Abstract
Mercury pollution has emerged as a major problem in industrialized zones and presents a serious threat to environment and health of local communities. Effectiveness and wide distribution of mer operon by horizontal and vertical gene transfer in its various forms among large community of microbe reflect importance and compatibility of this mechanism in nature. This review specifically describes mer operon and its generic molecular mechanism with reference to the central role played by merA gene and its related gene products. The combinatorial action of merA and merB together maintains broad spectrum mercury detoxification system for substantial detoxification of mercurial compounds. Feasibility of mer operon to coexist with antibiotic resistance gene (amp r, kan r, tet r) clusters enables extensive adaptation of bacterial species to adverse environment. Flexibility of the mer genes to exist as intricate part of chromosome, plasmids, transposons, and integrons enables high distribution of these genes in wider microbial gene pool. Unique ability of this system to manipulate oligodynamic property of mercurial compounds for volatilization of mercuric ions (Hg2+) makes it possible for a wide range of microbes to tolerate mercury-mediated toxicity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteria are known for exceptional level of adaptation to their environment. Even though they are ancient life-forms, their genetic and morphological flexibility along with immense variability in physiology enable them to survive in most extremist of the environmental conditions. From early origins of life, this very genetic flexibility and continuous course of evolution have enabled them to develop countless mechanisms relating to survival, proliferation, tolerance, and utilization of diverse resources. Environment itself is a global hub where intricate interactions among bacterial groups with themselves and with other organisms help regulate complex biogeochemical cycles and every group of organism has their own ecological niche (Fenchel et al. 1999; Vetriani et al. 2005). Mercury biogeochemical cycles are not an exception in support to this statement. Heavy metal-mediated toxicity has always remained one of the greatest barriers against survival of microbes (Sillanpää and Oikari 1996; Sorvari and Sillanpää 1996; Sillanpää et al. 2001). However, as single cellular organisms, bacteria have evolved multiple mechanisms to deal with this impediment (Baath 1989). Heavy metals and their compounds exert inhibitory effects on the functioning of bacterial enzymes and proteins thus rendering them useless. Phylogenetic and gene sequence analysis indicates that mer-related genes first originated among thermophilic microorganisms during the changes in geothermal environments (Wang et al. 2009).
In this review, we mainly focus on the mercurial compounds-mediated bacterial toxicity and describe the generic molecular models for its detoxification. The genetic system evolved as “mer operon” is in fact the only well-known bacterial metal resistance system with high yield transformation of its toxic target (Schaefer et al. 2002) into volatile non-toxic forms. Originally believed to be evolved in narrow groups of ancient species, efficacy and applicability of mer genes have enabled it to transfer and flourish in gene pools of wider microbial community. Basically, cysteine residues in proteins are the most vulnerable targets for Hg2+-based toxicity as it has high affinity toward this site. However, protein products of mer genes efficiently utilize this very characteristic of mercurial compounds for their interactions, enzymatic degradation, and transportation (Barnes and Seward 1997; Moore 1960; Sadhukhan et al. 1997; Schelert et al. 2004). Bioavailability of the mercurial compounds has direct influence on the levels of bacterial-mediated volatilization of the mercury. This in turn is dependent on the nature and concentration of the binding phase controlled by redox status of surrounding environment (Kim et al. 2006). Thus, high efficacy of mer system has generated major interest in scientific community for its detailed studies and possible utilization as a biotechnological vehicle for employing such genetic resistance mechanisms for the remediation of mercury-related environmental pollution.
Toxicity of mercury
Mercury is a toxic heavy metal and is ranked at sixth position among the top ten hazardous elements (Nascimento and Chartone-Souza 2003). Areas contaminated with mercury pose threat to both inhabitants and their environment (Virkutyte and Sillanpää 2006; Huang et al. 2008, 2009, 2011; Shrestha et al. 2010). Mercury exists in nature mainly as cinnabar ores (Barnes and Seward 1997), and several of its compounds enter aquatic environment through leaching, washing of soils sediments and rocks by rain (Shrestha and Sillanpää 2008; Sillanpää 2009). In addition, artificial mode of mercury pollution includes leakage from landfills, sludge applications, and byproducts from chemical industries (Vilhunen and Sillanpää 2009; Vilhunen et al. 2009, 2011; Rassaei et al. 2009; Sillanpää and Rämö 2009). These byproducts and chemical waste are responsible for massive amounts of organic as well as inorganic forms of mercurial compounds released into environment as indicated in the report of National Research Council (2000).
