Abstract
Chromobacterium violaceum is a free-living bacterium that inhabits low-nutrient environments such as the Amazon basin. Bacteria respond to phosphate (Pi) shortage by expressing a range of genes involved in Pi uptake and assimilation, known as the PHO regulon. Several PHO regulon genes have been annotated in the genome of C. violaceum. Here we show that C. violaceum is extremely well adapted to low-Pi conditions. Remarkably, this bacterium is able to grow in media containing only traces of Pi. The PHO regulon genes are induced upon Pi depletion, but the bacteria continued to grow under these conditions. Unlike other Proteobacteria hitherto analyzed, neither PstS nor PhoU play a role in the repression of the PHO regulon under Pi excess.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Free-living organisms frequently face shortage of macronutrients in natural environments, which are often poor in basic resources, such as carbon, nitrogen and phosphorus (Smil 2000). Phosphorus occurs in nature in its maximal oxidized state (PO4 3− or Pi), which is also the preferred phosphorus source. Once Pi is taken up by bacteria, it is directly incorporated into phosphorylated monomers, primarily as ATP (Wanner and Wilmes-Riesenberg 1992). Given the importance of Pi and the irregular availability of this nutrient in natural habitats, bacteria must be able to sense changes in Pi concentration in the environment and to modify the pattern of gene expression accordingly. When the external concentration of Pi is below a minimum threshold that supports bacterial growth (ca. 4 μM), a set of genes involved in scavenging alternative sources of this nutrient, the PHO regulon, is activated (Wanner F1996). The most well-studied PHO genes are phoA, the pst operon and the phoBR operon that, respectively, encode the periplasmic enzyme alkaline phosphatase, the high-affinity Pi-transport system Pst and the two-component system that controls PHO expression.
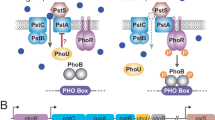
The histidine kinase PhoR autophosphorylates in response to low Pi in the medium and transfers the Pi moiety to PhoB, which in turn binds to the PHO-boxes, that are specific sequences in the regulatory regions of all PHO genes in place of the standard −35 sequence. PhoB then interacts with the RNA polymerase associated with σ70, thus initiating transcription (Makino et al. 1989). Under Pi excess, PhoR dephosphorylates PhoB, thereby repressing the expression of the PHO genes (Carmany et al. 2003).
The Pst transporter is a typical ATP-binding cassette system (ABC) comprised of four proteins: PstS or PiBP (Pi-binding protein), a periplasmic protein whose function is to scavenge Pi molecules in the periplasm and to present them to the channel formed by the integral proteins PstC and PstA. PstB is an ATPase that energizes the transport process (Surin et al. 1985). The Pst system of most Proteobacteria is encoded by an operon composed of five genes—pstS, pstC, pstA, pstB and phoU, transcribed as a single polycistronic mRNA (Aguena et al. 2002, 2009). The most distal gene of the operon, phoU, encodes a protein that does not participate in Pi transport and which exact function is unknown (Steed and Wanner 1993). A complicating factor in the regulation of PHO is that the five proteins encoded by the pst operon act as repressors of PHO transcription when Pi is in excess. Null mutations in phoU or in any other pst gene cause the constitutive expression of the PHO regulon irrespectively of Pi concentration in the medium (Wanner 1996). The mechanism through which the Pst proteins repress PHO expression is unclear, but it has recently been shown that under Pi-excess conditions PhoU physically interacts with PstB and PhoR (Gardner et al. 2014).
Chromobacterium violaceum is a free-living β-proteobacterium that inhabits the soil and water of tropical environments and that thrives in the acidic and low-nutrient waters of the Rio Negro in the Amazon basin. A proteomic analysis revealed that 3 % of C. violaceum proteins are involved in inorganic ion transport and metabolism (Ciprandi et al. 2013). Several putative PHO regulon genes have been annotated in the genome of C. violaceum (Consortium 2003). In the present study, we investigated the growth of C. violaceum under Pi limitation, its ability to take up Pi and the pattern of induction of the PHO regulon. We show that this bacterium is well adapted to conditions of Pi scarcity.
