Abstract
Cellulomonas uda efficiently solubilized chitinous substrates with a simple chitinase system composed of an endochitinase, designated ChiA, which hydrolyzed insoluble substrates into long-chain chitooligosaccharides, and an as yet uncharacterized exochitinase activity. ChiA, isolated from culture supernatant fluids, was found to be a glycosylated endochitinase with an apparent molecular mass of approximately 70 kDa and pI of 8.5. The gene encoding ChiA was cloned in Escherichia coli and sequenced, revealing an open reading frame of 1,716 bp encoding a 571-amino-acid protein with a predicted molecular mass of 59.2 kDa. The region upstream of chiA included a conserved −35 hexamer flanked by two direct repeats analogous to those found in many Streptomyces chitinase promoters, and thought to function as binding sequences for regulatory proteins. Analysis of the deduced amino acid sequence showed a modular protein consisting of a signal peptide at its N terminus, a family 2 carbohydrate-binding module (CBM2) that was closely related to the substrate-binding domains of glycosyl hydrolases from distantly related bacteria, and a family 18 glycosyl hydrolase catalytic module related to Streptomyces chitinases. In contrast to the fibronectin type III domains of Streptomyces chitinases, the linker region between modules in ChiA consisted of a long proline- and threonine-rich module, thought to contribute to the glycosylation and flexibility of the mature protein.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chitin is an insoluble polymer of β-(1,4)-linked N-acetylglucosamine (NAG) residues that functions as a structural polysaccharide in the exoskeletons of arthropods and the cell walls of most fungi (Muzzarelli 1977). It is an abundantly produced polymer on Earth and, for this reason, its degradation contributes greatly to both the carbon and nitrogen cycles in the biosphere. The biodegradation of chitin is catalyzed by chitinases, glycosyl hydrolases that recognize and cleave β-(1,4) bonds in chitin molecules. Chitinases are produced by a vast array of microbes to meet nutritional requirements, since chitin may act as their source of carbon, nitrogen, and energy.
Most information on chitinases derives from analyses of cloned genes (Cohen-Kupiec and Chet 1998; Felse and Panda 1999). Chitinases are modular enzymes, generally comprising a substrate-binding module (carbohydrate-binding module or CBM) and a catalytic domain. CBMs are found not only in chitinases, but also in other glycosyl hydrolases; such as cellulases and xylanases; and, in some cases, they have been shown to bind to chemically related substrates (e.g., cellulose and chitin) with the same specificity (Henrissat 1999). Based on sequence and structural similarities, the catalytic domains of chitinases are classified into families 18 and 19 of glycosyl hydrolases (Henrissat and Bairoch 1993; Henrissat 1999). This classification also reflects a different mechanism of action in the cleavage of the β-(1,4) bond: a retaining mechanism by family 18 chitinases or an inverting mechanism by chitinases of family 19 (Koga et al. 1999). Most bacterial chitinases known to date belong to family 18 glycosyl hydrolases. Although family 19 was thought to be composed only of plant chitinases, it is now known that family 19 chitinases are widespread among species of Streptomyces (Ohno et al. 1996; Saito et al. 1999) and can be found in other bacteria (Ueda et al. 2003).
Chitin biodegradation has been studied extensively in marine environments, where vast amounts of chitin are synthesized annually. However, chitin is also abundant in terrestrial environments (Muzzarelli 1977). Advancing our understanding of the biodegradation of chitin in terrestrial environments is of considerable importance since chitin serves as a major source of combined carbon and nitrogen for soil bacteria. Investigations of chitinase diversity in soil pastures have shown that actinobacteria-like chitinase sequences are predominant, suggesting a major role for actinobacteria in the mineralization of chitin in terrestrial environments (Metcalf et al. 2002). While several chitinase sequences appear to match those of known enzymes from representative groups of actinobacteria, such as species of Streptomyces, a large number apparently are from as yet uncultured bacteria or from cultured bacteria whose chitinases have not yet been studied (Metcalf et al. 2002).
