Abstract
The entomopathogenic mushroom Cordyceps militaris is a storehouse of various medicinal compounds and pharmacological effects. However, the high frequency of strain degeneration during subculture and preservation severely limits the large-scale production of C. militaris. DNA methylation is an important epigenomic modification involved in gene regulation. In this study, we used bisulfite sequencing for DNA methylation profiling of wild-type and mutant C. militaris. The differentially methylated regions (DMRs) of the two types were analyzed using Gene Ontology (GO) clustering and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis. DNA methylation levels of the wild-type and mutant-type C. militaris were 0.48% and 0.56%, respectively. Methylation appeared at CHH dinucleotides in 58.62% and 58.20% of all methylated cytosine sites in the wild and mutant types, respectively. In all, 188 DMRs were identified from the wild and mutant types. Most of the DMRs ranged from 200 to 350 bp in length. KEGG pathways of the expression of DMR-related genes, which are involved in pyruvate metabolism, glycerophospholipid metabolism, DNA replication, and N-glycan biosynthesis. This contributes to the knowledge and understanding of the possible mechanisms of C. militaris strain degeneration.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cordyceps militaris is a type of edible and medicinal fungus that is considerably similar to Cordyceps sinensis. It can be used as a substitute of C. sinensis to treat many diseases, because they both have nearly identical chemical composition and pharmacological activities (Kim and Yun 2005; Li et al. 2006). C. militaris exhibits several desirable biological activities, including antitumor (Lin and Chiang 2008; Park et al. 2009; Chen et al. 2015), antioxidant (Yu et al. 2007), anti-inflammatory (Won and Park 2004), and hypotensive effects (Chiou 2000). The effective biological constituents are cordycepin (3′-deoxyadenosine), adenosine, polysaccharides, and cordycepic acid. Recently, the biosynthesis mechanism of the main pharmacological component, cordycepin, was revealed (Xia et al. 2017).
Owing to its large market demand, natural C. militaris is in seriously short supply. This shortage has been eliminated by establishing artificial cultivation of C. militaris thereby realizing its large-scale industrial production. However, strain degeneration occurs during preservation and subculture, which markedly affects C. militaris yield (Lu et al. 2016). Therefore, determining the reasons for this degeneration of C. militaris is important to overcome them and meet the increasing demands.
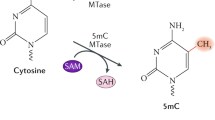
DNA methylation refers to the transfer of methyl groups to a specific base by DNA methyltransferase, where S-adenosylmethionine acts as the methyl donor (Sulewska et al. 2007). DNA methylation plays a key role in eukaryotic gene silencing and expression, phylogeny, cell differentiation, and other processes (Lee et al. 2010). Numerous studies have shown that DNA methylation can cause changes in chromatin structure, DNA conformation, DNA stability, and DNA–protein interaction, thereby affecting gene expression.
DNA methylation has recently become crucial in the field of fungal epigenetics. Compared to that in other eukaryotes, the DNA methylation ratio in fungi is relatively low, but has important effects on degeneration during repeated cultures. The degree of DNA methylation differs across species and changes dynamically during different growth and development stages of fungi. In fungi, the highest level of DNA methylation has been reported in Neurospora crassa (1.5% methylated cytosine [mC] of total cytosines, throughout the life cycle) (Selker et al. 2003). About 0.22% of cytosines are methylated in Magnaporthe grisea (mycelia) (Jeon et al. 2015). However, no mC has been detected in the asexual stage of Saccharomyces cerevisiae (Colot and Rossignol 1999) or Aspergillus flavus (Liu et al. 2012).
Epigenetic modification in most eukaryotes, including genome regulation and development, has been suggested to play crucial roles in the mechanisms modulating gene transcription, and thus affecting various cellular processes (Cichewicz 2010). However, whether metabolic processes underlying strain degeneration in C. militaris are related to epigenetic variation, particularly DNA methylation, is not yet known.
