FXR), pregnane X receptor (PXR), and small heterodimer partner (SHP)]. This induces an array of transcriptional changes to facilitate removal of BAs and their products (Vinken et al. 2013).
AOPs are to be considered as living documents that evolve over time as new knowledge becomes available (Villeneuve et al. 2014). In this respect, the present study was set up to verify the relevance and reliability of the available AOP for predicting different types of cholestatic liver injury in vitro and in vivo as well as to generate new information that could be used during further AOP optimization. Intrahepatic cholestasis was mimicked by human hepatoma HepaRG cells that were exposed to different cholestatic drugs, while extrahepatic cholestasis was surgically induced in mice by means of bile duct ligation (BDL). Cell culture and liver tissue samples were subjected to transcriptomic analysis with subsequent pathway analysis, and results were benchmarked against the available AOP on cholestatic liver injury.
Materials and methods
Animals and treatment
Male 8-weeks-old Sv129 mice were purchased form Harlan (Horst, The Netherlands). Mice were housed in the animal facility of the Faculty of Medicine and Health Sciences at Ghent University, Belgium, and acclimatized 1 week prior the experiments under controlled conditions. Care was given in accordance with Federation for Laboratory Animal Science Associations guidelines and the national guidelines for animal protection. The animal protocols used in this study were evaluated and approved by the Ethical Committee of Experimental Animals at the Faculty of Medicine and Health Sciences, Ghent University, Belgium (ECD 15/36). Cholestatic liver injury was induced by performing BDL as previously described (Van Campenhout et al. 2019). Briefly, mice were anaesthetized with isoflurane inhalation (Isoflo, Abbott, Belgium), a midline abdominal incision was made, and the common bile duct was isolated and ligated with two knots of non-resorbable suture (Silkan 5/0, Braun Aesculap, Germany). The first ligature was made below the junction of the hepatic ducts and the second was made above the entrance of the pancreatic duct. The common bile duct was resected between the two ligatures, after which the abdomen was closed by suturing the abdominal muscle and skin in two separate layers. Control mice were sham operated, whereby the common bile duct was isolated, but not ligated. Liver samples were collected 6 weeks post-surgery.
Cell cultures and treatment
Cryopreserved differentiated HepaRG cells (Biopredic International, France) were thawed and seeded following manufacturer’s instructions with basal hepatic cell medium (Williams’ E basal medium with GlutaMAX containing phenol red, Biopredic International, France) supplemented with thaw seed and general purpose medium (Biopredic International, France). Hereafter, HepaRG cells were maintained in basal hepatic cell medium supplemented with maintenance and metabolism medium (Biopredic International, France) and refreshed every 2–3 days. Stock solutions of atazanavir (ATA), cyclosporin A (CsA), and nefazodone (NEFA) were made in dimethyl sulfoxide (DMSO) (Sigma-Aldrich, Belgium). The final drug solutions were prepared ex tempore by diluting the concentrated stock solutions with basal hepatic cell medium supplemented with induction serum-free medium (Biopredic International, France) and contained a final DMSO concentration of 0.25% v/v. A 50 × concentrated mixture of 5 BAs (i.e. 66 µM glycochenodeoxycholic acid, 20 µM deoxycholic acid, 19.5 µM chenodeoxycholic acid, 19 µM glycodeoxycholic acid, and 17.5 µM glycocholic acid, Sigma-Aldrich, Belgium) was included in the cell culture medium of HepaRG cells from day 7 after seeding in combination with the drug. Incubations with drugs were routinely carried out for 72 h with daily renewal of cell culture media, including the BA mix and drugs.
Cell viability assessment
HepaRG cell viability was assessed by means of a 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) viability assay (Sigma-Aldrich, Belgium) (Mosmann 1983). MTT was dissolved in phosphate-buffered saline at a concentration of 5 mg/mL (pH 7.65). A concentration of 0.5 mg/mL was obtained by performing a 10 × dilution with Williams’ E medium without phenol red. HepaRG cells were seeded in 96-well plates following the manufacturer’s instructions. At day 7, HepaRG cells were exposed for 72 h to a range of concentrations of ATA (i.e. 5–100 µM), CsA (i.e. 5–50 µM), and NEFA (i.e. 5–50 µM). Thereafter, the cells were rinsed with phosphate-buffered saline and incubated with MTT solution for 1.5 h at 37 °C in a 5% CO2 atmosphere. The formed formazan crystals were dissolved in DMSO by shaking the multi-well plates on an orbital shaker (VWR, Belgium) for 10 min at room temperature and protected from light. Finally, the absorbance of each well of the multi-well plate was measured with a spectrophotometer (PerkinElmer, Belgium) at 560 ± 10 nm and conversely correlated with the viability. Cholestatic index (CIx) values were determined for every drug. CIx values reflect the functionality or viability of hepatocytes co-exposed to a cholestatic drug plus the BA mix compared to exposure to the cholestatic drug alone (Chatterjee et al. 2014; Hendriks et al. 2016). In the present study, data acquired from the MTT assays were used to calculate CIx values.
Compounds were considered to possess a cholestatic potential when CIx values were below or equal to 0.8 (Hendriks et al. 2016).
Determination of bile salt efflux pump activity
HepaRG cells were seeded in 96-well plates following manufacturer’s instructions. At day 7, HepaRG cells were incubated for 30 min with 5 µM of the specific BSEP probe cholyl-l-lysyl-fluorescein (CLF) dissolved in Hanks' balanced salt solution (excitation/emission wavelength 450/530 nm), while being exposed to cholestatic concentrations of ATA, CsA and NEFA or the BA mix. After 30 min of incubation, cells were rinsed, and nuclei were stained with Hoechst 33342 (excitation/emission wavelength 365/420 nm). The cell culture dishes were imaged using a Nikon Eclipse Ti2. Fluorescence images were collected at × 200 magnification.
Quantification of bile acids
HepaRG cells were seeded in 24-well plates following manufacturer’s instructions. At day 7, HepaRG cells were exposed to cholestatic concentrations of ATA, CsA, and NEFA with or without the BA mix added to the cell culture medium. After 72 h of exposure, HepaRG cells were rinsed with cold Hanks’ balanced salt solution (Thermo Fisher, Belgium) and subsequently collected using 250 µL cold methanol per well. Quantification of BAs was performed as previously described (Dewaele et al. 2019). The cells were kept at − 20 °C until further analysis. A total of six samples were pooled per condition and centrifuged at 20,168×g for 15 min at 4 °C with a Mikro 220R centrifuge (Hettich, The Netherlands). Furthermore, samples were evaporated using a Savant Speedvac concentrator (Thermo Scientific, USA) and reconstituted in 100 µL 50/50 MeOH/ammonium buffer (5 mM adjusted to pH 3.6 with acidic acid). Finally, the samples were transferred into autosampler vials for liquid chromatography and mass spectrometry/mass spectrometry analysis and cholic acid (CA), chenodeoxycholic acid (CDCA), deoxycholic acid (DCA), glycocholic acid (GCA), glycochenodeoxycholic acid (GCDCA) and glycodeoxycholic acid (GDCA) were quantified.