Mercury pollution in marine sediments and its effects through bioaccumulation in food chain are very serious emerging problems. Toxicity of mercury toward microbes is mainly through its oligodynamic effects (Hattemer 1954). Affinity of mercury toward organic molecules generally results in the formation of recalcitrant and highly toxic organomercurial complexes. Highly reactive mercuric ions are attributed to its binding to sulphydryl groups of the cysteine residue in essential enzymes and proteins, thus rendering them inactive and blocking vital cellular functions. The toxicity of Hg2+ ion is very swift and lethal as it is lipid soluble and readily binds to thiol group of proteins. Metallic and organomercurials can pass through biological membranes, and compounds like methylmercury (MeHg) can cause irreversible damage to nucleic acids, thereby altering normal configuration and biological activity of the cell. Mercury has been reported to react with the amino-, carboxyl-, phosphate-, and imidazole-group and diminish or inhibit (Grier 1977) the activities of vital enzymes like lactate dehydrogenase and glutathione peroxidase.
Possible mechanism of mercury detoxification
Constant exposure to mercurial compounds has enabled bacterial community to develop various types of resistance mechanism which allows them to resist the adverse effects of mercury-mediated toxicity (Osborn et al. 1997). Due course of time, evolution and enrichment of metal resistant organisms have added to diversity in tolerance mechanisms (Barkay 1987; Müller et al. 2001; Rasmussen and Sørensen 2001). Generally, detoxification of the mercury compound takes place by the volatilization or by putative entrapment (De et al. 2008). Development of mer operon and other related genetic system (Schaefer et al. 2004) is the outcome of such events. Significant levels of dissolved gaseous mercury (DGM) were detected in various types of coastal water bodies under dark condition (Fantozzi et al. 2009), which were assumed to be products of bacterial-mediated Hg2+ detoxification. Recently, genes in the conjugative transposon Tn6009 that contained Tn916 element (Soge et al. 2008) were found to resemble closely to the mer operon of Gram-positive bacteria like S. aureus which contain merA, merB, merR, and merT gene responsible for the detoxification of the mercury compound. Strangely, purified cytochrome c oxidase from Acidithiobacillus ferooxidans was also reported to show detoxification activity against mercurial compounds after intracellular transport (Sugio et al. 2010). In addition, natural phenomenon of horizontal gene transfer (HGT) has contributed to wider spreading of such genes among diverse groups (Rasmussen and Sørensen 2001) of microbial communities. In general, mechanisms for heavy metal tolerance can be classified as: (1) Blocking, in which the toxic ion is prevented from entering the cell, (2) Active efflux of the metal ion from the cell by highly specific system encoded by resistant gene, (3) Intracellular physical sequestration of the metal by binding proteins, (4) Extracellular sequestration, often by extracellular polysaccharides on the cell wall, and (5) Enzymatic conversion of the metal to less toxic or volatile forms. In nature, role of mercury resistant microorganism is significant to mercury biogeochemistry as it plays a key role in degrading MeHg and reducing Hg2+ into volatile Hg0 forms. This statement is supported by correlation among MerA activity (Siciliano et al. 2002), transcript abundance (Schaefer et al. 2004), and flux of intracellular Hg0 to the atmosphere.
Exact mechanisms and complexity among the ecological niche of mercury resistant microbes are still not fully described. Some bacteria like Cupriavidus metallidurans whose MSR33 and CH34 strains contained polyhydroxybutyrate (PHB) granules after exposure to the mercury indicating that they contain gene for PHB synthesis which activates to tolerate the stress generated by mercury (Janssen et al. 2010). Recent advancement in biotechnological techniques is helping shift the focus toward implementation of various microbial process for bioremediation and bioaccumulation (Ruta et al. 2010). In accordance to this statement, expression of the bacterial polyphosphate kinase gene (ppk) in transgenic tobacco resulted in the increased accumulation of the Hg2+ from mercury-contaminated soil without releasing mercury vapor into the ambient, thereby protecting tobacco from its toxicity (Nagata et al. 2006). Some strains of Enterobacter sp. were found to bioaccumulate and simultaneously synthesize uniformly sized mercury nanoparticles (2–5 nm). These nanoparticles were recoverable and also prevented the vaporization of mercury back into environment (Sinha and Khare 2010). Since this article mainly focuses in genetic mechanism (mer operon) for detoxification of mercurial compounds, we will be considering genetic models for mercury tolerance in bacterial community.
Bacterial mer operon
Bacteria resistant to inorganic and organic mercury compound along with resistance to penicillin was first reported in clinical samples (Moore 1960). Prolonged exposure to Hg2+ increases likelihood of bacterial strain to tolerate high level of mercury contamination.
