Abstract
Methanogens growing on C-1 substrates synthesize 2-carbon acetyl groups in the form of acetyl-CoA for carbon assimilation using the multienzyme complex acetyl-CoA decarbonylase/synthase (ACDS) which contains five different subunits encoded within an operon. In species growing on acetate ACDS also functions to cleave the acetate C-C bond for energy production by methanogenesis. A number of species of Methanosarcina that are capable of growth on either C-1 compounds or acetate contain two separate ACDS operons, and questions have been raised about whether or not these operons play separate roles in acetate synthesis and cleavage. Methanosarcina thermophila genomic DNA was analyzed for the presence of two ACDS operons by PCR amplifications with different primer pairs, restriction enzyme analyses, DNA sequencing and Southern blot analyses. A single ACDS operon was identified and characterized, with no evidence for more than one. MALDI mass spectrometric analyses were carried out on ACDS preparations from methanol- and acetate-grown cells. Peptide fragmentation patterns showed that the same ACDS subunits were present regardless of growth conditions. The evidence indicates that a single form of ACDS is used both for acetate cleavage during growth on acetate and for acetate synthesis during growth on C-1 substrates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Methanogens are a group of microorganisms within the domain of the Archaea that derive their energy by reducing a limited number of compounds to methane. In addition to simple C-1 compounds, which are used by most methanogens, species of the genera Methanosarcina and Methanosaeta are also capable of reducing the methyl group of acetate to methane—a process that requires cleavage of the C–C bond and oxidation of the carboxyl group to CO2. Acetate is first converted to acetyl-CoA, and thereafter cleaved in a reaction in which the methyl group is donated to tetrahydrosarcinapterin (H4SPt) to form N 5-methyltetrahydrosarcinapterin (CH3-H4SPt), as shown in Eq. 1,
in which Fdox and Fdred are the oxidized and reduced forms of ferredoxin, respectively. Cleavage of acetyl-CoA is catalyzed by the acetyl-CoA decarbonylase/synthase (ACDS) multienzyme complex (Grahame 1991) which contains five subunits α, β, γ, δ, ɛ in equimolar amounts (Terlesky et al. 1986).
The molecular mass of the ACDS complex is approximately 2 MDa (Kocsis et al. 1999), indicating eight copies of each subunit, and the complex can be dissociated into three subcomponent proteins that carry out separate partial reactions contributing to Eq. 1 (Grahame and DeMoll 1996.) The dissociated α2ɛ2 unit contains Ni and several Fe/S clusters and possesses carbon monoxide dehydrogenase activity (Krzycki and Zeikus 1984); the γ δ unit contains an Fe/S cluster and a corrinoid cofactor (Abbanat and Ferry 1991) and catalyzes methyl group transfer reversibly between the methylated B12 cofactor and the substrate H4SPt (Grahame and DeMoll 1996); and the isolated β subunit protein contains a binuclear Ni–Ni center bridged to an Fe/S cluster, binds to acetyl-CoA and catalyzes acetyl group transfer and C–C bond cleavage and synthesis (Grahame and DeMoll 1996; Gencic and Grahame 2003). It has been established that Eq. 1 is freely reversible. Footnote 1
When grown on acetate, two-carbon units in the form of acetyl-CoA are readily available for anabolic processes of carbon assimilation, which take place following carboxylation of acetyl-CoA to form pyruvate. However, when grown on C-1 substrates, methanogens must first generate two-carbon units by condensation of two C-1 precursors to produce acetyl-CoA for biosynthetic purposes. Therefore, within a single organism, such as Methanosarcina thermophila, which can reduce methanol as well as the methyl group of acetate to methane (Zinder and Mah 1979), a single form of the ACDS complex could function in both catabolic and biosynthetic processes, depending on the growth conditions. Nevertheless,the idea has been introduced that two different ACDS complexes are synthesized by methanogens: one for acetyl-CoA synthesis, when growing on one-carbon compounds such as methanol, and one for acetyl-CoA cleavage, when growing on acetate (Terlesky et al. 1986; Lindahl and Chang 2001).
The structural genes encoding the five ACDS subunits were first sequenced and their arrangement in an operon structure (along with one other accessory protein) was established in 1996 (Maupin-Furlow and Ferry 1996a, b). The finding of two ACDS operons in the genomes of Methanosarcina species that have been sequenced, Methanosarcina acetivorans C2A (Galagan et al. 2002) and Methanosarcina mazei Gö1 (Deppenmeier et al. 2002), or partially sequenced, Methanosarcina barkeri Fusaro (U.S. Department of Energy Joint Genome Institute at http://genome.jgi-psf.org/microbial/), has further fuelled the arguments for the possibility of two forms of the complex.
Herein we present evidence supporting the conclusion that M. thermophilaencodes and expresses only a single ACDS complex under conditions of growth on either methanol or acetate. The methods used for this include restriction enzyme analysis of various PCR products, DNA sequencing, Southern blot analyses and mass spectrometry. The results strongly suggest that M. thermophila uses a single operon-encoded form of ACDS both anabolically and catabolically for the synthesis and cleavage of acetyl-CoA.