Total RNA isolation
HepaRG cells were seeded in 24-well plates following manufacturer’s instructions. At day 7, HepaRG cells were exposed to cholestatic concentrations of ATA, CsA, and NEFA with or without the BA mix added to the cell culture medium. After 72 h of exposure, samples were collected for RNA isolation by aspirating the cell culture medium and adding lysis buffer directly to the well (Qiagen, Belgium). Total RNA extraction (Qiagen, Belgium) was done according to manufacturer’s instructions. Quantification and purity of the isolated RNA were determined by means of spectrophotometric analysis with a Nanodrop spectrophotometer (ThermoFisher Scientific, Belgium).
Microarray analysis
Whole genome expression analysis was performed using microarray technologies from Affymetrix (Germany) as previously described (Rodrigues et al. 2018). For this purpose, 100 ng total RNA per sample was amplified using a GeneChip 3′IVT Express Kit following manufacturer’s instructions (Affymetrix, Germany). Amplified RNA was purified with magnetic beads and 15 mg biotin-amplified RNA was treated with fragmentation reagent. Then, 12.5 µg of fragmented amplified RNA was hybridized to Affymetrix Human genome U133 plus 2.0 GeneChip and Affymetrix Mouse Genome 430 2.0 GeneChip. Subsequently, the chips were placed in a GeneChip Hybridization Oven 645 (Affymetrix, Germany) following manufacturer’s instructions. After incubation, the arrays were washed with GeneChip Fluidics Station 450 (Affymetrix, Germany) and stained with Affymetrix HWS kit. Thereafter, stained arrays were scanned via an Affymetrix GeneChip Scanner 3000 7G. Hybridization controls were performed using Affymetrix GCOS software. Normalization quality controls, such as scaling factors, background intensities, noise and raw Q-values, average intensities and present calls were done with robust multiarray analysis and Affymetrix Transcriptome Analysis Console software and were all within the acceptable limits of all used chips. Functional toxicological analyses were performed using QIAGEN’s Ingenuity Pathway Analysis (IPA) software.
Statistics
The number of batches (i.e. in vitro experiments) and replicates (i.e. in vitro and in vivo experiments) used for each type of analysis were specified in the results section. Data were expressed as mean ± standard deviation (SD) or as fold change compared to untreated HepaRG cells or sham-operated mice. Results of the BA transporter activity and BA quantification assays were statistically processed by one-way and two-way ANOVA followed by post hoc tests with Bonferroni correction using GraphPad Prism7 software with a p value ≤ 0.05 considered as significant. Transcriptomic analysis was performed with both IPA and Transcriptome Analysis Console software, which used the Fisher’s exact test with Benjamini–Hochberg correction or z-scores and one-way ANOVA with Benjamini–Hochberg correction, to identify significantly modified genes.
Results
Assessment of working concentrations of cholestatic drugs in vitro
Human hepatoma HepaRG cells were cultured in monolayer configuration and exposed to well-known cholestatic drugs for 72 h in the presence and absence of a BA mix. Serum BA pool concentrations typically increase by 30–50-fold in cholestasis patients (Humbert et al. 2012; Tagliacozzi et al. 2003). Therefore, a 50 × concentrated BA mix of the five most abundant BAs present in human serum was added to HepaRG cells together with the cholestatic drugs. This setup enabled more close resemblance to the in vivo situation during cholestasis, to detect BA-selective sensitization towards toxic effect of cholestatic drugs, as well as to distinguish cholestatic hepatotoxicity from non-cholestatic hepatotoxicity (Sharanek et al. 2017; Gijbels et al. 2019). In this study, the cholestatic drugs ATA, CsA, and NEFA were selected based on literature data (Oorts et al. 2016; Rakotondravelo et al. 2012). The working concentrations of the cholestatic drugs were determined by calculating CIx values. The CIx value has been previously introduced and successfully applied to assess cholestatic liability of chemicals in spheroid cultures of primary human hepatocytes (Hendriks et al. 2016). An identical indicator, called the drug-induced cholestasis index, has been used to assess cholestatic potential in sandwich cultures of primary hepatocytes, albeit using a functionality parameter (i.e. urea formation) rather than cell viability (Chatterjee et al. 2014). CIx values or drug-induced cholestasis index values equal to or lower than 0.8 designated cholestatic properties (Chatterjee et al. 2014; Hendriks et al. 2016). For the purpose of this study, a viability assay was preferred for early detection of hepatotoxicity. Preliminary experiments revealed that the MTT reductase assay is more sensitive than the adenosine triphosphate assay to assess the cell viability (data not shown). CIx values were calculated of a range of concentrations of ATA (i.e. 5–100 µM), CsA (i.e. 5–50 µM) and NEFA (i.e. 5–50 µM). The lowest concentration yielding a CIx value lower than or equal to 0.8 was selected for further experiments, namely 60 µM for ATA, 20 µM for CsA, and 30 µM for NEFA (Table 1). Similar concentrations for CsA and NEFA have been associated with drug-induced cholestasis in primary human hepatocytes (Chatterjee et al. 2014; Oorts et al. 2016), yet such data seems unavailable for ATA. To translate CIx values obtained in vitro into relevant concentrations in vivo, a safety margin was further introduced. The latter was calculated by dividing the lowest concentration yielding a CIx value lower than or equal to 0.8 by the total therapeutic peak plasma concentration, retrieved from literature. Safety margin values of ATA, CsA, and NEFA were all below 30, which have been previously found to be indicative for cholestatic risk (Oorts et al. 2016).
Induction of molecular initiating event of cholestasis in vitro
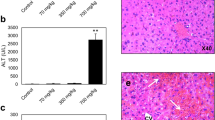
The available AOP on cholestatic liver injury mainly focuses on the inhibition of the BSEP transporter as molecular initiating event (Vinken et al. 2013). The activity and cellular localization of BSEP in differentiated human hepatoma HepaRG cell cultures were previously determined and showed proper functionality and localization at the bile canalicular pole, supporting the suitability of these cells to study hepatobiliary transporters (Bachour-El Azzi et al. 2015; Rodrigues et al. 2018). Expression of the ABCB11 gene, which codes for BSEP, was significantly elevated when HepaRG cells were treated solely with the BA mix, while this was not the case when cells were exposed to the BA mix together with cholestatic drugs or cholestatic drugs alone (Fig. 1a). Upregulation of ABCB11 expression by the BA mix could be attributed to an adaptive response to cholestasis, which strives to counteract accumulation of BAs in hepatocytes by increasing their export. On the other hand, it has been reported that ABCB11 expression becomes strongly reduced in HepaRG cells, human HepG2 cells, and precision-cut human liver slices when treated with cholestatic drugs (Qu et al. 2018; Rodrigues et al. 2018; Vatakuti et al. 2017). Similar observations were made in the present study. Moreover, an accumulation of the BSEP probe CLF could be visualized in cells treated with cholestatic drugs (Fig. 1b). CsA acts as a potent inhibitor of BSEP with IC50 values ranging between 0.1 and 0.88 µM in primary human hepatocytes (Morgan et al. 2010; Zhang et al. 2016). NEFA was also reported to inhibit BSEP, albeit less potent than CsA with an IC50 value of 20 µM in primary human hepatocytes (Kostrubsky et al. 2006). No IC50 values for ATA-induced BSEP inhibition are available in literature. Nevertheless, it is important to keep in mind that although the AOP depicts BSEP inhibition as the main molecular initiating event of cholestatic liver injury, other molecular initiating events may play a role as well (Vinken et al. 2013). As a consequence of BSEP inhibition or induction of other molecular initiating events, BAs will accumulate in hepatocytes. This was verified in the HepaRG cell cultures in this study by means of liquid chromatography and mass spectrometry/mass spectrometry analysis to quantify the BAs CA, CDCA, DCA, GCA, GCDCA, and GDCA. Interestingly, significant lower concentrations of CDCA, DCA, GCDCA, and GDCA were observed in cell lysates of HepaRG cells treated with ATA, CsA, and NEFA together with the BA mix (Fig. 1c). This is in line with findings from a previous report that showed a rapid reduction in BA levels in HepaRG cells exposed to CsA (Sharanek et al. 2015). This has also been shown for the cholestatic drug bosentan, which was found to reduce concentrations of CA, GCA, and GCDCA in HepaRG cell cultures (Burbank et al. 2017; Lepist et al. 2014; Rodrigues et al. 2018). It should be stressed that these concentrations represent the total BA abundance, hence no distinction could be made between intracellular accumulation and bile pocket accumulation.