Methods
Strains, plasmids and oligomers
The strains, plasmids and oligomers used in this study are described in Table 1.
Culture media and growth conditions
C. violaceum and E. coli cells were routinely cultured at 37 °C in lysogenic broth medium (LB) and L-agar (Miller 1992). Medium A (0.12 M Tris, 80 mM NaCl, 20 mM KCl, 20 mM NH4Cl, 3 mM Na2SO4, 1 mM MgCl2, 0.2 mM CaCl2, 2 μM ZnCl2, 0.5 % glucose, 0.5 % bacto-peptone, pH 7.5) is a semi-rich Pi-limited medium (Spira et al. 2010). MM medium is modified MOPS minimal medium (Neidhardt et al. 1974) (9.52 mM NH4Cl; 0.28 mM K2SO4; 0.009 mM CaCl2; 0.52 mM MgCl2; 50 mM NaCl; 0.01 mM FeSO4; 40 mM MOPS; 3 × 10−6 mM (NH4)6(Mo7)24; 4 × 10−4 mM H3BO3; 3 × 10−5 mM CoCl2; 10−5 mM CuSO4; 8 × 10−5 MnCl2; 10−5 mM ZnSO4) supplemented with 0.2 % glucose and variable Pi concentrations. When required, C. violaceum cultures were supplemented with kanamycin (25 μg/ml), tetracycline (5 μg/ml) or ampicillin (100 μg/ml). Bacterial growth rate (μ) was calculated according to the formula: μ = ln(N/N 0)/(T), where N and N 0, respectively, correspond to two points at the exponential growth phase and t is the time course of the growth curve.
Detection of PHO-boxes in silico
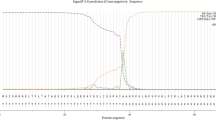
The upstream regions of putative PHO regulon genes of C. violaceum were scanned for PHO boxes using the consensus sequence 5′-CTGTCATAAATCTGTCAT with the Fuzznuc program from the EMBOSS software package (Rice et al. 2000); up to nine mismatches were allowed. Scores were attributed to the sequence matches based on their similarity to the PHO-box consensus as described by Yuan et al. (2006).
Alkaline phosphatase (AP) assay
Alkaline phosphatase quantitative assays were performed as described (Galbiati et al. 2014). The specific activity of AP was calculated according to the formula: A 410 min−1 OD −1660 , where A 410 is the absorbance of enzymatic products after removal of the cells by centrifugation, min is the reaction time and OD660 is the turbidity of the bacterial culture.
β-Galactosidase assay
β-Galactosidase assays were performed as described (Miller 1992), except that the cell density was measured at 660 nm. The specific activity was calculated according to the formula: A420 min−1 OD −1660 1000. The variables are as described above for the AP assay.
Pi uptake
Pi-uptake assays were performed as described by de Almeida et al. with some modifications (de Almeida et al. 2015). Briefly, bacteria were cultivated in MM medium containing 10 mM Pi (KH2PO4) for 48 h at 37 °C, centrifuged at 5000×g for 10 min, washed twice, suspended in Pi-free medium at an OD660 of 0.1 and grown for 6 h at 37 °C to induce phosphate starvation. Pi transport was assayed by adding a mix of 0.5 mM K2HPO4 + 10 μCi [32P] (IPEN, São Paulo, Brazil). One hundred microliter samples were withdrawn every 30 s, applied to nitrocellulose membrane disks (Millipore, MA) on a manifold and immediately washed with PBS (137 mM NaCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, 2.7 mM KCl, pH 7.4.). The filters were transferred to vials containing 5-ml scintillation cocktail (Perkin-Elmer, Waltham, MA) and read in a scintillation counter (Beckman Coulter, Fullerton, CA).
Microdetermination of Pi
Pi concentration in the culture medium was measured as described (de Almeida et al. 2015). The supernatants of the bacterial cultures were collected, diluted ten times and mixed with the same volume of the working reagent (1 volume of 167 mM H2SO4, 1 volume of 2.5 % (NH4)2MoO4, 1 volume of 10 % ascorbic acid and 2 volumes of deionized water) and incubated at 37 °C for 2 h. Following the incubation period, samples were measured in a spectrophotometer at 820 nm. Calibration solutions were prepared with the following concentrations of NaH2PO4: 0.16, 0.08, 0.04, 0.02 and 0.01 mM.
