Abstract
Microbial mats are prokaryotic communities that provide model systems to analyze microbial diversity and ecophysiological interactions. Community diversity of microbial mat samples was assessed at 8:00 a.m. and 3:00 p.m. in a combined analysis consisting of 16S rRNA-denaturing gradient gel electrophoresis (DGGE) and phospholipid fatty acid (PLFA) profiles. The divergence index determined from PLFA and DGGE data showed that depth-related differences have a greater influence on diversity than temporal variations. Shannon and Simpson indices yielded similar values in all samples, which suggested the stable maintenance of a structurally diverse microbial community. The increased diversity observed at 3:00 p.m. between 2.5 and 4 mm can be explained mainly by diversification of anaerobic microorganisms, especially sulfate-reducing bacteria. In the afternoon sampling, the diversity index reflected a higher diversity between 4 and 5.5 mm depth, which suggested an increase in the diversity of strict anaerobes and fermenters. The results are consistent with the conclusion that hypersaline microbial mats are characterized by high degree of diversity that shifts in response to the photobiological adaptations and metabolic status of the microbial community.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial mats are prokaryotic communities that are thought to represent the present-day analogues of the first ecosystems on Earth. The close spatial relationships between their members facilitate the establishment of microscale biochemical gradients and microniches, which, in turn, lead to a more complete recycling of nutrients, diversification of the microbiota, and closer community interactions over a range of temporal and spatial scales (Paerl et al. 2000). Microbial mats are highly diverse ecosystems characterized by diel variation of oxygen, sulfide, temperature, and salinity conditions. They are also a source of not-yet characterized microorganisms that are well adapted in the microbial community (Guerrero et al. 2002). Although microbial mats have been intensively studied as model microbial ecosystems, uncertainties remain concerning their biochemical cycles, cooperative associations, and the identities of their resident microorganisms.
Previous studies have quantified the microbial diversity of certain microbial mat populations by cultivation-independent approaches based on morphology, carotenoid content and 16S rRNA analysis (Nübel et al. 1999). However, our knowledge of mat community diversity is still limited due to methodological problems and to the lack of taxonomic information. Molecular techniques have been used to overcome the limitations of culture-based methods; however, they are also subject to restrictions (Kirk et al. 2004). White and Findlay (1988) developed a community-level approach to characterize the structures of microbial communities based on shifts in phospholipid fatty acids (PLFAs) obtained from environmental samples. Different groups of bacteria are characterized by specific PLFA profiles; therefore, a change in the phospholipid pattern implies a change in bacterial composition. This approach has resulted in the identification and quantification of both viable biomass and community structure in sediments (Ibekwe et al. 2001; Ringelberg et al. 1988) and microbial mats (Navarrete et al. 2000; Navarrete et al. 2004). Nonetheless, despite its versatility, PLFA analysis is limited to the study of bacteria (White and Ringelberg 1997) and has therefore been complemented by nucleic acid-based analyses, such as denaturing gradient gel electrophoresis (DGGE) (Macnaughton et al. 1999; Stephen et al. 1999). In previous studies, the combined use of DNA-based and lipid analyses provided a quantitative means of estimating microbial diversity in various environments, and the ability to relate microbial community structure to environmental conditions (Fromin et al. 2002; Torsvik and Øverås 2002; Torsvik et al. 2002). Although microbial community fingerprinting methods include a variety of well-known PCR biases (Wintzingerode et al. 1997), they provide comprehensive information on global patterns of microbial diversity and have thus proved useful in the study of factors that govern microbial diversity, ecology, and function in numerous habitats (Casamayor et al. 2002; Tankéré et al. 2002).
The hypersaline microbial mats investigated in this study are located in the pre-concentration pond in the Salin-de-Giraud solar salt works on the Mediterranean French coast in the Rhône Delta (Camargue, France; Caumette et al. 1994). These cyanobacterial mats are composed of diatoms and unicellular cyanobacterial in the surface layer. Below, filamentous cyanobacteria, mainly Microcoleus chthonoplastes, form a cohesive band. For a more detailed microbial composition in depth zonation see Fourçans et al. (2004) and Fourçans et al. (2006). In addition, Camargue mats are characterized by high sulfate reduction rates and high iron content (Wieland et al. 2005), and they have also been studied because of their potential for biodegradation of crude oil (Benthien et al. 2004).
