Abstract
A novel arsenic (As)-resistant, arsenate-respiring, alkane-metabolizing bacterium KAs 5-22T, isolated from As-rich groundwater of West Bengal was characterized by physiological and genomic properties. Cells of strain KAs 5-22T were Gram-stain-negative, rod-shaped, motile, and facultative anaerobic. Growth occurred at optimum of pH 6.0–7.0, temperature 30 °C. 16S rRNA gene affiliated the strain KAs 5-22T to the genus Rhizobium showing maximum similarity (98.4 %) with the type strain of Rhizobium naphthalenivorans TSY03bT followed by (98.0 % similarity) Rhizobium selenitireducens B1T. The genomic G + C content was 59.4 mol%, and DNA–DNA relatedness with its closest phylogenetic neighbors was 50.2 %. Chemotaxonomy indicated UQ-10 as the major quinone; phosphatidylethanolamine, phosphatidylglycerol, and diphosphatidylglycerol as major polar lipids; C16:0, C17:0, 2-OH C10:0, 3-OH C16:0, and unresolved C18:1 ɷ7C/ɷ9C as predominant fatty acids. The cells were found to reduce O2, As5+, NO3 −, SO4 2− and Fe3+ as alternate electron acceptors. The strain’s ability to metabolize dodecane or other alkanes as sole carbon source using As5+ as terminal electron acceptor was supported by the presence of genes encoding benzyl succinate synthase (bssA like) and molybdopterin-binding site (mopB) of As5+ respiratory reductase (arrA). Differential phenotypic, chemotaxonomic, genotypic as well as physiological properties revealed that the strain KAs 5-22T is separated from its nearest recognized Rhizobium species. On the basis of the data presented, strain KAs 5-22T is considered to represent a novel species of the genus Rhizobium, for which the name Rhizobium arsenicireducens sp. nov. is proposed as type strain (=LMG 28795T=MTCC 12115T).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Geogenic arsenic (As) in alluvial groundwater of West Bengal, Bangladesh and several other parts of Southeast Asia has created severe problems in drinking water resources, affecting millions of people. Geomicrobiological activities that form the integral components of As biogeochemical cycle in subsurface environment have been attributed to the prevalence of toxic As level in groundwater (Malasarn et al. 2008; Blum et al. 2009; Ohtsuka et al. 2013; Zhu et al. 2014). Bacterial transformation of sediment bound As (As5+) and/or Fe (Fe3+) of host minerals [Fe (Mn/Al)-oxides/hydroxides, phyllosilicates, and arseno-pyrites] is considered as the most feasible mechanism for As mobilization in sub-oxic alluvial groundwater (Paul et al. 2015). Some inhabitant bacteria use the sediment associated As5+ as a terminal electron acceptor during their anaerobic metabolism reducing it to more soluble As3+ species and are often categorized as dissimilatory As5+-reducing bacteria (DARB) (Newman et al. 1998; Saltikov and Newman 2003; Kudo et al. 2013; Osborne et al. 2015). Although, the anaerobic reduction of sediment bound As5+ by DARB has been a subject of considerable interest to decipher the role of such processes in As biogeochemistry and in As release within the alluvial environment, until today thorough characterization of bacterial strains isolated from As-rich alluvial groundwater and capable of such transformation remains mostly elusive.
Alluvial aquifer of West Bengal is characterized to be of low nutrient content. Presence of alkane hydrocarbons in As-rich aquifer of West Bengal and organisms capable of utilizing many of them as sole carbon source and respiring As5+ have been recently reported (Ghosh et al. 2015; Paul et al. 2015). Under anaerobic condition, very low concentrations of metabolizable substrates (mostly in the form of aromatics, long-chain alkanes, hopanes and steranes as well as geochemically driven simpler organic molecules) have been found to be the key factors driving the dissimilatory (respiratory) As5+ reduction (Lear et al. 2007; Paul et al. 2015). With respect to taxonomic and physiological characterization of DARB, up to now 21 cultivable representatives have been studied (Saltikov et al. 2003; Saltikov and Newman 2003; Kudo et al. 2013; Osborne et al. 2015). Interestingly, except the strain WB-3 Desulfuromonas/Pelobacter sp. affiliated to the class Gammaproteobacteria (Osborne et al. 2015), no other strain was isolated from As-rich alluvial environment. No pure culture has been isolated from As-contaminated groundwater and studied for its taxonomic characterization, overall physiology and As biotransformation ability (respiratory function) under anaerobic condition. Although many studies have shown the abundance of members of the class Alphaproteobacteria (mostly Rhizobium species) within As-rich aquifer and investigated their capability of using As5+ anaerobically (Lear et al. 2007; Fan et al. 2008; Kudo et al. 2013; Sarkar et al. 2014; Drewniak et al. 2015; Paul et al. 2015), none of them have been taxonomically described. The present study was carried out to highlight the taxonomic description of an As5+-respiring strain KAs 5-22T isolated from As-contaminated groundwater of India.