Construction of transcriptional fusions
lacZ transcriptional fusions were constructed in plasmid pRKlacZ290. DNA fragments encompassing the promoter regions of phoB and phoU were amplified by PCR using, respectively, the phoB-prom and phoU-prom pairs of primers and cloned in plasmid pTZR57 (Fermentas-Thermo) according to the manufacturer’s instructions. The resulting plasmids containing PphoU or PphoB DNA sequences were digested with EcoRI or EcoRI–HindIII, respectively, and ligated to plasmid pRKlacZ290 digested with the same enzymes resulting in plasmids pFN13 and pFN15. Cloning was confirmed by DNA sequencing using the lacZ reverse primer. Plasmids pFN13 and pFN15 were transformed into E. coli S17-1 and mobilized to C. violaceum.
Construction of the pstS and phoU knockout mutants
Knockout mutants of pstS and phoU were obtained by inserting the suicide plasmid pKNOCK-Km into C. violaceum chromosome. DNA fragments of 461 and 470 bp corresponding, respectively, to internal regions of phoU and pstS ORFs were amplified by PCR using phoU-int or pstS-int sets of primers. The amplicons were cloned in vector pTZ57R/T. The phoU fragment was digested with XbaI and HindIII, while the pstS fragment was digested only with PstI. Both phoU and pstS fragments were subcloned in the corresponding sites of pKNOCK-Km. The resulting plasmids were transformed into E. coli S17-1 and mobilized by conjugation to C. violaceum, where they underwent insertion in the bacterial chromosome. Bacteria were selected for resistance to Kanamycin. The phoU and pstS insertions in C. violaceum chromosome were confirmed by PCR using phoU-ext or pstS-ext sets of primers.
Genetic transfer and molecular biology techniques
Standard bacterial genetics and molecular biology procedures, such as electroporation, conjugation, DNA restriction and cloning, were performed as described (Miller 1992; Sambrook and Russell 2001).
Results and discussion
C. violaceum growth under low-Pi conditions and induction of the PHO regulon
C. violaceum thrives in Pi-deficient habitats, such as the Rio Negro basin (McClain et al. 2001; Hungria et al. 2005). To investigate how C. violaceum responds to Pi availability, we monitored the growth of bacterial cultures in minimal medium supplemented with a range of Pi concentrations and compared it to the growth of the prototrophic E. coli strain MG1655 (Fig. 1). C. violaceum grew better than E. coli under all tested Pi concentrations. Even when no Pi was added C. violaceum managed to present some growth, possibly by using Pi contaminants as a P source and/or by mobilizing internal Pi stores. C. violaceum also formed colonies of normal size (>1.5 mm) when grown in minimal medium containing Pi concentrations ranging from 1 μM to 10 mM, while E. coli did not grow at all in 1 μM Pi medium (not shown). These results suggest that C. violaceum is well adapted to the conditions of Pi scarcity, where even very versatile bacterial species, such as E. coli, are unable to grow.
Growth of E. coli and C. violaceum under different Pi concentrations. E. coli K-12 strain MG1655 (dark gray bars) and C. violaceum (light gray bars) are grown for 24 h in MOPS minimal medium supplemented with 0, 0.001, 0.01, 0.1 or 1 mM KH2PO4. Culture turbidity is measured at OD660. Bars represent the mean ± SEM of three independent cultures
To correlate bacterial growth with the induction of the PHO regulon, a growth curve in medium A, a semi-rich medium with limited Pi concentration (~0.1 mM) used to induce Pi starvation and to study the expression of the PHO genes (Spira and Yagil 1999), was conducted (Fig. 2a). Interestingly, bacteria inoculated in medium A and in medium A supplemented with 1 mM Pi showed nearly identical growth curves. During the first 5 h, the growth rate of both cultures was about 0.4 h−1 and after that decreased to 0.15 h−1. The similarity in the growth pattern of the cultures growing under Pi excess or starvation suggests that C. violaceum can grow equally well under high- and low-Pi concentrations and that the reduction in growth rate after 5 h was caused by factors unrelated to Pi concentration in the medium. However, only in the low-Pi medium AP was induced, evidencing the onset of the Pi-starvation phase. This indicates that, unlike E. coli, where AP induction occurs concomitantly with growth arrest (Spira et al. 1995), the PHO regulon of C. violaceum was activated, while bacteria were still growing, albeit at a lower rate. It is worth noticing that the continued increase in OD660 under Pi starvation was not caused by enlargement of bacterial cells or by any other phenomenon that might have artificially increased the turbidity of the culture, but was due to bacterial replication (Fig. S1).