Materials and methods
Strains and cell growth
Methanosarcina thermophila strain TM-1 (Zinder et al. 1985; Murray and Zinder 1985) was obtained from the Oregon Collection of Methanogens (OCM 12). Growth on acetate was carried out at 50°C under strictly anaerobic conditions. The basal medium was prepared essentially as described previously (Grahame and Stadtman 1987) and was supplemented with 0.15 g/l Bacto yeast extract and 0.3 mg/l p-aminobenzoic acid. The pH of the medium was adjusted with acetic acid, and cultures were continuously maintained between pH 6.0–6.8 by periodic addition of glacial acetic acid during growth. Growth on methanol (1–2% v/v) was carried out in MS medium (OCM; Boone et al. 1989).
ACDS operon sequence analysis
Genomic DNA was isolated from M. thermophila strain TM-1 by grinding cells under liquid nitrogen with a mortar and pestle followed by proteinase K digestion in the presence of 0.5% SDS and standard phenol–chloroform extraction and ethanol precipitation (Sambrook et al. 1989). In early plans to carry out heterologous expression of the ACDS β subunit protein, PCR amplification of the ACDS β subunit gene (cdhC) was done using primers designed to be identical to sequences previously published (Maupin-Furlow and Ferry 1996b). A fragment of the expected length, ca. 1.4 kb, was obtained; however, the sequence was not identical to the published cdhC gene sequence (overall 84.4% match, and 9 bp longer), as determined by direct DNA sequence analysis of the PCR product itself as well as by sequencing of separate clones. Therefore, subsequently, three additional PCR primer sets were designed to obtain overlapping fragments covering the entire ACDS operon. The PCR fragments indicated in Fig. 1 were subcloned in the pCRII-TOPO vector (Invitrogen), and sequence analysis of the cloned plasmid DNAs yielded the assembled sequence of 8189 bp (GenBank accession number AF173830) as shown in Fig S1 under Electronic Supplementary Information. Automatic sequencing was performed using the method of dideoxynucleotide termination with the ABI PRISM Big-Dye Terminator cycle sequencing kit v2.0 with AmpliTaq DNA polymerase FS (Applied Biosystems).
Organization of the ACDS operon from M. thermophila TM-1 and the location of PCR products used for sequence analysis. The location of each of the five ACDS subunit genes (cdhA-E) and the accessory protein ACDS ORF is indicated. The overlapping PCR products shown (#1, 1,419 bp; #2, 2,196 bp; #3, 2,378 bp; and #4, 4,367 bp) were subcloned and sequenced. As an additional control, the PCR product #5 corresponding to the entire operon (8,189 bp) was partially sequenced and restriction analysis was performed to confirm its identity. Restriction sites are indicated for: NcoI (N), EcoRI (E), BamHI (B), HindIII (H) and PstI (P)
Long PCR amplification of the entire ACDS operon was carried out using the Expand Long Template PCR System (Roche) with buffer 1 or buffer 2 (1.75 or 2.75 mM MgCl2 final concentration, respectively) under conditions suggested by the manufacturer. Genomic DNA was used as a template with the forward primer 5′ GATGCCATGGCCAAACTAACTACCGGGAGTTTTTC 3′ containing an NcoI site (which introduces one of two mismatches in the second codon) and a reverse primer of 5′ GAGGTCTAGATTATTTGGGTGGCCAGTTCTTCTC 3′ containing an XbaI site.
Southern blot analysis
For Southern blot analyses, 10 μg of genomic DNA was used for digestion with EcoRI, BamHI, HindIII or PstI, and subjected to electrophoresis on a 1% agarose gel. Thereafter, the gel was prepared for transfer by in situ depurination with 0.125 M HCl followed by alkaline denaturation and partial cleavage of the fragments. After neutralization, 20X SSC transfer buffer was used for blotting onto a Hybond N nylon membrane (Sambrook et al. 1989). The full-length 1.4 kb cdhC PCR fragment (product–1, Fig. 1) was 32P-labeled by the random oligonucleotide primer method (Feinberg and Vogelstein 1983) with the RadPrime DNA Labeling System (Invitrogen), using [α-32P] dCTP. Hybridization was performed at 42°C in a buffer containing 50% formamide followed by low stringency washes in order to detect potentially weakly hybridizing fragments.
Purification of ACDS
Purification of the ACDS complex was performed using cells harvested in the late exponential growth phase as described previously for the enzyme complex from M. barkeri (Grahame 1991), with modifications to scale-down the original procedure to be applicable for use inside a Coy anaerobic chamber. For the ACDS complex from methanol-grown cells, ion exchange chromatography on Q-Sepharose was used as an additional purification step due to the lower abundance of ACDS relative to other proteins in those cells.
Mass spectrometric analyses
Trypsin (Promega, Trypsin Gold, mass spectrometry grade) was used for digestion of the ACDS complex purified from M. thermophila grown on acetate or methanol. Peptides of the ACDS complex were prepared by the following procedures that were all performed anaerobically. Purified ACDS was incubated in the presence of 10 mM dithiothreitol, 100 mM NH4HCO3 at 56°C for 60 min, and after cooling to room temperature, an equal volume of 55 mM iodoacetamide, 100 mM NH4HCO3 solution was added. The alkylation proceeded for 45 min at room temperature in the dark, after which the iodoacetamide solution was inactivated by addition of excess 2–mercaptoethanol. Trypsin (25 μg in 1 mM HCl) and CaCl2 (final concentration 1 mM) were added, and the samples were digested at 37°C overnight.
