Abstract
Nitrosomonas europaea, as an ammonia-oxidizing bacterium, has a high Fe requirement and has 90 genes dedicated to Fe acquisition. Under Fe-limiting conditions (0.2 μM Fe), N. europaea was able to assimilate up to 70% of the available Fe in the medium even though it is unable to produce siderophores. Addition of exogenous siderophores to Fe-limited medium increased growth (final cell mass). Fe-limited cells had lower heme and cellular Fe contents, reduced membrane layers, and lower NH3- and NH2OH-dependent O2 consumption activities than Fe-replete cells. Fe acquisition-related proteins, such as a number of TonB-dependent Fe-siderophore receptors for ferrichrome and enterobactin and diffusion protein OmpC, were expressed to higher levels under Fe limitation, providing biochemical evidence for adaptation of N. europaea to Fe-limited conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ammonia-oxidizing bacteria, such as Nitrosomonas europaea, are important participants in the global N cycle because they transform ammonia (NH3) to nitrite (NO −2 ), the first step in nitrification. As lithotrophs, these bacteria derive all their energy and reductant necessary for growth from the oxidation of NH3. As autotrophs, these bacteria can meet their C requirements solely using carbon dioxide (CO2). N. europaea is a member of the β-proteobacteria and has a single circular chromosome of 2,812,094 base pairs with 2,460 protein-encoding genes (Chain et al. 2003). In N. europaea, the genes encoding transporters of inorganic ions are plentiful, but those encoding transporters of organic compounds are scant, consistent with the life style of this bacterium.
Iron (Fe) is an essential cofactor for the transfer of electrons in many enzymes participating in energy-generating pathways. However, biological availability of Fe is typically low because in nature Fe exists mostly in the form of insoluble oxy-hydroxide complexes, and in the cell Fe is present mostly in iron-binding proteins (Andrews et al. 2003). To cope with the low biological Fe supply, bacteria have evolved high-affinity Fe transport systems for scavenging Fe from the environment, ways to store Fe intracellularly, and mechanisms to down-regulate certain high Fe-containing proteins to economize Fe needs (Andrews et al. 2003).
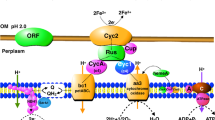
In bacteria there are two common mechanisms for efficient Fe acquisition (Kammler et al. 1993): (a) direct uptake through ferrous (Fe2+) uptake systems or through heme and hemoprotein uptake systems, and (b) a more prevalent mechanism, the production and excretion of siderophores that bind free ferric iron (Fe3+) or sequester Fe3+ from other Fe3+-binding proteins (Braun and Killmann 1999; Clarke et al. 2001; Faraldo-Gomez and Sansom 2003). The uptake of Fe3+-loaded siderophores in Gram-negative bacteria requires a sophisticated transport system. Outer membrane (OM) receptors capture Fe3+-loaded siderophores and translocate them into the periplasmic space. This process requires energy (proton-motive force) that is mediated by the TonB/ExbBD complex located in the cytoplasmic membrane. Fe-loaded siderophores are then transported across the periplasm by periplasmic-binding proteins, and finally into the cytoplasm by ATP-dependent membrane transporters (Clarke et al. 2001; Faraldo-Gomez and Sansom 2003).
With the exception of genes for the synthesis of citrate, N. europaea notably lacks genes for siderophore production (Chain et al. 2003). In contrast, up to 4% of the N. europaea genes code for Fe-related transport systems, including the uptake of Fe3+-loaded siderophores. Congruent with the high number of genes related to Fe acquisition, over 2% of the genes code for heme synthesis and heme-containing proteins and for proteins with Fe–S centers, reflecting the importance of Fe to the energy metabolism of N. europaea (Whittaker et al. 2000). How N. europaea acquires sufficient Fe for its metabolism without the production and excretion of siderophores is intriguing. We hypothesize that N. europaea has multiple avenues for Fe acquisition, including ferric and possibly ferrous uptake systems and the uptake of siderophores produced by other organisms. Despite the importance of Fe to ammonia-oxidizing bacteria, Fe nutrition has not been studied in this group of bacteria. In this work we determined the Fe requirements for N. europaea and its physiological response to Fe stress.
Materials and methods
Bacterial culture
Nitrosomonas europaea (ATCC 19178) was cultured as described (Ensign et al. 1993; Stein and Arp 1998) and harvested in mid to late exponential phase. The standard (Fe-replete) medium contained 10 μM Fe3+ complexed with EDTA. Medium made from reagent grade chemicals without any addition of Fe salts still contained about 0.2 μM Fe (Fe-limited medium). The Fe-free medium was made by treatment with the metal chelator Chelex (Sigma, St. Louis, MO, USA) followed by the addition of MgSO4, CaCl2 and CuSO4 to the concentrations of untreated medium. Cells grew normally when Fe (10 μM) was added to the Fe-free medium. All glassware was soaked in 1.0% HNO3 overnight and rinsed thoroughly with double deionized water. Because medium containing 0.1 μM Fe could not provide sufficient cell material for analyses, medium containing 0.2 μM Fe was adopted for all Fe-limiting experiments. Nitrite (NO −2 ) concentration was determined colorimetrically with the Griess reagent (sulfanilamide and N-naphthylethylenediamine) (Hageman and Hucklesby 1971). The accumulation of NO −2 is consistently proportional to the increase in cell mass during growth.