Induction of AP and Pi consumption by wild-type C. violaceum cells. a Bacteria grown overnight are resuspended in non-supplemented medium A (~0.1 mM Pi) or in medium A supplemented with 1 mM Pi and grown for another 11 h. Growth (OD660) (dashed lines) and AP activity (solid lines) are followed hourly. (Square) non-supplemented medium A; (circle) medium A supplemented with 1 mM Pi. b Bacteria suspended in non-supplemented medium A are assayed for Pi consumption. Samples are taken hourly to assay Pi concentration in the medium and cell turbidity (inset). Each point corresponds to the mean ± SEM of three independent experiments
In another experiment, the consumption of Pi of C. violaceum growing in non-supplemented medium A (~0.1 mM Pi) was assessed (Fig. 2b). Bacteria growing in this medium consumed all available Pi in 5 h. During this period, the bacterial population doubled twice (from 0.16 to 0.68 OD660 units). After 4 h, the growth rate dropped from 0.41 to 0.21 h−1, but the population kept growing. The reduction in growth rate in both Pi-starved and non-starved cultures was likely to be caused by scarcity of another macronutrient. The fact that cells in the Pi-starved culture continued to grow irrespectively of Pi concentration in the medium suggests that C. violaceum can mobilize internal Pi reservoirs to support growth.
The PHO regulon of C. violaceum
Twenty-three PHO regulon genes, such as phoB-phoR, the pstSCAB and ugpBAEC operons, the phn gene cluster, phoA and phoU, were annotated in the genome of C. violaceum. Sequences compatible with the PHO-box consensus (5′-CTGTCATAAATCTGTCAT) were found upstream of phoU, phnG, ugpB, phoB and pstS ORFs (Table 2). The sequences immediately upstream to phoB, phoU and ugpB showed a high degree of similarity to the PHO-box consensus. The PHO-box upstream to pstS displayed the lowest score.
Expression of phoB-lacZ and phoU-lacZ
Transcriptional fusions between the putative promoters of two PHO genes—phoB and phoU with lacZ, were constructed and used to test the pattern of expression of these genes under different growth conditions. Plasmids pFN15 (PphoB-lacZ) and pFN13 (PphoU-lacZ) were transferred by conjugation to C. violaceum. The exconjugants were grown in medium A and sampled hourly for bacterial growth, β-galactosidase and AP activity (Fig. 3). Both cultures entered the Pi-starvation phase after 4 h of growth in medium A, as shown by the sharp increase in AP activity. Bacteria continued growing even after AP induction, as observed previously (Fig. 2). Around this time point, β-galactosidase activity of both phoB-lacZ and phoU-lacZ also started to increase. phoU-lacZ and phoB-lacZ activities increased eightfold (from 260 to 2200 Miller units) and threefold (from 4000 to 12,000 Miller units), respectively, throughout the course of the experiment. The β-gal activity of the phoB-lacZ fusion was from the start considerably higher than that of phoU-lacZ, even though both promoters carry PHO boxes with similar scores (Table 2). Similarly, the phoB gene of E. coli also showed a higher basal level and lower level of induction than other PHO genes (Shinagawa et al. 1983). Overall, these results demonstrate that both phoB and phoU respond to Pi starvation, as expected for genes that belong to the PHO regulon.