Gram-negative bacteria are found to be more extensively studied in terms of their mer operon as compared to Gram-positive bacteria even though both have similar sets of mer genes and are arranged in similar order. The mer locus is found to be widely distributed among eubacterial lineages, and mer-like sequence has been identified in several archea genomes such as Sulfolobus solfataricus, Thermoplasma volcanicum, and Halobacterium species (Barkay et al. 2003). Variation in structure and organization of mer operon are reported (Bogdanova et al. 1992) among different isolates, indicating mosaic nature of this operon. Few characteristic differences regarding mer genes exist between Gram-negative and Gram-positive bacteria. However, the merB gene is more common to Gram-negative mer operons than in Gram-positive (Barkay et al. 2003). Analysis of various sequence of mer operon revealed that most of mer operons consist of merR gene as a regulatory gene at one terminus that is subjected to be transcribed from the structural gene of mer Operator/Promoter (O/P) region.
A number of transport function encoding genes lie proximal to the mer O/P along with merT and merP genes. Likely, in some bacterial operon, merC and orfF have been attributed for encoding transport function proteins due to its homology to merT gene. MerC is typically a membrane bound protein showing high affinity to Hg2+ ions. This is supported by the findings (Inoue et al. 1996) which shows that increased uptake of 203Hg2+ is dependent upon increasing levels of merC induction in E. coli. Studies conducted on S. solfataricus show that there is a presence of two additional mer genes namely merH and merI which are found to be present on either side of merA gene (Schelert et al. 2006). However, the exact mechanism of their activity is unknown. Bacterial community exposed to mercury contamination was found to have abundance merA gene and IncP-1 plasmid as compared to those in non-exposed environment. In addition, the plasmid IncP-1 and merA were the responsible factors for the acclimatization of microbial communities both in surface and sub-surface to mercury-contaminated areas (de Lipthay et al. 2008). Hence, HGT may have played a key role in the selection and dispersal of such plasmids and corresponding mer genes to the wider microbial community.
Mercury reductase has central role in mercury volatilization
The mer operon is one of the most widely distributed Hg2+ detoxification genetic system. Various genes are involved in mer operon, which include merR/merD for detection, merP/merT/merC for transportation or mobilization, and finally merB/merA for enzymatic detoxification of inorganic and organic mercury compounds in bacteria (Schelert et al. 2004). Though these clusters of genes are present in bacteria, it remains vestigial until it gets exposed to mercurial compounds. These clusters of genes are generally under the regulation of the merR which gets activated during Hg2+ exposure. Upon transcription, the product of this gene activates other genes including mercury reductase enzyme (MerA). Mercury reductase, a flavin oxidoreductase (Summers and Sugarman 1974), is fundamentally responsible for the reduction in highly toxic ionic Hg2+ into less toxic and volatile Hg0 in a NAD(P)H-dependent reaction. Finally, this volatile Hg0 is fluxed out from cytosolic region into outer periplasm. Amino terminal domain of MerA is found to be homologous with small periplasmic mercury-binding protein MerP which transfers Hg2+ to MerT. Exact mechanism by which MerT transfers Hg2+ into cytosol is not clearly understood but it is predicted that a pair of cysteine residue is involved in the process.
Activities of merA in anaerobic environment significantly affect MeHg production by competing for Hg2+ with methylating microbes, including sulfate-reducing bacteria (Barkay et al. 2003). MerA and its activities were well documented among strict anaerobes, and formation of Hg0 in anoxic sediments has also been investigated with significant results (Rudrick et al. 1985; Weber et al. 1998). Comparative studies of mer operon and its related gene products in denitrifying soil bacteria suggest that the activity of mer genes is induced at higher concentration of Hg2+ during anaerobic as compared to aerobic conditions. However, analysis from mer-lacZ gene fusion experiment suggests that the level of Hg2+ intake into bacterial cytosol decreases with the lowering of redox activity in mercuric ions (Schaefer et al. 2002).
Deinococcus/Thermus phylum is the deepest-branching bacterial lineage that was found to have homolog of merA gene responsible for the production of mercury reductase (Wang et al. 2009). Recently, bacterial mercury reductase has been used in various industrial processes for the removal of Hg2+ which also included strategies involving the construction of bioreactor that contained immobilized MerA enzyme (or resistant bacteria) or by the overexpression of merA gene in bacteria, algae, or plants (Lyyra et al. 2007). Similarly, merA gene from Bacillus megaterium strain MB1 was used for the transformation of eukaryotic microalga, Chlorella sp. DT, which was then able to encode MerA in the algae (Huang et al. 2006). Hence, such scientific achievements show feasibility of bacterial genetic mechanism to detoxify mercurial compounds for biotechnological use.