Determination of Fe, siderophore and citrate concentrations
Fe content was determined by the ferrozine assay following digestion of cells or cellular fractions in 5% HNO3 overnight at 100°C (Carter 1971). Concentrations lower than 10 μM Fe were determined by Inductively Coupled Plasma Mass Spectrometry (ICP-MS) as described (Houk 1994). The quantification of total siderophore content in culture supernatants was done by the universal siderophore assay as described (Schwyn and Neilands 1987; Payne 1994). Citrate was determined using the Citric Acid Bioanalysis Kit (R-Biopharm, Darmstadt, Germany).
Electron microscopy
Washed N. europaea cells were fixed in glutaraldehyde and formaldehyde, and post-fixed in OsO4. After the bacterial cells were embedded and sectioned, the grids were stained with uranyl acetate and Pb citrate or as previously described (Murray and Watson 1965). The thin sections of cells were examined on a Philips CM-12 STEM transmission electron microscope, operated at 60 kV.
Enzyme activity determinations
Whole cell NH3-dependent and hydroxylamine-dependent O2 uptake activities were measured as described (Shiemke et al. 2004). HAO in vitro activity was determined by the hydroxylamine-dependent 2,6-dichlorophenol–indophenol (dcPIP; the electron acceptor) reduction assay (Hooper 1969). Malate dehydrogenase activity in membrane-free extracts was determined by monitoring NADH oxidation in the presence of oxaloacetic acid. NADH concentration changes were monitored on a Beckman DU640 Spectrophotometer using the kinetics mode at 340 nm and the extinction coefficient 6.22 l mmol−1 cm−1 (Thompson et al. 1998).
Cell fractionation, membrane preparation, and heme and protein quantification
Cell membranes were prepared by breaking cells via sonication at 30% energy output for 10 s on Ultrasonic Processor XL202 sonicator (Heat Systems-Ultrasonics, Inc., Farmingdale, NY, USA), followed by ultracentrifugation at 40,000 rpm (rotor SW50-1) for 1 h on a Beckman L8-70 Ultracentrifuge (Beckman Instrument, Inc., Fullerton, CA, USA). Crude membrane preparations were washed by thorough homogenization in Tris buffer (0.1 M; pH 7.8) containing 1 M KCl, collected again by ultracentrifugation and resuspended in Tris buffer (50 mM; pH 7.8). Simultaneously with the membrane preparation, soluble fractions were collected for heme content determination, and for protein content and peptide composition analysis. Heme content was quantified after extraction with pyridine as described (Berry and Trumpower 1987). Protein content was estimated by the Micro BCA Protein Assay Kit (Pierce, Rockford, IL, USA).
Protein gel electrophoresis and mass spectrometry analysis
Peptide composition was analyzed using polyacrylamide gel electrophoresis (PAGE) as described (Hyman and Arp 1993). For protein identification by tandem mass spectrometry (MS/MS), selected peptide bands (stained with Coomassie Brilliant Blue R 250) were excised and destained in water followed by 50% acetonitrile-50 mM bicarbonate. Gel slices were dehydrated with 100% acetonitrile and dried under vacuum. Gel slices were rehydrated with trypsin solution (Promega, Madison, WI, USA) and digested at 37°C for 16 h. After digestion, peptides were extracted three times with 250 μl 0.1% trifluoroacetic acid-60% acetonitrile. The extracts were pooled for mass spectrometric analysis. The data were acquired using data-dependent HPLC-MS/MS. Peptide extracts were run on a Waters Q-TOF (Time of Flight) Ultima Global (Milford, MA, USA). The peptides were eluted from a PicoFrit column (New Objective Inc., Cambridge, MA, USA) with a slow increase in concentration of acetonitrile in 0.1% formic acid. As the peptides eluted from the column, the mass and charge state was determined by the data system, and instrument conditions were adjusted to fragment the peptide “parent” ion. The data in the MS/MS spectra were searched using Mascot (Matrix Science, London, UK) against a general protein database and the N. europaea sequence database (Chain et al. 2003), and the theoretical MS/MS spectra were compared to the experimental spectra (significance threshold was 0.03 and Mascot score of 25).
Results
Fe requirements and physiological responses to Fe limitation
Nitrosomonas europaea was grown in media containing a wide range of Fe concentrations to define its Fe requirements. The maximum cell mass attained at 0.1 and 0.2 μM Fe was about 30 and 60%, respectively, of the cell mass attained in standard medium containing 10 μM Fe (Fig. 1a). N. europaea acquires Fe at these low Fe concentration, even though it does not have the genes necessary for siderophore synthesis. An Fe concentration below 1 μM is generally considered a low Fe condition, and Fe at 0.1 μM, an Fe-deficient condition (Neilands 1981). Concentrations ranging from 10 to 250 μM Fe resulted in normal growth of N. europaea as judged by culture optical densities. Growth was inhibited with 1,200 μM Fe, probably due to Fe toxicity and consequent oxidative damage to the cells (Keyer et al. 1995).