Taxonomic description of the genus Rhizobium denotes its affiliation to the family Rhizobiaceae, order Rhizobiales, class Alphaproteobacteria, and phylum Proteobacteria. Since its first description by Frank (1889), 94 validly named species (LPSN, http://www.bacterio.net/) were affiliated to the genus Rhizobium. Members of the genus Rhizobium are characterized as Gram-stain-negative, non-spore forming and chemo-organotrophic rod-shaped bacteria with C18:1 ɷ7c as the major fatty acid and DNA G + C content between 57 and 66 mol% (Tighe et al. 2000; Young et al. 2001). Members are well distributed in soil habitat and endowed with immense environmental as well as agricultural significance for their ability to fix nitrogen (N2) in legume crops. They have been recognized for their ability to form root nodules on legumes and fix nitrogen (Viteri and Schmidt 1987; Young et al. 2001). Although most of the members have been isolated from nodules of leguminous plants (Mnasri et al. 2007), in recent years, an increasing number of new members have been isolated from diverse non-legume niches including sand dunes, effluent treatment plant, activated sludge, bioreactor, pesticide-contaminated sites, freshwater river, and sea water (Kaur et al. 2011; Ramana et al. 2013; Liu et al. 2015; Sheu et al. 2015). Among the recently described members, presence of diverse metabolic functions like naphthalene degradation (R. naphthalenivorans; Kaiya et al. 2012), selenite reduction (R. selenitireducens; Hunter et al. 2007), utilization of hydrocarbons for exopolysaccharides production (R. alamii; Berge et al. 2009), degradation of aniline (R. borbori; Zhang et al. 2011), polycyclic aromatic hydrocarbon (R. petrolearium; Zhang et al. 2012), triazophos (R. flavum Gu et al. 2014), etc. has been reported. During our earlier survey on cultivable microbial diversity of highly As-contaminated groundwater of West Bengal, one such Rhizobium strain, designated as KAs 5-22, was isolated (Sarkar et al. 2013). Polyphasic taxonomic approach including phenotypic, chemotaxonomic, phylogenetic, genotypic, as well as in vitro physiological tests and functional gene-based analysis was carried out to investigate the species novelty and metabolic potential of the strain.
Materials and methods
Bacterial strains and growth conditions
Strain KAs 5-22T (MTCC 12115T, BCCM 28795T) was previously isolated from high As content groundwater (N 22°47.750 and E 88°44.121), of West Bengal, India, in the month of August, 2009 (Sarkar et al. 2013). Unless otherwise indicated, the strain was maintained and routinely subcultured on Luria Bertani (LB) agar or minimal salt medium (MSM) (Kazy et al. 1999). Reference strains of Rhizobium (R. daejeonense DSM 17795T, R. aggregatum DSM 1111T, R. selenitireducens LMG 1111T, R. radiobacter IAM 12048T, R. rossetiformans MTCC 9454T, R. naphthalenivorans KCTC 23252T), Escherichia coli K-12, and Shewanella sp. ANA-3 were obtained, grown in appropriate media and used in different experiments.
Phylogenetic and molecular analysis
Genomic and plasmid DNA were extracted according to the standard methods (Sambrook and Russel 2001). Amplification of genes encoding 16S rRNA, arrA, bssA like, nifH and nodA was performed by using either universal specific or degenerate primers (Table S1). PCR amplicons were gel purified, cloned in pTZ57R/T vector (InsTA clone kit, Thermo scientific), and sequenced. For sequencing the nearly complete stretch of 16S rRNA gene, internal primers were used, and individual sequences were assembled, edited (BioEdit version 7.1.11), and subjected to similarity search in NCBI Genbank, Ribosomal Database Project (RDP), and EzTaxon-e server. Phylogenetic dendrograms were constructed using neighbor-joining (NJ) and maximum-likelihood (ML) methods with bootstrap analyses based on 1000 replications. Nucleotide sequences obtained for the coding genes were translated using ExPASy tools, where the appropriate reading frame was selected. The amino acid sequences were searched for protein similarity in non-redundant protein database (BLASTP), and functional domain similarity through conserved domain database (CDD). The NJ tree was constructed by using deduced amino acid sequences of the genes with respective similar sequences.
DNA G + C content and DNA–DNA hybridisation
Molar G + C content of the strain KAs 5-22T was determined using the thermal denaturation method (De Ley et al. 1970) where Escherichia coli K-12 NCIM 2563 was used as an internal standard. DNA–DNA hybridization was carried out between strain KAs 5-22T and its closest neighbors using the SyBr green binding fluorimetry-based method (Gonzalez and Saiz-Jimenez 2005). Optimum renaturation temperature (T OR ) was calculated from the melting curve analysis. Hybridization was performed in 2X sodium chloride sodium citrate (SSC) buffer incorporating 0.1X SyBr dye following the real-time PCR program as described by Gonzalez and Saiz-Jimenez (2005). For each cycle, the relative fluorescence unit (RFU) (with an interval of 0.2 s) was plotted and ∆T m as well as binding percentage (B d%) was calculated following the equation as described by De Ley et al. (1970). The difference in ∆T m values between homologous and hybrid DNA of 5 °C or higher was considered the cutoff value to discriminate between bacterial species (Wayne et al. 1987; Rosselló-Mora and Amann 2001). All experiments were conducted in triplicate, and the mean was indicated as sample values.
