Abstract
Prephenate dehydratase is a key regulatory enzyme in the phenylalanine-specific pathway of Corynebacterium glutamicum. PCR-based random mutagenesis and functional complementation were used to screen for m-fluorophenylalanine (mFP)-resistant mutants. Comparison of the amino acid sequence of the mutant prephenate dehydratases indicated that Ser-99 plays a role in the feedback regulation of the enzyme. When Ser-99 of the wild-type enzyme was replaced by Met, the specific activity of the mutant enzyme was 30% lower than that of the wild-type. The Ser99Met mutant was active in the presence of 50 μM phenylalanine, whereas the wild-type enzyme was not. The functional roles of the eight conserved residues of prephenate dehydratase were investigated by site-directed mutagenesis. Glu64Asp substitution reduced enzyme activity by 15%, with a 4.5- and 1.7-fold increase in K m and k cat values, respectively. Replacement of Thr-183 by either Ala or Tyr resulted in a complete loss of enzyme activity. Substitution of Arg-184 with Leu resulted in a 50% decrease of enzyme activity. The specific activity for Phe185Tyr was more than 96% lower than that of the wild-type, and the K m value was 26-fold higher. Alterations in the conserved Asp-76, Glu-89, His-115, and Arg-236 residues did not cause a significant change in the K m and k cat values. These results indicated that Glu-64, Thr-183, Arg-184, and Phe-185 residues might be involved in substrate binding and/or catalytic activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phenylalanine is an important amino acid in terms of its applications in the pharmaceutical, food, and cosmetics industries. Microorganisms differ in the arrangement of enzymes for the biosynthesis of l-phenylalanine via chorismate. Escherichia coli, Pseudomonas stutzeri, and many other gram-negative bacteria utilize a bifunctional enzyme (P protein), chorismate mutase (EC 5.4.99.5)-prephenate dehydratase (EC 4.2.1.51), to catalyze the conversion of chorismate to phenylpyruvate (Ahmad and Jensen 1986). Phenylpyruvate is subsequently transaminated via aromatic amino acid aminotransferase to phenylalanine. Phenylalanine biosynthesis in E. coli is regulated by attenuation of transcription of the prephenate dehydratase gene (pheA) (Yanofsky 1988) and by feedback inhibition of the prephenate dehydratase (PDT) and chorismate mutase (CM) activities of the bifunctional enzyme (Dopheide et al. 1972). The active form of E. coli CM-PDT (P-protein) is a homodimer (Gething and Davidson 1976; Baldwin et al. 1981) and the increase in phenylalanine concentrations will shift the enzyme from a dimer to a mixture of dimer, tetramer, and higher-order species (Gething and Davidson 1976; Baldwin et al. 1981; Ahmad et al. 1988). It has been reported that the mutase, dehydratase, and regulatory activities reside in discrete domains of E. coli P-protein (Zhang et al. 1998). Corynebacterium glutamicum, Bacillus subtilis, and other gram-positive bacteria also synthesize phenylalanine via phenylpyruvate, producing phenylalanine from chorismate in consecutive steps catalyzed by the monofunctional enzymes CM and PDT (Fazel and Jensen 1980; Pierson and Jenson 1974). In B. subtilis, the CM reaction is carried out by a bifunctional protein (CM:DAHPS) or by a monofunctional CM (Byng and Jensen 1983). However, the CM of Brevibacterium flavum is reported to be bifunctional, with CM and 3-deoxy-d-arabino-heptulosonate-7-phosphate (DAHP) synthase (Shiio and Sugimota 1979).
C. glutamicum belongs to the actinomycetes subdivision of gram-positive bacteria (Liebl et al. 1991) and has a genome size of 3,082 kbp (Bathe et al. 1996). It is widely used in the industrial production of amino acids (Yoshinga and Nakamori 1983) and nucleotides (Kuninka 1986). Several reports have described the use of phenylalanine analogues to select for C. glutamicum mutants in which PDT is resistant to feedback inhibition by phenylalanine (Hagino and Nakatama 1974; Chan and Hsu 1996). Phenylalanine production by a metabolically engineered strain with deregulated DAHP synthase, CM, and PDT reached 28 g/l fermentation broth (Ikeda and Katsumata 1992). Nonetheless, current knowledge about the allosteric control of C. glutamicum PDT is limited, and a crystal structure of PDT is not yet available. In the present study, we describe the isolation and characterization of PDT mutants that exhibit resistance to feedback inhibition at high phenylalanine concentration. In addition, we conducted site-directed mutagenesis on eight conserved residues (Glu-64, Asp-76, Glu-89, His-115, Thr-183, Arg-184, Phe-185, and Arg-236) and compared the kinetic properties of selected mutant proteins to those of the wild-type enzyme. Our findings revealed that Glu-64 may play an essential role in the catalytic mechanism of the enzyme from C. glutamicum. Furthermore, the ring carboxylate and both the carbonyl and carboxyl groups on the pyruvyl side-chain of prephenate may interact with Thr-183, Arg-184, and Phe-185.