Our previous finding, that free-living species of cellulomonads might play a pivotal role in both the aerobic and anaerobic decomposition of chitin in terrestrial ecosystems (Reguera and Leschine 2001), and the scarce information available on the chitinolytic properties of this group of actinobacteria prompted us to further investigate the chitinase system of a representative member of the group. While most chitinolytic bacteria known to date efficiently degrade crystalline forms of chitinous substrates through the synergistic action of a battery of chitinolytic enzymes (Svitil et al. 1997; Saito et al. 2001), we found that the chitinase system of Cellulomonas uda lacked this complexity and that a single endochitinase, designated ChiA, was predominantly responsible for the solubilization of insoluble chitinous substrates. In the present study, we purified and characterized ChiA, and we cloned and sequenced the encoding gene to better understand the biochemistry of the enzyme system that C. uda employs for the efficient degradation of insoluble chitinous substrates. Our results emphasize the importance of investigating the biochemistry of chitin degradation in relevant soil bacteria, such as C. uda, in order to advance our understanding of the diversity of chitinase systems in terrestrial environments.
Materials and methods
Bacterial strains and culture conditions
C. uda ATCC 21399 (formerly Cellulomonas sp. strain ATCC 21399; Stackebrandt and Kandler 1979) was cultured aerobically at 32 °C with gentle shaking as previously described (Reguera and Leschine 2001) in medium GS2 (Cavedon et al. 1990) or nitrogen-limited medium *GS2 (Reguera and Leschine 2001), containing 0.2% (w/v) colloidal chitin (Hsu and Lockwood 1974) as growth substrate. In some experiments, 0.2% (v/v) chitobiose, cellobiose, or glucose replaced colloidal chitin in growth media. Escherichia coli strains were grown at 37 °C in Luria-Bertani (LB) medium containing ampicillin (100 μg/ml).
Enzyme and protein assays
Carboxymethylcellulase (CMCase) activity was determined by measuring the production of reducing sugars from carboxymethylcellulose as described previously (Reguera and Leschine 2001). One unit of CMCase activity was defined as the amount of enzyme that released 10 nmol reducing sugar (as glucose equivalents) per hour. Chitinase activity was determined by a modification of the method of Jeuniaux (1966), with colloidal chitin substrate, as previously described (Reguera and Leschine 2001). Reaction mixtures were prepared in a total volume of 5 ml with colloidal chitin (2.4 mg) and sodium phosphate buffer, pH 7.2 (0.37 mmol), and incubated at 50 °C for 4 h. In some experiments colloidal chitin was replaced by ball-milled chitin as a more crystalline chitinous substrate. Ball-milled chitin was prepared by milling practical-grade chitin flakes in H2O for 3 days following the method described for the preparation of ball-milled filter paper (Leschine and Canale-Parola 1983). A decrease in turbidity of the suspension (absorbance at 660 nm) was used as a measure of chitinase activity. One unit of activity was defined as the amount of enzyme that hydrolyzed 10 μg substrate per hour under the assay conditions.
N-acetylglucosaminidase (chitobiase) activity was assayed using 4-methylumbelliferyl-N-acetylglucosamine (4MU-NAG) and endo- and exo-chitinase activities were assayed using 4-methylumbelliferyl-chitotrioside (4MU-(NAG)3) or 4-methylumbelliferyl-chitobioside (4MU-(NAG)2), respectively. All three assays were carried out as modifications of the procedure described by O’Brien and Colwell (1987). Reaction mixtures were prepared by mixing 1.99 ml 0.1 M sodium phosphate buffer (pH 7.2) and 10 μl enzyme preparation (approximately 0.4 μg purified ChiA, or 10 μg protein in concentrated culture supernatant fluids) in a glass cuvette. Substrate (0.05 nmol) was added to reaction mixtures and mixed by inversion. Reaction mixtures were incubated at room temperature for 1 min and release of fluorescence was immediately measured using a Hoefer DyNa Quant 200 Fluorometer (Hoefer Amersham Biosciences, San Francisco, Calif., USA) equipped with a fixed excitation bandpass source of 365 nm and a fixed emission bandpass filter of 460 nm. The fluorochrome concentration was calculated by using a standard curve generated with 4-methylumbelliferone (4MU, Sigma, St. Louis, Mo., USA) following the protocol described by the fluorometer’s manufacturer. One unit of activity was defined as the amount of enzyme that released 1 pmol 4MU per min under assay conditions.