Expression of phoB and phoU under Pi starvation. Wild-type C. violaceum cells carrying either phoB-lacZ (circle) or phoU-lacZ (square) fusions are grown in non-supplemented medium A and assayed for a β-gal activity and b growth and AP activity (inset). Each point represents the mean ± SEM of three independent experiments
phoU and pstS are not involved in PHO repression
In most Proteobacteria hitherto analyzed phoU is the most distal gene of the pst operon. In contrast, in C. violaceum phoU forms a separate transcriptional unit together with rpiA, ppx and corA, that, respectively, encode a ribose 5-phosphate isomerase, an exopolyphosphatase and a protein involved in magnesium/cobalt transport. This genomic organization can also be found in other β-Proteobacteria. An analysis with the String program (Franceschini et al. 2013) showed that 12 out of 72 β-Proteobacteria species with sequenced genomes have the phoU gene separated from the pst operon, while only two out of 117 alpha and one out of 238 γ-Proteobacteria display this genomic arrangement. All δ and ε Proteobacteria carry phoU integrated in the pst operon. This suggests that phoU segregation from the pst operon is a relatively recent event in the evolution of C. violaceum, which may have been selected in response to severe low-Pi conditions.
phoU encodes a protein that in E. coli and in other Proteobacteria acts as a repressor of the PHO regulon under Pi-excess conditions (Wanner 1996). In these bacteria, null mutations in phoU result in the constitutive expression of the PHO regulon. To analyze the role of PhoU in PHO repression in C. violaceum, a phoU knockout was constructed. The phoU mutant and the wild-type strain were grown in medium A and assayed for AP activity (Fig. 4). Under Pi-limiting conditions, AP was induced in both wild-type and phoU mutant strains, while under Pi excess the level of AP was significantly lower (repressed) in both strains. This suggests that C. violaceum’s PhoU is not involved in PHO repression. In contrast, the phoU mutant of E. coli (Muda et al. 1992; Steed and Wanner 1993) and P. aeruginosa (de Almeida et al. 2015) express AP constitutively even under Pi-excess conditions. The exact function of PhoU in E. coli and in other bacteria is unknown, but it is well established that PhoU plays a role in PHO repression (Wanner 1996; Gardner et al. 2014). Interestingly, the firmicute B. subtilis does not carry a copy of phoU and its Pst system is not involved in PHO regulation (Qi et al. 1997). In contrast, C. violaceum does carry phoU, which is inducible by Pi starvation, but it is not located in the pst operon. This different genomic arrangement suggests that PhoU plays a different and still unknown role in this bacterium.
The role of pstS, the most proximal gene of the pst operon, on PHO repression was also analyzed. Similarly to what has been observed with the phoU mutant, the AP activity of the pstS mutant was repressed under Pi-excess conditions, suggesting that pstS also does not participate in PHO regulation. It is interesting to note that induction of AP by Pi starvation (the ratio of AP activity in the Pi-starved and Pi-excess cultures) was significantly lower (p = 0.045) in the pstS mutant (AP induction = 2.5) than in the wild-type strain (AP induction = 7.0). On the other hand, the AP activities of the Pi-starved wild-type strain and phoU mutant were not statistically different (p = 0.20), implying that AP induction in the phoU mutant was not significantly lower than in the wild-type strain.