Analyses of the tryptic digests of the ACDS proteins were performed at the University of Kentucky Mass Spectrometry Facility by MALDI MS-MS. The proteins were identified by computational reference through a search of the SwissProt protein database by Mascot Search (Matrix Science). The search results were obtained by ranking candidate peptides in the database according to their match with experimentally obtained mass values and C- and N-terminal sequence information provided by ion trapping and MS–MS fragmentation analysis of the peptides.
Results
PCR amplification of ACDS genes and DNA sequencing
In previous studies on characterization of the Ni-containing β subunit of the ACDS complex from M. thermophila strain TM-1 (Gencic and Grahame 2003), it was found that the DNA sequence of the cdhC gene, obtained by PCR amplification of genomic DNA, did not match the sequence published earlier (Maupin-Furlow and Ferry 1996b). As a possible explanation for this, we considered that more than one ACDS operon might be present in M. thermophila. To address this possibility, we carried out additional PCR amplification reactions to obtain sequence information for the other ACDS genes. Despite the fact that all primers designed to yield fragments 1–4 (Fig. 1) were based (originally) on the sequence information previously published (Sowers et al. 1993; Maupin-Furlow and Ferry 1996a, b; GenBank accession numbers L20952, U30484, and U66032, respectively, stated to be from the same strain, M. thermophila TM–1), all of our PCR products showed significant sequence variation from the originally published sequences on both the nucleotide and the amino acid levels (Table 1). Restriction digestion of the cdhC PCR product-1 showed no evidence for the two PstI sites, or the single NcoI and EcoRI sites present in the previously published sequence. “Heterozygosity” that might have arisen by PCR amplification of two different cdhC loci, also was not observed. Footnote 2
The nucleotide sequences of all overlapping regions of PCR products 1–4 (Fig. 1) matched exactly (with exceptions for deviations in the original primers), allowing for the assembly of the entire 8,189 bp ACDS operon, see Fig S1 under Electronic Supplementary Information (GenBank accession number AF173830). As shown in Fig. 2, PCR amplification of the entire operon was also carried out. The results showed a single band following electrophoresis on a 1% agarose gel in a region slightly larger than 8 kb. In addition, the 8.2 kb PCR product was digested separately with BamHI and EcoRI (not shown), which produced fragment patterns exactly as predicted by the sequence determined here, also without evidence for more than one ACDS operon amplified.
PCR amplification of the ACDS operon from M. thermophila TM-1 genomic DNA generates a single product slightly larger than 8 kb. A 1% agarose gel was loaded with samples from the original PCR reaction mixture (lanes 1 and 2, containing 1 and 1.75 X relative amounts, respectively). Lane 3 was loaded with a larger amount of the product obtained following agarose gel purification. Lane M contains the 1 kb ladder (New England BioLabs) marker, with sizes indicated at the left
Southern blot analyses
Additional evidence suggesting a single ACDS operon in M. thermophila TM-1 was obtained by Southern blot analyses. In the first analysis, genomic DNA was digested with EcoRI, BamHI, HindIII, or PstI, and hybridized using the full-length 1.4 kb PCR-amplified cdhC gene as a probe. As shown in Fig. 3, the number and sizes of the hybridizing fragments matched with those predicted on the basis of the operon sequence determined here. A markedly different fragment pattern would be expected from the previously published operon sequence (Maupin-Furlow and Ferry 1996a, b). From that sequence, two hybridizing EcoRI fragments are expected (975 and 1,236 bp) due to a centrally located internal EcoRI site in cdhC, which is not present in the sequence determined here. The single hybridizing BamHI fragment observed is larger than that of 6,529 bp predicted based on the previously published sequence. In addition, the sizes of two HindIII fragments of 2,588 and > 4,656 bp and one PstI fragment of 3,602 bp predicted from the previous ACDS operon data also vary from those observed in Fig. 3. In a separate experiment, the same cdhC probe, under the same low stringency washing conditions, also readily detected a number of specific fragments using genomic DNA from M. barkeri strain NIH (OCM #265) digested with several different restriction enzymes (not shown). Thus, if a second ACDS operon existed in M. thermophila TM-1 with a level of similarity comparable to the previously published sequence, then it should have also been detected with this probe.
Southern blot analysis of M. thermophila TM-1 genomic DNA using the 1.4 kb cdhC PCR product-1 as probe. Genomic DNA, 10 μg, was digested with the indicated restriction enzymes and analyzed on a 1% agarose gel. The positions and the lengths (bp) of λ HindIII marker fragments are indicated at left. Calculated DNA lengths of the hybridizing fragments are shown at right (observed) compared with the fragment sizes deduced from the ACDS operon sequence (sequence–deduced). * Minimum size limit based on the single BamHI site within the operon with an unknown BamHI site lying outside of the 8.19 kb region of sequence data
Theoretically, if a highly identical, duplicated ACDS operon existed in the genome, then it might not be detected using an internally located probe such as cdhC. Therefore, in a second Southern blot analysis, we used a probe corresponding to the first 290 bp from the 5′ end of cdhA, a region of that gene that is highly conserved, in order to detect genomic fragments produced from restriction sites located well outside the region of the ACDS operon. Restriction enzymes were chosen such that none cut within the region covered by the probe, and each had one known site located not too far from the 3′ end of the probe. As shown in Fig. 4., each restriction enzyme produced only a single DNA fragment hybridizing with the cdhA 5′ end probe. This result is consistent with a single cdhA gene and with a single ACDS operon.