Protein concentrations were determined with the Pierce bicinchoninic acid protein assay reagent (Smith et al. 1985) (Pierce, Rockford, Il., USA), using bovine serum albumin as protein standard.
Purification and characterization of ChiA
C. uda cultures in GS2-chitin medium that had been incubated until most insoluble substrate was degraded were used to inoculate (10% (v/v) inoculum) *GS2-chitin medium, which was then incubated until all the insoluble substrate was degraded (2–3 days). Culture supernatant fluids were obtained by pelleting the cells by centrifugation (20 min, 4,000×g, 4 °C). NaN3 was added to supernatant fluids to a final concentration of 0.025% (w/v) before they were concentrated using a tangential flow Ultrasette (Pall Filtron, Northborough, Mass., USA) equipped with a 10,000-M r -cutoff membrane (Pall Filtron). Anion-exchange chromatography of concentrated supernatant fluids was carried out using a Mono Q HR5/5 column (5×0.5 cm) and a Fast Protein Purification Liquid Chromatography (FPLC) System (Amersham Biosciences, Uppsala, Sweden) operated at room temperature. The column was equilibrated with 20 mM bis-Tris propane buffer at a pH of 7.2. Elution of supernatant proteins was effected in equilibration buffer, at a flow rate of 1 ml/min, with a 0–1 M NaCl step-wise gradient program (Fig. 1A). The peak threshold level was set to 1% full scale and the sample size was 1 ml. Seven protein peaks were detected by absorbance at 280 nm and the two major ones were designated F1 and F2. Peak samples (3 ml each) from several injections were collected and pooled for further analyses. Pooled fractions were desalted, their buffer exchanged with 100 mM sodium phosphate buffer (pH 7.2, 0.025 (w/v) NaN3), and further concentrated using an Omegacell (Pall Filtron) stirred cell equipped with a 3,000-M r -cutoff membrane (Pall Filtron).
Purification of ChiA of Cellulomonas uda. a Separation by anion-exchange chromatography of extracellular proteins produced by C. uda when grown in a chitin-containing medium. The two major protein peaks were designated F1 and F2. b Samples of F1, the only chitinase-active fraction (F1) and of concentrated supernatants fluids (S) were analyzed by SDS-PAGE. A single polypeptide with an apparent molecular mass of 70 kDa, designated ChiA, was the only protein band found in fraction F1. Numbers at left are molecular masses in kDa. c Protein in fraction F1 (F1) and in concentrated supernatants fluids (S) separated by SDS-PAGE were transferred to a nitrocellulose membrane. Protein glycosylation was detected directly on the membrane in a colorimetric immunoassay. ChiA gave a positive reaction in the assay, suggesting it is a glycoprotein
Proteins from culture supernatants (1 μg) or purified ChiA (0.1 μg) were subjected to PAGE (Laemmli 1970) under nondenaturing and denaturing conditions using, respectively, 4–15% and 10–15% linear gradient PhastGels in a PhastSystem (Amersham Biosciences). Proteins in gels were silver-stained following the method of Heukeshoven and Dernick (1985) as modified by the manufacturer (Amersham Biosciences). Molecular weight standards (broad range) were obtained from BioRad. Where indicated, proteins separated by SDS-PAGE were subjected to renaturation following the method of Williams et al. (2000), except that all renaturation steps were done at 4 °C. Isoelectric points were determined using a PhastSystem and IEF 3–9 PhastGels (Amersham Biosciences) following the manufacturer’s recommendations. Standards were from Sigma (pI, 3–9.5). Proteins from polyacrylamide gels were eluted using Nanosep Centrifugal Devices (Pall Gelman Laboratory, Ann Arbor, Mich., USA) equipped with a 300,000-M r -cutoff Omega membrane, as previously described (http://www.pall.com/laboratory/lifesa/fast_and_efficient.asp). For glycoprotein detection, proteins were separated by SDS-PAGE and transferred to a polyvinyledene difluoride Immobilon transfer membrane (Millipore, Bedford, Mass., USA) using a MINI-Trans-Blot electrophoretic transfer cell (BioRad) operated at 50 V (constant voltage) for 30 min (Towbin et al. 1979). Glycosylated proteins were detected using the Böhringer Mannheim (Roche Molecular Biochemicals, Indianapolis, Ind., USA) DIG Glycan Detection Kit following procedures recommended by the manufacturer.