Materials and methods
Bacterial strains, and growth conditions
Bacterial strains, plasmids, and phages used in this study are listed in Table 1. E. coli strains were grown at 37 °C either in Luria-Bertani (LB) medium or on minimal (M9) agar plates (Sambrook et al. 1989). When required, 100 µg ampicillin/ml was added.
DNA manipulations
Plasmid DNA was prepared from E. coli by standard procedures (Sambrook et al. 1989). DNA fragments were purified from agarose by using Geneclean III kit (Bio 101, Calif., USA). Site-directed mutagenesis was carried out with the Muta-Gene phagemid in vitro mutagenesis kit (Bio-Rad, Richmond, Calif., USA) (Kunkel et al. 1987) on a HindIII–BamHI fragment of C. glutamicum wild-type pheA inserted into M13mp18. The resulting plasmid was then introduced into E. coli CJ236 via transformation to produce uracil-containing single-stranded DNA ,which was used as template in site-directed mutagenesis. The oligonucleotide primers are listed in Table 2 and the altered nucleotides are in italics. All mutations were verified by DNA sequencing using a Sequenase R version 2 sequencing kit (United States Biochemical, Ohio, USA).
Expression and purification of the recombinant PDTs
To increase the expression level of pheA, the gene was cloned into pQE-30 under the control of T5 promoter. The pheA coding region from pARW was amplified using primers 1 and 2, and high-fidelity Vent DNA polymerase. The amplification reaction was done in a total volume of 100 μl containing 1 ng pARW, 10 mM of each deoxynucleotide triphosphate, 10 pmol of each primer, and 1 U Vent DNA polymerase. The reaction proceeded as follows: 94 °C for 5 min; 35 cycles of 94 °C for 3 min, 60 °C for 1 min, and 73 °C for 1 min; followed by 72 °C for 10 min. The amplified 0.95-kb BamHI–HindIII fragment was cloned into the corresponding sites of pQE-30 to create pQE-pheA for high-level expression of the N-terminal His6-tagged PDT.
E. coli JM109 harboring pQE-pheA was propagated at 37 °C in LB medium containing 100 μg ampicillin/ml. Overnight cultures were diluted 1/100 into the same medium, grown to OD600 of 0.7~0.9, induced with 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG), and cultivated for an additional 6 h. Cells were collected, resuspended in 10 ml of binding buffer (5 mM imidazole, 0.5 M NaCl, 20 mM Tris-HCl; pH 7.9), disrupted by sonication, and clarified by centrifugation. The crude extract was applied to a Ni2+-nitrilotriacetic acid agarose column (His-Bind resin, Novagen, Madison, Wis., USA) and chromatographed according to the supplier’s instructions.
Selection of mFP-resistant PDT
PCR-based random mutagenesis was carried out according to the procedure described by Cadwell and Joyce (1992). The mutagenic reaction mixture (100 μl) contained 1 ng pQE30-pheA, 10 mM of each deoxynucleotide triphosphate, 10 pmol of each primer, 0.25 mM MnCl2, 5 mM MgCl2, and 2.5 U Taq DNA polymerase. The PCR condition was as described above. The PCR product was purified on agarose gel, digested with BamHI and HindIII, and subcloned into pQE-30. The recombinant plasmid was introduced into E. coli CCRC51737 and the transformants were selected by their ability to grow on complete medium plates (Follettie and Sinskey 1986) containing 100 µg ampicillin/ml. Ampicillin-resistant transformants were then replicated on minimal (M9) agar plates containing 7 mM m-fluorophenylalanine.