Role of MerB gene in organomercurial mercury volatilization
MerB gene generally code for the organomercury lyase which is one of the key enzyme for the detoxification and bioremediation of the organomercurial compound. The processed products by organomercury lyase are finally volatilized by MerA gene. The merB gene is considered as an ancillary component of the mer operon (Mei-Fang Chien et al. 2010). In most cases, merB gene was found to be mapped immediately downstream of merA gene. Phylogenetic analysis of various bacteria shows that MerB is one of the unique enzymes whose homolog forms are not known (Barkay et al. 2003). MerB catalyzes the protonolysis of carbon–mercury bound, thereby releasing less toxic and less mobile Hg2+ species which is further acted upon by MerA enzyme for complete volatilization of organomercurial compounds (Murtaza et al. 2005).
Crystallography studies of MerB enzyme revealed two conserved cysteines residue namely Cys-96 and Cys-159 that are considered as substrate binding region. This region plays a crucial role in cleavage of the carbon–mercury bond, thereby releasing ionic Hg2+ form of mercury. Similarly, Asp-99 residue of MerB enzymes was found to play active role in proton transfer during protonolysis cleavage (Vanasse et al. 2008) of carbon–mercury bond.
MerA and MerB together act as broad spectrum mercury detoxification system
Mainly two types of mercury resistant mechanism are prevalent in nature: (1) narrow spectrum and (2) broad spectrum. In narrow spectrum, only merA gene is present and resistance mechanism is limited to enzymatic detoxification of only inorganic mercury compound. In case of broad spectrum, tolerance is exhibited to organic as well as inorganic mercurial compounds by converting both forms of compounds to their volatile forms (Sadhukhan et al. 1997).
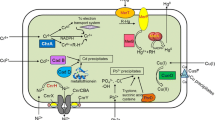
Broad spectrum mercury-tolerant bacteria (Fig. 1) contain extra gene merB which codes organomercurial lyase (Griffin et al. 1987; Silver and Phung 1996) for the cleavage of carbon–mercury bond in organomercurial compounds. In general, narrow spectrum mercury-resistant operon (e.g., merRTPADE) confers resistance to only inorganic mercurial compounds, while the board spectrum mercury-resistant operon (e.g., merRTPAGBDE) confers resistant to both inorganic and organic mercurial compounds (Rojas et al. 2011).
Generic mer operon in typical Gram-negative mercury-resistant bacteria. RSH represents low molecular mass compounds and X is a nucleophilic solvent. RSHgSR and CH3HgSR are reaction intermediates. Both organic (CH3HgX) and inorganic (HgX) forms of mercurial compounds passes via MerC and MerT inner-membrane proteins into cytosol where the action of enzyme MerA or both MerA and MerB results in volatilization and cellular release of only Hg0 or both Hg0 and CH4, respectively
A typical periplasmic protein MerG in Gram-negative bacteria is found to provide resistance against organomercurial in merB deficient strains (Barkay et al. 2003). Hence, presence of merA along with merG may still show the effect of broad spectrum mercury detoxification in such bacterial strains. Interestingly, nucleotide sequencing of Incp-1b plasmid isolated from mercury-contaminated river revealed mer genes existing as a part of transposon Tn50580 (Smalla et al. 2006). Likely, various species of floras including Arabidopsis, tobacco, and chlorella have been biotechnologically modified to incorporated merA and merB genes that carried out detoxification of mercury-contaminated soil (Ruiz and Daniell 2009) as a part of bioremediation. This further supports the flexibility and adaptability of bacterial mer operon as an inter-species compatible genetic mechanism for tolerance against mercurial toxicity.
Primers help identify mercury resistant determinants among bacterial population
Mercury resistance mechanisms have widely been distributed among bacterial populations and are even more common in Gram staining bacteria. Two separate set of primers are needed for Gram-negative and Gram-positive bacteria as the sequence of merA gene differs among species (Chatziefthimiou et al. 2007). The mer genes can be located in plasmids, chromosomes and have also been identified as components of transposons and integrons (Zeyaullah et al. 2010).
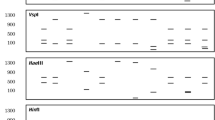
Multiple genes for detection, mobilization, and enzymatic detoxification of mercurial compounds are distributed among closely linked gene clusters. Within these clusters, the merA gene has remained in focus for primer design (Barkay et al. 2010) and detection of mercury resistant species via polymerase chain reaction (PCR). Use of primer enables exact identification and even helps to pinpoint the precise location of particular gene. Most widely used primers have been designed based on conserved regions of merA and merB genes. However, as suggested by multiple research data (Table 1), primers have been designed for different target genes within mer operon including merA, merB, merD, merP as well as transposons and integrons. The mer operon-related sequence homology studies conducted using PCR in thermophilic bacteria and other Hg2+ resistance microbes also support the statement that HGT may have played major role in wide distribution of mer operon (Zeyaullah et al. 2010; Lal and Lal 2010).