Chitinase activity in gels subjected to electrophoresis was detected by means of polyacrylamide gel zymograms that contained: glycol chitin (Trudel and Asselin 1989), 0.2 g; sodium phosphate buffer, pH 7.2, 6.1 mmol; polyacrylamide, 4.75 g; bis-acrylamide, 0.16 g; ammonium persulfate, 0.05 g; TEMED, 0.05 ml; and distilled water, in a total volume of 100 ml. The zymogram solution was cast on a GelBond film (FMC, Rockland, Me., USA) to a thickness of 1.0 mm and allowed to polymerize. The zymogram was then incubated in a humidity chamber for 1 h at 42 °C in contact with the gel that had been subjected to electrophoresis. After incubation, zymograms were stained for residual substrate with an aqueous solution of 0.1% (w/v) Congo red, destained with 1 M NaCl, and fixed with 5.0% acetic acid, in order to visualize areas where the glycol chitin had been hydrolyzed.
For amino acid sequencing, purified ChiA was proteolytically digested with trypsin and alkylated to ensure absence of linkage of multiple peptides through disulfide bonds before peptide separation by reverse-phase HPLC on an octadecyl silica gel (C18) column (PAN Facility, Stanford University Medical Center, Palo Alto, Calif., USA). The HPLC peaks were analyzed by MALDI-MS spectra and the peak that gave the clearest signal (at about 2,200 Da) was sequentially cleaved (Edman and Begg 1967) and the amino acids identified using an ABI Sequenator operated in liquid-pulse mode (PAN Facility). The ChiA internal amino acid sequence was analyzed for similarity to other protein sequences in the database using the NCBI BLAST program.
Biochemical properties of chitinase ChiA
ChiA and concentrated supernatant fluids from cultures grown in *GS2 medium supplemented with chitin were assayed for optimum pH and temperature for chitinase activity. To measure optimum temperature, chitinase assay reaction mixtures were incubated at 25, 30, 40, 45, 50, 55, 60, 65, 75, and 85 °C. Optimum pH was determined as in the chitinase assay except that different buffers, ranging from pH 3.0 to 9.0, were used. These buffers were 0.1 M citrate-phosphate buffer (pH 3.0–6.6), 0.1 M sodium phosphate buffer (pH 6.0–8.0), and 0.1 M Tris/HCl buffer (pH 8.0–9.0). All buffers were prepared according to the method of Gomori (1955).
The binding properties of ChiA were studied in a binding assay mixture that contained excess substrate (10 mg), enzyme (4 μg), and H2O to a final volume of 600 μl. Reaction mixtures were incubated at 32 °C for 2.5 h before they were subjected to microcentrifugation at maximum speed for 6 min. Chitinase activity present in supernatant fractions was determined as a measure of activity that did not bind to the substrate. Substrates tested were: chitin from crab shells, both practical-grade and purified powder (Sigma); ball-milled chitin and colloidal chitin, both prepared as described above; chitosan (Sigma); ball-milled filter paper (Leschine and Canale-Parola 1983); Avicel (microcrystalline α-cellulose; FMC); Solka floc (BW 200; Brown, Berlin, N.H., USA); and Sephadex G-10 (cross-linked dextran; Sigma).
For HPLC analysis of chitin hydrolysis products, chitinase-active supernatant fluids or purified ChiA were incubated at 50 °C with colloidal chitin in standard chitinase assay reaction mixtures, as described above, except that reactions were incubated for 4 and 21 h. Soluble products were analyzed by means of HPLC using a Model 2300 HPLC pump (ISCO, Lincoln, Neb., USA) connected to a refractive index detector (SpectraPhysics, San Jose, Calif., USA). Chitooligosaccarides were separated out at room temperature using an aminopropyl silica-based column (SUPELCOSIL LC-NH2; Supelco, Bellefonte, Pa., USA) with a particle size of 5 μm and a mobile phase of acetonitrile:water (65:35) at a flow rate of 0.8 ml/min.