a Bile salt export pump (BSEP|ABCB11) gene expression in treated and untreated HepaRG cells. HepaRG cells were exposed for 72 h to 60 µM ATA, 20 µM CsA or 30 µM NEFA with or without the BA mix. Subsequently, samples were collected and subjected to microarray analysis. ABCB11 gene expression in treated HepaRG cells were normalized to untreated HepaRG cells (control). Dotted line represents untreated HepaRG cells. (N = 1; n = 3). b Live staining of fluorescent BSEP probe cholyl-l-lysyl-fluorescein (CLF) in treated and untreated HepaRG cells. HepaRG cells treated with 60 µM ATA, 20 µM CsA, 30 µM NEFA (or the BA mix) were exposed to BSEP probe CLF (excitation/emission wavelength 450/530 nm). After 30 min of incubation, cells were rinsed, and nuclei were stained with Hoechst 33342 (excitation/emission wavelength 365/420 nm). Fluorescence images were taken at × 200 magnification. (N = 6; n = 1). c Liquid chromatography and mass spectrometry/mass spectrometry quantification of BAs in treated HepaRG cell lysates. HepaRG cells were exposed for 72 h to the BA mix with or without co-exposure to 60 µM ATA, 20 µM CsA or 30 µM NEFA. Subsequently, cell lysates from six samples were pooled and subjected to liquid chromatography and mass spectrometry/mass spectrometry analysis for CDCA, DCA, GCA, GCDCA and GDCA (ng/mL) quantification (N = 1; n = 3). a–c Results were analyzed by one-way or two-way ANOVA followed by post hoc tests using Bonferroni’s correction. Data were expressed as mean ± SD (*p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; ****p ≤ 0.0001). (ATA atazanavir, BA bile acid, BSEP bile salt export pump, CDCA chenodeoxycholic acid, CLF cholyl-l-lysyl-fluorescein, CsA cyclosporin A, DCA deoxycholic acid, GCA glycocholic acid, GCDCA glycochenodeoxycholic acid, GDCA glycodeoxycholic acid, NEFA nefazodone)
Transcriptomic analysis of established key events of cholestasis in vitro
HepaRG cells were cultured in monolayer configuration and exposed for 72 h to ATA, CsA, and NEFA in the presence and absence of the BA mix followed by transcriptomic analysis. A principle component analysis was performed on the transcriptomic data sets and showed no differences between tested HepaRG cell batches. The most pronounced variation was originating from different cholestatic drugs used to induce cholestasis and the presence or absence of the BA mix (Fig. S1). A Venn diagram of the differentially expressed genes was generated to visualize the variation (Fig. S2A and B). 7087 genes were differentially expressed when cells were treated with cholestatic drugs compared to control treatment, increasing to 9809 genes when cells were co-treated with BA. This confirms increased sensitivity upon co-exposure of the HepaRG cells to the cholestatic drugs and the BA mix. Comparing cholestatic drug treatment with the combination of cholestatic drugs and the BA mix resulted in 2655 differentially expressed genes (Fig. S2C).
The AOP on cholestasis describes two biological responses. The deteriorative response is featured by inflammation, the opening of the mitochondrial membrane permeability pore, oxidative stress, and cell death (Vinken et al. 2013). In the present study, inflammation could be evidenced by the enhanced expression of colony stimulating factor 1 (CSF1), jun proto-oncogene (JUN), mitogen-activated protein kinase-activated protein kinase 3 (MAPKAPK3), and serpin E1 (SERPINE1) in HepaRG cells treated with cholestatic drugs both in absence and in presence of the BAs (Fig. 2). The expression of C–C chemokine receptor type 2 (CCR2) was also increased upon treatment with CsA and NEFA. Noxious BAs trigger formation of the mitochondrial permeability pore, resulting in mitochondrial impairment. Mitochondrial dysfunction will, in turn, lead to increased production of reactive oxygen species and thus to oxidative stress (Vinken et al. 2013). To protect against oxidative stress, the nuclear-related factor 2 signaling pathway is activated, which regulates a high number of genes (Table 2) that encode detoxifying or anti-oxidative proteins (Vatakuti et al. 2017). In line with this, nuclear-related factor 2 signaling pathway was especially activated in HepaRG cells treated with cholestatic drugs CsA and NEFA in presence of the BAs, and to a lesser extent in HepaRG cells treated with ATA. Oxidative stress ultimately burgeons into cell death. However, there is still some discussion regarding the nature of this type of cell death. Apoptosis has been associated with cholestasis in rats. By contrast, necrosis seems the main cell death mechanism during cholestasis in humans and mice (Woolbright and Jaescke 2012). Genes involved in both necrosis and apoptosis were induced in CsA-treated HepaRG cells, while no induction nor suppression could be observed in the other treatment groups (Table 3).