Single fragment pattern in Southern blot analysis using a 290 bp probe from the 5′ end of cdhA. A Southern blot of M. thermophila TM-1 genomic DNA was carried out as indicated in Fig. 3, except that the probe consisted of a PCR product generated by amplification of the first 290 bp at the 5′ end of cdhA. Marker positions (1 kb ladder, New England BioLabs) are indicated at the left. For the restriction enzymes used, restriction sites are first encountered in the ACDS operon at the following positions: EcoRI (1607), BamHI (2620), KpnI (900), NcoI (1164), SphI (345), BspEI (929), PstI (1576)
Mass spectrometry
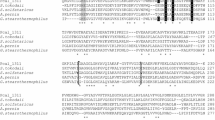
Mass spectrometry experiments were carried out in order to provide direct identification of specific ACDS subunit proteins present in cells utilizing different carbon sources. Trypsin digestion reactions were performed on ACDS preparations obtained from cells grown with either methanol or acetate, and the complex product mixtures were subjected to analysis by MALDI MS–MS. As indicated in Fig. 5, the procedure was able to resolve the peptide products to the extent that all of the subunits encoded by the ACDS operon sequence determined here were identified, except for the small ɛ subunit (CdhB), both in cells grown on C-1 or C-2 substrates, i.e., methanol or acetate. The highest Mascot scores were 189, 181, 157, and 82 for CdhD, CdhA, CdhC, and CdhE, respectively, with a value above 63 corresponding to P < 0.001 in the identification. Furthermore, these four specific subunits were the only proteins in the database that matched to any reasonable degree of significance, with the next highest score of 62 found for an unrelated bacterial protein. By coincidence, two of the ACDS peptides observed in the δ subunit (IIMDPTTAALGYGLDYAYTNMER and TVEEVLQAVDVPIAIGGSGNPQK) have the same sequence as in the previously published work (Maupin-Furlow and Ferry 1996a); however, these exact peptide sequences are also present in the δ subunits from M. acetivorans, M. barkeri and M. mazei, and this by itself does not signify the presence of any other ACDS δ subunit proteins other than the one identified here. The fact that 3 of the total 17 ACDS peptides found in acetate-grown cells were not detected in the ACDS preparation from cells grown on methanol (one missing in each subunit α, β, and δ, as shown in Fig. 5, double underlined peptides) is likely due to differences in digestion conditions or detection sensitivity encountered with a complex subunit mixture, as opposed to single polypeptide analysis.
Tryptic peptides found by MALDI MS–MS in the identification of ACDS subunits in acetate- and methanol-grown cells of M. thermophila TM-1. Solid single underlines indicate peptides found in ACDS complexes isolated under both conditions of growth. Additional, double underlined peptides, were found in ACDS isolated from acetate-grown cells, but were not detected in the analysis of ACDS from cells grown on methanol
Discussion
A single ACDS operon in M. thermophila TM-1
The complete genome sequences have been published for six methanogens: Methanocaldococcus jannaschii, 1.74 Mb (Bult et al. 1996); Methanothermobacter thermoautotrophicus, 1.75 Mb (Smith et al. 1997); Methanopyrus kandleri, 1.69 Mb (Slesarev et al. 2002); M. mazei, 4.1 Mb (Deppenmeier et al. 2002); M. acetivorans, 5.75 Mb (Galagan et al. 2002); and Methanococcus maripaludis, 1.66 Mb (Hendrickson et al. 2004). In addition, there are two others for which partial genome sequences are available: M. barkeri, 4.8 Mb (DOE Joint Genome Institute) and Methanococcoides burtonii, 2.8–3 Mb (DOE Joint Genome Institute; Goodchild et al. 2004). The ACDS subunit genes are found in all of these, almost always arranged altogether in an operon, with the exception of Methanocaldococcus jannaschii in which the operon has been split into two clusters, with the α, ɛ and β subunit genes separated from the γ and δ pair. In addition, the complete genome sequence of Archaeoglobus fulgidus, a non–methanogen that exhibits close taxonomic relationship to the Methanosarcinae, also contains genes encoding ACDS subunits, with two α-ɛ pairs separated from a single [β-ACDS ORF-δ-γ] cluster (Klenk et al. 1997). The genomes of the thermophilic hydrogenotrophic methanogens (Methanocaldococcus jannaschii, Methanothermobacter thermoautotrophicus and Methanopyrus kandleri) contain only a single ACDS operon, as does the mesophilic hydrogenotroph Methanococcus maripaludis. So far, the available genomic sequences suggest that only one ACDS operon exists also in the psychrophilic methylotrophic Methanococcoides burtonii. It is reasonable to consider that in all of these cases a single ACDS complex functions for acetyl-CoA C–C bond formation, as needed for carbon assimilation from C-1 substrates.