Antibiotic resistance is generally linked to Hg2+ resistance
Antibiotic resistance is presumably one of the most common features of bacterial adaptation (Boni and Feldman 2005). However, co-transfer of mercury and antibiotic resistance genes have immensely been found in nature as well as experimental conditions. Both antibiotic and metal resistances can occur on same conjugative plasmids, chromosomes as well as transposons elements (Roberts et al. 2008). In addition, multiple antibiotic resistance carrying plasmids have been found to carry Hg2+ resistance genes (Foster 1983). Studies on mercury-tolerant Pseudomonas, Kleblessa, Enterobacteriaceae, and other Gram-negative bacteria (Summers et al. 1993) suggest a kind of genetic linkage that results in co-transfer of both traits. Multiple researches conducted on Hg2+-resistant microbes (Zeyaullah et al. 2010; Lal and Lal 2010) also suggest that HGT played a key role in high distribution of mer-related genes along with antibiotic resistance genes in microbial gene pool. Even though current findings does not present a generic conclusion regarding this linkage, existence of mer and antibiotic loci at similar or close proximity may have a key role in their co-transfer (Summers et al. 1993). Transformations of experimental bacterial strains with derivatives of antibiotic and Hg2+ sensitive natural isolates or competent cells as recipients have shown (Wireman et al. 1997) that most Hg2+ resistant strains co-transferred Hg2+ linked arrays of antibiotic resistance markers (amp r, kan r, tet r) along with the target Hg2+-resistant genes. IncP-1 plasmid is perhaps the only known genetic system which only consists of mercury resistance transposon element but no antibiotic resistance genes (Smalla et al. 2006).
Conclusions
Detoxification of mercurial compound mediated by mer operon is one of the oldest studied bacterial mechanisms against heavy metal toxicity. Even though multiple genes plays integral role in constituting the resistance, activity of merA has remained central to enzymatic transformation of mercurial compounds during detoxification process. Despite availability of huge information regarding the genes involved in this operon, new insights into gene regulation and enzymology are constantly emerging. Presence of merA and merB in its various forms among wide range of microbes as a primary mechanism for mercury detoxification reflects the adaptability and importance of this mechanism in natural world. Relations between Hg2+ and antibiotics resistance are not clearly defined and are a subject of further studies. As multiple ongoing researches relating to microbe-mediated hazardous metal detoxification mechanisms and their possible applications in bioremediation are being considered, better understanding of the mer operon and their gene products becomes essential.
Abbreviations
- DGM:
-
Dissolved gaseous mercury
- HGT:
-
Horizontal gene transfer
- MeHg:
-
Methylmercury
- O/P:
-
Operator/promoter
- PCR:
-
Polymerase chain reaction
- PHB:
-
Polyhydroxybutyrate
References
Abou-Shanab RA, van Berkum P, Angle JS (2007) Heavy metal resistance and genotypic analysis of metal resistances genes in gram-positive and gram-negative bacteria present in Ni-rich serpentine soil and in the rhizosphere of Alyssum murale. Chemosphere 68:360–367
Baath E (1989) Effects of heavy metals in soil on microbial processes and populations. Water Air Soil Pollut 47:335–379
Barkay T (1987) Adaptation of aquatic microbial communities to Hg+2 stress. Appl Environ Microbiol 53:2725–2732
Barkay T, Miller SM, Summers AO (2003) Bacterial mercury resistance from atoms to ecosystems. FEMS Microbiol Rev 27:355–384
Barkay T, Kritee K, Boyd E, Geesey G (2010) A thermophilic bacterial origin and subsequent constraints by redox, light and salinity on the evolution of the microbial mercuric reductase. Environ Microbiol 12:2904–2917
Barnes HL, Seward TM (1997) Geothermal systems and mercury deposits. In: Barnes HL (ed) Geochemistry of hydrothermal ore deposits, 3rd edn. Wiley, NewYork, pp 699–736