Expression of genes involved in the inflammatory response in treated HepaRG cells. HepaRG cells were exposed for 72 h to 60 µM atazanavir (ATA), 20 µM cyclosporin A (CsA) and 30 µM nefazodone (NEFA) with or without the bile acid (BA) mix. Samples were collected and subjected to microarray analysis. Subsequently, transcriptomic analysis was performed by means of the Transcriptome Analysis Console software. Gene selection relevant for inflammation was based on the gene-level fold change ≤ − 2 or ≥ 2 and p value ≤ 0.05, calculated via one-way ANOVA followed by post hoc tests using Bonferroni’s correction and Benjamini–Hochberg correction. Normalized gene expression (Log2) of the selected genes in treated and untreated HepaRG cells were expressed as mean ± SD (**p ≤ 0.01; ***p ≤ 0.001; ****p ≤ 0.0001). (N = 1; n = 3). (ATA atazanavir, BA bile acid, CCR2 C–C chemokine receptor type 2, CsA cyclosporin A, CSF1 colony stimulating factor 1, CTL control, JUN jun proto-oncogene, MAPKAPK3 mitogen-activated protein kinase-activated protein kinase 3, NEFA nefazodone, SERPINE1 serpin E1)
The adaptive response in the cholestasis AOP can be characterized as a hepato-protective mechanism aimed to counteract BA accumulation to alleviate cholestatic liver damage. This protective mechanism relies on the activation of several NRs namely, FXR (NR1H4), PXR (NR1I2), CAR (NR1I3), and SHP (NR0B2), which coordinate a plethora of transcriptional modifications in view of reducing BA levels (Table 4). A number of transcriptional changes were reproduced in compliance with the scenario, namely repressed expression of cytochrome P450 7A1 (CYP7A1), which encodes the rate-limiting enzyme of BA biosynthesis, organic anion transporting peptide 1B1 (OATP1B1) and sodium-taurocholate co-transporting polypeptide |solute carrier family 10 member 1 (NTCP |SLC10A1) (Table 4). Other genes described in the adaptive response appeared unchanged or differentially regulated than predicted. Literature indicates that another basolateral transporter could be involved in the adaptive response, namely the multidrug resistance-associated protein 4 (MRP4|ABCC4) (Zollner et al. 2006). Gene expression of this transporter also showed upregulated in HepaRG cells treated with cholestatic drugs in presence and absence of the BA mix, especially with NEFA.
Transcriptomic analysis of established key events of cholestasis in vivo
To challenge the robustness of the AOP on cholestatic liver injury, which was initially designed for drug-induced cholestasis, the BDL mouse model was used as second experimental model of cholestasis. Besides different species (i.e. human versus mouse) and setting (i.e. in vitro versus in vivo), this model thus also differed from the HepaRG cell culture system in terms of the nature of trigger (i.e. chemical induced versus surgical induced). This implies that other molecular initiating events could be involved, but, it can be anticipated that the key events of cholestasis will remain unchanged. In this respect, the occurrence of inflammation in the liver of BDL mice was evidenced by significantly enhanced expression of genes Ccr2, Csf1, Jun, Mapkapk3, and Serpine1, identical to the in vitro setting (Fig. 3). Oxidative stress became manifested by upregulation of fos proto-oncogene (Fos), glutathione S-transferase α5/μ3 (Gstα5/μ3), and NAD(P)H quinone dehydrogenase 1 (Nqo1). It should be mentioned that, although these two key events, i.e. inflammation and oxidative stress, occurred both in the in vitro and in vivo settings, the genes involved may differ. Indeed, expression of genes relevant to inflammation, such as interleukin 1 receptor-like 1 (Il1rl1), interleukin 6/8 (Il6/8), and interleukin 1 receptor-associated kinase 2 (Irak2) remained unaffected in BDL mice, but were highly upregulated in HepaRG cells treated with cholestatic drugs. Vice versa, genes Csf2, Il7, and Irak3 appeared highly upregulated in BDL mice, yet not in HepaRG cells exposed to cholestatic drugs (data not shown). Likewise, 64 genes relevant for oxidative stress were differentially expressed in drug-induced cholestasis in HepaRG cells (Table 2), while this was only the case for 13 genes in BDL mice (Table 5). Of note, we did not identify significantly differential expression of genes involved in cell death types apoptosis and necrosis in BDL mice, which is unlike the in vitro setting (data not shown). A number of events linked to the adaptive response could be recapitulated in BDL mice, including upregulation of Mrp4|Abcc4, Ostβ|Slc51b and, downregulation of solute carrier organic anion transporter family 1B2 (Slco1b2) (human analogue OATP1B1), and Slc10a1 (Ntcp) (Table 6). Human counterparts of Slco1b2 and Slc10a1 also showed downregulated in HepaRG cells treated with cholestatic drugs (Table 4). This was not the case for Cyp7a1, which was upregulated in BDL mice, and remarkebly downregulated in the cholestatic in vitro system. Besides the different etiology of cholestasis, dissimilarities in gene expression between human HepaRG cells and BDL mice may, of course, also be explained by interspecies differences and in vitro–in vivo settings. For example, HepaRG cells can differentiate into just two cell types including hepatocyte-like cells and cholangiocyte-like cells (Parent et al. 2004), while the liver samples additionally consist out of Kupffer cells, stellate cells, liver sinusoidal endothelial cells, etc. (Kmiéc 2001).
Expression of genes involved in inflammation in bile duct ligation (BDL) and sham mice. Mice underwent BDL surgery or sham surgery (control, CTL). Samples were collected and subjected to microarray analysis. Subsequently, transcriptomic analysis was performed by means of the Transcriptome Analysis Console software. Gene selection relevant for inflammation was based on the gene-level fold change ≤ − 2 or ≥ 2 and p value ≤ 0.05, calculated via one-way ANOVA followed by post hoc tests using Bonferroni’s correction and Benjamini–Hochberg correction. Normalized gene expression (Log2) of the selected genes in BDL mice and CTL mice were expressed as mean ± SD (*p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; ****p ≤ 0.0001). (n = 6). (BDL bile duct ligation, Ccr2 C–C chemokine receptor type 2, Csf1 colony stimulating factor 1, CTL control, Jun jun proto-oncogene, Mapkapk3 mitogen-activated protein kinase-activated protein kinase 3, Serpine1 serpin E1)
Transcriptomic characterization of potential new key events in vitro and in vivo
An AOP is by definition a living document that should be continuously updated by feeding in new information. In this regard, the AOP on cholestasis, which is still the only one in its kind, has been introduced in 2013. Since that time, several reports have been published suggesting additional key events, such as endoplasmic reticulum stress/unfolded protein response (Burban et al. 2018), autophagy (Gao et al. 2014; Manley et al 2014), and necroptosis (Afonso et al. 2016). The transcriptomic analysis performed in the current study allows to verify the occurrence of these novel key events. During pathological conditions, including oxidative stress, endoplasmic reticulum homeostasis can become compromised and protein folding processes hampered, which is defined as endoplasmic reticulum stress. Consequently, an unfolded protein response is induced to restore endoplasmic reticulum homeostasis and cell survival (Bhat et al. 2017; Malhi and Kaufman 2011). Accordingly, genes indicative of endoplasmic reticulum stress and the consecutive unfolded protein response showed significantly upregulated in ATA-, CsA-, and NEFA-induced cholestasis, namely activation transcription factor 4/6 (ATF4/6) and CCAAT–enhancer-binding protein homologous protein (CHOP) (Fig. 4a). No significant modulation of these genes could be observed in BDL mice. This is in agreement with a previous study that found acute endoplasmic reticulum stress responses in BDL mice after 1 day, which returned to normal after 3 days (Liu et al. 2018). In drug-induced cholestasis, on the other hand, the role of endoplasmic reticulum stress still remains elusive. Endoplasmic reticulum stress has been suggested to play a critical role in the initiation and progression of drug-induced cholestasis, yet endoplasmic reticulum stress may equally participate in a rescue mechanism to promote the removal of excess BAs by suppressing the BA synthetic pathway (Burban et al. 2018; Henkel et al. 2017). Furthermore, in case of excessive endoplasmic reticulum stress, a switch may occur from a prosurvival response to a prodeath mode along with the initiation of apoptosis or autophagy (Bhat et al. 2017). In addition, necroptosis may occur (Afonso et al. 2016). At least three genes related to autophagy and necroptosis appeared significantly upregulated in ATA-induced, CsA-induced, and NEFA-induced cholestasis. These include microtubule associated protein 1 light chain 3β (MAP1LC3β), SH3 domain containing GRB2 like, endophilin B1 (SH3GLB1), and sequestosome 1 (SQSTM1) for autophagy, and cylindromatosis (CYLD), mixed lineage kinase domain-like pseudokinase (MLKL) and receptor interacting serine/threonine kinase 1 (RIPK1) for necroptosis (Fig. 4b, c). Similarly, in BDL mice, analogous genes related to autophagy and necroptosis were found significantly upregulated (Fig. 4b, c).