Recombinant DNA techniques
Genomic DNA from C. uda cultures growing exponentially in GS2-glucose or cellobiose medium was extracted using the G NOME extraction kit (BIO 101, Vista, Calif., USA) following the manufacturer’s recommendations. Basic recombinant techniques were carried out as described elsewhere, (Sambrook et al. 1989) using commercially available kits. Plasmid pEMU723R, carrying a 1.9-kb fragment containing chiC (GenBank accession no. D12647) from Streptomyces lividans TK64 (Fujii and Miyashita 1993), was kindly provided by Dr. Miyashita and Dr. Saito (National Institute of Agro-Environmental Sciences, Ibaraki, Japan) and was used as template in PCR reactions to prepare a 221-bp digoxigenin (DIG)-labeled probe with left (5′-ACCACGTGAAGAACCTGGTC-3′) and right (5′-GTACTTGGCCTTCAGCTTGC-3′) primers. DIG was incorporated during PCR using the PCR DIG Probe Synthesis Kit (Roche Molecular Biochemicals) following the protocol specified by the manufacturer.
For the cloning of chiA, genomic DNA was digested overnight with several restriction endonucleases that recognized 6- and 8-mer sequences. Products of restriction digestion were separated in a 1% agarose gel and transferred onto a positively charged nylon membrane (Roche Molecular Biochemicals) to identify a 7-kb NotI fragment that hybridized with the probe (Sambrook et al. 1989). NotI fragments from C. uda genomic DNA between 6 and 8 kb were isolated after agarose gel electrophoresis and ligated to dephosphorylated NotI-digested pBlueScript II SK(+) vector (Stratagene, La Jolla, Calif., USA), and introduced into E. coli XL1-Blue MRF′ supercompetent cells (Stratagene). Transformants were selected, replica-plated, and screened by colony hybridization (Sambrook et al. 1989) using the 221-bp DIG-labeled probe.
The nucleotide sequences of cloned inserts were determined using a subcloning strategy and “gene walking” procedure with custom sequencing primers. DNA was sequenced using an ABI 377XL automated DNA sequencer in conjunction with the ABI PRISM BigDye Terminator cycle sequencing reaction kit. Due to the high G+C content of the DNA sequenced (76.3 mol%), reaction mixtures were generally prepared with DMSO (8% (v/v), final concentration) or Q solution (Qiagen). Computer analyses of nucleic acid and predicted amino acid sequences were carried out with the software available at the SDSC Biology Workbench (University of California San Diego) and the BLAST search of the National Center for Biotechnology Information (NCBI). ORF analyses were done with the NCBI program ORF-Finder (http://www.ncbi.nlm.nih.gov/gorf/gorf.html). The domain structure of chiA was analyzed using the NCBI Conserved Domain Search (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) and information found at the CAZy website (http://afmb.cnrs-mrs.fr/CAZY/).
Nucleotide sequence accession number
The nucleotide sequence of chiA has been deposited in the GenBank database under accession number AY008839.
Results
Purification and properties of ChiA
Proteins in supernatant fluids from C. uda cultures grown in *GS2-colloidal chitin medium were separated by means of anion-exchange chromatography as described in Materials and methods. Two major protein fractions (F1 and F2) were detected, as well as several minor protein peaks (Fig. 1A). Most of the chitinase activity was present in a single fraction, F1, which eluted before the NaCl gradient was applied, while no chitinase activity was detected in F2 (Fig. 1A). A second peak with low levels of chitinase activity eluted after protein fraction F2. However, the limited amount of protein in this fraction prevented further characterization. Samples of fraction F1 from several chromatography injections were pooled and analyzed by SDS-PAGE. After silver staining, a single protein band appeared on gels that corresponded to a polypeptide with an apparent molecular mass of approximately 70 kDa (Fig. 1B). Protein in fraction F1 and proteins in whole supernatant fluids separated by SDS-PAGE were transferred to a nitrocellulose membrane and the presence of glycoproteins was examined after oxidative treatment and labeling with DIG. The results shown in Fig. 1C indicate that the 70-kDa polypeptide reacted with a DIG-specific antibody, suggesting that it was a glycoprotein.