In the majority of bacterial species hitherto analyses—E. coli (Wanner 1996), P. aeruginosa (Nikata et al. 1996), S. meliloti (Geiger et al. 1999), P. mirabilis (Jacobsen et al. 2008), P. putida (Wu et al. 1999) and Synechocystis sp. (Burut-Archanai et al. 2009), the five pst genes play a repressive role in the expression of the PHO regulon under Pi-excess conditions. The Pst system of C. violaceum might have lost the ability to repress the PHO regulon concomitantly with the rearrangement of the phoU locus as a result of bacterial adaptation to poor Pi environments, where the lack of a PHO repressor might have conferred a selective advantage. In E. coli, the signal of Pi availability passes through the Pst system to PhoR (Gardner et al. 2014), as the PhoR protein of E. coli does not present a large periplasmic domain that senses the concentration of Pi (Scholten and Tommassen 1993). In contrast, according to an in silico analysis with the program Cello (Yu et al. 2006) C. violaceum’s PhoR appears to have a larger periplasmic domain and might, therefore, receive the signal of Pi availability directly from the environment. In bacteria such as E. coli that colonize habitats with different levels of Pi concentration, such as the intestines of mammals and birds (relatively high Pi) and water bodies and soil (low Pi), repression by the Pi-transport system is required to avoid overconsumption of Pi that can be toxic. On the other hand, in C. violaceum and other bacterial species that usually inhabits Pi-deficient places and rarely face abundance of Pi, an extra level of PHO repression by Pst would be unnecessary. In fact, it has recently been suggested that PhoU does not repress the PHO regulon of the oligotrophic α-Proteobacterium Caulobacter crescentus that similarly to C. violaceum dwells in low-nutrient environments (Lubin et al. 2015). More broadly, bacteria may adapt to low-Pi conditions in at least three different ways: (1) by selection of a very efficient high-affinity Pi-uptake system; or (2) an efficient Pi storage system, which could be readily used to promote growth under severe Pi starvation; or (3) by incorporating genetic modifications in the regulatory system that controls the PHO regulon. The latter being more common as evolutionary jumps often involve altered patterns of gene regulation (Wang et al. 2010).
pstS is essential for Pi uptake
PstS is an integral part of the Pst transport system acting as a periplasmic Pi-binding protein. Null mutations in pstS abolish Pi transport altogether (Wanner 1996; Luz et al. 2012; Qi et al. 1997; Braibant et al. 1996; Poole and Hancock 1984; Diaz et al. 2005). To test whether C. violaceum’s pstS participates in Pi transport, a Pi-uptake assay was conducted (Fig. 5). Pi starvation was induced in the wild-type strain and in the pstS mutant by growing the bacteria in MOPS minimal medium lacking Pi. The bacteria were then exposed to the addition of 0.5 mM 32Pi, and samples were withdrawn every 30 s. The wild-type bacteria took up Pi at a Vmax of 9.1 nmol Pi OD −1660 min−1, while the pstS mutant rate of Pi uptake was ten times lower (0.89 nmole Pi OD −1660 min−1). It can be concluded that pstS plays a fundamental role in Pi uptake in C. violaceum.
In summary, C. violaceum is well adapted to low-Pi conditions. It is able to grow with traces of Pi in the medium and does not stop growing upon Pi depletion. The PHO genes, phoA, pstS and phoB, were found to respond to Pi limitation, but unlike most Proteobacteria, pstS and phoU do not play a role in PHO repression under Pi excess.
References
Aguena M, Yagil E, Spira B (2002) Transcriptional analysis of the pst operon of Escherichia coli. Mol Genet Genomics 268(4):518–524
Aguena M, Ferreira GM, Spira B (2009) Stability of the pstS transcript of Escherichia coli. Arch Microbiol 191(2):105–112
Alexeyev MF (1999) The pKNOCK series of broad-host-range mobilizable suicide vectors for gene knockout and targeted DNA insertion into the chromosome of gram-negative bacteria. Biotechniques 26(5):824–826