Bogdanova E, Mindlin S, Pakrava E, Kocur M, Rouch D (1992) Mercuric reductase in enviromental gram-positive bacteria sensitive to mercury. FEMS Microbiol Lett 97:95–100
Boni MF, Feldman MW (2005) Evolution of antibiotic resistance by human and bacterial niche construction. Evolution 59:477–491
Chatziefthimiou AD, Crespo-Medina M, Wang Y et al (2007) The isolation and initial characterization of mercury resistant chemolithotrophic thermophilic bacteria from mercury rich geothermal springs. Extremophiles 11:469–479
Chien MF, Narita M, Lin KH, Matsui K, Huang CC, Endo G (2010) Organomercurials removal by heterogeneous merB genes harboring bacterial strains. J Biosci Bioeng 110:94–98
de Lipthay JR, Rasmussen LD, Oregaard G, Simonsen K, Bahl MI, Kroer N, Sørensen SJ (2008) Acclimation of subsurface microbial communities to mercury. FEMS Microbiol Ecol 65:145–155
De J, Ramaiah N, Vardanyan L (2008) Detoxification of toxic heavy metals by marine bacteria highly resistant to mercury. Mar Biotechnol 10:471–477
Fantozzi L, Ferrara R, Frontini FP, Dini F (2009) Dissolved gaseous mercury production in the dark: evidence for the fundamental role of bacteria in different types of Mediterranean water bodies. Sci Total Environ 407:917–924
Fenchel T, King GH, Blackburn TH (1999) Bacterial biogeochemistry. The ecophysiology of mineral cycling. Int Microbiol 2:201–204
Foster TJ (1983) Plasmid-determined resistance to antimicrobial drugs and toxic metal ions in bacteria. Microbiol Rev 47:361–409
Grier N (1977) Mercurials-inorganic and organic in disinfection, sterilization, and preservation, 2nd edn. Lea and Febiger, Philadelphia, pp 361–385
Griffin HG, Foster TJ, Silver S, Misra TK (1987) Cloning and DNA sequence of the mercuric- and organomercurial-resistance determinant of plasmid pDU1358. Proc Natl Acad Sci USA 84:3112–3116
Hattemer AJ (1954) Oligodynamic effects of heavy metals. Zahnarztl Rundsch 63:431–436
Huang CC, Chen MW, Hsieh JL, Lin WH, Chen PC, Chien LF (2006) Expression of mercuric reductase from Bacillus megaterium MB1 in eukaryotic microalga Chlorella sp. DT: an approach for mercury phytoremediation. Appl Microbiol Biotechnol 72:197–205
Huang X, Sillanpää M, Duo B, Gjessing ET (2008) Water quality in the Tibetan plateau: metal contents of four selected rivers. Environ Pollut 156:270–277
Huang X, Sillanpää M, Gjessing E, Vogt RD (2009) Water quality in the Tibetan plateau: major ions and trace elements in the headwaters of four major Asian rivers. Sci Tot Environ 407:6242–6254
Huang X, Sillanpää M, Gjessing ET, Peräniemi S, Vogt RD (2011) Water quality in the southern Tibetan plateau: chemical evaluation of the river Yarlung Tsangpo (Brahmaputra). River Res Appl 27:113–121
Inoue C, Kusano T, Silver S (1996) Mercuric ion uptake by Escherichia coli cells producing Thiobacillus ferrooxidans MerC. Biosci Biotechnol Biochem 60:1289–1292
Janssen PJ, van Houdt R, Moors H, Monsieurs P, Morin N et al (2010) The complete genome sequence of Cupriavidus metallidurans strain CH34, a master survivalist in harsh and anthropogenic environments. PLoS ONE 5:e10433. doi:10.1371/journal.pone.0010433
Kim EH, Mason RP, Porter ET, Soulen HJ (2006) The impact of resuspension on sediment mercury dynamics, and methylmercury production and fate: a mesocosm study. Mar Chem 102:300–315
Lal D, Lal R (2010) Evolution of mercuric reductase (merA) gene: a case of horizontal gene transfer. Mikrobiologiia 79:524–531
Liebert CA, Wireman J, Smith T, Summers AO (1997) Phylogeny of mercury resistance (mer) operons of gram-negative bacteria isolated from the fecal flora of primates. Appl Environ Microbiol 63:1066–1076
Lyyra S, Meagher RB, Kim T et al (2007) Coupling two mercury resistance genes in eastern cottonwood enhances the processing of organomercury. Plant Biotechnol J 5:254–262
Martins AS, Silva de Jesus M, Lacerda M, Moreira JC, Filgueiras ALL, Barrocas PRG (2008) A conservative region of the mercuric reductase gene (merA) as a molecular marker of bacterial mercury resistance. Braz J Microbiol 39:307–310
Moore B (1960) A new screen test and selective medium for the rapid detection of epidemic strains of Staphylococcus aureus. Lancet 2:453–458