In the present survey, we complemented a previous study based on a combined PLFA/nucleic approach to monitor changes in the physiological status, biomass, and community composition in a hypersaline mat (Villanueva et al. 2004), giving emphasis on the spatial and temporal microbial diversity variations at a microscale level. Thus, PLFA and DGGE were performed in samples obtained at various depths (0–8.75 mm depth) from a hypersaline microbial mat. This study was based on daytime measurements to gain insight into the microbial diversity changes induced by the close coupling between autotrophs and heterotrophs in hypersaline microbial mats (Wieland and Kühl 2000), and for this reason, samples were taken at two selected times during the day (8:00 a.m., 3:00 p.m.) as representative of the beginning and height of the photosynthetic period. Ecological diversity is a function of the number of different classes (richness) and the relative distribution of elements between them (evenness; Begon et al. 1990). In this case, diversity indices and divergence between samples were calculated, which allowed classification of the PLFA and DGGE data according to clustering methods. These studies revealed the high diversity of microbial mats and the importance of combining different analytical approaches for estimating diversity in complex microbial communities.
Material and methods
Sampling and lipid analysis
Cyanobacterial microbial mats were sampled in a pre-concentration pond of the Salin-de-Giraud solar salt works (Camargue, France) in April 2002 at two selected times during the day (8:00 a.m. and 3:00 p.m.; salinity ranged between 70 and 80‰; for a more detailed description see Caumette et al. 1994). Samples A were obtained at 8:00 a.m. GMT + 1:00 and samples B at 3:00 p.m. GMT + 1:00. Each sample was cut on a microtome into 50-μm thick slices, and then ten cuts grouped to form each sample group (total depth per layer, ca. 500-μm thick for sample group 1–15; group 16 contained 25 slices 50-μm thick, total depth 8.75 mm). Duplicate samples were extracted according to the modified method of Bligh and Dyer (1959) described by White et al. (1979) and the resulting total lipid extracts were fractionated into neutral, glyco-, and polar lipids by silicic acid chromatography. Polar-lipid fractions were then transesterified to fatty acid methyl esters (Guckert et al. 1985) and analyzed by gas chromatography/mass spectrometry (for a more detailed description on analytical method and nomenclature see Navarrete et al. 2000).
DNA purification and DGGE analysis
Nucleic acids were precipitated directly from the PLFA aqueous phase as described in Chang et al. (1999). 16S rRNA gene fragments were PCR-amplified with primers targeted eubacterial 16S regions corresponding to Escherichia coli nucleotide positions 341–534 (Brosius et al. 1981). DGGE was performed by using a D-Code 16/16-cm gel system with a 1.5-mm gel width (Bio-Rad, Hercules, CA, USA). Gradients were formed between 30 and 65% denaturant (with 100% denaturant defined as 7 M urea plus 40% [v/v] formamide). Gels were run at 35 V for 16 h as described by Muyzer et al. (1993). Excised DGGE bands were used as templates in PCR reactions, and the purified PCR products were sequenced. Amplification products that failed to directly generate legible sequence were cloned into the pGEM-T Easy system II (Promega, WI, USA) cloning vector according to the manufacturer’s instructions.
Sequence and phylogenetic analysis
Sequences were compared with the GenBank database using the BlastN facility of the National Center for Biotechnology information (Altschul et al. 1997) and aligned in ClustalW. Matrices of evolutionary distances were computed from the sequence alignment using MEGA 3.0 software (Kumar et al. 2004). Distance matrices were done according to the neighbor-joining algorithm and with the Jukes and Cantor model. In the comparative analysis the pairwise deletion was selected. To validate the reproducibility of the branching pattern, a bootstrap analysis was performed (1,000 replicates).