By contrast, two ACDS operons are found in certain species of Methanosarcina (M. acetivorans, M. mazei and M. barkeri) which, as well as being able to grow on C-1 compounds, are all capable of utilizing acetate as sole carbon and energy source. For M. thermophila TM–1, the SwissProt database currently designates two copies for each of the five ACDS subunits (see Table I), which makes it appear that this species of Methanosarcina contains these two operons. However, the results of the present study show this is not the case, that these two ACDS operon sequences clearly are not present together within the genome of the organism studied here. Furthermore, our results suggest that M. thermophila TM-1 actually possesses only one ACDS operon. The most likely explanation for why the operon sequence here differs from that published earlier is that DNA from a different strain of Methanosarcina—different from the strain used here, OCM 12—was used in the previous studies. Our results from PCR amplifications with various different primer pairs, restriction enzyme analyses of PCR products, DNA sequencing and Southern blot analyses show no evidence for another ACDS operon. Taken as a whole, this suggests that only one ACDS operon is present in M. thermophila.
For those methanogens that do harbor two ACDS operons, the question arises why. Evidence here that a single operon suffices for the synthesis and cleavage of acetate in M. thermophila suggests that the reason may be more subtle than previously considered. In the case of M. acetivorans, a recent gene duplication event is most likely, since the two operons in that organism exhibit a very high degree of sequence identity (95% identical over a 6.6 kb stretch, including a subsection of nearly 4 kb with only six nucleotide changes) (Galagan et al. 2002). Therefore, possibilities for any hypothetically unique functions of the encoded proteins are limited. On the whole, methanogens that contain only a single operon tend to be those with genomes restricted to around 2 Mb or less, while those containing two operons are all larger than 4 Mb. Thus, species that are unable to maintain a large genome size may not have the luxury of carrying more than one copy of large operons such as ACDS.
One form of ACDS responsible for acetate cleavage and synthesis
The existence of a single ACDS operon in M. thermophila, supported by molecular genetic analyses presented here, would suggest that only one ACDS complex could be expressed in M. thermophila, regardless of the physiological growth conditions. In fact, our results from mass spectrometric analyses confirm that this is the case. In an organism such as M. acetivorans that possesses two ACDS operons, elevated levels of proteins encoded by one of these (MA3860-3865) were clearly evident when cells were grown on acetate (Li et al. 2005a). However, the only evidence suggesting the apparent expression of genes from the other operon (MA1011-1016) was based on MS–MS detection of a single 11-amino acid peptide derived from the 2-D electrophoresis spot containing the α subunit(s). It should be noted that three of the six ACDS genes (subunits γ, δ and the ACDS accessory protein) cannot be distinguished by Li et al. (2005a, b) because the proteins are either perfectly identical or differ by only one amino acid (MA3865=MA1011, MA3864≅MA101, MA3863=MA1013),Footnote 3, making them impossible or nearly impossible to distinguish. The speculation that expression of both operons in M. acetivorans might be needed to produce sufficient amounts of ACDS required for growth on acetate (Li et al. 2005b) is not supported by the situation of sufficiently high levels of ACDS produced from a single operon in M. thermophila. In M. mazei, which possesses two ACDS operons that are more readily distinguishable, there is still no evidence available for significant expression of more than one, let alone differential expression of separate sets of ACDS genes under different growth conditions. Northern blot analyses have been previously used to examine M. mazei for expression of the two ACDS operons (Eggen et al. 1996). Using probes selective for the two cdhA genes, transcription of one of the operons was clearly identified; however, no evidence for the expression of the other could be obtained, regardless of whether acetate or methanol was used as growth substrate.
The first evidence that ACDS expression in Methanosarcina is regulated at the level of transcription was obtained by Sowers et al. (1993). On Northern blots, a large decrease in the signal corresponding to cdhA mRNA was observed within minutes upon addition of methanol to cells actively growing on acetate. However, in those studies, as well as in experiments with cells grown on methanol or trimethylamine, no new signals were observed that might signify expression of a different form of ACDS. The finding of two peaks of CO dehydrogenase in protein chromatographic separations (Terlesky et al. 1986; Grahame 1991) cannot be interpreted as evidence for two forms of the ACDS complex, because a small amount of the α2ɛ2 protein, devoid of acetyl-CoA cleavage and synthesis activity, is usually present in a form not associated with the complex.
In the Archaea, the active site for C–C bond synthesis and cleavage resides on the β subunit of the ACDS complex (Grahame and DeMoll 1996; Gencic and Grahame 2003), whereas, in the anaerobic acetogenic bacteria the active site for acetate C-C bond synthesis resides on the α subunit (so-called α metallosubunit) of a 310 kDa α2β2 bifunctional enzyme known as carbon monoxide dehydrogenase/acetyl-CoA synthase (Xia and Lindahl 1996; Doukov et al. 2002; Darnault et al. 2003). Theoretically, acetate synthesis in methanogens might alternatively utilize a bacterial-like α metallosubunit homolog in combination with a CO dehydrogenase protein. However, other than the ACDS operon-associated β subunits, α metallosubunit homologues are generally not found in the Archaea. A Blastp search for hits to the α metallosubunit from Moorella thermoacetica returns only one such protein in the Archaea, that being in Methanocaldococcus jannaschii, which encodes a single bacterial α metallosubunit-like protein MJ0152 of undetermined function in addition to its complete set of ACDS subunits, including β.