a Expression of genes involved in endoplasmic reticulum stress/unfolded protein response in treated and untreated HepaRG cells (left) and, in bile duct ligation (BDL) and sham mice (right). b Genes involved in autophagy in treated and untreated HepaRG cells (left) and in BDL and sham mice (right). c Genes involved in necroptosis in treated and untreated HepaRG cells (left) and in BDL and sham mice (right). a–c HepaRG cells were exposed for 72 h to 60 µM atazanavir (ATA), 20 µM cyclosporin A (CsA), and 30 µM nefazodone (NEFA) with or without the bile acid (BA) mix. Mice underwent BDL surgery or sham surgery. Samples were collected and subjected to microarray analysis. Subsequently, transcriptomic analysis was performed by means of the Transcriptome Analysis Console software. In addition, functional toxicological analysis was executed by means of Ingenuity Pathway Analysis (IPA). Gene selections relevant in endoplasmic reticulum stress/unfolded protein response (a), autophagy (b) and necroptosis (c) were based on the gene-level fold change ≤ − 2 or ≥ 2 and p value ≤ 0.05, calculated via one-way ANOVA followed by post hoc tests using Bonferroni’s correction and Benjamini–Hochberg correction. Normalized gene expression (Log2) of the selected genes in treated and untreated HepaRG cells and, BDL mice and control mice were expressed as mean ± SD (*p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; ****p ≤ 0.0001). (in vitro N = 1; n = 3 and in vivo n = 6) (ATA atazanavir, ATF4/6 activation transcription factor 4/6, ATG7 autophagy-related 7, BA bile acid, BDL bile duct ligation, CHOP CCAAT-enhancer-binding protein homologous protein, CsA cyclosporin A, CYLD cylindromatosis, GRP78 glucose-regulated protein 78, MAP1LC3B microtubule-associated protein 1 light chain 3β, MLKL mixed lineage kinase domain-like pseudokinase, NEFA nefazodone, RIPK1/3 receptor interacting serine/threonine kinase 1/3, SH3GLB1 SH3 domain containing GRB2 like, endophilin B1, SQSTM1 sequestosome 1)
Discussion
This study was set up to test the robustness of an available AOP on cholestatic liver injury (Vinken et al. 2013), and additionally generate new information for further AOP optimization. For this purpose, intrahepatic cholestasis was mimicked by treating monolayer cultures of human hepatoma HepaRG cells with cholestatic drugs ATA, CsA or NEFA and a concentrated BA mix for 72 h. HepaRG cells are widely used as an adequate alternative to primary human hepatocytes to study diverse types of drug-induced liver injuries, including drug-induced cholestasis (Anthérieu et al. 2013; McGill et al. 2011; Sharanek et al. 2014). In parallel, extrahepatic cholestasis was induced in mice by performing a BDL surgery. BDL mice is a well-known model of cholestasis, reliable in reproducing cholestasis in vivo (Tag et al. 2015). Cell culture and liver tissue samples were collected and subjected to transcriptomic analysis and results were tested for mechanistic anchorage in the existing AOP on cholestatic liver injury. The AOP on cholestatic liver injury particularly focused on BSEP inhibition as a major molecular initiating event (Vinken et al. 2013). This complied with the observations of the present study, which showed suppression of BSEP activity by ATA, NEFA, and CsA in HepaRG cell cultures, albeit no altered gene expression of the transporter was observed. Nevertheless, it should be emphasized that besides BSEP inhibition, other molecular initiating events underlie cholestatic liver injury, including effects on alternative transporters, hepatocellular changes, and bile canalicular changes (Gijbels et al. 2019). In this regard, NEFA has been reported to inhibit OATP(s) and multidrug resistance protein 1 (MDR1) transporter (Dragovic et al. 2016; Kolaríc et al. 2019). It has been suggested that UDP-glucuronosyltransferase 1A1 inhibition can be involved in the onset of ATA-induced cholestasis (Bissio and Lopardo 2013; Zhang et al. 2005). In the case of CsA-induced cholestasis, disruption of the cytoskeleton and altered membrane fluidity were earlier characterized as additional molecular initiating events of cholestasis (Roman et al. 2003; Yasumiba et al. 2001). These triggering factors normally result in BA accumulation (Vinken et al. 2013). Surprisingly, several studies, including the present one, showed higher levels of intracellular BAs in HepaRG cells solely treated with the BA mix compared to HepaRG cells treated with cholestatic drugs and the BA mix (Burbank et al. 2017; Lepist et al. 2014; Rodrigues et al. 2018; Sharanek et al. 2015). Moreover, a rapid clearance (i.e. 24 h) was reported in accumulating BAs from HepaRG cell culture layers into the cell culture medium when treated with CsA. It could be argued that reduced function of NTCP and BA synthesis enzymes, such as CYP7A1, along with induced alternative basolateral transporters could be causing this rapid decrease (Sharanek et al. 2015). This aligns with the results from the present study, which showed downregulation of CYP7A1 and SLC10A1 in HepaRG cells treated with cholestatic drugs and the BA mix, while the basolateral transporter ABCC4, which encodes basolateral transporter MRP4, was positively affected. As predicted by the AOP (Vinken et al 2013), disturbance in BA homeostasis resulted in inflammation, which often progresses into oxidative stress along with increasing levels of mitochondrial reactive oxygen species (Arduini et al. 2012). Inflammation and increased expression of anti-oxidants appeared both in intrahepatic cholestasis (i.e. cholestasis induced by ATA, CsA, NEFA (this study), and bosentan (Rodrigues et al. 2018)) and extrahepatic cholestasis (BDL mice), albeit the modulated genes differed. Additionally, the number of differentially expressed genes related to oxidative stress was lower in vivo compared to in vitro (13 versus 62) (Tables 2 and 5). Relative to the response in humans, mice are believed more resilient to inflammatory challenge (Seok et al. 2013), which probably results in less oxidative stress. According to the AOP, inflammation and oxidative stress can ultimately burgeon into the onset of cell death (Vinken et al. 2013). The latter is mainly necrotic cell death (Woolbright and Jaescke 2012), but an apoptotic mechanism may also be involved (Botla et al. 1995; Gores et al. 1998; Schoemaker et al. 2004; Vinken et al. 2013). According to the transcriptomic data obtained in this study, apoptosis and necrosis were activated in HepaRG cells treated with CsA in the absence and presence of the BA mix, while necrosis seemed induced after bosentan and CsA treatment of those cells (Rodrigues et al. 2018). Neither apoptosis nor necrosis was observed in HepaRG cells treated with ATA or NEFA nor in BDL mice. These inconsistencies may also be explained by differences in the etiology profile (i.e. triggering factor) and differences in the severity level of the acquired cholestasis. Furthermore, interspecies differences (Woolbright and Jaeschke 2012), differences in cell culture configuration (e.g. monolayer configuration vs spheroid or sandwich configuration) or different analysis methods (e.g. gene expression analysis via microarray) could play a role in which type of cell death could be observed. Concomitant with the deteriorative response, the adaptive response commenced to counteract accumulating BAs. In this respect, a number of genes involved in BA metabolism and BA transport were regulated to decrease the amount of BAs. Indeed, CYP7A1, OATP1B1, and SCL10A1 appeared downregulated in the different types of drug-induced cholestasis in HepaRG cells. OSTα/β was also modulated, but their regulation varied across the different cholestatic drugs. Extrahepatic cholestasis in BDL mice was accompanied by modulation of Abcc4, Ostβ, Slco1b2, and Slc10a1 in agreement with the AOP (Vinken et al. 2013). AOP compliance with the in vitro model (i.e. intrahepatic drug-induced cholestasis) and in vivo model (i.e. extrahepatic cholestasis) was compared and summarized in Table 7. Interestingly, in agreement with the AOP, gene expression of OATP1B1 and SLC10A1 showed downregulated during cholestasis, independent of the etiology. Additionally, unlike what was predicted in the AOP, downregulation of the gene expression of UGT2B4 and CYP2B6 also showed common among the two different types of cholestasis. It may be interesting to consider these modulated genes as potential novel biomarkers of cholestasis.