Gel electrophoresis and immunoblot analysis
SDS-PAGE was done with gels cast using the method of Laemmli (1970). Proteins were transferred to Immobilon-P Transfer Membrane (Millipore, Bedford, Mass., USA) using a Mini Trans-Blot Electrophoretic Transfer Cell (Bio-Rad), and unbound membrane sites were blocked with a blocking buffer containing 50 mM Tris-HCl (pH 7.4), 150 mM NaCl, 1 mM EDTA, 0.05% Tween 20, and 5% nonfat milk. The blots were probed with rabbit polyclonal antibodies against PDT. Alkaline-phosphatase-conjugated antibodies raised in goats against rabbit immunoglobulin G (Bio-Rad) were used to visualize the PDT protein.
Enzyme assay and kinetic characterization
PDT activity was determined according to the method of Dopheide et al. (1972). The reaction mixture, containing 50 μl of 8 mM prephenate, 200 μl of 100 mM Tris-HCl (pH 7.5), and 50 μl of appropriately diluted enzyme solution was incubated at 30 °C for 5 min. The reaction was stopped by the addition of 800 μl of 1 N NaOH and the absorbance at 320 nm was then measured. One unit of activity was defined as the amount of enzyme that catalyzed the formation of 1 μM phenylpyruvate per min under the assay conditions. Protein concentrations were determined with Bio-Rad Protein Assay kit (Bio-Rad) using bovine serum albumin as the standard.
The K m and k cat values were estimated by measuring phenylpyruvate production in 0.4-ml reaction mixtures containing various concentrations of the substrate (0.33~2.0 K m) in 100 mM Tris-HCl buffer, pH 7.5, and a suitable amount of enzyme such that reaction rates were linear over the 5-min time course of the assays. The K m and k cat values were calculated from the rate of phenylpyruvate production using the Michaelis-Menten equation and the Grafit 5 software (Sigma).
Circular dichroism spectroscopy
Circular dichroism experiments (CD) were done on an Aviv Circular Dichroism Spectropolarimeter (model 61DS) equipped with a single-position thermoelectric cuvette holder. CD spectra were recorded at 20 °C in phosphate buffer with a protein concentration of 1 μM and a path-length of 0.2 cm. Spectra were obtained by averaging five wavelength scans from 200 to 260 nm in 0.5-nm steps, with a signal averaging time of 2 s and a bandwidth of 1.5 nm.
Chemicals
All chemicals used were of the highest quality available and were obtained from Merck (Gibbstown, N.J., USA) or Sigma (St. Louis, Mo., USA). Restriction enzymes and other DNA-modifying enzymes were acquired from Boehringer Mannheim (Mannheim, Germany), New England Biolabs (Beverly, Mass., USA) and Promega (Madison, Wis., USA). Goat-anti-rabbit IgG was purchased from Bio-Rad. The resin for protein purification was obtained from Novagen. Oligonucleotides were acquired from Quality Inc. (Taipei, Taiwan).
Results
Expression and characterization of His6-tagged PDT
C. glutamicum pheA was cloned into pQE30 to generate pQE-pheA, in which the expression of pheA was driven from the E. coli phage T5 promoter containing two lac operater sequences, so that the production of PDT was induced by IPTG. The PDT purified from cell extract of E. coli JM109 (pQE-pheA) exhibited a specific activity of 9.48 U/mg protein (Table 5). As a control, no significant level of PDT activity was detected in the cell extract of E. coli JM109 harboring pQE30. SDS-PAGE revealed a prominent band at approximately 39 kDa in the pQE-pheA-harboring strain but it was not observed in the control cells (data not shown). These results indicated that the His6-tagged enzyme was produced in high quantity and was apparently active.
The overexpressed wild-type PDT was purified in a single step by Ni2+-chelate affinity chromatography. The purified His6-tagged enzyme was found to be sensitive to feedback inhibition by l-phenylalanine. The addition of 50 μM l-phenylalanine to the reaction mixture completely inhibited the activity of His6-tagged PDT. In contrast, l-tyrosine (50 µM) caused about 3.4-fold increase in relative activity (Table 4). Tryptophan, histidine, threonine, and leucine had only relatively minor effects on PDT activity (data not shown). Kinetic evaluation revealed that K m, k cat, and k cat/K m of purified His6-tagged PDT were 0.07 mM, 0.12 min−1, and 1.71 min−1 mM−1, respectively (Table 5).