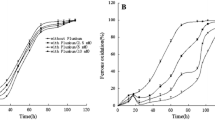
Nitrosomonas europaea growth as affected by various Fe concentrations and by the siderophore desferal. a Growth (optical density) in media containing 10 (open circle), 0.2 (filled circle), 0.1 (open inverted triangle), and < 0.01 (filled inverted triangle) μM Fe (Chelex treated medium). The 10 μM Fe culture was inoculated with Fe-replete cells, the three other cultures were inoculated with cells grown in Fe-limited medium. b Effect of desferal on the growth of N. europaea in Fe-limited medium. Growth curves shown are in Fe-limited (0.2 μM) medium without desferal (filled circle) and Fe-limited medium containing 5.0 μM desferal (open circle). The data presented are the average of duplicates with < 10%variation
The growth rates and maximum cell masses obtained for N. europaea in media initially made with 10 μM non-complexed ferric iron (FeCl3, Fe3+), or Fe3+-EDTA, or ferrous iron (FeSO4, Fe2+), were similar. Addition of up to 40 μM α, α′-dipyridyl (Sigma), a strong Fe2+ chelator commonly used to create Fe deficient conditions (Chart et al. 1986), did not inhibit N. europaea growth in medium containing 0.2 μM Fe. This result indicates that N. europaea can use Fe3+, the predominant form of Fe in the medium under aerobic condition at near neutral pH, rather than rely on trace amounts of Fe2+ for growth. However, this result does not rule out the possibility of N. europaea taking up Fe2+.
Fe concentrations below 1.0 μM often trigger siderophore synthesis in other bacteria (Neilands 1995). Early stationary phase culture supernatants were collected from N. europaea and analyzed for siderophore content. As expected, no siderophores were detected in the media under Fe-limiting or Fe-replete conditions. As a positive control, the same method was successfully applied to the detection and quantification of siderophores in an Azotobacter vinelandii Fe-limiting culture (data not shown).
Since N. europaea has the necessary genes to produce citrate, we considered citrate a candidate Fe siderophore. Citrate was detected in culture filtrates from both Fe-limited and Fe-replete cultures. Culture filtrates from Fe-sufficient cultures had a higher concentration of citrate (∼5 μM) than those from low-Fe cultures (∼3 μM). Citrate concentrations were approximately proportional to the cell densities of the two cultures.
Promotion of N. europaea growth by exogenous siderophores
Addition of the siderophore desferal (desferrioxamine mesylate; Sigma) to the Fe-limited medium led initially to a slower growth rate (Fig. 1b). However, after 3 days, the growth rate increased, and by the fifth day cell densities in the desferal-containing medium exceeded those in the medium without desferal (Fig. 1b). Addition of 20 μM desferal had a similar effect as 5 μM. Siderophore ferrichrome (10 μM) (Sigma) increased N. europaea growth (final cell mass) in Fe-limited medium to a similar extent as desferal but without a long lag phase (data not shown). Growth was also observed in Fe-limited medium containing 10 μM enterobactin, a catechol-type siderophore (data not shown). Addition of citric acid to either Fe-limited or Fe-replete media did not lead to significant increases in growth (data not shown). The increases in cell densities in the presence of desferal, ferrichrome and enterobactin suggest that N. europaea has the potential to utilize external siderophores for Fe uptake under Fe limitation.
Physiological and ultrastructural changes of N. europaea upon Fe limitation
Medium with 0.2 μM Fe yielded about 60% cell mass of the Fe-replete medium and affected cell appearance and physiology. The color of cells harvested from Fe-limited media was lighter and the heme content of the Fe-limited cells was over threefold lower than that of Fe-replete cells (Table 1). Furthermore, Fe-limited cells had nearly ten-fold less Fe than Fe-replete cells (Table 1). About 70% of the Fe initially in the medium was taken up and incorporated into Fe-limited cells during growth (Table 1). Analyses of the spent media and cellular Fe contents by ICP-MS confirmed this result. Heme-bound Fe accounted for about 85% of Fe in the soluble fraction of Fe-limited cells and 59% in Fe-replete cells (Table 1). Fe limitation influenced both the amount of soluble cytochromes produced and the proportion of Fe distributed to cytochromes.
As indicators of the overall cell activity, NH3- and NH2OH-dependent O2 uptake rates and in vitro HAO and malate dehydrogenase activities were measured. The NH3- and NH2OH-dependent O2 uptake (AMO and HAO) activities in whole cells were about twofold lower in Fe-limited cells than in Fe-replete cells (Table 2). HAO activity in cell-free extracts was also lower in cells from Fe-limited medium than cells from normal medium. Malate dehydrogenase was chosen as a representative of non Fe-containing enzymes. In contrast to the decrease of NH3- and NH2OH-dependent O2 uptake activities, specific malate dehydrogenase activity of Fe-limited cells was about fourfold higher than that of Fe-replete cells (Table 2).
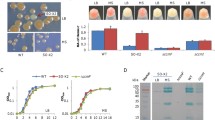
Nitrosomonas cells are characterized by extensive membrane invaginations (Hooper et al. 1972). AMO is located in the cell’s inner membranes (Arp et al. 2002). Many other Fe-containing proteins such as cytochromes and electron transport enzymes are also localized in the membranes. We hypothesized that Fe would be an important factor affecting N. europaea membrane content and structure. To test this, Fe-limited and Fe-replete cells were fixed, stained, and sectioned for electron microscopic examination. The internalized membranes of the cells from Fe-limited medium had fewer layers and were less regularly stacked than cells cultured in Fe-replete medium (Fig. 2). Many of the Fe-limited cells were irregular in shape (images not shown).