Statistical analysis of PLFA profiles and DGGE bands
The PLFA data were analyzed using the Microsoft Excel software package and Statgraphics Plus 5.1 for Windows (StatPoint, Inc., VA, USA). Reproducibility of PLFA analysis between mat cores replicates was tested by analysis of variance and standard deviation (two sampling events × four replicate mat cores; standard deviation between samples was ± 10%). Differences in the community structure of microbial mats over time and space were estimated by interpreting PLFA profile data. The divergence index (Div) (Hiraishi 1999; Iwasaki and Hiraishi 1998; Krebs 1985) was used to estimate differences between samples based on PLFA content or DGGE band intensities. Div was calculated according to Eq. 1:
where X ki , X kj ≥ 0.01, ∑X ki = ∑ X kj = 100, and X ki and X kj indicate the levels (expressed as mol% or P i of a DGGE band in a gel lane, see below for explanation) of the PLFA or DGGE band k in samples i and j, respectively. The neighbor-joining algorithm (Saitou and Nei 1987) was used to construct a dendrogram based on the Div matrix data obtained using Mega 3.0 software. Scanned DGGE gels were analyzed with the Scion Image software package for Windows (NIH Image, Scion, USA) as previously described in Eichner et al. (1999) and Sekiguchi et al. (2002). Community diversity was measured by means of the Shannon–Wiener index (also known as the Shannon–Weaver index) (H) that represents the uncertainty in predicting the species of an individual chosen at random from a community, which increases with richness as well as with evenness (Krebs 1985; Ludwig Reynolds 1988; Shannon and Weaver 1963) and was calculated using the following function: H = −∑P i log P i, where P i is the importance probability of the bands in a gel lane or the mol% data of each PLFA in a sample. P i was calculated as follows: P i = n i/N, where n i is the band intensity for individual bands or the pmol g−1 of a certain PLFA, and N is the sum of intensities of bands in a lane or the sum of all PLFAs in a sample, expressed as pmol g−1. Evenness was calculated from H/H max, where H max is equal to ln(S) and S is the total number of phylotypes (Dunbar et al. 1999). The Simpson index (λ) (Simpson 1949) was calculated for PLFA data and for DGGE data (D = 1 − λ = 1 − ∑P 2i ). Since the scale for D varies between communities according to S, the measure was standardized by dividing by D max. The latter was calculated from [1 − 1/(S)] and provides an evenness index similar to the one derived from the Shannon–Wiener function. The PLFA and DGGE data were normalized to a common analytical sensitivity in order to compare their diversity indices (Hedrick et al. 2000).
Results and discussion
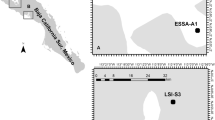
As it was described in Villanueva et al. (2004), PCR-DGGE analysis of bacterial community structure was carried out with DNA recovered from the aqueous phase after total lipid extraction of microbial mat samples (Fig. 1a, b; a phylogenetic tree of the DGGE bands and closest relatives is detailed in Fig. 2a, b). At the top of the mat (a depth of 0–2.5 mm), the A and B samples generated complex banding patterns with several similarities. For example, the prominent band A2A, which was strongly present until a depth of 2 mm and then weakened or disappeared, was also found in B samples at the same relative position (B1A). Band A3E was brighter between 0.5 and 2 mm, and its corresponding band in the gel containing the B samples (gel B), B2B, was found under the same conditions and at the same intensity. Sequence analysis of bands obtained from this part of the mat showed their high homology with Marinobacter sp. The sequences recovered from the top of the mat suggested the presence of Flavobacteriaceae (B1A, A2A, B1D, B5A) and the presence of γ-Proteobacteria (B2B, A3E), members of the phylum Firmicutes (Halanaerobiales, A5A, A5B), cyanobacteria (B2C), members of the Cytophaga-Flavobacterium-Bacteroides phylum, and spirochaetes (A3D).
The middle part of the mat (samples 6–10) yielded an increased number of bands (A8E, A8F, A9B, A8H, and A9D). Those derived from the gel containing the A samples (gel A) remained strong between 2 and 5.5 mm (A6B, A6E, A8E–H), except for bands A6D-E, A9B, A9D, and A10A, which were present along the entire vertical profile. The sequence derived from band A6D showed 100% similarity with Halanaerobium saccharolyticum. Finally, DGGE analysis of samples collected from the bottom of the mat (samples 11–16, corresponding to a depth of 5–8.75 mm) showed a greater intensity of the sample B bands than of the sample A bands.