The present study also aimed to identify novel potential key events that could fit in the AOP. Accordingly, transcriptomic data from HepaRG cells treated with ATA, CsA, and NEFA suggested the involvement of the endoplasmic reticulum stress and the unfolded protein response in drug-induced cholestasis. This is in agreement with a number of other studies, which indicated endoplasmic reticulum stress to play a role in both the initiation and progression of cholestasis as in the protective mechanism by removing excess BAs (Burban et al. 2018; Henkel et al. 2017). Moreover, modulated genes involved in two additional types of cell death, namely autophagy and necroptosis, could be recognized in both drug-induced cholestasis in HepaRG cells as well as in extrahepatic cholestasis in BDL mice. Autophagy and necroptosis were already found associated with cholestasis in earlier studies (Afonso et al. 2016; Gao et al. 2014; Manley et al 2014).
In conclusion, the results of this study show that the available AOP on cholestatic liver injury seems fit for predicting intrahepatic drug-induced cholestasis yet requires optimization. In this respect, a simplified AOP, including the various molecular initiating events as well as novel key events, is proposed in Fig. 5. The outcome of this study further suggests that some established key events described in the AOP should be omitted or adjusted, different AOPs should be developed for alternative types of cholestatic insults, and that the applicability domain of AOPs should be well defined.
Updated version of the AOP on cholestatic liver injury. Drug-induced cholestasis is currently typified by three different types of molecular initiating events (MIE), including transporter changes, hepatocellular changes, and bile canalicular changes (Gijbels et al. 2019). These triggering factors evoke noxious bile acid (BA) accumulation (i.e. intracellular, extracellular or both), which activates two cellular responses, namely a deteriorative response and an adaptive response. The deteriorative response starts with the occurrence of inflammation (Woolbright and Jaescke 2012) and mitochondrial impairment (Begriche et al. 2011), which lead to oxidative stress (Copple et al. 2010). Oxidative stress, in turn, may lead to endoplasmic reticulum stress concomitant with the unfolded protein response (particularly in intrahepatic drug-induced cholestasis) (Burban et al. 2018). These events can further burgeon into cell death (i.e. apoptosis, autophagy or necro(pto)sis) (Afonso et al. 2016; Gao et al. 2014; Manley et al. 2014; Vinken et al. 2013; Woolbright and Jaescke 2012). Simultaneously, the adaptive response strives to counteract BA accumulation, hence the deteriorative response, by activating several nuclear receptors, which regulate genes involved in BA homeostasis (Halilbasic et al. 2013), albeit regulation of these genes were found divergent between different types of cholestasis and/or in discrepancy with the AOP. New suggested key events (i.e. endoplasmic reticulum stress, autophagy, and necroptosis) were marked in orange. (ABCC2/3 ATP-binding cassette C2/3, AO(P) adverse outcome (pathway), BA bile acid, CAR constitutive androstane receptor, CYP cytochrome P450, FXR farnesoid X receptor, MIE molecular initiating event, NR nuclear receptor, OATP1B1 organic anion transporting peptide 1B1, OSTα/β organic solute transporter α/β, PXR pregnane X receptor, SHP short heterodimer partner, SLC10A1 solute carrier family 10 member 1, SULT2A1 sulfotransferase 2A1, UGT2B4 UDP glucuronosyltransferase 2B4)
Abbreviations
- AOP:
-
Adverse outcome pathway
- ABC:
-
ATP-binding cassette family
- ATA:
-
Atazanavir
- ATF:
-
Activation transcription factor
- BA(s):
-
Bile acid(s)
- BDL:
-
Bile duct ligation
- BSEP:
-
Bile salt export pump
- CA:
-
Cholic acid
- CAR:
-
Constitutive androstane receptor
- CCR:
-
C–C chemokine receptor type
- CDCA:
-
Chenodeoxycholic acid
- CHOP:
-
CCAAT-enhancer-binding protein homologous protein
- CIx:
-
Cholestatic index
- CLF:
-
Cholyl-l-lysyl-fluorescein
- CsA:
-
Cyclosporin A
- CSF:
-
Colony stimulating factor
- CYLD:
-
Cylindromatosis
- CYP:
-
Cytochrome P450
- DILI:
-
Drug-induced liver injury
- DMSO:
-
Dimethyl sulfoxide
- DCA:
-
Deoxycholic acid
- Fos:
-
Fos proto-oncogene
- FXR:
-
Farnesoid X receptor
- GCA:
-
Glycocholic acid
- GCDCA:
-
Glycochenodeoxycholic acid
- GDCA:
-
Glycodeoxycholic acid
- Gst:
-
Glutathione S-transferase
- Il(1rl1):
-
Interleukin (1 receptor-like 1)
- IPA:
-
Ingenuity Pathway Analysis
- IRAK:
-
Interleukin 1 receptor-associated kinase
- JUN:
-
Jun proto-oncogene
- MAP1LC3β:
-
Microtubule-associated protein 1 light chain 3β
- MAPKAPK:
-
Mitogen-activated protein kinase-activated protein kinase
- MDR:
-
Multidrug resistance protein
- MLKL:
-
Mixed lineage kinase domain-like pseudokinase
- MRP:
-
Multidrug resistance-associated protein
- MTT:
-
3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide
- NEFA:
-
Nefazodone
- NR(s):
-
Nuclear receptor(s)
- NTCP:
-
Sodium-taurocholate co-transporting polypeptide
- Nqo:
-
NAD(P)H quinone dehydrogenase
- OATP(s):
-
Organic anion transporting peptide(s)
- OST:
-
Organic solute transporter
- PXR:
-
Pregnane X receptor
- RIPK:
-
Receptor interacting serine/threonine kinase
- SD:
-
Standard deviation
- SERPINE1:
-
Serpin E1
- SHP:
-
Small heterodimer partner
- SH3GLB1:
-
SH3 domain containing GRB2 like, endophilin B1
- SLC(O):
-
Solute carrier (organic anion transporter) family
- SQSTM:
-
Sequestosome
- UGT:
-
UDP-glucuronosyltransferase
References
Afonso MB, Rodrigues PM, Simão AL et al (2016) Activation of necroptosis in human and experimental cholestasis. Cell Death Dis 7:e2390