Morphological, biochemical, and physiological characterization
Cell morphology was studied using bright-field optical microscopy, fluorescence microscopy, and electron microscopy. For all microscopy, mid log phase cells were harvested, washed with normal saline (0.85 % w/v), fixed [formaldehyde or glutaraldehyde (0.2 %, v/v) in 0.1 mM phosphate buffer saline (PBS), at 4 °C, as appropriate, for 12 h], dehydrated (for SEM), stained appropriately, and viewed. Fluorescence microscopy was performed by staining cells with acridine orange (AO) [100 µg/mL (v/v)] and incubating in the dark for 20 min at 4 °C. For SEM, the cell suspension was fixed with glutaraldehyde; fixed cells were serially dehydrated with increasing concentration of ethanol (30–100 %) (v/v), placed on poly-l-lysine-coated cover glass, coated with gold, and viewed under SEM (SEM, JEOL JSM5800) using a Cu grid. Gram reaction was studied by using Gram’s staining kit (Hi-Media). Motility was tested by the flagella staining protocol of Kodaka et al. (1982). Strain KAs 5-22T along with all the taxonomically related type strains was tested for catalase, oxidase, and other general biochemical properties (viz. nitrate reduction, utilization of gelatin, esculin, citrate, urea, etc.) using either appropriate kits (Hi-Media or Bio-Merieux) as per the manufacturer’s instructions, or following the standard procedures (Cowan and Steel 1965; Kelly and Fulton 1953; Smibert and Krieg 1994). Assimilation of carbon, nitrogen sources, and enzyme production was examined by API 20NE kit (Bio-Merieux) and Biolog GEN-III microplate following manufacturer’s instructions. Temperature, pH, and salt (NaCl %) tolerance was monitored by allowing cell growth in MSM medium. Anaerobic growth was performed by growing cells on LB agar which was purged with a mixture of CO2/N2/H2 [(90:5:5, (%)] and amended with L-Cys-HCl (0.05 %, v/v) and Na2S (2 %, v/v of 70 mM stock solution) using an anaerobic work station (Coy Laboratories, USA). Inoculated plates were incubated in anaerobic gas jar with anaero gas-pak system. Antibiotic susceptibility was tested on Mueller-Hinton agar (Hi-Media) using disk diffusion method. The antibiotics disks used were as follows: ceftriaxone (30 µg), cefixime (5 µg), amikacin (30 µg), cefotaxime (30 µg), chloramphenicol (30 µg), ofloxacin (5 µg), polymyxin-B (300 units), tetracycline (30 µg), ciprofloxacin (5 µg), and erythromycin (15 µg). Arsenic resistance was tested following bacterial growth in LB broth amended with graded concentrations of As3+ (NaAsO2; 0.1–30 mM) and As5+ (Na2HAsO4; 1–300 mM). The lowest concentration of As species, sufficient to completely inhibit bacterial growth, was considered as minimum inhibitory concentration (MIC).
Chemotaxonomic characterization
Cellular fatty acid methyl esters (FAMEs) and isoprenoid quinones were analyzed using cells from the mid log phase of growth. Cells mass collected through centrifugation was subjected to saponification followed by methylation and esterification of fatty acids (Miller 1982; Kuykendall et al. 1988). FAMEs were separated using a gas chromatograph (GC, CLARUS 500, PerkinElmer) fitted with Omega-Wax capillary column (30 mm × 0.25 mm, d f 0.25 µ) and detected by flame ionization detector (FID). FAMEs were identified by comparing with the bacterial fatty acid mixture standard (Sigma). Isoprenoid quinones were extracted from cells and analyzed by reverse phase high-performance liquid chromatography (HPLC) using Sorbax C18 reverse phase column (Agilent) (Komagata and Suzuki 1987). A mixture of methanol: isopropanol (2:1, v/v), using a flow rate of 1 mL min−1, was used for elution of quinones which were subsequently identified by liquid chromatography–mass spectrometry (LC–MS) using positive ionization mode (Hiraishi et al. 1996). Polar lipids were extracted and analyzed by two-dimensional (2-D) thin-layer chromatography (TLC) (Komagata and Suzuki 1987).
Test for use of different electron and carbon donors, electron acceptors and As respiratory growth
Utilization of different electron and carbon donors was tested under both aerobic and anaerobic conditions using MSM supplemented with different sugars (at a concentration of 30 mM) and hydrocarbons (at a concentration of 100 mgL−1). Bacterial growth was monitored by measuring the culture turbidity at 600 nm with a UV–Vis spectrophotometer (Cary 50, Varian). Anaerobic utilization of substrates was tested with As5+ (5 mM) as terminal electron acceptor. Utilization of different terminal electron acceptors (TEAs) was assessed by adding a range of test compounds (As5+, Fe3+, NO3 −, etc.) into MSM with either glucose (30 mM) or lactate (30 mM) or nonadecane (100 mgL−1) as sole carbon–electron source. Anaerobic growth was performed in serum vials (50 ml) containing MSM added with L-Cys-HCl (0.5 g L−1) as reducing agent and resazurin (0.01 %, w/v) as redox indicator. Vials were crimp sealed keeping 30 % (v/v) head space filled with filtered N2 and autoclaved. All anaerobic experiments were set up within anaerobic work station (Coy laboratory, USA) purged with N2. Concentrations of TEAs during growth were measured by standard spectrophotometric methods. Concentrations of glucose and nonadecane were estimated by a modified phenol sulfuric acid method (Albalasmeh et al. 2013) and GC-FID, respectively. Concentration of As was measured by inductively coupled plasma–mass spectrometer (ICP-MS) (Varian 810 ICP-MS System). All experiments were done in triplicate, and mean was quoted as sample values.