Müller AK, Westergaard K, Christensen S, Sorensen SJ (2001) The effect of long-term mercury pollution on the soil microbial community. FEMS Microbiol Ecol 36:11–19
Murtaza I, Dutt A, Mushtaq D, Ali A (2005) Molecular cloning and genetic analysis of functional merB gene from Indian isolates of Escherichia coli. Curr Microbiol 51:297–302
Nagata T, Kiyono M, Pan-Hou H (2006) Accumulation of mercury in transgenic tobacco expressing bacterial polyphosphate. Biol Pharm Bull 29:2350–2353
Nascimento AMA, Chartone-Souza E (2003) Operon mer: bacterial resistance to mercury and potential for bioremediation of contaminated environments. Genet Mol Res 2:92–101
National Research Council (2000) Toxicological effects of methylmercury. National Academy Press, Washington, pp 147–246
Osborn AM, Bruce KD, Strike P, Ritchie DA (1997) Distribution, diversity and evolution of the bacterial mercury resistance (mer) operon. FEMS Microbiol Rev 4:239–262
Ramond JB, Berthe T, Duran R, Petit F (2009) Comparative effects of mercury contamination and wastewater effluent input on gram-negative merA gene abundance in mudflats of an anthropized estuary (Seine, France): a microcosm approach. Res Microbiol 160:10–18
Rasmussen LD, Sørensen SJ (2001) Effects of mercury contamination on the culturable heterotrophic, functional and genetic diversity of the bacterial community in soil. FEMS Microbiol Ecol 36:1–9
Rassaei L, Sillanpää M, Edler KJ, Marken F (2009) Electrochemically active mercury nanodroplets trapped in a carbon nanoparticle—chitosan matrix. Electroanalysis 21:261–266
Roberts MC, Leroux BG, Sampson J, Luis HS, Bernardo M, Leitão J (2008) Dental amalgam and antibiotic- and/or mercury-resistant bacteria. J Dent Res 87:475–479
Rojas LA, Yanez C, Gonzalez M, Lobos S, Smalla K, Seeger M (2011) Characterization of the metabolically modified heavy metal-resistant Cupriavidus metallidurans strain MSR33 generated for mercury bioremediation. PLoS ONE 6(3):e17555. doi:10.1371/journal.pone.0017555
Rudrick JT, Bawdon RE, Guss SP (1985) Determination of mercury and organomercurial resistance in obligate anaerobic bacteria. Can J Microbiol 31:276–281
Ruiz ON, Daniell H (2009) Genetic engineering to enhance mercury phytoremediation. Curr Opin Biotechnol 20:213–219
Ruta L, Paraschivescu C, Matache M et al (2010) Removing heavy metals from synthetic effluents using ‘‘kamikaze’’ Saccharomyces cerevisiae cells. Appl Microbiol Biotechnol 85:763–771
Sadhukhan PC, Ghosh S, Chaudhuri J, Ghosh DK, Mandal A (1997) Mercury and organomercurial resistance in bacteria isolated from freshwater fish of wetland fisheries around Calcutta. Environ Pollut 97:71–78
Schaefer JK, Letowski J, Barkay T (2002) mer mediated resistance and volatilization of Hg(II) under anaerobic conditions. Geomicrobiol J 19:87–102
Schaefer JK, Yagi J, Reinfelder JR, Cardona T, Ellickson KM, Tel-Or S, Barkay T (2004) Role of the bacterial organomercury lyase (MerB) in controlling methylmercury accumulation in mercury-contaminated natural waters. Environ Sci Technol 38:4304–4311
Schelert J, Dixit V, Hoang V, Simbahan J, Drozda M, Blum P (2004) Occurrence and characterization of mercury resistance in the hyperthermophilic archaeon Sulfolobus solfataricus by use of gene disruption. J Bacteriol 186:427–437
Schelert J, Drozda M, Dixit V, Dillman A, Blum P (2006) Regulation of mercury resistance in the crenarchaeote Sulfolobus solfataricus. J Bacteriol 188:7141–7150
Shrestha RA, Sillanpää M (2008) Influence of Eh/pH-Barriers on releasing/accumulation of manganese and iron at sediment-water interface. Res J Chem Environ 12:7–13
Shrestha R, Kafle B, Sillanpää M (2010) Water quality of Dhulikhel area, Nepal. Res J Chem Environ 14:36–38
Siciliano SD, O’Driscoll NJ, Lean DRS (2002) Microbial reduction and oxidation of mercury in freshwater lakes. Environ Sci Technol 36:3064–3068
Sillanpää M (2009) Occurrence, interaction with heavy metals and behaviour of complexing agents in the environment: a review. Res J Chem Environ 13:99–103