In summary, the evidence here indicates that a single form of ACDS serves for both acetate cleavage and synthesis in M. thermophila, and perhaps in other species of Methanosarcina as well. Previous enzymological and thermodynamic studies have shown that formation of acetate is favorable under physiological redox conditions during growth on C-1 substrates, while acetate C–C bond cleavage is favorable under redox conditions prevailing during growth on acetate. The possibility that other modifications might be used somehow to modulate the activity of a single complex under different physiological conditions cannot be excluded. However, since the reaction is freely reversible, it is unclear at the present time why such modifications would be needed.
Notes
Radioisotopic exchange techniques were first used to demonstrate the reversibility of acetyl C–C bond formation (Raybuck et al. 1991), net synthesis of acetyl-CoA from CO + CoA + CH3–H4SPt was shown (Grahame 1993), and the equilibrium thermodynamics describing the reversibility of acetyl-CoA synthesis and cleavage have been characterized by the use of CO2 derived from bicarbonate and a reducing system coupled to ferredoxin (Grahame and DeMoll 1995).
To ensure that the characteristics of M. thermophila TM-1 (i.e., OCM 12) maintained in our laboratory had not been altered somehow, separate samples were obtained from the Oregon Collection of Methanogens (OCM), one of which was mass cultured for us by the OCM and used directly for genomic DNA isolation without further growth. PCR amplifications followed by restriction analyses were performed with several restriction enzymes (EcoRI, NcoI, PstI and Sau3AI) chosen specifically to distinguish between the cdhC gene sequences of Maupin-Furlow and Ferry (1996b) and the present work. The results showed identical digestion patterns regardless of the cultivation history of the cells, with no evidence for changed characteristics.
Perfect match of the two M. acetivorans ACDS accessory protein sequences is evident after inspection of the original genomic DNA sequences. The annotated version in the database assumes different translation start sites, apparently without valid reason.
References
Abbanat DR, Ferry JG (1991) Resolution of component proteins in an enzyme complex from Methanosarcina thermophila catalyzing the synthesis or cleavage of acetyl-CoA. Proc Natl Acad Sci 88:3272–3276
Bhaskar B, DeMoll E, Grahame DA (1998) Redox-dependent acetyl transfer partial reaction of the acetyl-CoA decarbonylase/synthase complex: kinetics and mechanism. Biochemistry 13:14491–14499
Boone, DR, Johnson, RL, Liu, Y (1989) Diffusion of the interspecies electron carriers H2 and formate in methanogenic ecosystems and its implications in the measurement of K m for H2 or formate uptake. Appl Environ Microbiol 55:1735–1741
Bult CJ, White O, Olsen GJ, Zhou L, Fleischmann RD, Sutton GG, Blake JA, FitzGerald LM, Clayton RA, Gocayne JD, Kerlavage AR, Dougherty BA, Tomb JF, Adams MD, Reich CI, Overbeek R, Kirkness EF, Weinstock KG, Merrick JM, Glodek A, Scott JL, Geoghagen NS, Weidman JF, Fuhrmann JL, Nguyen DT, Utterback T, Kelley JM, Peterson JD, Sadow PW, Hanna MC, Cotton MD, Hurst MA, Roberts KM, Kaine BB, Borodovsky M, Klenk HP, Fraser CM, Smith HO, Woese CR,Venter JC (1996) Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science 273:1058–1073
Dai YR, Reed DW, Millstein JH, Hartzell PL, Grahame DA, DeMoll E (1998) Acetyl–CoA decarbonylase/synthase complex from Archaeoglobus fulgidus. Arch Microbiol 169:525–529
Darnault C, Volbeda A, Kim EJ, Legrand P, Vernède X, Lindahl PA, Fontecilla-Camps JC (2003) Ni-Zn-[Fe4S4] and Ni-Ni-[Fe4S4] clusters in closed and open α subunits of acetyl-CoA synthase/carbon monoxide dehydrogenase. Nat Struct Biol 4:271–279
Deppenmeier U, Johann A, Hartsch T, Merkl R, Schmitz RA, Martinez-Arias R, Henne A, Wiezer A, Baumer S, Jacobi C, Bruggemann H, Lienard T, Christmann A, Bomeke M, Steckel S, Bhattacharyya A, Lykidis A, Overbeek R, Klenk HP, Gunsalus RP, Fritz HJ, Gottschalk G (2002) The genome of Methanosarcina mazei: evidence for lateral gene transfer between bacteria and archaea. J Mol Microbiol Biotechnol 4:453–461