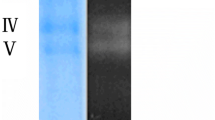
Attempts to renature protein(s) in fraction F1 or chitinase-active supernatants following SDS-PAGE resulted in the recovery of some activity in areas of the gel where the 70-kDa polypeptide was expected to migrate; however, diffusion of proteins during the renaturation procedure prevented precise localization of the activity (data not shown). For this reason, protein in F1 was also analyzed by nondenaturing-PAGE (Fig. 2). A single band was revealed after silver staining, which migrated slowly into the gel (Fig. 2A) and showed chitinase activity on glycol-chitin zymograms (Fig. 2B). This chitinase-active band (Fig. 2B) was eluted from nondenaturing gels, concentrated ten-fold and subjected to SDS-PAGE, confirming the presence of a 70-kDa protein band in the chitinase-active band (data not shown). The 70-kDa-protein, purified ten-fold by ultrafiltration and anion-exchange chromatography (Table 1), was therefore a chitinase, and was designated ChiA.
Nondenaturing PAGE and zymogram analyses of fraction F1 and concentrated culture supernatant fluids from C. uda. a Proteins present in samples of fraction F1 (F1) and in concentrated culture supernatants (S) were analyzed by nondenaturing PAGE. b Glycol-chitin zymogram analysis of proteins present in (a) showing areas of chitinase activity due to ChiA
ChiA had a molecular mass of approximately 70 kDa and a pI of 8.5. Maximum ChiA activity against colloidal chitin was observed at 60 °C, and higher temperatures resulted in a significant loss of activity. The optimum temperature for chitinase activity in culture supernatant fluids was 55 °C. Optimum pH for ChiA activity was approximately 7.0. This was also the optimum pH for chitinase activity of culture supernatants.
ChiA preferentially bound to crystalline chitinous substrates, such as purified chitin powder and ball-milled chitin (100% of activity bound to the substrate) and, to a lesser extent, to Sephacryl (67%), colloidal chitin (61%), ball-milled cellulose (61%), Sephadex (54%), and chitosan (46%). ChiA did not bind to practical-grade chitin. This commercial form of chitin (dehydrated ground crab shells) differs significantly from native chitin found in nature, inasmuch as bound H2O molecules help maintain the highly ordered structure of native chitin and are thought to be essential for binding and hydrolysis by chitinases (Koga et al. 1999).
ChiA was assayed for activity against chitinous substrates with different degrees of crystallinity (ball-milled chitin, colloidal chitin, and glycol chitin) and also with cellulosic substrates. Highest levels of activity (5,647 U per mg protein) were detected when colloidal chitin was used as the substrate in enzyme assays, but ChiA also showed activity (183 U per mg protein) against ball-milled chitin, a crystalline form of chitin. Activity against glycol chitin was not detected following standard assays based on the detection of NAG as a hydrolysis product (Jeuniaux 1966), but could be demonstrated by means of glycol-chitin zymograms (Fig. 2B). ChiA did not exhibit CMCase activity.
The activity of ChiA was also examined using 4MU-linked substrates as analogues of chitotetraose, chitotriose, and chitobiose (Table 2). Highest activities for ChiA were detected with 4MU-(NAG3) (analogue of chitotetraose), suggesting that ChiA was an endo-acting chitinase. No activity was detected against 4MU-NAG (analogue of chitobiose) and only low levels of activity were detected with 4MU-(NAG2) (analogue of chitotriose) as substrate. Endochitinase was also the predominant activity present in culture supernatant fluids, but exochitinase activity also was present. Only very low levels of chitobiase were detected in culture supernatant fluids, suggesting that chitin was degraded extracellularly to chitobiose, which was assimilated intracellularly.