Transmission electron microscopic images of sections of N. europaea cells grown in a 10 μM Fe medium and b 0.2 μM Fe medium. Note the internalized membranes or invaginations. Transmission electron microscope images were taken at ×72,500 magnification. Images shown are representative of dozens examined. The line in the picture represents a length of 0.1 μm (micron)
Genomic analysis of N. europaea Fe acquisition genes
Nitrosomonas europaea has 90 genes (∼4% of the total genes) related to Fe uptake. Of these, 42 encode OM siderophore receptors (Table 3), a relatively large number compared to 11 genes in the chemolithoautotroph Acidithiobacillus ferroxidans (Quatrini et al. 2005), or six in the heterotroph E. coli and 35 genes in the heterotroph Pseudomonas aeruginosa (Andrews et al. 2003). There are 22 sets of genes with the organization similar to the Fe regulatory and uptake system, i.e. fecI/fecR/fecA (σ factor/anti σ factor/TonB-dependent OM receptor gene) (Table 3). All genes in this group, with the exception of NE1086/85/87, exist in the order fecI/fecR/fecA. Some have intervening, unrelated genes in between the Fe uptake genes. This gene organization is similar to the fecIR and fecABCDE (Fe-dicitrate) systems in E. coli (Visca et al. 2002), and to the fpvIR and fpvA/pvd (pyoverdin) system in P. aeruginosa (Ravel and Cornelis 2003). Unlike the fec system in E. coli, there are no Fe ABC transporter genes in these N. europaea operons. Twenty additional Fe-siderophore OM receptor genes are not immediately adjacent to σ-factor/anti σ-factor genes, although several of these genes (e.g. NE1088, NE1089, Table 3) are in proximity to other OM receptor genes that are adjacent to σ- and anti σ-factor genes. Of the 42 putative OM receptor genes, 13 are either truncated, interrupted by IS elements, or are pseudogenes due to frameshifts (Table 3). Surprisingly, N. europaea lacks genes with high similarity to the E. coli fecA (OM ferric dicitrate receptor); the closest one, NE2433, has only 24% similarity, but this gene is more similar to ferrichrome or catechol receptor gene fiuA. A wide plethora of putative siderophore receptors are encoded in N. europaea: some are receptors for siderophores such as ferrichrome, ferrioxamine, coprogen, pyoverdin, and many others for catechol/catecholate-type siderophores (for siderophore types, see Matzanke 1991). Similar to other Gram-negative bacteria (Andrews et al. 2003), genes for Fe ABC transporter and the energy-transducing TonB–ExbBD complex are present in the genome. Other genes related to Fe nutrition include those encoding Fe-storage protein bacterioferritin, bacterioferritin comigratory protein, and ferric uptake regulators (Fur). N. europaea appears to have genes to encode the many components necessary for efficient Fe uptake and storage.
Expression of membrane receptor proteins under Fe-limiting condition
To study whether the Fe acquisition genes are expressed under Fe starvation condition, membrane fractions of cells grown in Fe-limited and Fe-replete media were isolated and their protein compositions compared by SDS-PAGE. Several proteins were expressed about tenfold higher in cells grown in Fe-limited than in Fe-replete media (gels not shown). The comparison was also made between the proteins of the soluble fractions of the Fe-replete cells and Fe-limited cells, but no major differences were observed. The membrane proteins differentially expressed were excised from the gels and subjected to HPLC/MS/MS analysis. Of the top 12 proteins identified by tandem MS, seven are clearly related to Fe acquisition (Table 4). Six of the proteins that were highly up-regulated under the Fe-limiting condition are OM siderophore receptors. Three are putative receptors for catechol-type siderophores, one is for ferrichrome (NE1089), one similar to heme receptor, and one is for unidentified siderophores. The expression of the ferrichrome receptor in absence of any siderophore can explain why there was no lag phase for N. europaea grown in medium containing ferrichrome. In addition, an OM protein OmpC (a general diffusion Gram-negative porin), a multicopper oxidase, and a type II secretion pathway protein were also identified among the highly expressed proteins in Fe-limited cells (Table 4). Only one siderophore receptor, encoded by NE1532, was detected uniquely in Fe-replete cells, albeit a much larger amount of total membrane protein was required for isolating enough samples for the MS/MS detection.
Discussion
Despite its lack of siderophore production, N. europaea can grow moderately well at low Fe concentrations (0.2 μM) (Fig. 1a). In order to grow at 0.2 μM Fe, many other microorganisms rely on siderophores for Fe uptake. For instance, in E. coli (Neilands 1995) and Fusarium venenatum (Wiebe 2002) siderophore synthesis was turned on when Fe was below 1 μM. In some species that require high Fe availability, siderophore synthesis is turned on even when Fe was well above 1 μM. For example, when Fe level was around 5 μM, A. vinelandii and Magnetospirillum magneticum AMB-1 started to produce siderophores (Tindale et al. 2000; Calugay et al. 2003). Pseudomonas species usually become Fe-deficient at 1–2 μM (Hofte et al. 1993; Mossialos et al. 2000), even though most of them can produce siderophores.
Nitrosomonas europaea has a high cellular Fe and heme content (Table 1) relative to E. coli. This is consistent with N. europaea’s many heme proteins that include HAO, heme/copper type cytochrome oxidase, cytochrome peroxidase, and other cytochromes (e.g. c554, cm552, p460) (Whittaker et al. 2000; Upadhyay et al. 2003). N. europaea cultured in Fe-limited medium had a cellular Fe concentration 19-fold higher (1.88 mM) than the level reported for E. coli (0.1 mM) cultured in minimal medium (∼0.1 μM Fe) (Outten and O’Halloran 2001). N. europaea grown in Fe-replete medium had a cellular Fe concentration 80-fold higher (16.3 mM) than that for E. coli cells (0.2 mM) grown in LB medium (∼2.0 μM Fe) (Outten and O’Halloran 2001). The heme C content (0.9 nmol/unit OD600 ml) in the soluble fraction of N. europaea from Fe-replete medium (Table 1) was much greater than ∼0.1 (nmol/unit OD600 ml) detected in E. coli grown in full Fe medium (Verderber et al. 1997). This high heme content is evident in the distinct reddish-brown color of N. europaea cells, compared to the pale yellowish color of E. coli cells.