Screening mFP-resistant pheA
In the presence of 7 mM MgCl2, the average error rate for Taq DNA polymerase has been estimated to be 6.6×10-3. Using PCR-based random mutagenesis, six mFP-resistant mutants, M1, M17, M24, M25, M36, and M41, were obtained from 61,035 colonies, indicating that the average error rate for the modified PCR on pheA encoding the mFP-resistance phenotype was 9.8×10−5. A protein band exhibiting an apparent molecular mass of 39 kDa was detected in the purified enzymes from mutant strains (Fig. 1A). Western blot analysis confirmed the presence of the recombinant PDTs in transformed E. coli (Fig. 1B). To localize those mutations responsible for the mFP-resistance phenotype of each isolate, pheA was sequenced. A comparison of the aligned amino acid sequence between these mutants and the wild-type PDT indicated amino acid sequence changes in the mutants (Table 3). With the exception of PDT-M1, an A-to-T transversion in pheA led to the replacement of Ser-99 with Met in five other mutants. In the presence of 50 μM phenylalanine, the wild-type enzyme lost all PDT activity. Protein variant M1 was strongly resistant to feedback inhibition at a much higher concentration of phenylalanine (500 μM) with more than 30% retention of enzymatic activity (Table 4). Like protein variant M1, variants M36 and M41 were active under the same condition.
The role of Ser-99 in C. glutamicum PDT
To investigate the functional role of Ser-99, a set of mutants was constructed and the enzymatic properties of the resultant proteins were examined. Replacement of Ser-99 with Lys or Tyr yielded proteins that were expressed more poorly than wild-type enzyme as judged by SDS-PAGE and immunoblots (data not shown). Moreover, the purified mutant proteins did not exhibit any detectable PDT activity, indicating that structural changes in the protein may have led to their inactivity and instability. In contrast, the well-expressed Ser99Met, Ser99Thr, and Ser99Cys proteins had activities comparable to that of wild-type enzyme (Table 5). The kinetic parameters of wild-type and mutant enzymes are listed in Table 5. The Ser99Met mutant, which displayed ~70% wild-type PDT activity, showed a 1.6-fold increase in K m, and k cat was unchanged. Replacement of Ser-99 by Ala, Cys, or Leu resulted in a significant increase in the feedback resistance of the enzyme (Table 4) and indicated that Ser-99 plays an important role in its feedback regulation. It has been reported that l-tyrosine stimulates the PDT activity of Amycolatopsis methanolica (Euverink et al. 1995). Similarly, we noted that tyrosine markedly stimulated PDT activity of both wild-type and mutant enzymes (Table 4).
Site-specific mutagenesis and kinetic studies
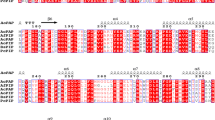
The tertiary structure of PDT has not been reported. However, amino acid sequences from a variety of organisms have now become available. The alignments of several bacterial PDTs (Fig. 2) indicated that Glu-64, Asp-76, Glu-89, His-115, Thr-183, Arg-184, Phe-185, and Arg-236 of C. glutamicum PDT are conserved among the enzymes compared. These conserved residues were targeted for site-directed mutagenesis to determine whether they play a role in catalysis or substrate binding. The effect of charge was examined by substituting the cationic group with a polar or neutral side-chain, while steric factors were addressed by replacing the residue for another positively charged group of a different size. For kinetic studies, the PDT activity in the crude extracts was purified by nickel-chelate chromatography. The purified proteins had an apparent molecular mass of approximately 39 kDa. The purification step resulted in a yield of 37–56% activity and a 27–62-fold increase in specific activity. The Glu64Asp enzyme retained 85% of its PDT activity, while the kinetic parameters of the reactions were remarkably different from those of the wild-type enzyme. The K m of the Asp-64 mutant increased 3.2-fold and the k cat 0.8-fold, while the k cat/K m was 2.5-fold lower than that of the wild-type enzyme. Glu64Val, Glu64Gln, and Glu64Ser proteins were well expressed but protein variants were inactive under the assay conditions (Table 5), indicating that the carboxyl group of Glu plays an important role in the catalytic activity.