Doukov TI, Iverson TM, Seravalli J, Ragsdale SW, Drennan CL (2002) A Ni–Fe–Cu center in a bifunctional carbon monoxide dehydrogenase/acetyl-CoA synthase. Science 298:567–572
Eggen RIK, van Kranenburg R, Vriesema AJM, Geerling ACM, Verhagen MFJM, Hagen WR, de Vos WM (1996) Carbon monoxide dehydrogenase from Methanosarcina frisia Gö1. Characterization of the enzyme and the regulated expression of two operon-like cdh gene clusters. J Biol Chem 271:14256–14263
Feinberg AP, Vogelstein B (1983) A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 132:6–13
Galagan JE, Nusbaum C, Roy A, Endrizzi MG, Macdonald P, FitzHugh W, Calvo S, Engels R, Smirnov S, Atnoor D, Brown A, Allen N, Naylor J, Stange-Thomann N, DeArellano K, Johnson R, Linton L, McEwan P, McKernan K, Talamas J, Tirrell A, Ye W, Zimmer A, Barber RD, Cann I, Graham DE, Grahame DA, Guss AM, Hedderich R, Ingram-Smith C, Kuettner HC, Krzycki JA, Leigh JA, Li W, Liu J, Mukhopadhyay B, Reeve JN, Smith K, Springer TA, Umayam LA, White O, White RH, Conway de Macario E, Ferry JG, Jarrell KF, Jing H, Macario AJ, Paulsen I, Pritchett M, Sowers KR, Swanson RV, Zinder SH, Lander E, Metcalf WW, Birren B (2002) The genome of M. acetivorans reveals extensive metabolic and physiological diversity. Genome Res 12:532–542
Gencic S, Grahame DA (2003) Nickel in subunit beta of the acetyl-CoA decarbonylase/synthase multienzyme complex in methanogens. Catalytic properties and evidence for a binuclear Ni-Ni site. J Biol Chem 278:6101–6110
Goodchild A, Raftery M, Saunders NFW, Guilhaus M, Cavicchioli R. (2004) Biology of the cold adapted archaeon, Methanococcoides burtonii determined by proteomics using liquid chromatography-tandem mass spectrometry. J Proteome Res 3:1164–1176
Grahame DA (1991) Catalysis of acetyl-CoA cleavage and tetrahydrosarcinapterin methylation by a carbon monoxide dehydrogenase-corrinoid enzyme complex. J Biol Chem 266:22227–22233
Grahame DA (1993) Substrate and cofactor reactivity of a carbon monoxide dehydrogenase-corrinoid enzyme complex: stepwise reduction of iron-sulfur and corrinoid centers, the corrinoid Co2+/1+ redox midpoint potential, and overall synthesis of acetyl-CoA. Biochemistry 32:10786–10793
Grahame DA, DeMoll E (1995) Substrate and accessory protein requirements and thermodynamics of acetyl-CoA synthesis and cleavage in Methanosarcina barkeri. Biochemistry 34:4617–4624
Grahame DA, DeMoll E (1996) Partial reactions catalyzed by protein components of the acetyl-CoA decarbonylase synthase enzyme complex from Methanosarcina barkeri. J Biol Chem 271:8352–8358
Hendrickson EL, Kaul R, Zhou Y, Bovee D, Chapman P, Chung J, Conway de Macario E, Dodsworth JA, Gillett W, Graham DE, Hackett M, Haydock AK, Kang A, Land ML, Levy R, Lie TJ, Major TA, Moore BC, Porat I, Palmeiri A, Rouse G, Saenphimmachak C, Söll D, Van Dien S, Wang T, Whitman WB, Xia Q, Zhang Y, Larimer FW, Olson MV, Leigh JA (2004) Complete genome sequence of the genetically tractable hydrogenotrophic methanogen Methanococcus maripaludis. J Bacteriol 186:6956–6969
Jablonski PE, DiMarco AA, Bobik TA, Cabell MC, Ferry JG (1990) Protein content and enzyme activities in methanol- and acetate-grown Methanosarcina thermophila. J Bacteriol 172:1271–1275
Klenk HP, Clayton RA, Tomb JF, White O, Nelson KE, Ketchum KA, Dodson RJ, Gwinn M, Hickey EK, Peterson JD, Richardson DL, Kerlavage AR, Graham DE, Kyrpides NC, Fleischmann RD, Quackenbush J, Lee NH, Sutton GG, Gill S, Kirkness EF, Dougherty BA, McKenney K, Adams MD, Loftus B, Peterson S, Reich CI, McNeil LK, Badger JH, Glodek A, Zhou L, Overbeek R, Gocayne JD, Weidman JF, Donald L, Utterback T, Cotton MD, Spriggs T, Artiach P, Kaine BP, Sykes SM, Sadow PW, D’Andrea KP, Bowman C, Fujii C, Garland SA, Mason TM, Olsen GJ, Fraser CM, Smith HO, Woese CR, Venter JC (1997) The complete genome sequence of the hyperthermophilic, sulphate-reducing archaeon Archaeoglobus fulgidus. Nature 390:364–370
Kocsis E, Kessel M, DeMoll E, Grahame DA (1999) Structure of the Ni/Fe-S protein subcomponent of the acetyl-CoA decarbonylase/synthase complex from Methanosarcina thermophila at 26-Å resolution. J Struct Biol 128:165–174
Krzycki JA, Zeikus JG (1984) Characterization and purification of carbon monoxide dehydrogenase from Methanosarcina barkeri. J Bacteriol 158:231–237