Considering the lack of information on DNA methylation in C. militaris, in this study, bisulfite sequencing (BS-Seq), which is considered as the gold standard for detecting DNA methylation, was used for DNA methylation profiling. To our knowledge, this is the first study to reveal a clear and comprehensive DNA methylation profile of C. militaris. The genomic DNA from the mycelia of wild and mutant types of C. militaris was extracted and sequenced using BS-SEq. The differentially methylated regions (DMRs) were analyzed using Gene Ontology (GO) clustering and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis. Our findings might contribute to the understanding of the DNA methylation status of C. militaris and its effect on strain degeneration. Furthermore, the use of BS-Seq to detect DNA methylation in species with low DNA methylation might provide a basis for future scientific studies in other hypomethylated species.
Materials and methods
Cultivation of C. militaris mycelium
A colony of C. militaris with abundant mycelial growth on agar slants was selected and inoculated into 100 mL of broth medium(Wu et al. 2012). The inoculum was incubated under agitation in the dark at 25 °C and 130–140 rpm for 5 days to obtain C. militaris mycelia. The C. militaris mycelia were subcultured for the acquisition of the second, third, fourth, fifth, and sixth generations, and designated as CmG2, CmG3, CmG4, CmG5, and CmG6, respectively. Mycelia from all six generations were co-cultured in rice medium. In CmG6 only, completely degenerated mycelia without fruiting bodies were noted. Thus, this was used as the mutant type (Yin et al. 2017a). Mycelia from CmG1 and CmG6 were selected and collected for DNA extraction.
Genomic DNA extraction
Excess water was removed from 5-day-old mycelium of C. militaris using a vacuum pump. Dried mycelium (0.25 g) was rapidly ground into powder in liquid nitrogen. The genomic DNA of C. militaris was extracted using a modified method by Mayjonade et al. (2016). The integrity of the extracted genomic DNA was determined using 0.8% agarose gel electrophoresis, and its purity was determined on the basis of the OD260/OD280 ratio.
Bisulfite library construction and sequencing of C. militaris
An unmethylated DNA fragment (~ 50 kb) extracted from transfected Escherichia coli was used to determine the bisulfite conversion efficiency. For this, 25 ng of unmethylated DNA was added to 5 ng genomic DNA of C. militaris. Next, the mixed DNA was fragmented by sonication to 100–300 bp fragments, followed by end blunting, deoxyadenosine addition at the 3′ end, and adapter ligation. Unmethylated cytosines were converted to uracils by bisulfite treatment using the EZ DNA Methylation-Gold kit (Zymo Research, Irvine, CA). DNA fragments in the size range of 180–260 bp were gel-purified for sequencing. BS-Seq and data processing were performed as described previously (Lou et al. 2014). The C. militaris DNA was bisulfite converted using a modified NH4SO4-based protocol (Hayatsu et al. 2006), and the DNA was amplified by conducting 12 cycles of PCR. Ultra-high-throughput 50 bp pair-end sequencing was performed using the Illumina Solexa GA sequencer (Illumina, San Diego, CA, USA), according to manufacturer’s instructions. All raw data were processed using the Illumina pipeline (v1.3.1; Illumina, San Diego, CA, USA).
The cleaned reads generated were aligned to the reference genomic sequence of C. militaris using BSMAP (version: 2.90; Xi and Li 2009). Since DNA methylation is strand-specific, the two strands of the reference C. militaris genome were modified separately in silico to convert all cytosines to thymines to generate a combined 6 Gbp target genome for aligning reads after bisulfite conversion. All the reads mapped to unique locations with minimum mismatches. Clear strand information was defined as uniquely matched reads, which were used to determine the methylated cytosines. The sequence that matched to the genomic sequence of C. militaris was selected for bioinformatics analysis. All information on the bases and methylated cytosines in the entire genome were obtained.
Data processing and analysis
Before sequencing, raw data were filtered to obtain high-quality clean data. Clean reads were then assembled de novo into longer contigs based on overlapping regions using the Trinity platform (http://trinityrnaseq.sourceforge.net/; Mortazavi et al. 2008). All subsequent analyses were performed using clean data. For annotation, clean data were mapped to the C. militaris genomic data (BioProject accession no., PRJNA225510) from the NCBI transcriptome reference database.