Alignment of deduced amino acid sequence of Corynebacterium glutamicum (Cgl) prephenate dehydratase with some equivalent sequences. Eco, E. coli CM-PDT (TrEMBL Q8FEZ6); Ehe, Erwinia herbicola CM-PDT (TrEMBL Q02286); Hin, Haemophilus influenzae CM-PDT (TrEMBL P43900); Mja, Methanococcus jannascihii PDT (TrEMBL Q58054); Ame, Amycolatopis methanolica PDT (TrEMBL Q44104); Bsu, Bacillus subtilis PDT (TrEMBL P21203); Pst, Pseudomonas stutzeri CM-PDT (TrEMBL Q9RI01); Ath, Arabidopsis thaliana CM-PDT (TrEMBL Q8LFI1). Highly conserved residues among the enzyme are shown on a gray background. Residues subjected to site-directed mutagenesis are indicated by solid triangles
Amino acid substitution at position 183 had dramatic effects on the enzymatic activity (Table 5). The change from an hydroxyl-containing Thr side-chain to an aromatic (Tyr) or nonpolar (Ala) group resulted in a complete loss of catalytic activity. Compared with the wild-type enzyme, the catalytic activity of the Thr183Ser mutant was also severely compromised, with a 92% decrease in k cat, indicating the importance of the Thr residue in catalysis. The K m obtained with prephenate as the substrate was increased by a factor of about 5.6. Since the hydroxyl group of threonine side-chain is thought to be important for PDT activity and for maintaining the structure of E. coli CM-PDT (Baldwin and Davidson 1981), the corresponding Thr-183 residue, located within the TRF motif, of C. glutamicum PDT could play a similar role. It is interesting that the Thr183Ser mutant was very sensitive to inhibition by phenylalanine, and that an eight-fold increase in specific activity was observed after the addition of 500 μM tyrosine (Table 4).
To address the importance of Arg-184 in prephenate binding, two variant proteins, Arg184Lys and Arg184Leu, were examined. Although the positive charge was conserved in the lysine substitution, the resulting mutant (Arg184Lys) was inactive and poorly expressed in a crude cell extract (data not shown). Arg184Leu, by contrast, yielded a highly expressed and active PDT. Kinetic analysis of the Leu-184 variant showed a 7.7-fold increase in the K m for prephenate without a significant change in the k cat value. Therefore, this conserved Arg residue may be important in the binding of prephenate but is not an essential catalytic residue.
The removal of the aromatic side-chain at position 185 and its replacement with Leu resulted in an enzyme that is catalytically inactive. The Phe185Tyr mutant only retained about 3% of its enzymatic activity compared to the wild-type PDT (Table 5). Kinetic parameters were, however, obtained for this mutant using assays containing larger amounts of enzyme (0.5 mg per assay compared with 6.2 μg wild-type enzyme in the same assay). The resulting kinetic data showed a 91% decrease in k cat and a 23.7-fold increase in K m. Mutagenesis of Phe-185 affected K m much more strongly than k cat, suggesting that Phe-185 is important for substrate binding.
The kinetic parameters of Asp76Ala, Glu89Val, His115Phe, and Arg236Leu showed only small variations in K m and k cat values (data not shown) and implied that these residues play a minor or indirect role in the catalytic mechanism of PDT. However, Arg236Leu retained 24% PDT activity in the presence of 100 μM phenylalanine (Table 4), indicating that this residue is also involved in feedback inhibition of the enzyme. The far-UV CD spectra of mutant enzymes listed in Table 5 were essentially superimposable with that of the wide-type PDT (data not shown), demonstrating that no significant disruption in global secondary structure was caused by the point mutations.
Discussion
Phenylalanine analogues have been employed in the isolation of deregulated PDT enzymes in various organisms. In this study, six mutants with altered regulatory properties were obtained by PCR-based mutagenesis coupled to mFP selection. Sequence analysis of mFP-resistant mutants of C. glutamicum (Table 3) revealed that Ser-99 plays a role in feedback inhibition, and a subsequent mutagenesis study confirmed this fact. Studies using crude extracts or partially purified PDT of C. glutamicum (Sugimoto and Shiio 1974; Fazel and Jensen 1980; Follettie and Sinskey 1986) have demonstrated that these enzymes are activated by tyrosine. Although PDT activities of the mutant enzymes are not affected by low levels of phenylalanine, they are inhibited at higher phenylalanine concentrations—similar to the wild-type enzyme. It has been shown that chemical modification of Gly-309 of the bifunctional CM-PDT from E. coli (P-protein) reduced the extent of feedback inhibition by phenylalanine (Nelms et al. 1992). Fluorescence emission spectra and binding assays indicated that the interaction of phenylalanine with P-protein is localized to a C-terminal 12-kDa fragment, and deletion of this fragment leads to a fully functional enzyme that is insensitive to phenylalanine (Zhang et al. 1998). Deletion studies on the gene encoding CM-PDT in Erwinia herbicola showed that the regulatory domain of PDT is localized in the C-terminus of the bifunctional enzyme (Xia et al. 1992). Our earlier work also identified two amino acids, Arg-202 and Gly-224, that are important for feedback resistance of C. glutamicum PDT; these residues are located in the C-terminal region of this enzyme (Chan and Hsu 1996). Interestingly, each residue or both were included in the amino acid changes in mutants M1, M17, M25, and M36, confirming that Arg-202 and Gly-224 residues play an important role in phenylalanine inhibition of PDT. Based on the above findings, we conclude that Ser-99 is not directly involved in the binding of phenylalanine to the putative allosteric regulation site in C. glutamicum. Instead, this residue may be important in the maintenance of the proper conformation of the allosteric site in order to allow feedback regulation of the enzyme. Studies on the mechanism of phenylalanine binding and feedback inhibition in E. coli P-protein suggested that Arg-331 directly interacts with the acidic group of Phe to induce inhibition (Pohnert et al. 1999). A similar phenomenon could also occur at the equivalent residue, Arg-236, of C. glutamicum PDT since Leu substitution at this position yielded a mutant enzyme that was resistant to feedback inhibition by phenylalanine.