Li Q, Li L, Rejtar T, Karger BL, Ferry JG (2005a) Proteome of Methanosarcina acetivorans part I: an expanded view of the biology of the cell. J Proteome Res 4:112–128
Li Q, Li L, Rejtar T, Karger BL, Ferry JG (2005b) Proteome of Methanosarcina acetivorans part II: comparison of protein levels in acetate- and methanol-grown cells. J Proteome Res 4:129–135
Lindahl PA, Chang B (2001) The evolution of acetyl-CoA synthase. Origins of life and evolution of the biosphere 31:403–434
Maupin-Furlow JA, Ferry JG (1996a) Characterization of the cdhD and cdhE genes encoding subunits of the corrinoid/iron-sulfur enzyme of the CO dehydrogenase complex from Methanosarcina thermophila. J Bacteriol 178:340–346
Maupin-Furlow JA, Ferry JG (1996b) Analysis of the CO dehydrogenase/acetyl-coenzyme A synthase operon of Methanosarcina thermophila. J Bacteriol 178:6849–6856
Murray PA, Zinder SH (1985) Nutritional requirements of Methanosarcina sp strain TM-1. Appl Environ Microbiol 50:49–55
Nelson MJK, Ferry JG (1984) Carbon monoxide-dependent coenzyme M methylreductase in acetotrophic Methanosarcina spp. J Bacteriol 160:526–532
Raybuck SA, Ramer SE, Abbanat DR, Peters JW, Orme-Johnson WH, Ferry JG, Walsh CT (1991) Demonstration of carbon-carbon bond cleavage of acetyl coenzyme A by using isotopic exchange catalyzed by the CO dehydrogenase complex from acetate-grown Methanosarcina thermophila. J Bacteriol 173:929–932
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Slesarev AI, Mezhevaya KV, Makarova KS, Polushin NN, Shcherbinina OV, Shakhova VV, Belova GI, Aravind L, Natale DA, Rogozin IB, Tatusov RL, Wolf YI, Stetter KO, Malykh AG, Koonin EV, Kozyavkin SA (2002) The complete genome of hyperthermophile Methanopyrus kandleri AV19 and monophyly of archaeal methanogens. Proc Natl Acad Sci 99:4644–4649
Smith DR, Doucette-Stamm LA, Deloughery C, Lee HM, Dubois J, Aldredge T, Bashirzadeh R, Blakely D, Cook R, Gilbert K, Harrison D, Hoang L, Keagle P, Lumm W, Pothier B, Qiu D, Spadafora R, Vicaire R, Wang Y, Wierzbowski J, Gibson R, Jiwani N, Caruso A, Bush D, Safer H, Patwell D, Prabhakar S, McDougall S, Shimer G, Goyal A, Pietrovski S, Church GM, Daniels CJ, Mao JI, Rice P, Nolling J, Reeve JN (1997) Complete genome sequence of Methanobacterium thermoautotrophicumΔH: functional analysis and comparative genomics. J Bacteriol 179:7135–7155
Sowers KR, Thai TT, Gunsalus RP (1993) Transcriptional regulation of the carbon monoxide dehydrogenase gene (cdhA) in Methanosarcina thermophila. J Biol Chem 268:23172–23178
Terlesky KC, Nelson MJK, Ferry JG (1986) Isolation of an enzyme complex with carbon monoxide dehydrogenase activity containing corrinoid and nickel from acetate-grown Methanosarcina thermophila. J Bacteriol 168:1053–1058
Xia J, Lindahl PA (1996) Assembly of an exchange-coupled [Ni:Fe4S4] cluster in the α metallosubunit of carbon monoxide dehydrogenase from Clostridium thermoaceticum with spectroscopic properties and CO-binding mimicking those of the acetyl-CoA synthase active site. J Am Chem Soc 118:483–484
Zinder SH, Mah RA (1979) Isolation and characterization of a thermophilic strain of Methanosarcina unable to use H2-CO2 for methanogenesis. Appl Environ Microbiol 38:996–1008
Zinder SH, Sowers KR, Ferry JG (1985) Methanosarcina thermophila sp nov, a thermophilic, acetotrophic, methane-producing bacterium. Int J Syst Bacteriol 35:522–523
Acknowledgements
We gratefully acknowledge the University of Kentucky Mass Spectrometry Facility. This research was supported by grants from the U.S. Department of Energy, Division of Biological Energy Research, Office of Basic Energy Sciences (#DE-FG02-00ER15108 to DAG; #DEGG02-97ER20270 to ED) and from the National Science Foundation, Program for Metabolic Biochemistry (#MCB-0215106 to DAG).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Grahame, D.A., Gencic, S. & DeMoll, E. A single operon-encoded form of the acetyl-CoA decarbonylase/synthase multienzyme complex responsible for synthesis and cleavage of acetyl-CoA in Methanosarcina thermophila . Arch Microbiol 184, 32–40 (2005). https://doi.org/10.1007/s00203-005-0006-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-005-0006-3