Taken together, the results are consistent with a major contribution in the upper layers of the mat of aerobic heterotrophic bacteria (Psychroflexus sp., Sphingobacterium sp., Marinobacter sp.) belonging to the Cytophaga-Flavobacterium-Bacteroides group and γ-Proteobacteria. Other sequences retrieved from DGGE bands of this layer showed 100% similarity with the Halanaerobium genus. Important contributions of members of the phylum Bacteroidetes, which are fermentative bacteria seemingly adapted to high salinities, were also identified. Moreover, the detection of several bands related to Chloroflexus-like species suggests an important role of this genus in microbial mats. Indeed, the presence of these microbial groups in hypersaline microbial mats, where they are major contributors to carbon cycling, was recently reported (Fourçans et al. 2004). Fourçans et al. (2004), performed an extensive analysis of the microbial composition of Camargue microbial mats based on morphological and molecular analysis, and demonstrated the importance of Chloroflexus-like members in surface layers as well as the presence of purple anoxygenic bacteria, members of the Rhodobacteraceae family and Thiomicrospira genus, and also of sufate-reducing bacteria (Desulfovibrio, Desulfobacter and Desulfonema genera) in the underlying layers. Similarly, Sørensen et al. (2005) detected 16S rRNA gene sequences in a hypersaline endoevaporitic microbial community in Eilat (Israel) with high similarity with microbial members described by Fourçans et al. (2004) and Ley et al. (2006). In fact, Ley et al. (2006) suggested a symbiotic or antagonist process between Chloroflexi and cyanobacteria as an explanation for the presence of Chloroflexus-derived sequences throughout the vertical profile of a hypersaline microbial mat at Guerrero Negro (Baja California, Mexico).
Phospholipid fatty acid data describing community composition of the analyzed samples (included as a supplementary data) reported a higher proportion of gram-positive bacteria in the middle layers and at the deepest samples in the morning as well as a similar concentration of PLFA representative of anaerobic microorganisms along the vertical profile. On the contrary, in the afternoon the proportion of PLFA indicative of anaerobic microorganisms was higher and increased with depth. In order to compare the PLFA and DGGE patterns of samples obtained from different depths and at two different times during the day, the Div was calculated. The Div can be used to determine the extent of differences among samples from microbial community structures. Thus, the unweighted pair-group method with an arithmetic mean (UPGMA) algorithm was used to create a dendogram describing pattern similarities (Figs. 3, 4). The results suggested that, with respect to community diversity, depth-related differences were greater than temporal differences based on both DGGE and PLFA data. Cluster analysis based on PLFA data revealed a grouping pattern with cluster 1 comprising samples obtained at 3:00 p.m. (B samples) at a depth of 4–7.5 mm, while cluster 2 consisted of samples obtained at 8:00 a.m. at a depth of 4–8.75 mm and at 3:00 p.m. at a depth of 7.5–8.75 mm. Finally, cluster 3 grouped A and B samples taken from a depth of 2.5–4 mm and cluster 4 grouped the most surface samples of both sampling times. Likewise, cluster analysis based on DGGE data showed a similar tendency of grouping superficial samples (cluster 3) as well as those recovered from the deepest layers of the mat (cluster 4), and it also confirms a stronger influence of the depth-related differences. However, comparing both cluster analysis we can observe a more defined grouping based on PLFA data that can be attributed to the basis of the method itself and it demonstrates the convenience of combining both methods for a more accurate analysis. The predominant role of depth-related differences suggests that the PLFA and DGGE profiles of mat populations do not change in response to diurnal cycles but instead reflect the community composition of established microniches.
Shannon and Simpson diversity indices were calculated from the 16S rRNA-DGGE data and from the PLFA data of the mat community. The Shannon index (H) takes into account the number and relative intensities of bands in a gel strip, H (DGGE), and the type and mol% PLFA in a sample, H (PLFA). The Simpson index (λ) was subtracted from 1 to give a D value that ranged from 0 to 1. Table 1 presents the diversity value H at the two sampling times and in the vertical profile. At 8:00 a.m., H (PLFA) and H (DGGE) were higher in samples taken from a depth of 2.5–7 mm, with the exception of the 3–3.5 mm samples, in which the overall diversity was slightly decreased. As Hedrick et al. (2000) observed in previous studies, the species richness calculated based on DGGE agreed well with that calculated from the PLFA profiles. Nevertheless, when using a combination of methods to measure microbial diversity, the data should be adjusted to a common sensitivity to avoid differences in biomass between samples (Kirk et al. 2004).