GO was used to predict the possible functions of all differentially expressed genes (DEGs). GO annotations for each DEG were retrieved by mapping to GO terms in the database at http://www.geneontology.org. Pathway analysis was performed using the Molecular Homological Description System 2.0 (MAS, 2.0, http://www.capitalbio.com) developed by CapitalBio Corporation. KEGG orthology terms for DEGs were retrieved from the KEGG pathway database (http://www.genome.jp/kegg/). Cluster analysis of gene expression patterns was performed using cluster software (Hoon et al. 2004) and Java TreeView software (Saldanha 2004). Based on KEGG, pathways with Q values ≤ 0.05 were defined as significantly enriched pathways. The Bonferroni corrected P values were calculated for the relative transcript level of genes.
Results
Genome-wide DNA methylation level in the C. militaris genome
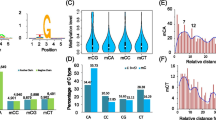
DNA methylation refers to the methylation of the carbon atom at position 5 of a cytosine (m5C), which mostly occurs within CpG (CG), CpHpG (CHG), and CpHpH (CHH) nucleotide patterns in eukaryotes (Cokus et al. 2008). The methylation density measure mCG/CG has been commonly used in various methylome studies to quantify the DNA methylation level (Lister et al. 2009).
The DNA methylation of C. militaris was determined by performing whole-genome BS-Seq of wild-and mutant-type C. militaris. Different numbers of mC sites were identified in the genome of each sample, although the average methylation level (defined as the number of mC reads divided by the number of total reads covering the site) of individual mC sites remained between 20 and 30% across chromosomes and samples. Results showed that the DNA methylation level in wild-type C. militaris was 0.48%, of which 0.47%, 0.48%, and 0.49% were at CG, CHG, and CHH sites, respectively. In the mutant, the DNA methylation level was 0.56%, of which 0.55%, 0.55%, and 0.57% were at CG, CHG, and CHH sites (Table 1). No significant differences were found in the %mC of all genomic cytosines; however, mCs between wild- and mutant-type C. militaris showed considerable differences. The methylation level of the mutant type was significantly higher than that of the wild type. This implies that the DNA methylation level might be closely linked to the degradation of C. militaris.
DNA methylation patterns in wild-and mutant-type C. militaris
We explored the global patterns of DNA methylation in wild- and mutant-type C. militaris. Methylation appeared at CHH dinucleotides, with 58.62% and 58.20% of all mC sites methylated in the wild and mutant types, respectively. Methylation levels at CG and CHG dinucleotides were 22.66% and 23.45%, and 18.72% and 18.35% of all mC sites in wild-and mutant-type C. militaris, respectively. No significant differences in CG, CHG, or CHH nucleotide patterns were noted between the two types (Fig. 1).
Analyses of DMR length
DMRs were identified based on the differential methylation levels of mC sites from the same region of the two genomes. Between the wild and mutant types, 188 DMRs, with an average length of 299 bp, were identified. Most DMRs ranged from 200 to 350 bp in length. Only 2 and 4 DMRs ranged from 500 to 550 bp, respectively (Fig. 2).
GO analysis of DMR-related genes in the 2 kb upstream and downstream regions and gene body
To ascertain the relationship of DMR distribution between wild- and mutant-type C. militaris, the 188 DMR-related genes were subjected to GO clustering, with P ≤ 0.05 as the critical value. We found that the DMR-related genes in the 2 kb downstream region were enriched in five cell components, participated in 12 functional molecular processes, and were involved in 26 biological processes. DMR-related genes in the gene body region were enriched in one cell component, participated in seven molecular functions, and were involved in 19 biological processes. DMR-related genes in the 2 kb upstream region were enriched in five cell components, participated in six molecular functions, and were involved in 21 biological processes (Table 2). The cell component genes mainly included the reverse transcription complex and SWI/SNF superfamily complex, among others. The molecular functions were mainly phosphoric diester hydrolase activity, nucleic acid-binding transcription factor activity, and sequence-specific DNA binding, among others. Further, the main biological processes were lipid metabolism, citrate metabolism, and tricarboxylic acid metabolism (Fig. 3).