Sillanpää M, Oikari A (1996) Assessing the impact of complexation by EDTA and DTPA on heavy metal toxicity using microtox bioassay. Chemosphere 32:1485–1497
Sillanpää M, Rämö J (2009) Metal analysis of pulp: ICP-AES, XRF and ISE methods and their on-line feasibility. Res J Chem Environ 13:63–67
Sillanpää M, Orama M, Rämö J, Oikari A (2001) The importance of ligand speciation in environmental research: a case study. Sci Tot Environ 267:23–31
Silver S, Phung LT (1996) Bacterial heavy metal resistance; new surprises. Ann Rev Microbiol 50:753–789
Sinha A, Khare SK (2010) Mercury bioaccumulation and simultaneous nanoparticle synthesis by Enterobacter sp. cells. Bioresour Technol 102:4281–4284
Smalla K, Haines AS, Jones K, Krögerrecklenfort E, Heuer H, Schloter M, Thomas CM (2006) Increased abundance of IncP-1beta plasmids and mercury resistance genes in mercury-polluted river sediments: first discovery of IncP-1beta plasmids with a complex mer transposon as the sole accessory element. Appl Environ Microbiol 72(11):7253–7259
Soge OO, Beck NK, White TM, No DB, Robert M (2008) A novel transposon, Tn6009, composed of a Tn916 element linked with a Staphylococcus aureus mer operon. J Antimicrob Chemother 62:674–680
Sorvari J, Sillanpää M (1996) Influence of metal complex formation on heavy metal and free EDTA and DTPA acute toxicity determined by D. magna. Chemosphere 33:1119–1127
Sugio T, Komoda T, Okazaki Y, Takeda Y, Nakamura S, Takeuchi F (2010) Volatilization of metal mercury from organomercurial by highly mercury-resistant Acidithiobacillus ferrooxidans MON-1. Biosci Biotechnol Biochem 74:1007–1012
Summers AO, Sugarman LI (1974) Cell-free mercury(II)-reducing activity in a plasmid-bearing strain of Escherichia coli. J Bacteriol 119:242–249
Summers AO, Wireman J, Vimy MJ, Lorscheider FL, Marshall B, Levy SB, Bennett S, Billard L (1993) Mercury released from dental “silver” fillings provokes an increase in mercury—and antibiotic-resistant. Antimicrob Agents Chemother 37:825–834
Vanasse JL, Lefebvre M, Lello PD, Sygusch J, Omichinsk JG (2008) Crystal structures of the organomercurial lyase MerB in its free and mercury-bound forms. J Biol Chem 284:938–944
Vetriani C, Chew YS, Miller SM, Yagi J, Coombs J, Lutz RA, Barkay T (2005) Mercury adaptation among bacteria from a deep-sea hydrothermal vent. Appl Environ Microbiol 71:220–226
Vilhunen SH, Sillanpää MET (2009) Ultraviolet light emitting diodes and hydrogen peroxide in the photodegradation of aqueous phenol. J Hazard Mater 161:1530–1534
Vilhunen S, Särkkä H, Sillanpää M (2009) Ultraviolet light emitting diodes in water disinfection. Environ Sci Pollut Res 16:439–442
Vilhunen S, Puton J, Virkutyte J, Sillanpää M (2011) Efficiency of hydroxyl radical formation and phenol decomposition by using UV light emitting diodes and H2O2. Environ Technol 32:865–872
Virkutyte J, Sillanpää M (2006) Chemical evaluation of potable water in eastern Qinghai province, China: human health aspects. Environ Int 32:80–86
Wang Y, Freedman Z, Lu-Irving P, Kaletsky R, Barkay R (2009) An initial characterization of the mercury resistance (mer) system of the thermophilic bacterium Thermus thermophilus HB27. FEMS Microbiol Ecol 67:118–129
Weber JH, Evans R, Jones SH, Hines ME (1998) Conversion of mercury(II) into mercury(0), monomethylmercury cation, and dimethylmercury in saltmarsh sediment slurries. Chemosphere 36:1669–1687
Wireman J, Liebert CA, Smith T, Summers AO (1997) Association of mercury resistance with antibiotic resistance in the gram-negative fecal bacteria of primates. Appl Environ Microbiol 63:4494–4503
Zeyaullah Md, Islam B, Arif Ali A (2010) Isolation, identification and PCR amplification of merA gene from highly mercury polluted Yamuna river. AJB 9:3510–3514
Conflict of interest
Authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Rights and permissions
About this article
Cite this article
Mathema, V.B., Thakuri, B.C. & Sillanpää, M. Bacterial mer operon-mediated detoxification of mercurial compounds: a short review. Arch Microbiol 193, 837–844 (2011). https://doi.org/10.1007/s00203-011-0751-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-011-0751-4