Differences in internal membrane content and structure were apparent between Fe-limited and Fe-replete cells. An early study showed that Nitrosomonas cell membranes contain a large proportion of Fe-containing proteins (A-type cytochromes and electron transport enzymes) (Hooper et al. 1972). Cellular Fe content of Fe-limited cells was much lower than that of Fe-replete cells (Table 1). This lower Fe content probably reflects lower Fe storage in addition to the lower heme content. However, the cell density in Fe-limited medium was not affected to the same degree as the Fe and heme contents. These results suggest that when Fe supply is limited, N. europaea allocates a greater proportion of total Fe to the most critical molecules such as hemes. The lower specific activities of AMO and HAO in Fe-limited cells than in Fe-replete cells (Table 2) might be due to the lowered available heme in Fe-limited cells (Table 1) because HAO requires 24 hemes per enzyme for function (Arciero and Hooper 1993). The reason for the elevated malate dehydrogenase activity in Fe-limited cells is not clear at present, but this result indicated that not all metabolic activities were down-regulated under Fe limitation.
Other studies have showed the increased production of certain OM proteins upon Fe deficiency in other bacteria (see e.g. Page and von Tigerstrom 1982; Poole et al. 1991), but the availability of the genome sequence and HPLC/MS/MS enabled us to reliably determine the identities of the proteins differentially expressed under Fe-limited and Fe-replete conditions. HPLC/MS/MS can identify multiple proteins in one sample and distinguish proteins with similar molecular mass and pI. N. europaea appears to respond to Fe limitation by making more proteins that can potentially participate in Fe-uptake (Table 4). The six membrane proteins include OM receptors for several different types of siderophores. This result provides biochemical evidence for the possible role of the genes and their protein products in Fe acquisition and hence in the growth and survival of N. europaea under Fe-limiting conditions. In addition the siderophore feeding experiments (Fig. 1b) indicate the functionality of the siderophore receptor genes in N. europaea. One of the membrane proteins is similar to a heme receptor, though heme, an Fe source for animal pathogenic microbes, seems an unlikely Fe source for N. europaea, a chemoautotrophic soil and wastewater species.
Negative regulation of the expression of Fe uptake systems by ferric uptake regulator (Fur), Fur box, and Fe levels is well characterized in other species such as E. coli and P. aeruginosa (Escolar et al. 1999; Visca et al. 2002; Ravel and Cornelis 2003). In E. coli, FecI is an extracytoplasmic function (ECF) family σ-factor; FecR is an anti σ factor (Mahren and Braun 2003). The ferric uptake regulator is at the top of the regulatory hierarchy in the Fe acquisition system. These components together regulate the expression of the ferric citrate uptake system in E. coli (Braun et al. 2003). Three putative fur genes are identified in the N. europaea genome using BLAST (Chain et al. 2003). Of the 29 intact OM siderophore receptor/transducer genes in the genome, three genes (NE1089/1205/1540) are preceded by sequences similar to the consensus Fur box. These three genes were among those highly induced in Fe-limited cells (Table 4). A putative Fur box is also present upstream of NE2563 (porin OmpC). Based on the MS/MS results and the genome information, a negative regulatory mechanism involving Fur is likely the reason for the elevated expression of the siderophore receptors under Fe limitation. The HPLC/MS/MS results are also interesting in that only six OM siderophore receptors were highly expressed in cells grown in siderophore-free, Fe-limited medium (Table 4). All of them are encoded by genes that do not have cognate σ-factor/anti σ-factor genes (Table 3) and none belongs to the OM siderophore transducer family which has an N-terminal extension that interacts with the anti σ-factor to effect the regulatory pathways (Schalk et al. 2004). This result suggests that the expression of the majority of the 29 genes is likely dependent on the induction by their respective siderophores.
The increased production of the OM siderophore receptors upon lack of Fe and the ability of N. europaea to use common siderophores desferal, ferrichrome and enterobactin for its Fe acquisition supports our hypothesis that N. europaea can obtain Fe in its natural habitats by competing for Fe3+-loaded siderophores made by other organisms. The prolonged lag phase of N. europaea when grown in the presence of desferal suggests that the expression of the ferrioxamine uptake system needs to be induced. In contrast, at least one ferrichrome receptor (NE1089) was expressed constitutively (in absence of ferrichrome, Table 4), consistent with the observation that no induction was needed when N. europaea was grown in medium containing ferrichrome. The growth of N. europaea in medium where Fe was bound by enterobactin, a polycatechol siderophore, also indicates that the catechol siderophore receptor genes (Table 3) are functional.