KEGG pathway analyses of DMR-related genes
DMR-related genes were annotated using KEGG pathway information (Table 3; Fig. 4). Pathway enrichment analysis showed that, in the mutant type, eight genes were significantly enriched in four pathways in the 2 kb downstream region; eight other genes were significantly enriched in four different pathways in the gene body region; and six genes were significantly enriched in two pathways in the 2 kb upstream region. The findings suggested that the DMR-related genes were involved mainly in ten pathways, including pyruvate metabolism pathway, glycerophospholipid metabolism pathway, DNA replication pathway, and N-glycan biosynthesis pathway.
Discussion
Owing to the similarities between C. militaris and C. sinensis, C. militaris is widely used in clinical treatment as a valuable traditional Chinese medicine and often as a substitute of C. sinensis. The production scale of C. militaris has increased considerably, and it is being industrialized for supplying to Asian countries such as China, Korea, and Japan. However, strain degeneration frequently occurs during subculture and preservation, leading to remarkable economic losses. Several factors might lead to fungal strain degeneration, including DNA methylation, gene mutations, mating type change, and viral infections (Hu and Lv 2015). In a previous study, we found that mutations in 18S and mating-type genes, as well as the downregulation of CmMAT expression levels, might play important roles in the degeneration of C. militaris in culture (Yin et al. 2017a). Transcriptome-wide analyses revealed 51 DEGs between wild-and mutant-type C. militaris. Strain degeneration of C. militaris is associated with genes involved in toxin biosynthesis, energy metabolism, DNA methylation, and chromosome remodeling (Yin et al. 2017b). Recently, Sun et al. (2017) suggested that this degradation was not attributable to the DNA changes identified by RAPD and SRAP, and may be caused by the inhibition or in harmony of metabolite synthesis of metabolic regulation.
Some studies have shown that DNA methylation in fungal species is phylogenetically widespread and ancient in origin, but shows considerable variation, which is reflected in both genome methylation patterns and diverse regulation mechanisms (Zemach et al. 2010). BS-Seq allows DNA methylation calls at a single-base resolution and in a quantitative manner. It not only estimates the global DNA methylation level but also reveals the DNA methylation pattern in the context of different sequences, thereby providing important insights into the distribution and function of DNA methylation. Despite the potential importance of DNA methylation in fungi, the relationship between the genome-wide methylation patterns and fungal degeneration has remained poorly understood. In this study, we determined the average levels of DNA methylation using deep BS-Seq in wild-and mutant-type C. militaris, and the DMRs of these two types were analyzed.
Our results showed that the DNA methylation levels in wild type were 0.48% (Table 1). This methylation level was lower than that of N. crassa (1.5% mC of C, Selker et al. 2003), but higher than that of M. grisea (0.22% mC of C, Jeon et al. 2015). In differentiated mammalian cells, methylation appears predominantly at CpG dinucleotides, with methylation at about 60–90% of all CpG sites(Bird 1986). In this study, C. militaris methylation was noted in CHH dinucleotides at 58.62% and 58.20% of all mC sites in the wild-and mutant-type C. militaris, respectively (Fig. 1). Our findings are similar to those of Bird (1986). The methylation level of the mutant type (0.56%) was significantly higher than that of the wild type (0.48%, Table 1). This implies that the DNA methylation level might be closely linked to the degradation of C. militaris.
Methylation modification and DNA recombination can alter a strain’s genotype and thus induce strain degeneration (Wang et al. 2015). In this study, we characterized and manually annotated the functions of DMRs involved in methylation modification using GO and KEGG pathway analysis. We found that the pyruvate metabolism, glycerophospholipid metabolism, ubiquitin-mediated proteolysis, and N-glycan biosynthesis pathways might be related to the degeneration of C. militaris.