Results and discussion
Phylogenetic and molecular analysis
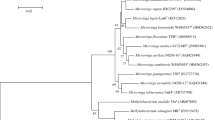
Comparison of the nearly complete 16S rRNA gene sequence (1440 nucleotides; Genbank accession number JX173993) indicated the taxonomic affiliation of strain KAs 5-22T to the genus Rhizobium, with highest sequence similarity to R. naphthalenivorans TSY03bT (98.4 %) and R. selenitireducens B1T (98.0 %) strains while similarities with all other species of Rhizobium were in the range of 93.0–97.5 %. The NJ phylogenetic analysis showed that the strain KAs 5-22T formed a coherent cluster with R. naphthalenivorans TSY03bT and R. selenitireducens B1T by a bootstrap value of 97.0 % (Fig. 1) and confirmed the affiliation of strain KAs 5-22T within the genus Rhizobium. Similar tree topology was also inferred through ML method suggesting robustness of the phylogenetic tree (Supplementary Fig. S1).
Neighbor-joining tree based on 16S rRNA gene sequences showing phylogenetic relationship between strain KAs 5-22T and its closely related phylogenetic neighbors, constructed through MEGA version 5.0. Numbers at nodes represent bootstrap values obtained with 1000 replications. The GenBank accession numbers for 16S rRNA gene sequences are shown in parentheses. Bar 0.01 indicates 1 % nucleotide substitution
The genomic G + C content of the strain KAs 5-22T was found to be 59.4 ± 1.0 %, a value that fits well within the 57–63 % GC content range reported for the members of Rhizobium genus and also with that of its closest taxonomic neighbors R. naphthalenivorans TSY03bT and R. selenitireducens B1T (Table 1). As reported by Gonzalez and Saiz-Jimenez (2005), members of the genus Rhizobium have G + C mol% that may vary up to 10 %, whereas members of the same species have G + C mol% values that are within 3–5 % of each other. DNA–DNA hybridization analysis of strain KAs 5-3T with two close taxonomic neighbors indicated 50.2 ± 1.5 % (∆T m of 6.6 °C) of relatedness with both R. selenitireducens B1T and R. naphthalenivorans TSY 03bT. In relation to other related type members, DNA level relatedness of 48.5 ± 1.0 % (∆T m of 6.8 °C), 47.0 ± 1.2 % (∆T m; 6.8 °C), and 42.0 % (∆T m 7.2 °C) was observed with R. daejeonense L61T and R. aggregatum IFAM 003T and R. rossetiformans W3T, respectively. Since the difference in ∆T m between strain KAs 5-22T and closest type strains is higher than 5 °C (maximum of 7.2 °C and minimum of 6.6 °C) and hybridization data indicated <70 % similarity at DNA level, the strain KAs 5-22T may be considered as a novel species (Wayne et al. 1987; Stackebrandt and Goebel 1994). Thus, the phylogenetic analysis and DNA–DNA relatedness (% hybridization) clearly indicated that strain KAs 5-22T belongs to the genus Rhizobium and represents a distinct species.
Morphological, biochemical, and physiological characterization
All phenotypic properties of strain KAs 5-22T and its taxonomic neighbors are presented in Table 1. Colonies of strain KAs 5-22T were observed to be circular, convex, translucent, and entire with gram stain negative, rod-shaped (1.2–1.5 µm × 0.6–0.8 µm dimensions), motile cells capable of facultative anaerobic growth (Supplementary Fig. S2). Positive responses of strain KAs 5-22T toward nitrate, citrate, β-galactosidase, N-acetyl glucosamine, l-ornithine, reduction of tetrazolium salts and inability to use glucose (fermentative), arabinose, mannose, manitol, gluconate, caprate, adipate, PAC are in line with similar results for R. naphthalenivorans TSY03bT and R. selenitireducens B1T. The strain showed its ability to utilize various D-sugars, deoxy sugars, amino sugars, sugar acids, and sugar phosphates, but inability to use di- or oligo-sugars and sugar alcohols (except glycerol) (data not shown). Along with the motile nature of the strain KAs 5-22T, its ability to grow at 28–30 °C, pH 6–8 and 1–3 % NaCl; use of glucose, NAG, with few organic acids and inability to use cellulose is in good agreement with the common traits of Rhizobium members (Young et al. 2001). Ability to utilize l-proline, serine, and reduction of methylene blue could be attributed to non-nodulating nature of the strain (Kaur et al. 2011). Strain KAs 5-22T also showed high resistance toward As, compared to its reference strains, among which none of them could be able to tolerate As concentrations higher than 0.5 mM for As3+ and 2 mM for As5+ (Table 1).