The D (PLFA) value calculated from the Simpson index was higher in the 2.5–3.5-mm samples; however, the D (DGGE) values were similar in all samples. Indeed, the Simpson index is relatively less sensitive to richness than the Shannon diversity index and is more sensitive to differences in species comprising the community, thus placing more weight on common species (Simpson 1949). Moreover, the evenness index of PLFA in the morning was higher in samples from a depth of 5–8.75 mm and 3–3.5 mm. This finding was coincident with the lower values of H (PLFA) and H (DGGE), and the higher values of D (PLFA), which, in turn, are related to a decrease in richness (variety, number of species) and an increase in the frequency of certain phylotypes (relative distribution). The increase in evenness could be assessed in the DGGE gels (Fig. 1), where there was a reduction in the predominant bands as well as an ‘unresolved’ smear of DNA fragments consistent with a more even distribution of microorganisms in the sample. By contrast, the evenness index calculated based on DGGE data did not agree with the PLFA index, since similar values were reported for all samples expect for a reduction in those acquired at 5–8.75 mm.
In the afternoon (3:00 p.m.), there was good agreement between H (PLFA) and H (DGGE) in the 2.5–4 mm samples, and both indices indicated increased richness (number of phylotypes). The H (DGGE) value was higher than the H (PLFA) value at 6.5–7.5 mm due to the increasing dominance of certain bands in the gel and the appearance of others that were not recovered for sequencing purposes (unresolved patterns and overlapping of bands). Moreover, D (DGGE) was not informative because the values were similar in all samples and in the 8:00 a.m. profile. However, D (PLFA) values were higher in samples taken at the very top of the mat, which was indicative of a more even distribution of the members in the system. A similar observation was made for samples taken at 3:00 p.m. and analyzed by DGGE (Fig. 1b), i.e., rather than a relative predominance of bands there were several bands with a comparatively more even abundance. The evenness of the PLFA and DGGE data at this sampling time was higher at the bottom layers of the mat and also at 2.5–4 mm, coincident with an increase in the D/D max values for PLFA. This confirms the convenience of using the D/D max index as a measure to provide information comparable to that resulting from the Shannon–Wiener function, and the index showed good agreement of the DGGE and PLFA diversity values in most cases. It is also essential to note the importance of complementary diversity indices in reflecting richness and evenness in this kind of study, in order to avoid the conclusion that rare species strongly influence the magnitude of the diversity index itself (Margalef 1958).
The Shannon PLFA index was indicative of the contribution of anaerobic microorganisms (branched monoenoics and mid-branched saturated fatty acids) (Table 2). The diversity of anaerobes at 4–5.5 mm in the B samples was high, whereas the A samples indicated a similar diversity in all samples with a moderately increased H (PLFA anaerobes) at the topmost layers and in the deepest samples. In this case, the diversity data were coincident with the anaerobic character of the mat system at night and early in the day followed by the diversification of strict anaerobes and fermenters, which contribute to the carbon cycle by recycling the photosynthates that derive from autotrophic members of the mat (Ollivier et al. 1994). The similarity of the H and D diversity indices in all samples suggested the stable maintenance of a structurally diverse microbial community. Previous studies performed by Wieland et al. (2005) in Camargue mat samples taken over a diel cycle detected higher sulfate reduction rates during the day and below 1 mm depth. Those studies also observed anoxic conditions from 1.5 mm depth at 16:33 p.m. as well as an important decay in the pH values from 9.25 to 6.7 from 1.5 to 2.5 mm depth at the same sampling time. In our case, the increased diversity observed at 3:00 p.m. of 2.5–4 mm can be mostly explained by diversification of anaerobic populations, which is coincident with the higher sulfate reduction rates detected in the studies mentioned above. Besides, changes in the pH gradient might contribute to a minor extent inducing reduction in the microbial diversity overlying 2.5 mm depth and then diversification of the anaerobic populations in the deepest layers, which remain anoxic and close to neutral pH until the following morning.
Conclusions
These results demonstrate that depth-related differences determined by PLFA and DGGE divergence indices have a greater influence on diversity than temporal variations and reflected established microniches. Moreover, diversity indices data suggested the stable maintenance of a structurally diverse microbial community. Apart from that, DGGE analysis of microbial mat samples detected temporal differences of certain microbial groups as well as vertical migrations over time. In general, the data presented in this study suggest that microbial mat diversity remain apparently stable over a period of hours during the daily cycle with exception of certain microorganisms experiencing vertical migrations and changes in the abundance of other microbial groups especially after events of intense photosynthetic activity inducing the stratification of the community (Garcia-Pichel et al. 1994).