Ankley GT, Bennett RS, Erickson RJ et al (2010) Adverse outcome pathways: a conceptual framework to support ecotoxicology research and risk assessment. Environ Toxicol Chem 29:730–741
Anthérieu S, Bachour-El Azzi P, Dumont J et al (2013) Oxidative stress plays a major role in chlorpromazine-induced cholestasis in human HepaRG cells. Hepatology 57:1518–1529
Arduini A, Serviddio G, Tormos AM et al (2012) Mitochondrial dysfunction in cholestatic liver diseases. Front Biosci 4:2233–2252
Bachour-El Azzi P, Sharanek A, Burban A et al (2015) Comparative localization and functional activity of the main hepatobiliary transporters in HepaRG cells and primary human hepatocytes. Toxicol Sci 145:157–168
Bale SS, Vernetti L, Senutovitch N et al (2014) In vitro platforms for evaluating liver toxicity. Exp Biol Med 239:1180–1191
Begriche K, Massart J, Robin MA et al (2011) Drug-induced toxicity on mitochondria and lipid metabolism: mechanistic diversity and deleterious consequences for the liver. J Hepatol 54:773–794
Bhat TA, Chaudhary AK, Kumar S et al (2017) Endoplasmic reticulum-mediated unfolded protein response and mitochondrial apoptosis in cancer. Biochim Biophis Acta 1867:58–66
Bissio E, Lopardo GD (2013) Incidence of hyperbilirubinemia and jaundice due to atazanavir in a cohort of hispanic patients. AIDS Res Hum Retroviruses 29(3):415–417
Botla R, Spivey JR, Aguilar H et al (1995) Ursodeoxycholate (UDCA) inhibits the mitochondrial membrane permeability transition induced by glycochenodeoxycholate: a mechanism of UDCA cytoprotection. J Pharmacol Exp Ther 272:930–938
Burban A, Sharanek A, Guguen-Guillouzo C et al (2018) Endoplasmic reticulum stress precedes oxidative stress in antibiotic-induced cholestasis and cytotoxicity in human hepatocytes. Free Radic Biol Med 115:166–178
Burbank MG, Sharanek A, Burban A et al (2017) Mechanistic insights in cytotoxic and cholestatic potential of the endothelial receptor antagonists using HepaRG cells. Toxicol Sci 157:451–464
Chatterjee S, Richert L, Augustijns P et al (2014) Hepatocyte-based in vitro model for assessment of drug-induced cholestasis. Toxicol Appl Pharmacol 274:124–136
Copple BL, Jaeschke H, Klaassen CD (2010) Oxidative stress and the pathogenesis of cholestasis. Semin Liver Dis 30:195–204
Dawson S, Stahl S, Paul N et al (2012) In vitro inhibition of the bile salt export pump correlates with risk of cholestatic drug-induced liver injury in humans. Drug Metab Dispos 40:130–138
Dewaele D, Annaert P, Hoeben E (2019) LC-MS/MS analysis of bile acids in in vitro samples. Methods Mol Biol 1981:15–23
Dragovic S, Vermeulen NPE, Gerets HH et al (2016) Evidence-based selection of training compounds for use in the mechanism-based integrated prediction of drug-induced liver injury in man. Arch Toxicol 90:2979–3003
Gao L, Lv G, Guo X et al (2014) Activation of autophagy protects against cholestasis-induced hepatic injury. Cell Biosci 4:47
Gijbels E, Vilas-Boas V, Deferm N et al (2019) Mechanisms and in vitro models of drug-induced cholestasis. Arch Toxicol 93:1169–1186
Goldberg DS, Forde KA, Carbonari DM, Lewis JD, Leidl KBF, Reddy KR, Haynes K, Roy J, Sha D, Marks AR, Schneider JL, Strom BL, Corley DA, Lo Re V (2015) Population-representative incidence of drug-induced acute liver failure based on an analysis of an integrated health care system. Gastroenterology 148(7):1353–1361.e3
Gores GJ, Miyoshi H, Botla R et al (1998) Induction of the mitochondrial permeability transition as a mechanism of liver injury during cholestasis: a potential role for mitochondrial proteases. Biochim Biophys Acta 1366:167–175
Halilbasic E, Baghdasaryan A, Trauner M (2013) Nuclear receptors as drug targets in cholestatic liver diseases. Clin Liver Dis 17:161–189
Hendriks DFG, Puigvert LF, Messner S et al (2016) Hepatic 3D spheroid models for the detection and study of compounds with cholestatic liability. Sci Rep 6:35434
Henkel AS, LeCuper B, Olivares S et al (2017) Endoplasmic reticulum stress regulated hepatic bile acid metabolism in mice. Cell Mol Gastroenterol Hepatol 3:261–271
Humbert L, Maubert MA, Wolf C et al (2012) Bile acid profiling in human biological samples: comparison of extraction procedures and application to normal and cholestatic patients. J Chromatogr B 899:135–145
Jones SC, Kortepeter C, Brinker AD (2018) Postmarketing surveillance of drug-induced liver injury. Drug-induced liver toxicity. Methods Pharmacol Toxicol. https://doi.org/10.1007/978-1-4939-7677-5_22
Kmiéc Z (2001) Cooperation of liver cells in health and disease. Adv Anat Embryol Cell Biol 161:1–151
Kolarić TO, Ninčević V, Smolić R et al (2019) Mechanisms of hepatic cholestatic drug injury. J Clin Transl Hepatol 7:86–92
Kostrubsky SE, Strom SC, Kalgutkar AS et al (2006) Inhibition of hepatobiliary transport as a predictive method for clinical hepatotoxicity of nefazodone. Toxicol Sci 90:451–459
Laverty HG, Antoine DJ, Benson C et al (2010) The potential of cytokines as safety biomarkers for drug-induced liver injury. Eur J Clin Pharmacol 66:961–976
Lee WM (2013) Drug-induced acute liver failure. Clin Liver Dis 17:575–586
Lepist EI, Gillies H, Smith W et al (2014) Evaluation of the endothelin receptor antagonists ambrisentan, bosentan, macitentan, and sitaxsentan as hepatobiliary transporter inhibitors and substrates in sandwich- cultured human hepatocytes. PLoS ONE 9:e87548
Liu R, Li X, Huang Z et al (2018) C/EBP homologous protein-induced loss of intestinal epithelial stemness contributes to bile duct ligation-induced cholestatic liver injury in mice. Hepatology 67:1441–1457
Malhi H, Kaufman RJ (2011) Endoplasmic reticulum stress in liver disease. J Hepatol 54:795–809
McGill MR, Yan HM, Ramachandran A et al (2011) HepaRG cells: a human model to study mechanisms of acetaminophen hepatotoxicity. Hepatology 53:974–982