Chemotaxonomic characteristics
The predominant quinone of the strain KAs 5-22T was found to be UQ-10, along with minor percentage of UQ-8 (Table 2), which was consistent with other members of the genus Rhizobium (Tighe et al. 2000). UQ-10 was earlier reported to be the major quinone of R. naphthalenivorans TSY03bT (Kaiya et al. 2012). Among the predominant fatty acids (>5 % of the total fatty acid content) of KAs 5-22T, C10:0 2-OH (11.3 %) and C16:0 3-OH (10.25 %), were not detected in any of the closest relatives tested, while C16:0 (5.8 %) and C17:0 (6.56 %) were found to be present in others, but at relatively lower proportions. Unresolved C18:1 ɷ7C/ɷ9C found to be present at higher proportion (26.8 %) seems to be signature for Rhizobium members. Polar lipid profile of strain KAs 5-22T showed a similar composition to that of all the strains compared, particularly with respect to presence of PE, PG, and DPG, while absence of PL1, PL2 and GL1 was observed in both the closest taxonomic neighbors R. naphthalenivorans TSY03bT and R. selenitireducens B1T (Table 2).
Use of different e-donors, e-acceptors and As respiratory growth
The strain KAs 5-22T was found to be capable of using various sugar, sugar acids and hydrocarbons including both long-chain alkanes and poly aromatics (Table 3). Carbon or electron preference of the closest taxonomic type strains with that of the strain KAs 5-22T was compared (Table 3). A good agreement on both utilization and non-utilization of several of the test substrates is found between strain KAs 5-22T and its closest neighbors R. naphthalenivorans TSY03bT and R. selenitireducens B1T. With respect to metabolism of hydrocarbons, observed performance of strain KAs 5-22T is in line with R. naphthalenivorans TSY03bT. Notably, with respect to utilization of sugars except lactate and citrate none of the other carbon sources are utilized by the two closest relatives under anaerobic condition. Similar to strain KAs 5-22T, R. naphthalenivorans TSY03bT showed its ability to utilize pentadecane and nonadecane under anaerobic condition.
Anaerobically, strain KAs 5-22T was found to use As5+, Fe3+, SO4 2−, NO3 − and NO2 − as TEAs with an order of preference (As5+> NO3 − > Fe3+> SO4 2− > NO2 −). As5+ respiratory growth was further characterized with As5+ as the sole TEA and glucose or nonadecane as C and energy source, and it was found that anaerobic growth rate was low compared to that of aerobic growth. Anaerobically, strain KAs 5-22T showed a faster growth rate and higher cell yield for glucose [final cell yield (CFU/ml) of 1 × 109 (nonadecane) and 1 × 1012 (glucose)] (Fig. 2). Within 60 h, ~95 % of the added As5+ was reduced to As3+ with glucose, while at ~80 h, complete reduction was achieved with both the substrates. Addition of nitro-phenol (as respiratory inhibitor) confirmed its As5+ respiratory metabolism.
Hydrocarbon metabolizing ability of KAs 5-22T, reducing As5+ is in line with its natural habitat, i.e., alluvial aquifer enriched with geogenic As (mainly as As5+ sorbed on Fe/Mn(Al) oxides-hydroxide minerals) that has been found to contain low concentration of alkane and other hydrocarbons (Ravenscroft et al. 2001; McArthur et al. 2004; Rowland et al. 2006; Ghosh et al. 2015). Alkane metabolism by bacterial strains isolated from As-rich groundwater of West Bengal has been recently reported (Paul et al. 2014, 2015).
Functional gene-based analysis
Genetic potential of strain KAs 5-22T to metabolize hydrocarbons and respire with As5+ was evaluated by analyzing genes encoding benzyl succinate synthase (bssA-like gene) and As5+ respiratory reductase (arrA). Anaerobic oxidation of hydrocarbons proceeds through fumarate addition (as a primary reaction step) (catalyzed by benzyl succinate synthase family of proteins), followed by β-oxidation pathway (Aitken et al. 2013). Amplification of 450 bp of putative bssA-like gene (accession KX011179) and its phylogenetic analysis with similar genes (belonging to pyruvate formate lyase superfamily [like bssA, nmsA and assA]) confirmed its affiliation (Supplementary Fig. S3). Similarly, the strain’s ability to use As5+ as respiratory substrate was complemented by detection of arrA gene stretch (560 bp; accession KR340465). Similarity search showed highest identity (90 %) of this sequence with molybdopterin-binding superfamily (mopB) of membrane bound respiratory reductases. NJ tree of deduced amino acid sequence further confirmed its evolutionary closeness with the molybdopterin-binding protein of DMSO transmembrane reductase superfamily (Supplementary Fig. S4). Amplification of genes for nitrogen fixation (nifH) and nodulation (nodA) was further attempted. Presence of nifH gene could be detected only from the plasmid, while nodA could not be amplified (both genomic and plasmid DNA), indicating the strain’s nitrogen fixing non-nodulating behavior.
In conclusion, strain KAs 5-22T showed a close taxonomic resemblance with members of the genus Rhizobium, thereby providing evidence on its affiliation to the same genus. The observed distinctiveness of strain KAs 5-22T in terms of taxonomic and physiological characters clearly indicated its uniqueness, which might be attributed to strategies for competitive survival of the organism (strain KAs 5-22T) in As-rich oligotrophic aquifer environment. Based on the results obtained, it is proposed that strain KAs 5-22T represents a novel species of the genus Rhizobium, with the name Rhizobium arsenicireducens.