Nitrosomonas europaea carries out aerobic oxidation of NH3 most efficiently at neutral to slightly alkaline pH. Natural environments with these properties are often Fe-limited owing to the extremely low solubility of ferric iron (10−18 M at pH 7), the predominant form of Fe in the presence of O2 at neutral to alkaline pH; hence, siderophores are necessary for Fe acquisition. The genome makeup of N. europaea suggests an Fe acquisition strategy that is adapted to both low energy yield and high Fe demand associated with NH3 oxidation. Siderophore biosyntheses may require as many as a dozen genes (see e.g. Ravel and Cornelis 2003) and can be a significant draw on the carbon and energy budget of a bacterium. In N. europaea, limited resources are not invested in the synthesis of siderophores but in many siderophore receptors whose production is negatively regulated by Fe levels. This strategy is effective, provided that there are siderophores in the environment. In the natural growth environments of N. europaea, other common soil microorganisms produce siderophores, possibly including ferrioxamine and other hydroxamate siderophores, pyoverdine, azotobactin, and catecholate-type siderophores (Meyer 2000; Page and Huyer 1984; Yang and Leong 1982; Frederick et al. 1981; Powell et al. 1983; Loper and Henkels 1999). Some studies suggest that soil pore water might contain siderophores at concentrations up to 240 μM (Hersman et al. 1995), and there is evidence that these siderophores are utilized by other bacteria (Winkelmann 1991) some of which are non-siderophore-producers (Champomier-Verges et al. 1996).
In the experiments without added siderophores, it seems unlikely that the highly expressed siderophore receptors are involved in Fe uptake in pure cultures since no siderophore was detected except low levels of citrate (3–5 μM). It should be pointed out that these citrate concentrations are at the lower limit level of detection for the method used. However, given that the N. europaea genome has no genes with high similarity to the known fecA, and that the addition of citrate to Fe-limited cultures did not increase growth significantly, the role, if any, of citrate in Fe acquisition for N. europaea seems limited. Nonetheless, cultures grew moderately well in Fe-limited medium. This observation raises the possibility of a siderophore-independent Fe uptake pathway in N. europaea. One possibility, the uptake of Fe2+, seems unlikely under oxic and neutral pH conditions where Fe2+ contributions would be negligible. Furthermore, dipyridyl, an Fe2+ chelator, did not inhibit N. europaea growth. However, N. europaea has most of the genes for potential siderophore-independent Fe3+ uptake. In yeast, membrane surface-localized reductases first generate Fe2+ from Fe3+; multicopper oxidases then convert Fe2+ to Fe3+ which is immediately transported by Ftr1 into the cytoplasm (Stearman et al. 1996; Hassett et al. 1998). While N. europaea genome only has genes with low similarity to the characterized ferric reductase genes, it has a gene (NE0294) that matches well to the genes for a high affinity Fe3+ transporter and has at least seven putative multicopper oxidase genes (Chain et al. 2003). The product of NE0294 is a cytochrome C type protein that is similar to the yeast high-affinity Fe3+ transporter Ftr1. NE0926, one of the multicopper oxidase genes, is similar to the yeast FET5 necessary for Fe uptake (Spizzo et al. 1997). Multicopper oxidases are essential for Fe acquisition in P. aeruginosa when the source is Fe2+ (Huston et al. 2002), and for Fe3+/Fe2+ uptake in Chlamydomonas reinhardtii (Herbik et al. 2002). Another Fe3+ uptake pathway was proposed in Helicobacter pylori (Velayudhan et al. 2000). In this system, flavins reduce Fe3+ to Fe2+ which is taken up through OM porin into periplasm, and finally by FeoB (inner membrane ferrous transporter) into cytoplasm. However, N. europaea genome has only one gene (NE1286) with low similarity to the known feoB. Nonetheless, a porin (OmpC, encoded by NE2563) and a multicopper oxidase (MnxG, by NE0315) were among the highly-expressed proteins under Fe-limited conditions (Table 4). Further studies are needed on the potential siderophore-independent Fe uptake systems in N. europaea, especially on OmpC, FeoB, and multicopper oxidases.
In summary, this study showed that N. europaea was able to grow in medium with relatively low Fe by adjusting its physiology, and it responded to Fe limitation by elevated production of OM siderophore receptors potentially useful for Fe uptake. In addition, growth of N. europaea was increased by the addition of siderophores to Fe-limited medium, which demonstrates that N. europaea can use heterologous siderophores for Fe acquisition. Recently it was shown that N. europaea mutants defective in ferrioxamine receptors failed to grow in media where Fe was chelated by Desferal (unpublished data). These results demonstrate the functionality of some of the siderophore receptor genes in N. europaea. The ability to respond to the changing Fe availability would enable N. europaea to compete successfully for limited Fe supply in its habitats. Further genetic and functional studies on N. europaea Fe uptake systems are warranted.