Another purpose of this investigation was to identify residues of C. glutamicum PDT that are essential for catalytic activity and/or substrate binding. Toward this goal, we conducted site-directed mutagenesis on eight conserved residues and compared the kinetic parameters of the reactions catalyzed by wild-type and selected mutant enzymes (Table 5). The results suggested that Glu-64 plays an important role in the dehydratase reaction of the enzyme, but Asp-76 and Glu-89 residues are not essential. By contrast, Zhang et al. (2000) demonstrated that the equivalent acidic residues (Glu-159, Asp-171, and Glu-183) of the E. coli P-protein dehydratase domain are not involved directly in the dehydration-decarboxylation of prephenate. It is generally accepted that protonation of the ring hydroxyl group triggers the concurrent loss of H2O and CO2 in a Grob-type fragmentation of the ionized prephenate. Since prephenate rapidly undergoes nonenzymatic acid-catalyzed aromatization (Zamir et al. 1983), PDT might need a weaker acid or perhaps only hydrogen bonding to promote the dehydration-decarboxylation reaction. Further insights concerning the role of Glu-64 awaits structural information on the active site of PDT.
Threonine was previously proposed to be important for the activity of E. coli K12 CM–PDT (Davidson et al. 1972). Moreover, Jetten and Sinskey (1995) speculated that Thr-183 in C. glutamicum PDT, which corresponds to Thr-278 of E. coli P-protein, may be important for enzyme activity. In order to assess the role of this conserved residue, we substituted the threonine with alanine, serine, and tyrosine. The Thr183Ser enzyme retained approximately 4% of wild-type PDT activity but showed a 5.6-fold increase in K m for prephenate (Table 5). In contrast, the other two mutant proteins were inactive. Like threonine, tyrosine also possesses a hydroxyl group; however, its bulky side-chain may alter the conformation of the active site through steric effects. Using computer graphics modeling, Xue and Lipscomb (1995) reported that a threonine hydroxyl group in the active site of Saccharomyces cerevisiae chorimate mutase can form a hydrogen bond with the carboxyl group of an inhibitory chorismate analogue. The data in the present study suggest that Thr-183 of C. glutamicum PDT might also hydrogen bond to prephenate or to active site residues involved in catalysis.
A number of studies have established a role for cation-π interactions in substrate-enzyme binding (Scrutton and Raine 1996; Lee et al. 2002) and protein-ligand interactions (Ting et al. 1998; Fermandez-Recio et al. 1999). For example, the Phe-54 and Leu-115 of B. subtilis CM appear to lock the reactive C-terminus in an orientation that promotes the rearrangement of chorismate to prephenate (Chook et al. 1994). Based on the results of our study, we believe that C. glutamicum PDT achieves the effect, in part, by lining the binding pocket with hydrophobic residues that anchor the substrate by means of short van der Waals interactions.
In summary, site-directed mutagenesis was conducted on eight conserved residues (Glu-64, Asp-76, Glu-89, His-115, Thr-183, Arg-184, Phe-185, and Arg-236) in order to identify those essential for the catalytic mechanism of C. glutamicum PDT. Glu-64, Thr-183, Arg-184, and Phe-185 of PDT were shown to play important roles during the dehydration-decarboxylation reaction of prephenate. Determination of the three-dimensional structure of C. glutamicum PDT, which is currently in progress, should help clarify the roles of these conserved residues in the mechanism of the enzyme.