To our knowledge, this is the first report on the DNA methylation level in C. militaris mycelium and provides an overview of methylation dynamics during fungal subculture. DNA methylation does not appear to have a direct association with the degeneration of C. militaris, but epigenetic control of certain pathways might lead to differences in gene expression between the wild- and mutant-type C. militaris. Changes of DNA methylation level could occur in C. militaris industrial production. However, notably, epigenetic regulation is multilayered in nature, and DNA methylation represents only one level; other modes of epigenetic regulation include histone modifications and non-coding RNAs. In the future, the epigenetic regulatory mechanisms should be investigated during the subculture of C. militaris.
References
Bird AP (1986) CpG-rich islands and the function of DNA methylation. Nature 321:209–213
Chen C, Wang LM, Jin C, Chen HJ, Li SH, Li SY, Dou XF, Jia JQ, Gui ZZ (2015) Cordyceps militaris polysaccharide triggers apoptosis and G0/G1 cells arrest in cancer cells. J Asia-Pac Entomol 18:433–438
Chiou WF, Chang PC, Chou CJ, Chen CF (2000) Protein constituent contributes to the hypotensive and vasorelaxant activities of Cordyceps sinensis. Life Sci 66:1369–1376
Cichewicz RH (2010) Epigenome manipulation as a pathway to new natural product scaffolds and their congeners. Nat Prod Rep 27:11–22
Cokus SJ, Feng S, Zhang X, Chen Z, Merriman B, Haudenschild CD, Pradhan S, Nelson SF, Pellegrini M, Jacobsen SE (2008) Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452:215–219
Colot V, Rossignol JL (1999) Eukaryotic DNA methylation as an evolutionary device. BioEssays 21:402–411
Hayatsu H, Tsuji K, Negishi K (2006) Does urea promote the bisulfite-mediated deamination of cytosine in DNA? Investigation aiming at speeding-up the procedure for DNA methylation analysis. Nucleic Acids Symp Ser 50:69–70
Hoon MJ, De L, Imoto S, Nolan J, Miyano S (2004) Open source clustering software. Bioinformatics 20:1453–1454
Hu XC, Lv AJ (2015) University JN. Discussion of the degenerate strains of Cordyceps militaris. Edible Fungi China 34:1–3
Jeon JH, Choi JY, Lee GW, Park SY, Huh A, Dean RA, Lee YH (2015) Genome-wide profiling of DNA methylation provides insights into epigenetic regulation of fungal development in a plant pathogenic fungus Magnaporthe oryzae. Sci Rep 5:8567
Kim HO, Yun JW (2005) A comparative study on the production of exopolysaccharides between two entomopathogenic fungi Cordyceps militaris and Cordyceps sinensis in submerged mycelial cultures. J Appl Microbiol 99:728–738
Lee TF, Zhai J, Meyers BC (2010) Conservation and divergence in eukaryotic DNA methylation. PNAS 107:9027–9028
Li SP, Yang FQ, Tsim KW (2006) Quality control of Cordyceps sinensis, a valued traditional Chinese medicine. J Pharmaceut Biomed 41:1571–1584
Lin YW, Chiang BH (2008) Anti-tumor activity of the fermentation broth of Cordyceps militaris cultured in the medium of Radix astragali. Process Biochem 43:244–250
Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo Q, Edsall L, Antosiewicz-Bourget J, Stewart R, Ruotti V, Millar AH, Thomson JA, Ren B, Ecker JR (2009) Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462:315–322
Liu SY, Lin JQ, Wu HL, Wang CC, Huang SJ, Luo YF, Sun JH, Zhou JX, Yan SJ, He JG, Wang J, He ZM (2012) Bisulfite sequencing reveals that Aspergillus flavus holds a hollow in DNA methylation. PLoS One 7:e30349
Lou SK, Lee HM, Qin H, Li JW, Gao ZB, Liu X, Chan LL, Lam VKL, So WY, Wang Y, Lok S, Wang J, Ma RCW, Tsu SKW, Chan JCN, Chan TF, Yip KY (2014) Whole-genome bisulfite sequencing of multiple individuals reveals complementary roles of promoter and gene body methylation in transcriptional regulation. Genome Biol 15:408