Abbreviations
- OM:
-
Outer membrane
- AMO:
-
Ammonia monooxygenase
- HAO:
-
Hydroxylamine oxidoreductase
- HPLC/MS/MS:
-
High performance liquid chromatography tandem mass spectrometry
References
Andrews SC, Robinson AK, Rodriguez-Quinones F (2003) Bacterial iron homeostasis. FEMS Microbiol Rev 27:215–237
Arciero DM, Hooper AB (1993) Hydroxylamine oxidoreductase from Nitrosomonas europaea is a multimer of an octa-heme subunit. J Biol Chem 268:14645–14654
Arp DJ, Sayavedra-Soto LA, Hommes NG (2002) Molecular biology and biochemistry of ammonia oxidation by Nitrosomonas europaea. Arch Microbiol 178:250–255
Berry EA, Trumpower BL (1987) Simultaneous determination of hemes a, b, and c from pyridine hemochrome spectra. Anal Biochem 161:1–15
Braun V, Killmann H (1999) Bacterial solutions to the iron-supply problem. Trends Biochem Sci 24:104–109
Braun V, Mahren S, Ogierman M (2003) Regulation of the FecI-type ECF sigma factor by transmembrane signalling. Curr Opin Microbiol 6:173–180
Calugay RJ, Miyashita H, Okamura Y, Matsunaga T (2003) Siderophore production by the magnetic bacterium Magnetospirillum magneticum AMB-1. FEMS Microbiol Lett 218:371–375
Carter P (1971) Spectrophotometric determination of serum iron at the submicrogram level with a new reagent (ferrozine). Anal Biochem 40:450–458
Chain P et al (2003) Complete genome sequence of the ammonia-oxidizing bacterium and obligate chemolithoautotroph Nitrosomonas europaea. J Bacteriol 185:2759–2773
Champomier-Verges MC, Stintzi A, Meyer JM (1996) Acquisition of iron by the non-siderophore-producing Pseudomonas fragi. Microbiology 142(Pt 5):1191–1199
Chart H, Buck M, Stevenson P, Griffiths E (1986) Iron regulated outer membrane proteins of Escherichia coli: variations in expression due to the chelator used to restrict the availability of iron. J Gen Microbiol 132:1373–1378
Clarke TE, Tari LW, Vogel HJ (2001) Structural biology of bacterial iron uptake systems. Curr Top Med Chem 1:7–30
Ensign SA, Hyman MR, Arp DJ (1993) In vitro activation of ammonia monooxygenase from Nitrosomonas europaea by copper. J Bacteriol 175:1971–1980
Escolar L, Perez-Martin J, de Lorenzo V (1999) Opening the iron box: transcriptional metalloregulation by the Fur protein. J Bacteriol 181:6223–6229
Faraldo-Gomez JD, Sansom MS (2003) Acquisition of siderophores in gram-negative bacteria. Nat Rev Mol Cell Biol 4:105–116
Frederick CB, Szaniszlo PJ, Vickrey PE, Bentley MD, Shive W (1981) Production and isolation of siderophores from the soil fungus Epicoccum purpurascens. Biochemistry 20:2432–2436
Hageman RH, Hucklesby DP (1971) Nitrate reductase in higher plants. Meth Enzymol 23:491–503
Hassett RF, Romeo AM, Kosman DJ (1998) Regulation of high affinity iron uptake in the yeast Saccharomyces cerevisiae. Role of dioxygen and Fe. J Biol Chem 273:7628–7636
Herbik A, Bolling C, Buckhout TJ (2002) The involvement of a multicopper oxidase in iron uptake by the green algae Chlamydomonas reinhardtii. Plant Physiol 130:2039–2048
Hersman L, Lloyd T, Sposito G (1995) Siderophore-promoted dissolution of hematite. Geochim Cosmochim Acta 59:3327–3330
Hofte M, Buysens S, Koedam N, Cornelis P (1993) Zinc affects siderophore-mediated high affinity iron uptake systems in the rhizosphere Pseudomonas aeruginosa 7NSK2. Biometals 6:85–91
Hooper AB (1969) Biochemical basis of obligate autotrophy in Nitrosomonas europaea. J Bacteriol 97:776–779
Hooper AB, Erickson RH, Terry KR (1972) Electron transport systems of Nitrosomonas: isolation of a membrane-envelope fraction. J Bacteriol 110:430–438
Houk RS (1994) Elemental and isotopic analysis by inductively coupled plasma mass spectrometry. Acc Chem Res 27:333–339
Huston WM, Jennings MP, McEwan AG (2002) The multicopper oxidase of Pseudomonas aeruginosa is a ferroxidase with a central role in iron acquisition. Mol Microbiol 45:1741–1750
Hyman MR, Arp DJ (1993) An electrophoretic study of the thermal-dependent and reductant-dependent aggregation of the 27 kDa component of ammonia monooxygenase from Nitrosomonas europaea. Electrophoresis 14:619–627
Kammler M, Schon C, Hantke K (1993) Characterization of the ferrous iron uptake system of Escherichia coli. J Bacteriol 175:6212–6219
Keyer K, Gort AS, Imlay JA (1995) Superoxide and the production of oxidative DNA damage. J Bacteriol 177:6782–6790
Loper JE, Henkels MD (1999) Utilization of heterologous siderophores enhances levels of iron available to Pseudomonas putida in the rhizosphere. Appl Environ Microbiol 65:5357–5363
Mahren S, Braun V (2003) The FecI extracytoplasmic-function sigma factor of Escherichia coli interacts with the beta’ subunit of RNA polymerase. J Bacteriol 185:1796–1802
Matzanke BF (1991) Structures, coordination chemistry and functions of microbial iron chelates. In: Winkelmann G (eds) Handbook of microbial iron chelates. CRC Press Inc., Boca Raton, pp 15–64
Meyer JM (2000) Pyoverdines: pigments, siderophores and potential taxonomic markers of fluorescent Pseudomonas species. Arch Microbiol 174:135–142
Mossialos D et al (2000) Quinolobactin, a new siderophore of Pseudomonas fluorescens ATCC 17400, the production of which is repressed by the cognate pyoverdine. Appl Environ Microbiol 66:487–492