References
Ahmad S, Jensen RA (1986) The evolutionary history of two bifunctional proteins that emerged in the purple bacteria. Trends Biochem Sci 11: 108–112
Ahmad S, Wilson AT, Jensen RA (1988) Chorismate mutase:prephenate dehydratase from Acinetobacter calcoaceticus. Purification, properties and immunological cross-reactivity. Eur J Biochem 116: 69–79
Baldwin GS, Davidson BE (1981) A kinetic and structural comparison of chorismate mutase/prephenate dehydratase from mutant strains of Escherichia coli K12 defective in the pheA gene. Arch Biochem Biophys 211:66-75
Baldwin GS, Mckenzie GH, Davidson BE (1981) The self-association of chorismate mutase/prephenate dehydratase from Escherichia coli K12. Arch Biochem Biophys 211: 76–85
Bathe B, Kalinowski J, Pühler A (1996) A physical and genetic map of the Corynebacterium glutamicum ATCC 13032 chromosome. Mol Gen Genet 252: 255–265
Byng GS, Jensen RA (1983) Impact of isozymes upon partitioning of carbon flow and regulation of aromatic biosynthesis in prokaryotes. In: Ratazzi MC, Scandalios JG, Whitt GS (eds) Isozymes, vol. 8, Alan R. Liss, New York, pp 115–140
Cadwell RC, Joyce CF (1992) Randomization of genes by PCR mutagenesis. PCR Methods Appl 2: 28–33
Chan MS, Hsu WH (1996) Cloning of m-fluorophenylalanine-resistant gene and mutational analysis of feedback-resistant prephenate dehydratase from Corynebacterium glutamicum. Biochem Biophys Res Comm 219: 537–542
Chook YM, Gray JV, Ke H, Lipscomb WN (1994) The monofunctional chorismate mutase from Bacillus subtilis. Structure determination of chorismate mutase and its complexes with a transition state analog and prephenate, and implications for the mechanism of the enzymatic reaction. J Mol Biol 240:476–500
Davidson BE, Blackburn EH, Dopheide TA (1972) Chorismate mutase-prephenate dehydratase from Escherichia coli K-12. I. Purification, molecular weight, and amino acid composition. J Biol Chem 247:4441–4446
Dopheide TA, Crewther P, Davidson BE (1972) Chorismate mutase-prephenate dehydratase from Escherichia coli K-12. II. Kinetic properties. J Biol Chem 247: 4447–4452
Dougherty DA (1996) Cation-pi interactions in chemistry and biology: a new view of benzene, Phe, Tyr, and Trp. Science 271:163–168
Euverink GJW, Wolters DJ, Dijkhuizen L (1995) Prephenate dehydratase of the actinomycete Amycolatopsis methanolica: purification and characterization of wild-type and deregulated mutant proteins. Biochem J 308: 313–320
Fazel RS, Jensen RA (1980) Regulation of prephenate dehydratase in coryneform species of bacteria by L-phenylalanine and by remote effectors. Arch Biochem Biophys 200: 165–176
Fermandez-Recio J, Romero A, Sancho J (1999) Energetics of a hydrogen bond (charged and neutral) and of a cation-pi interaction in apoflavodoxin. J Mol Biol 290:319–330
Follettie MT, Sinsky AJ (1986) Molecular cloning and nucleotide sequence of the Corynebacterium glutamicum pheA gene. J Bacteriol 167: 659–702
Gething MJH, Davidson BE (1976) Chorismate mutase/prephenate dehydratase from Escherichia coli K12. 2. Evidence for identical subunits catalyzing the two activities. Eur J Biochem 71: 327–336
Hagino H, Nakatama K (1974) L-Phenylalanine production by an analogue-resistant mutant of Corynebacterium glutamicum. Agric Biol Chem 38:157–161
Ikeda M, Katsumata R (1992) Metabolic engineering to produce tyrosine or phenylalanine in a tryptophan-producing Corynebacterium glutamicum strain. Appl Environ Microbiol 58: 781–785
Jetten MS, Sinsky AJ (1995) Recent advances in the physiology and genetics of amino acid-producing bacteria. Crit Rev Biotechnol 15:73–103
Kuninka A (1986) Nucleic acid, nucleotides and related compounds. In: Rehm HJ, Reed G (eds) Biotechnology vol. 4. Microbial production. Neenbeem VCH Verlagsyese II Schaff, Weinheim, Germany pp 71–117
Kunkel TA, Roberts JD, Zakour RA (1987) Rapid and efficient site-specific mutagenesis without phenotype selection. Methods Enzymol 154: 367–38