Manley S, Ni HM, Kong B et al (2014) Suppression of autophagic flux by bile acids in hepatocytes. Toxicol Sci 137:478–490
Mariotti V, Strazzabosco M, Fabris L et al (2017) Animal models of biliary injury and altered bile acid metabolism. Biochim Biophys Acta 1864:1254–1261
Morgan RE, Trauner M, van Staden CJ et al (2010) Interference with bile salt export pump function is a susceptibility factor for human liver injury in drug development. Toxicol Sci 118:485–500
Nguyen KD, Sundaram V, Ayoub WS (2014) Atypical causes of cholestasis. World J Gastroenterol 20:9418–9426
Noor F (2015) A shift in paradigm towards human biology-based systems for cholestatic-liver diseases. J Physiol 593:5043–5055
Oorts M, Baze A, Bachellier P et al (2016) Drug-induced cholestasis risk assessment in sandwich-cultured human hepatocytes. Toxicol In Vitro 34:179–196
Parent R, Marion MJ, Furio L et al (2004) Origin and characterization of a human bipotent liver progenitor cell line. Gastroenterology 126:1147–1156
Qu X, Zhang Y, Zhang S, Zhai J, Gao H, Tao L, Song Y (2018) Dysregulation of BSEP and MRP2 may play an important role in isoniazid-induced liver injury via the SIRT1/FXR pathway in rats and HepG2 cells. Biol Pharm Bull 41(8):1211–1218
Rakotondravelo S, Poinsignon Y, Borsa-Lebas F et al (2012) Complicated atazanavir-associated cholelithiasis: a report of 14 cases. Clin Infect Dis 55:1270–1272
Riede J, Poller B, Huwyler J et al (2017) Assessing the risk of drug-induced cholestasis using unbound intrahepatic concentrations. Drug Metab Dispos 45:523–531
Rodrigues RM, Kollipara L, Chaudhari U et al (2018) Omics-based responses induced by bosentan in human hepatoma HepaRG cell cultures. Arch Toxicol 92:1939–1952
Roman ID, Fernandez-Moreno MD, Fueyo JA et al (2003) Cyclosporin A induced internalization of the bile salt export pump in isolated rat hepatocyte couplets. Toxicol Sci 71:276–281
Schoemaker MH, Conde de la Rosa L, Buist-Homan M et al (2004) Tauroursodeoxycholic acid protects rat hepatocytes from bile acid-induced apoptosis via activation of survival pathways. Hepatology 39:1563–1573
Seok J, Warren HS, Cuenca AG et al (2013) Genomic responses in mouse models poorly mimic human inflammatory diseases. Proc Natl Acad Sci USA 110:3507–3512
Sharanek A, Bachour-El Azzi P, Al-Attrache H et al (2014) Different dose-dependent mechanisms are involved in early cyclosporine-A induced cholestatic effects in HepaRG cells. Toxicol Sci 141:244–253
Sharanek A, Burban A, Humbert L et al (2015) Cellular accumulation and toxic effects of bile acids in cyclosporine A-treated HepaRG hepatocytes. Toxicol Sci 147:573–587
Sharanek A, Burban A, Humbert L et al (2017) Progressive and preferential cellular accumulation of hydrophobic bile acids induced by cholestatic drugs is associated with inhibition of their amidation and sulfatation. Drug Metab Dispos 45:1292–1303
Tag CG, Sauer-Lehnen S, Weiskirchen S et al (2015) Bile duct ligation in mice: induction of inflammatory liver injury and fibrosis by obstructive cholestasis. J Vis Exp 96:52438
Tagliacozzi D, Mozzi AF, Casetta B et al (2003) Quantitative analysis of bile acids in human plasma by liquid chromatography-electrospray tandem mass spectrometry: a simple and rapid one-step method. Clin Chem Lab Med 41:1633–1641
Van Campenhout S, Van Vlierberghe H, Devisscher L (2019) Common bile duct ligation as model for secondary biliary cirrhosis. Methods Mol Biol 1981:237–247
Van den Hof WF, Ruiz-Aracama A, Van Summeren A et al (2015) Integrating multiple omics to unravel mechanisms of cyclosporin A induced hepatotoxicity in vitro. Toxicol In Vitro 29:489–501
Vatakuti S, Olinga P, Pennings JLA, Groothuis GMM (2017) Validation of precision-cut liver slices to study drug-induced cholestasis: a transcriptomics approach. Arch Toxicol 91(3):1401–1412
Villeneuve DL, Crump D, Garcia-Reyero N et al (2014) Adverse outcome pathway (AOP) development I: strategies and principles. Toxicol Sci 142:312–320
Vinken M, Landesmann B, Goumenou M et al (2013) Development of an adverse outcome pathway from drug-mediated bile salt export pump inhibition to cholestatic liver injury. Toxicol Sci 136:97–106
Woolbright BL, Jaeschke H (2012) Novel insight into mechanisms of cholestatic liver injury. World J Gastroenterol 18:4985–4993
Yasumiba S, Tazuma S, Ochi H et al (2001) Cyclosporin A reduces canalicular membrane fluidity and regulates transporter function in rats. Biochem J 354:591–596
Zhang D, Chango TJ, Everett DW et al (2005) In vitro inhibition of UDP glucuronosyltransferases by atazanavir and other HIV protease inhibitors and the relationship of this property to in vivo bilirubin glucuronidation. Drug Metab Dispos 33:1729–1739
Zhang J, Kan H, Cai L et al (2016) Inhibition of bile salt transport by drugs associated with liver injury in primary hepatocytes from human, monkey, dog, rat and mouse. Chem Biol Interact 255:45–54
Zollner G, Marschall HU, Wagner M et al (2006) Role of nuclear receptors in the adaptive response to bile acids and cholestasis: pathogenetic and therapeutic considerations. Mol Pharm 3:231–251
Zollner G, Trauner M (2006) Molecular mechanisms of cholestasis. Wien Med Wochenschr 156:380–385
Acknowledgements
This work was supported by grants of the Research Foundation Flanders, Belgium and the Scientific Fund Willy Gepts, Belgium and the Center for Alternatives to Animal Testing at Johns Hopkins University, USA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gijbels, E., Vilas‐Boas, V., Annaert, P. et al. Robustness testing and optimization of an adverse outcome pathway on cholestatic liver injury. Arch Toxicol 94, 1151–1172 (2020). https://doi.org/10.1007/s00204-020-02691-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-020-02691-9