Description of Rhizobium arsenicireducens sp. novel
Rhizobium arsenicireducens (ar.se.ni.ci.re.du’cens. L. n. arsenicon, arsenic; L. part. adj. reducens, N.L. part. adj. arsenicireducens, arsenic-reducing, referring to the ability of the organism to reduce arsenic during its growth).
Cells are rod-shaped, gram-stain-negative, motile, facultative anaerobic with catalase and oxidase positive activity. On agar plate, colonies are small, circular with entire margins, pale creamy, and approximately 1–2 mm in diameter. Optimum growth occurs at pH 6.0–8.0 with 1 % NaCl at 28–30 °C. It shows positive reaction for citrate, N-acetyl glucosamine, glucose, urea, D-galactose, d-fructose, L-fucose, d-glucose-6-phosphate, d-fructose-6-phosphate, dextrin, trehalose, inosine, glycerol, L-rhamnose, L-galactonic acid, L-gluconic acid, L-glucuronic acid, L-lactic acid, L-malic acid, lactate, methyl pyruvate, α-ketoglutarate, α-hydroxy butyrate, acetic acid, citric acid, bromo-succinic acid, acetoacetic acid, propionic acid, N-acetyl mannosamine, and N-acetyl neuraminic acid. It can also use l-alanine, L-aspartate, d-serine but not L-glutamate, D-aspartate, l-serine, l-arginine, glycyl proline, l-histidine, pyroglutamate and amino butyric acid as preferred N source. It shows resistance to erythromycin, troleandomycin, lincomycin, aztreonam and can metabolize tetrazolium dyes. The major cellular fatty acids are C16:0, C17:0, 2-OH C10:0, 3-OH C16:0, and unresolved C18:1 ɷ7C and/or ɷ9C, and the major quinone is UQ-10. The DNA G + C content is 59.4 mol%. The polar lipids include phosphatidylethanolamine, phosphatidylglycerol, diphosphatidylglycerol, unidentified aminolipid, glycolipid, and one choline-containing lipid. Presence of genetic determinants encoding mopB of arrA, PFL domain of bssA-like gene, and nifH along with apparent absence of nodA gene confirmed further the physiological novelty of the strain. Based on biochemical, chemotaxonomic, genotypic and physiological properties, the strain is proposed to be a novel representative species of the genus Rhizobium, for which the name Rhizobium arsenicireducens sp. nov. is proposed.
The type strain KAs 5-22T (=BCCM/LMG 28795T=MTCC 12115T) is isolated as an As5+-reducing and highly As-resistant bacterium, from As-contaminated groundwater of North 24 Parganas, West Bengal, India.
References
Aitken CM, Jones DM, Maguire MJ, Gray ND, Sherry A, Bowler BFJ et al (2013) Evidence that crude oil alkane activation proceeds by different mechanisms under sulfate-reducing and methanogenic conditions. Geochim Cosmochim Acta 109:162–174
Albalasmeh SA, Berhe AA, Ghezzehei TA (2013) A new method for rapid determination of carbohydrate and total carbon concentrations using UV spectrophotometry. Carbohydr Polym 97:253–261
Berge O, Lodhi A, Brandelet G, Santaella C, Roncato MA, Christen R, Heulin T, Achouak W (2009) Rhizobium alamii sp. nov., an exopolysaccharide-producing species isolated from legume and non-legume rhizospheres. Int J Syst Evol Microbiol 59:367–372
Blum JS, Han S, Lanoil B, Saltikov C, Witte B, Tabita FR, Langley S, Beveridge TJ, Jahnke L, Oremland RS (2009) Ecophysiology of “Halarsenatibacter silvermanii” Strain SLAS-1T, gen. nov., sp. nov., a Facultative Chemoautotrophic Arsenate Respirer from Salt-Saturated Searles Lake, California. Appl Environ Microbiol 75:1950–1960
Cowan ST, Steel KJ (1965) Manual for the identification of medical bacteria. Cambridge University Press, London
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Drewniak L, Stasiuk R, Uhrynowski W, Sklodowska A (2015) Shewanella sp. O23S as a driving agent of a system utilizing dissimilatory arsenate-reducing bacteria responsible for self-cleaning of water contaminated with arsenic. Int J Mol Sci 16:14409–14427
Fan H, Su C, Wang Y, Yao J, Zhao K, Wang Y, Wang G (2008) Sedimentary arsenite-oxidizing and arsenate-reducing bacteria associated with high arsenic groundwater from Shanyin, Northwestern China. J Appl Microbiol 105:529–539
Frank B (1889) U¨ ber die Pilzsymbiose der Leguminosen. Ber Dtsch Bot Ges 7:332–346 (in German)
Ghosh D, Routh J, Dario M, Bhadury P (2015) Elemental and biomarker characteristics in a Pleistocene aquifer vulnerable to arsenic contamination in the Bengal Delta Plain, India. Appl Geochem 61:87–98
Gonzalez JM, Saiz-Jimenez C (2005) A simple fluorimetric method for the estimation of DNA–DNA relatedness between closely related microorganisms by thermal denaturation temperatures. Extremophiles 9:75–79
Gu T, Sun LN, Zhang J, Sui XH, Li SP (2014) Rhizobium flavum sp. nov., a triazophos-degrading bacterium isolated from soil under the long-term application of triazophos. Int J Syst Evol Microbiol 64:2017–2022