Our results are consistent with the conclusion that depth-related differences have greater influence on diversity than temporal variations. Although the findings of this study provide insight into the changes in microbial diversity in mats, they also highlight the need for new diversity indices that can incorporate data derived from different methodological approaches (Hughes et al. 2001). The information presented here is subject to the potential limitations of the presented methods; nonetheless, it can be used for quantitative studies of mat ecosystems under different environmental conditions. Microbial mats were classically considered to be microbial ecosystems low in diversity (Des Marais 1990), but recent studies have revealed that they are made up of a large number of species and are therefore “hot spots” of microbial diversity (Ley et al. 2006). The circumstances giving rise to the great diversity detected in mat systems and the relationships between the various members should be investigated in future studies. The results will further our knowledge of the first stable microbial ecosystems on Earth (Allwood et al. 2006).
References
Allwood AC, Walter MR, Kamber BS, Marshall CP, Burch IW (2006) Stromatolite reef from the Early Archaean era of Australia. Nature 441:714–718
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Begon M, Harper JL, Towsend CR (1990) Ecology–individuals, populations, communities. Blackwell scientific publications, Oxford, UK
Benthien M, Wieland A, García de Oteyza T, Grimalt JO, Kühl M (2004) Oil-contamination effects on a hypersaline microbial mat community (Camargue, France) as studied with microsensors and geochemical analysis. Ophelia 58:135–150
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37:911–917
Brosius J, Dull TL, Sleeter DD, Noller HF (1981) Gene organisation and primary structure of a ribosomal RNA operon from Escherichia coli. J Mol Biol 148:107–127
Casamayor EO, Pedrós-Alió C, Muyzer G, Amann R (2002) Microheterogeneity in 16S ribosomal DNA-defined bacterial populations from stratified planktonic environment is related to temporal changes and to ecological adaptations. Appl Environ Microbiol 68:1706–1714
Caumette P, Matheron R, Raymond N, Relexans JC (1994) Microbial mats in the hypersaline ponds of Mediterranean salterns (Salins-de-Giraud, France). FEMS Microbiol Ecol 13:273–286
Chang YJ, Stephen JR, Richter AP, Venosa AD, Bruggemann J, Macnaughton SJ, Kowalchuk GA, Haines JR, Kline E, White DC (1999) Phylogenetic analysis of aerobic freshwater and marine enrichment cultures efficient in hydrocarbon degradation: effect of profiling method. J Microbiol Methods 40:19–31
Des Marais DJ (1990) Microbial mats and the early evolution of life. Trends Ecol Evol 5:140–144
Dunbar J, Takala S, Barns SM, Davis JA, Kuske CR (1999) Levels of bacterial community diversity in four arid soils compared by cultivation and 16S rRNA gene cloning. Appl Environ Microbiol 65:1662–1669
Eichner CA, Erb RW, Timmis KH, Wagner-Dögler I (1999) Thermal gradient gel electrophoresis analysis of bioprotection from pollutant shocks in the activated sludge microbial community. Appl Environ Microbiol 65:102–109
Fourçans A, García de Oteyza T, Wieland A, Solé A, Diestra E, van Bleijswijk J, Grimalt JO, Kühl M, Esteve I, Muyzer G, Caumette P, Duran R (2004) Characterization of functional bacterial groups in a hypersaline microbial mat community (Salins-de-Giraud, Camargue, France). FEMS Microbiol Ecol 51:55–70
Fourçans A, Sole A, Diestra E, Ranchou-Peyruse A, Esteve I, Caumette P, Duran R (2006) Vertical migration of phototrophic bacterial populations in a hypersaline microbial mat from Salins-de-Giraud (Camargue, France). FEMS Microbiol Ecol 57:367–377
Fromin N, Hamelin J, Tarnawski S, Roesti D, Jourdain-Miserez K, Forestier N, Teyssier-Cuvelle S, Gillet F, Aragno M, Rossi P (2002) Statistical analysis of denaturing gel electrophoresis (DGE) fingerprinting patterns. Environ Microbiol 4:634–643