Murray RG, Watson SW (1965) Structure of Nitrosocystis oceanus and comparison with Nitrosomonas and Nitrobacter. J Bacteriol 89:1594–1609
Neilands JB (1981) Iron absorption and transport in microorganisms. Annu Rev Nutr 1:27–46
Neilands JB (1995) Siderophores: structure and function of microbial iron transport compounds. J Biol Chem 270:26723–26726
Outten CE, O’Halloran TV (2001) Femtomolar sensitivity of metalloregulatory proteins controlling zinc homeostasis. Science 292:2488–2492
Page WJ, Huyer M (1984) Derepression of the Azotobacter vinelandii siderophore system, using iron-containing minerals to limit iron repletion. J Bacteriol 158:496–502
Page WJ, von Tigerstrom M (1982) Iron- and molybdenum-repressible outer membrane proteins in competent Azotobacter vinelandii. J Bacteriol 151:237–242
Payne SM (1994) Detection, isolation, and characterization of siderophores. Methods Enzymol 235:329–344
Poole K, Neshat S, Heinrichs D (1991) Pyoverdine-mediated iron transport in Pseudomonas aeruginosa: involvement of a high-molecular-mass outer membrane protein. FEMS Microbiol Lett 62:1–5
Powell PE, Szaniszlo PJ, Reid CP (1983) Confirmation of occurrence of hydroxamate siderophores in soil by a novel Escherichia coli bioassay. Appl Environ Microbiol 46:1080–1083
Quatrini R, Jedlicki E, Holmes DS (2005) Genomic insights into the iron uptake mechanisms of the biomining microorganism Acidithiobacillus ferrooxidans. J Ind Microbiol Biotechnol 32:606–614
Ravel J, Cornelis P (2003) Genomics of pyoverdine-mediated iron uptake in pseudomonads. Trends Microbiol 11:195–200
Schalk IJ, Yue WW, Buchanan SK (2004) Recognition of iron-free siderophores by TonB-dependent iron transporters. Mol Microbiol 54:14–22
Schmidt I, Look C, Bock E, Jetten MS (2004) Ammonium and hydroxylamine uptake and accumulation in Nitrosomonas. Microbiology 150:1405–1412
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160:47–56
Shiemke AK, Arp DJ, Sayavedra-Soto LA (2004) Inhibition of membrane-bound methane monooxygenase and ammonia monooxygenase by diphenyliodonium: implications for electron transfer. J Bacteriol 186:928–937
Spizzo T, Byersdorfer C, Duesterhoeft S, Eide D (1997) The yeast FET5 gene encodes a FET3-related multicopper oxidase implicated in iron transport. Mol Gen Genet 256:547–556
Stearman R, Yuan DS, Yamaguchi-Iwai Y, Klausner RD, Dancis A (1996) A permease-oxidase complex involved in high-affinity iron uptake in yeast. Science 271:1552–1557
Stein LY, Arp DJ (1998) Ammonia limitation results in a loss of ammonia-oxidizing activity in Nitrosomonas europaea. Appl Environ Microbiol 64:1514–1521
Thompson H, Tersteegen A, Thauer RK, Hedderich R (1998) Two malate dehydrogenases in Methanobacterium thermoautotrophicum. Arch Microbiol 170:38–42
Tindale AE, Mehrotra M, Ottem D, Page WJ (2000) Dual regulation of catecholate siderophore biosynthesis in Azotobacter vinelandii by iron and oxidative stress. Microbiology 146(Pt 7):1617–1626
Upadhyay AK, Petasis DT, Arciero DM, Hooper AB, Hendrich MP (2003) Spectroscopic characterization and assignment of reduction potentials in the tetraheme cytochrome C554 from Nitrosomonas europaea. J Am Chem Soc 125:1738–1747
Velayudhan J et al (2000) Iron acquisition and virulence in Helicobacter pylori: a major role for FeoB, a high-affinity ferrous iron transporter. Mol Microbiol 37:274–286
Verderber E, Lucast LJ, Van Dehy JA, Cozart P, Etter JB, Best EA (1997) Role of the hemA gene product and delta-aminolevulinic acid in regulation of Escherichia coli heme synthesis. J Bacteriol 179:4583–4590
Visca P, Leoni L, Wilson MJ, Lamont IL (2002) Iron transport and regulation, cell signalling and genomics: lessons from Escherichia coli and Pseudomonas. Mol Microbiol 45:1177–1190
Whittaker M, Bergmann D, Arciero D, Hooper AB (2000) Electron transfer during the oxidation of ammonia by the chemolithotrophic bacterium Nitrosomonas europaea. Biochim Biophys Acta 1459:346–355
Wiebe MG (2002) Siderophore production by Fusarium venenatum A3/5. Biochem Soc Trans 30:696–698
Winkelmann G (1991) Specificity of iron transport in bacteria and fungi. In: Winkelmann G (eds) CRC handbook of microbial iron chelates. CRC Press, Boca Raton, p 366
Yang CC, Leong J (1982) Production of deferriferrioxamines B and E from a ferroverdin-producing Streptomyces species. J Bacteriol 149:381–383
Acknowledgments
We thank Dr. B. Dubbels for his insightful discussions, B. Arbogast (Central Laboratory, OSU) for help with the HPLC/MS/MS analysis, and M. Nesson for help with electron microscopy (Electron Microscope Facility, Department of Botany and Plant Pathology, OSU). We appreciate the help of A. Ungerer (College of Oceanic and Atmospheric Sciences, OSU) for Fe determination by ICP-MS. This research was supported by grant DE-FG03-01ER63149 to DJA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wei, X., Vajrala, N., Hauser, L. et al. Iron nutrition and physiological responses to iron stress in Nitrosomonas europaea . Arch Microbiol 186, 107–118 (2006). https://doi.org/10.1007/s00203-006-0126-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-006-0126-4