Hiraishi A, Ueda Y, Ishihara J, Mori T (1996) Comparative lipo-quinone analysis of influent sewage and activated sludge by high performance liquid chromatography and photodiode array detection. J Gen Appl Microbiol 42:457–469
Hunter WJ, Kuykendall LD, Manter DK (2007) Rhizobium selenireducens sp. nov., a selenite-reducing α Proteobacteria isolated from a bioreactor. Curr Microbiol 55:455–460
Kaiya S, Rubaba O, Yoshida N, Yamada T, Hiraishi A (2012) Characterization of Rhizobium naphthalenivorans sp. nov. with special emphasis on aromatic compound degradation and multilocus sequence analysis of housekeeping genes. J Gen Appl Microbiol 58:211–224
Kaur J, Verma M, Lal R (2011) Rhizobium rosettiformans sp. nov., isolated from a hexachlorocyclohexane dump site, and reclassification of Blastobacter aggregatus Hirsch and Muller 1986 as Rhizobium aggregatum comb. nov. Int J Syst Evol Microbiol 61:1218–1225
Kazy SK, Sar P, Asthana RK, Singh SP (1999) Copper uptake and its compartmentalization in Pseudomonas aeruginosa strains: chemical nature of cellular metal. W J Microbiol Biotechnol 15:599–605
Kelly A, Fulton M (1953) Use of triphenyl tetrazolium in motility test medium. Am J Clin Pathol 23:512
Kodaka H, Armfield AY, Lombard GL, Dowell VR (1982) Practical procedure for demonstrating bacterial flagella. J Clin Microbiol 16:948–952
Komagata K, Suzuki K (1987) Lipid and cell wall analysis in bacterial systematics. Methods Microbiol 19:161–206
Kudo K, Yamaguchi N, Makino T, Ohtsuka T, Kimura K, Dong DT, Amachi S (2013) Release of arsenic from soil by a novel dissimilatory arsenate-reducing bacterium, Anaeromyxobacter sp. strain PSR-1. Appl Environ Microbiol 79:463–468
Kuykendall LD, Roy MA, O’Neill JJ, Devine TE (1988) Fatty acids, antibiotic resistance, and deoxyribonucleic acid homology groups of Bradirhizobium japonicum. Int J Syst Bacteriol 38:358–361
Lear G, Polya DA, Song B, Gault AG, Lloyd JR (2007) Molecular analysis of arsenate-reducing bacteria within cambodian sediments following amendment with acetate. Appl Environ Microbiol 73:1041–1048
Liu Y, Wang RP, Ren C, Lai QL, Zeng RY (2015) Rhizobium marinum sp. nov., a malachite green tolerant bacterium isolated from sea water. Int J Syst Evol Microbiol 65:4449–4454
Malasarn D, Keeffe HR, Newman DK (2008) Characterization of the arsenate respiratory reductase from Shewanella sp. strain ANA-3. J Bacteriol 190:135–142
McArthur JM, Banerjee DM, Hudson-Edwards KA, Mishra R, Purohit R (2004) Natural organic matter in sedimentary basins and its relation to arsenic in anoxic ground water: the example of West Bengal and its worldwide implications. Appl Geochem 19:1255–1293
Miller LT (1982) A single derivatization method for bacterial fatty acid methyl esters including hydroxy acids. J Clin Microbiol 16:584–586
Mnasri B, Mrabet M, Laguerre G, Aouani ME, Mhamdi R (2007) Salt-tolerant rhizobia isolated from a Tunisian oasis that are highly effective for symbiotic N -fixation with Phaseolus vulgaris constitute a novel biovar (bv. mediterranense) of Sinorhizobium meliloti. Arch Microbiol 187:79–85
Newman DK, Ahmann D, Morel FMM (1998) A brief review of microbial arsenate respiration. Geomicrobiol J 15:255–268
Ohtsuka T, Yamaguchi N, Makino T, Sakurai K, Kimura K, Kudo K, Homma E, Dong DT, Amachi S (2013) Arsenic dissolution from Japanese paddy soil by a dissimilatory arsenate-reducing bacterium Geobacter sp. OR-1. Environ Sci Technol 47:6263–6271
Osborne TH, Mc Arthur JM, Sikdar PK, Santini JM (2015) Isolation of an arsenate-respiring bacterium from a redox front in an arsenic-polluted aquifer in West Bengal, Bengal Basin. Environ Sci Technol 49:4193–4199
Paul D, Poddar S, Sar P (2014) Characterization of arsenite-oxidizing bacteria isolated from arsenic-contaminated groundwater of West Bengal. J Environ Sci Heal A 49:1481–1492
Paul D, Kazy SK, Gupta AK, Pal T, Sar P (2015) Diversity, metabolic properties and arsenic mobilization potential of indigenous bacteria in arsenic contaminated groundwater of West Bengal, India. PLoS ONE 10:1–40
Quan ZX, Bae HS, Baek JH, Chen WF, Im WT, Lee ST (2005) Rhizobium daejeonense sp. nov. isolated from a cyanide treatment bioreactor. Int J Syst Evol Microbiol 55:2543–2549
Ramana CV, Parag B, Girija KR, Ram BR, Ramana VV, Sasikala C (2013) Rhizobium subbaraonis sp. nov., an endolithic bacterium isolated from beach sand. Int J Syst Evol Microbiol 63:581–585
Ravenscroft P, McArthur JM, Hoque BA (2001) Geochemical and palaeohydrological controls on pollution of groundwater by arsenic. In: Chappell WR, Abernathy CO, Calderon R (eds) Arsenic exposure and health effects IV. Elsevier Science, Oxford, pp 53–78