Lu Y, Xia Y, Luo F, Dong C, Wang C (2016) Functional convergence and divergence of mating-type genes fulfilling in Cordyceps militaris. Fungal Genet Bio l88:35–43
Mayjonade B, Gouzy J, Donnadieu C, Pouilly N, Marande W, Callot C, Langlade N, Muños S (2016) Extraction of high-molecular-weight genomic DNA for long-read sequencing of single molecules. Biotechniques 61:203–205
Mortazavi A, Williams BA, Mccue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-SEq. Nat Methods 5:621–628
Park S, Yoo HS, Jin CY, Hong SH, Lee YW, Kim BW, Lee SH, Kim WJ, Cho CK, Choi HY (2009) Induction of apoptosis and inhibition of telomerase activity in human lung carcinoma cells by the water extract of Cordyceps militaris. Food Chem Toxicol 47:1667–1675
Saldanha AJ (2004) Java Treeview–extensible visualization of microarray data. Bioinformatics 20:3246–3248
Selker EU, Tountas NA, Cross SH, Margolin BS, Murphy JG, Bird AP, Freitag M (2003) The methylated component of the Neurospora crassa genome. Lett Nat 422:893–897
Sulewska A, Niklinska W, Kozlowski M, Minarowski L, Naumnik W, Niklinski J, Dabrowska K, ChyczewskiL (2007) DNA methylation in states of cell physiology and pathology. Folia Histochem Cyto 45:149–158
Sun SJ, Deng CH, Zhang LY, Hu KH (2017) Molecular analysis and biochemical characteristics of degenerated strains of Cordyceps militaris. Arch Microbiol 199(6):939–944
Wang YL, Wang ZX, Wang SB, Huang B (2015) Genome-wide analysis of DNA methylation in the sexual stage of the insect pathogenic fungus Cordyceps militaris. Fungal Biol 119:1246–1254
Won SY, Park EH (2004) Anti-inflammatory and related pharmacological activities of cultured mycelia and fruiting bodies of Cordyceps militaris. J Ethnopharmacol 96:555–561
Wu FY, Yan H, Ma XN, Jia JQ, Zhang GZ, Guo XJ, Gui ZZ (2012) Comparison of the structural characterization and biological activity of acidic polysaccharides from Cordyceps militaris cultured with different media. World J Microbiol Biotechnol. https://doi.org/10.1007/s11274-012-1005-6
Xi Y, Li W (2009) BSMAP: whole genome bisulfite sequence MAPping program. BMC Bioinform 10:232
Xia YL, Luo FF, Shang YF, Chen PL, Lu YZ, Wang CS (2017) Fungal cordycepin biosynthesis is coupled with the production of the safeguard molecule pentostatin. Cell Chem Biol. https://doi.org/10.1016/j.chembiol.2017.09.001
Yin J, Xin XD, Weng YJ, Li SH, Jia JQ, Gui ZZ (2017a) Genotypic analysis of degenerative Cordyceps militaris cultured in the pupa of Bombyx mori. Entomol Res. https://doi.org/10.1111/1748-5967.12246
Yin J, Xin XD, Weng YJ, Gui ZZ (2017b) Transcriptome-wide analysis reveals the progress of Cordyceps militaris subculture degeneration. PloS One. https://doi.org/10.1371/journal.pone.0186279
Yu R, Yang W, Song L, Yan C, Zhang Z, Zhao Y (2007) Structural characterization and antioxidant activity of a polysaccharide from the fruiting bodies of cultured Cordyceps militaris. Carbohyd Polym 70:430–436
Zemach A, McDaniel IE, Silva P, Zilberman D (2010) Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science 328:916–919
Funding
This research was funded by the Special Fund for Agro-scientific Research in the Public Interest of China (No. 201403064), and Postgraduate Research & Practice Innovation Program of Jiangsu Province (SJCX17_0610).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Communicated by Olaf Kniemeyer.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xin, X., Yin, J., Zhang, B. et al. Genome-wide analysis of DNA methylation in subcultured Cordyceps militaris. Arch Microbiol 201, 369–375 (2019). https://doi.org/10.1007/s00203-019-01621-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-019-01621-3