Braibant M, Lefèvre P, de Wit L, Peirs P, Ooms J, Huygen K, Andersen AB, Content JA (1996) Mycobacterium tuberculosis gene cluster encoding proteins of a phosphate transporter homologous to the Escherichia coli Pst system. Gene 176(1–2):171–176
Burut-Archanai S, Incharoensakdi A, Eaton-Rye JJ (2009) The extended n-terminal region of SphS is required for detection of external phosphate levels in Synechocystis sp. Pcc 6803. Biochem Biophys Res Commun 378(3):383–388
Carmany DO, Hollingsworth K, McCleary WR (2003) Genetic and biochemical studies of phosphatase activity of PhoR. J Bacteriol 185(3):1112–1115
Ciprandi A, da Silva WM, Santos AV, de Castro Pimenta AM, Carepo MSP, Schneider MPC, Azevedo V, Silva A (2013) Chromobacterium violaceum: important insights for virulence and biotechnological potential by exoproteomic studies. Curr Microbiol 67(1):100–106
Consortium BNGP (2003) The complete genome sequence of Chromobacterium violaceum reveals remarkable and exploitable bacterial adaptability. Proc Natl Acad Sci USA 100(20):11660–11665
de Almeida LG, Ortiz JH, Schneider RP, Spira B (2015) phoU inactivation in Pseudomonas aeruginosa enhances accumulation of ppGpp and polyphosphate. Appl Environ Microbiol 81(9):3006–3015
Diaz M, Esteban A, Fernandez-Abalos JM, Santamaria RI (2005) The high-affinity phosphate-binding protein PstS is accumulated under high fructose concentrations and mutation of the corresponding gene affects differentiation in Streptomyces lividans. Microbiology 151(Pt 8):2583–2592
Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C, Jensen LJ (2013) String v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res 41(Database issue):D808–D815
Galbiati HF, Taschner NP, Spira B (2014) The effect of the rpoSam allele on gene expression and stress resistance in Escherichia coli. Arch Microbiol 196(8):589–600
Gardner SG, Johns KD, Tanner R, McCleary WR (2014) The PhoU protein from Escherichia coli interacts with PhoR, PstB, and metals to form a phosphate-signaling complex at the membrane. J Bacteriol 196(9):1741–1752
Geiger O, Röhrs V, Weissenmayer B, Finan TM, Thomas-Oates JE (1999) The regulator gene phoB mediates phosphate stress-controlled synthesis of the membrane lipid diacylglyceryl-n, n, n-trimethylhomoserine in Rhizobium (Sinorhizobium) meliloti. Mol Microbiol 32(1):63–73
Gober JW, Shapiro L (1992) A developmentally regulated Caulobacter flagellar promoter is activated by 3′ enhancer and IHF binding elements. Mol Biol Cell 3(8):913–926
Hungria M, Astolfi-Filho S, Chueire LMO, Nicolás MF, Santos EBP, Bulbol MR, Souza-Filho A, Nogueira Assunção E, Germano MG, Vasconcelos ATR (2005) Genetic characterization of Chromobacterium isolates from black water environments in the Brazilian Amazon. Lett Appl Microbiol 41(1):17–23
Jacobsen SM, Lane MC, Harro JM, Shirtliff ME, Mobley HL (2008) The high-affinity phosphate transporter Pst is a virulence factor for Proteus mirabilis during complicated urinary tract infection. FEMS Immunol Med Microbiol 52(2):180–193
Lubin EA, Henry JT, Fiebig A, Crosson S, Laub MT (2015) Identification of the PhoB regulon and role of PhoU in the phosphate-starvation response of Caulobacter crescentus. J Bacteriol 198(1):187–200
Luz DE, Nepomuceno RSL, Spira B, Ferreira RCC (2012) The Pst system of Streptococcus mutans is important for phosphate transport and adhesion to abiotic surfaces. Mol Oral Microbiol 27(3):172–181
Makino K, Shinagawa H, Amemura M, Kawamoto T, Yamada M, Nakata A (1989) Signal transduction in the phosphate regulon of Escherichia coli involves phosphotransfer between PhoR and PhoB proteins. J Mol Biol 210(3):551–559
McClain ME, Victoria RL, Richey JE (2001) The biogeochemistry of the Amazon basin. Oxford University Press, New York
Miller JH (1992) A short course in bacterial genetics: a laboratory manual and handbook for Escherichia coli and related bacteria. Cold Spring Harbor Laboratory, Cold Spring Harbor
Muda M, Rao NN, Torriani A (1992) Role of PhoU in phosphate transport and alkaline phosphatase regulation. J Bacteriol 174(24):8057–8064
Neidhardt FC, Bloch PL, Smith DF (1974) Culture medium for enterobacteria. J Bacteriol 119(3):736–747