Garcia-Pichel F, Mechling M, Castenholz RW (1994) Diel migrations of microorganisms within a benthic, hypersaline mat community. Appl Environ Microbiol 60:1500–1511
Guckert JB, Antworth CP, Nichols PD, White DC (1985) Phospholipid, ester-linked fatty acid profiles as reproducible assays for changes in prokaryotic community structure of estuarine sediments. FEMS Microbiol Ecol 31:147–158
Guerrero R, Piqueras M, Berlanga M (2002) Microbial mats and the search for minimal ecosystems. Int Microbiol 5:177–188
Hedrick DB, Peacock A, Stephen JR, Macnaughton SJ, Brüggeman J, White DC (2000) Measuring soil microbial community diversity using polar lipid fatty acid and denaturing gradient gel electrophoresis data. J Microbiol Methods 41:235–248
Hiraishi A (1999) Isoprenoid quinones as biomarkers of microbial populations in the environment. J Biosci Bioeng 88:449–460
Hughes JB, Hellmann JJ, Ricketts TH, Bohannan BJM (2001) Counting the uncountable: statistical approaches to estimating microbial diversity. Appl Environ Microbiol 67:4399–4406
Ibekwe AM, Papiernik SK, Gan J, Yates SR, Yang CH, Crowley DE (2001) Impact of fumigants on soil microbial communities. Appl Environ Microbiol 67:3245–3257
Iwasaki M, Hiraishi A (1998) A new approach to numerical analyses of microbial quinone profiles in the environment. Microbes Environ 13:67–76
Kirk JL, Beaudette LA, Hart M, Moutoglis P, Klironomis JN, Lee H, Trevors JT (2004) Methods of studying soil microbial diversity. J Microbiol Methods 58:169–188
Krebs CJ (1985) Species diversity. In: Krebs CJ (ed) Ecology: the experimental analysis of distribution and abundance. Harper and Row, New York, pp 507–534
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Ley RE, Harris JK, Wilcox J, Spear JR, Miller SR, Bebout BM, Maresca JA, Bryant DA, Sogin ML, Pace NR (2006) Unexpected diversity and complexity of the Guerrero Negro hypersaline microbial mat. Appl Environ Microbiol 72:3685–3695
Ludwig JA, Reynolds JF (1988) Statistical ecology: I. Primer on methods and computing. Wiley-Interscience, New York, p 96
Macnaughton SJ, Stephen JR, Venosa AD, Davis GA, Chan YJ, White DC (1999) Microbial population changes during bioremediation of an experimental oil spill. Appl Environ Microbiol 65:3566–3574
Margalef R (1958) Information theory in ecology. Gen Syst 3:36–71
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Navarrete A, Peacock A, Macnaughton SJ, Urmeneta J, Mas-Castellà J, White DC, Guerrero R (2000) Physiological status and community composition of microbial mats of the Ebro delta, Spain, by Signature Lipid Biomarkers. Microb Ecol 39:92–99
Navarrete A, Urmeneta J, Cantu JM, Vegas E, White DC, Guerrero R (2004) Signature lipid biomarkers of microbial mats of the Ebro delta (Spain), Camargue and Étang de Berre (France): an assessment of biomass and activity. Ophelia 58:175–188
Nübel U, Garcia-Pichel F, Kühl M, Muyzer G (1999) Quantifying microbial diversity: morphotypes, 16S rRNA genes, and carotenoids of oxygenic phototrophs in microbial mats. Appl Environ Microbiol 65:422–430
Ollivier B, Caumette P, García JL, Mah RA (1994) Anaerobic bacteria from hypersaline environments. Microbiol Rev 58:27–38
Paerl HW, Pinckney JL, Steppe TF (2000) Cyanobacterial–bacterial mat consortia: examining the functional unit of microbial survival and growth in extreme environments. Environ Microbiol 2:11–26
Ringelberg DB, Davis JD, Smith GA, Pfiffner SM, Nichols PD, Nickels JS, Henson JM, Wilson JT, Yates M, Kampbell DH, Reed HW, Stocksdale TT, White DC (1988) Validation of signature phospholipids fatty acid biomarkers for alkaline-utilizing bacteria in soils and subsurface aquifer materials. FEMS Microbiol Ecol 62:39–50
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sekiguchi H, Watanabe M, Nakahara T, Xu B, Uchiyama H (2002) Succession of bacterial community structure along the Changjiang River determined by denaturing gradient gel electrophoresis and clone library analysis. Appl Environ Microbiol 68:5142–5150