Rosselló-Mora R, Amann R (2001) The species concept for prokaryotes. FEMS Microbiol Rev 25:39–67
Rowland HAL, Polya DA, Lloyd JR, Pancost RD (2006) Characterisation of organic matter in a shallow, reducing, arsenic-rich aquifer, West Bengal. Org Geochem 37:1101–1114
Saltikov CW, Newman DK (2003) Genetic identification of a respiratory arsenate reductase. Proc Natl Acad Sci USA 100:10983–10988
Saltikov CW, Cifuentes A, Venkateswaran K, Newman DK (2003) The ars Detoxification System Is Advantageous but Not Required for As(V) Respiration by the Genetically Tractable Shewanella Species Strain ANA-3. Appl Environ Microbiol 69:2800–2809
Sambrook J, Russel DW (2001) Rapid isolation of yeast DNA. In: Sambrook J, Russel DW (eds) Molecular cloning, a laboratory manual. Cold Spring Harbor Laboratory, New York, pp 631–632
Sarkar A, Kazy SK, Sar P (2013) Characterization of arsenic resistant bacteria from arsenic rich groundwater of West Bengal, India. Ecotoxicology 22:363–376
Sarkar A, Kazy SK, Sar P (2014) Studies on arsenic transforming groundwater bacteria and their role in arsenic release from subsurface sediment. Environ Sci Pollut Res 21:8645–8866
Sheu SY, Huang HW, Young CC, Chen WM (2015) Rhizobium alvei sp. nov., isolated from a freshwater river. In J Syst Evol Microbiol 65:472–478
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington DC, pp 607–654
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA–DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Tighe SW, de Lajudie P, Dipietro K, Lindstro MK, Nick G, Jarvis BD (2000) Analysis of cellular fatty acids and phenotypic relationships of Agrobacterium, Bradyrhizobium, Mesorhizobium, Rhizobium and Sinorhizobium species using the Sherlock Microbial Identification System. Int J Syst Evol Microbiol 50:787–801
Viteri SE, Schmidt EL (1987) Ecology of indigenous soil rhizobia: response of Bradyrhizobium japonicum to readily available substrates. Appl Environ Microbiol 53:1872–1875
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE et al (1987) International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Young JM, Kuykendall ID, Martínez-Romero E, Kerr A, Sawada HA et al (2001) A revision of Rhizobium Frank 1889, with an emended description of the genus, and the inclusion of all species of Agrobacterium Conn 1942 and Allorhizobium undicola de Lajudie et al. 1998 as new combinations: Rhizobium radiobacter, R. rhizogenes, R. rubi, R. undicola and R. vitis. Int J Syst Evol Microbiol 51:89–103
Zhang GX, Ren SZ, Xu MY, Zeng GQ, Luo HD, Chen JL, Tan ZY, Sun GP (2011) Rhizobium borbori sp. nov., aniline-degrading bacteria isolated from activated sludge. Int J Syst Evol Microbiol 61:816–822
Zhang X, Li B, Wang H, Sui X, Ma X, Hong Q, Jiang R (2012) Rhizobium petrolearium sp. nov., isolated from oil-contaminated soil. Int J Syst Evol Microbiol 62:1871–1876
Zhu YG, Yoshinaga M, Zhao FJ, Rosen BP (2014) Earth abides arsenic biotransformations. Annu Rev Earth Planet Sci 42:443–467
Acknowledgments
The work is financially supported by the grant from Council of Scientific and Industrial Research (CSIR), Govt. of India, project number 38/1314/11/EMR II, and the fellowship to BM is provided by INSPIRE fellowship scheme of Department of Science and Technology (DST), Govt. of India, fellowship number IF120832. Authors are thankful for the kind help of R. Lal (Professor, University of Delhi, North Campus, New Delhi, India) and D.K. Newman (Professor, California Institute of Technology, Pasadena, U.S.A), for providing the type strains. The authors express gratitude to S. Marqués (Professor, Consejo Superior de Investigaciones Científicas, Estación Experimental del Zaidín, Department of Environmental Protection, Granada, Spain) and H. S. Gehlot (Professor, Tejpur University, India) for providing the primers of bssA-like gene and nodA gene, respectively. We also acknowledge Prof A. Oren and Prof A. C. Parte for suggesting species epithet and etymology of the strain. The GenBank accession numbers for 16S rRNA, molybdopterin-binding site of As5+ respiratory reductase (arrA) and putative benzyl succinate synthase (bssA like) gene are JX173993, KR340465, and KX011179, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Erko Stackebrandt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mohapatra, B., Sarkar, A., Joshi, S. et al. An arsenate-reducing and alkane-metabolizing novel bacterium, Rhizobium arsenicireducens sp. nov., isolated from arsenic-rich groundwater. Arch Microbiol 199, 191–201 (2017). https://doi.org/10.1007/s00203-016-1286-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-016-1286-5