Nikata T, Sakai Y, Shibat K, Kato J, Kuroda A, Ohtake H (1996) Molecular analysis of the phosphate-specific transport (pst) operon of Pseudomonas aeruginosa. Mol Gen Genet 250(6):692–698
Poole K, Hancock RE (1984) Phosphate transport in Pseudomonas aeruginosa. involvement of a periplasmic phosphate-binding protein. Eur J Biochem 144(3):607–612
Qi Y, Kobayashi Y, Hulett FM (1997) The pst operon of Bacillus subtilis has a phosphate-regulated promoter and is involved in phosphate transport but not in regulation of the pho regulon. J Bacteriol 179(8):2534–2539
Rice P, Longden I, Bleasby A (2000) Emboss: the European molecular biology open software suite. Trends Genet 16(6):276–277
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, vol 999. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Scholten M, Tommassen J (1993) Topology of the phor protein of escherichia coli and functional analysis of internal deletion mutants. Mol Microbiol 8(2):269–275
Shinagawa H, Makino K, Nakata A (1983) Regulation of the pho regulon in Escherichia coli k-12. genetic and physiological regulation of the positive regulatory gene phoB. J Mol Biol 168(3):477–488
Simon R, Priefer U, Pühler A (1983) A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in gram negative bacteria. Nat Biotechnol 1(9):784–791
Smil V (2000) Phosphorus in the environment: natural flows and human interferences. Ann Rev Energy Environ 25:53–88
Spira B, Yagil E (1999) The integration host factor (IHF) affects the expression of the phosphate-binding protein and of alkaline phosphatase in Escherichia coli. Curr Microbiol 38(2):80–85
Spira B, Silberstein N, Yagil E (1995) Guanosine 3′,5′-bispyrophosphate (ppGpp) synthesis in cells of Escherichia coli starved for Pi. J Bacteriol 177(14):4053–4058
Spira B, Aguena M, de Castro Oliveira JV, Yagil E (2010) Alternative promoters in the pst operon of Escherichia coli. Mol Genet Genomics 284(6):489–498
Steed PM, Wanner BL (1993) Use of the rep technique for allele replacement to construct mutants with deletions of the pstSCAB-phoU operon: evidence of a new role for the PhoU protein in the phosphate regulon. J Bacteriol 175(21):6797–6809
Surin BP, Rosenberg H, Cox GB (1985) Phosphate-specific transport system of Escherichia coli: nucleotide sequence and gene-polypeptide relationships. J Bacteriol 161(1):189–198
Wang L, Spira B, Zhou Z, Feng L, Maharjan RP, Li X, Li F, C McKenzie, Reeves PR, Ferenci T (2010) Divergence involving global regulatory gene mutations in an Escherichia coli population evolving under phosphate limitation. Genome Biol Evol 2:478–487
Wanner BL (1996) Phosphorus assimilation and control of the phosphate regulon. In: Neidhardt NC, Curtiss R, III, Ingraham JL, Lin ECC, Low KB, Magasanik B, Reznikoff WS, Riley M, Schaechter M, Umbarger HE (eds) Escherichia coli and Salmonella: cellular and molecular biology, 2nd edn. American Society for Microbiology, Washington, DC, pp 1357–1381
Wanner BL, Wilmes-Riesenberg MR (1992) Involvement of phosphotransacetylase, acetate kinase, and acetyl phosphate synthesis in control of the phosphate regulon in Escherichia coli. J Bacteriol 174(7):2124–2130
Wu H, Kosaka H, Kato J, Kuroda A, Ikeda T, Takiguchi N, Ohtake H (1999) Cloning and characterization of Pseudomonas putida genes encoding the phosphate-specific transport system. J Biosci Bioeng 87(3):273–279
Yu CS, Chen YC, Lu CH, Hwang JK (2006) Prediction of protein subcellular localization. Proteins 64(3):643–651
Yuan ZC, Zaheer R, Morton R, Finan TM (2006) Genome prediction of PhoB regulated promoters in Sinorhizobium meliloti and twelve proteobacteria. Nucleic Acids Res 34(9):2686–2697
Acknowledgments
We are grateful to Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for supporting this study. F.N.V. was supported by FAPESP scholarship 2009/05265-7.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
da Costa Vasconcelos, F.N., Padilla, G. & Spira, B. Chromobacterium violaceum adaptation to low-phosphate conditions. Arch Microbiol 198, 269–277 (2016). https://doi.org/10.1007/s00203-016-1188-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-016-1188-6