Shannon CE, Weaver W (1963) The mathematical theory of communications. University of Illinois Press, Urbana
Simpson EH (1949) Measurement of diversity. Nature 163:688
Sørensen KB, Canfield DE, Teske AP, Oren A (2005) Community composition of a hypersaline endoevaporitic microbial mat. Appl Environ Microbiol 71:7352–7365
Stephen JR, Chang YJ, Gan YD, Peacock A, Pfiffner SM, Barcelona MJ, White DC, Macnaughton SJ (1999) Microbial characterization of a JP-4 fuel-contaminated site using a combined lipid biomarker/polymerase chain reaction-denaturing gradient gel electrophoresis (PCR-DGGE)-based approach. Environ Microbiol 1:231–241
Tankéré SPC, Bourne DG, Muller FLL, Torsvik V (2002) Microenvironments and microbial community structure in sediments. Environ Microbiol 4:97–105
Torsvik V, Øverås L (2002) Microbial diversity and function in soil: from genes to ecosystems. Curr Opin Microbiol 5:240–245
Torsvik V, Øverås L, Thingstad TF (2002) Prokaryotic diversity–magnitude, dynamics and controlling factors. Science 269:1064–1066
Villanueva L, Navarrete A, Urmeneta J, White DC, Guerrero R (2004) Combined phospholipid biomarker-16S rRNA gene denaturing gradient gel electrophoresis analysis of bacterial diversity and physiological status in an intertidal microbial mat. Appl Environ Microbiol 70:6920–6926
White DC, Bobbie RJ, Heron JS, King JD, Morrison SJ (1979) Biochemical measurements of microbial mass and activity from environmental samples. In: Costerton JW, Colwell RR (eds) Native aquatic bacteria: enumeration, activity and ecology, ASTM STP 695. American Society for Testing and Materials, Philadelphia, pp 69–81
White DC, Findlay RH (1988) Biochemical markers for measurements of predation effects on the biomass, community structure, nutritional status, and metabolic activity of microbial biofilms. Hydrobiologia 159:119–132
White DC, Ringelberg DB (1997) Utility of the signature lipid biomarker analysis in determining in situ microbial biomass, community structure and nutritional/physiological status of deep subsurface microbiota. In: Amy PS, Haldeman DL (eds) The microbiology of the terrestrial subsurface. CRC, Boca Raton, pp 117–134
Wieland A, Kühl M (2000) Irradiance and temperature regulation of oxygenic photosynthesis and O2 consumption in a hypersaline cyanobacterial mat (Solar Lake, Egypt). Mar Bio 137:71–85
Wieland A, Zopfi J, Benthien M, Kühl M (2005) Biogeochemistry of an iron-rich hypersaline microbial mat (Camargue, France). Microb Ecol 49:34–49
Wintzingerode FV, Göbel UB, Stackebrandt E (1997) Determination of microbial diversity in environmental samples: pitfalls of PCR-based rRNA analysis. FEMS Microbiol Rev 21:213–229
Acknowledgments
This paper is dedicated to the memory of David C. White: “Thank you for being a friend and mentor, we will always remember the wonderful times we spent together”. We thank Mercè Piqueras and Wendy Ran for useful suggestions. We are grateful to the Center for Biomarker Analysis (TN, USA) staff for advice and technical assistance. This research was supported by Spanish MCyT grant BOS2002-02944 and MEC CGL2005-04990, and by grant DE-FC02-96ER62278, from the Office of Biological and Environmental Research and the Natural and Accelerated Bioremediation Research Program. LV was recipient of a scholarship from the Spanish MECD (AP2001-0953).
Author information
Authors and Affiliations
Corresponding author
Additional information
Dedicated to the memory of David C. White.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Villanueva, L., Navarrete, A., Urmeneta, J. et al. Analysis of diurnal and vertical microbial diversity of a hypersaline microbial mat. Arch Microbiol 188, 137–146 (2007). https://doi.org/10.1007/s00203-007-0229-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-007-0229-6