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685
Lee S, Lin X, McMurray J, Sun G (2002) Contribution of an active site cation-pi interaction to the spectroscopic properties and catalytic function of protein tyrosine kinase Csk. Biochemistry 41:12107–12114
Liebl W, Ehrmann M, Ludwig W, Schleifer KH (1991) Transfer of Brevibacteria divaricatum DSM 20297, Brevibacteria flavum DSM 20412 and DSM 1412 and Corynebacterium lilium DSM 20137 to Corynebacterium glutamicum and their distinction by rRNA gene restriction patterns. Int J Syst Bacteriol 41: 225–260
Nelms J, Edwards RM, Warwick J, Fotheringham I (1992) Novel mutations in the pheA gene of Escherichia coli K-12 which result in highly feedback inhibition-resistant variants of chorismate mutase/prephenate dehydratase. Appl Environ Microbiol 58:2592–2598
Pierson DL, Jensen RA (1974) Metabolic interlock: control of an interconvertible prephenate dehydratase by hydrophobic amino acid in Bacillus subtilis. J Mol Biol 252: 5839–5846
Pohnert G, Zhang S, Husain A, Wilson DB, Ganem B (1999) Regulation of phenylalanine biosynthesis. Studies on the mechanism of phenylalanine binding and feedback inhibition in the Escherichia coli P-protein. Biochemistry 38:12212–12217
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laborotary manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York pp17.2–17.44
Scrutton NS, Raine AR (1996) Cation-pi bonding and amino-aromatic interactions in the biomolecular recognition of substituted ammonium ligands. Biochem J 319:1-8
Shiio I, Sugimota S (1979) Two components of chorismate mutase in Brevibacterium flavum. J Biochem (Tokyo) 86: 17–25
Studier FW, Moffatt BA (1986) Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol 189:113–130
Sugimoto S, Shiio I (1974) Regulation of prephenate dehydratase in Brevibacterium flavum. J Biochem (Tokyo) 76:1103–1111
Ting AY, Shin I, Lucero C, Schultz G (1998) Energetic analysis of an engineered cation-π interaction in staphylococcal nuclease. J Am Chem Soc 120:7135–7136
Xia T, Zhao G, Jensen RA (1992) Loss of allosteric control but retention of the bifunctional catalytic competence of a fusion protein formed by excision of 260 base pairs from the 3′ terminus of pheA from Erwinia herbicola. Appl Environ Microbiol 58:2792–2798
Xue Y, Lipscomb WN (1995) Location of the active site of allosteric chorismate mutase from Saccharomyces cerevisiae, and comments on the catalytic and regulatory mechanisms. Proc Natl Acad Sci USA 92:10595–10598
Yanofsky C (1988) Transcription attenuation. J Biol Chem 263: 609–612
Yoshinga F, Nakamori S (1983) Production of amino acids. In: Hermann KM, Somerville KL (eds) Amino acid, biosynthesis and genetic regulation. Addison Wesley, New York, pp 310–355
Zamir LO, Tiberio R, Jensen RA (1983) Differential acid-catalyzed aromatization of prephenate, arogenate, and spiroarogenate. Tetrahedron Lett 24:2815-2818
Zhang S, Pohnert G, Kongsaeree P, Wilson DB, Clardy J, Ganem B (1998) Chorismate mutase-prephenate dehydratase from Escherichia coli. Study of catalytic and regulatory domains using genetically engineered proteins. J Biol Chem 273:6248–6253
Zhang S, Wilson DB, Ganem B (2000) Probing the catalytic mechanism of prephenate dehydratase by site-directed mutagenesis of the Escherichia coli P-protein dehydratase domain. Biochemistry 39:4722–4728
Acknowledgements
This work was supported by grants NSC88–2311-B-005-030 and NSC89–2316-B-005–018 from the National Science Council of the Republic of China.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hsu, SK., Lin, LL., Lo, HH. et al. Mutational analysis of feedback inhibition and catalytic sites of prephenate dehydratase from Corynebacterium glutamicum . Arch Microbiol 181, 237–244 (2004). https://doi.org/10.1007/s00203-004-0649-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-004-0649-5