End products of hydrolysis of colloidal chitin by purified ChiA or chitinase-active supernatant fluids were analyzed by means of HPLC. Figure 3 shows HPLC elution profiles of chitin hydrolysis products following incubation of colloidal chitin with ChiA or culture supernatant fluids for 4 and 21 h. Incubation times longer than 21 h led to loss of chitinase activity in the samples and no changes in the pattern of digestion products from colloidal chitin were observed (data not shown). After a 4-h incubation of ChiA with colloidal chitin, all insoluble substrate was degraded producing large chitooligosaccarides that were further hydrolyzed into shorter chitooligosaccarides when incubation was extended for 21 h (Fig. 3B). No significant peaks of NAG, chitobiose, or chitotriose were detected, indicative of an endo-acting chitinase. When culture supernatant fluids were incubated with colloidal chitin, the chitooligosaccharide peak shifted from longer to shorter molecules between 4 and 21 h of incubation; however, in contrast to ChiA-catalyzed reactions, a peak of chitobiose was present (Fig. 3B). Thus, an additional activity, presumably an exochitinase that hydrolyzed the soluble chitooligosaccharides produced by ChiA to chitobiose, also was present in culture supernatants.
Cloning and sequence analysis of chiA
The amino (N)-terminus of native ChiA, purified from supernatant fluids of C. uda cultures grown on colloidal chitin, apparently was blocked; therefore, purified protein was digested with trypsin in order to obtain internal sequence information. A 2,050-Da peptide was subjected to Edman degradation and an 18 amino-acid N-terminal sequence (AYTADQSVDGVADTXDQP, where X corresponds to an undetermined amino acid) was identified. BLAST analyses of this ChiA internal amino acid sequence showed it was highly similar to regions in the catalytic domain of family 18 chitinases from Streptomyces species and a chitinase from Cellulomonas sp. strain GM13. The greatest similarity (94% identity out of the 18 amino acids) was found with chitinases from S. lividans, Streptomyces coelicolor, Streptomyces plicatus, and Streptomyces peucetius. In all homologous sequences, the amino acid at position “X” in ChiA was a conserved tryptophan (W), suggesting that this might be the undetermined amino acid in the ChiA sequence.
The Streptomyces chitinases with internal amino acid sequences similar to that of ChiA are encoded by highly conserved modular genes. Based on this information, a pair of primers was designed to amplify a 221-bp region of the catalytic domain of S. lividans chiC, which was used as a probe in the cloning of chiA from C. uda. In a NotI-digested C. uda chromosomal DNA preparation, a single fragment of about 7 kb hybridized to the probe. For this reason, NotI fragments of C. uda DNA between 6 and 8 kb were cloned in E. coli. After screening transformants by colony hybridization using the 221-bp probe, two positive clones, designated pSK5 and pSK17, were isolated.
Restriction endonuclease digestion and sequence analysis confirmed the presence of a 7-kb NotI-insert in pSK5 (Fig. 4A). The DNA inserted in pSK17 was found to be about 11 kb and it contained the same NotI 7-kb fragment found in pSK5 plus two additional flanking regions (Fig. 4A), suggesting that it was generated as a result of a partial digestion by the restriction endonuclease.
Genetic map of cloned 7-kb (pSK5) and 11-kb (pSK17) fragments from C. uda chromosomal DNA containing chiA. b Putative promoter region of chiA identified upstream of the starting ATG codon. The deduced amino acid sequence of the first codons identified in the chiA sequence is shown in italics. Upstream of the Shine-Dalgarno sequence (AGGA, boxed), a pair of direct (dotted arrows) and inverted (solid arrows) repeats were found that flanked a conserved −35 hexamer (bold). A putative −10 hexamer (bold) is also indicated
Sequence analyses of the inserts in pSK5 and pSK17 identified an ORF corresponding to a chitinase, designated chiA (Fig. 4A). Upstream of chiA, we identified two ORFs that were similar to an E. coli gene encoding a maltodextrin glucosidase, MalZ, (35% identity, 46% amino acid similarity) and a Staphylococcus aureus ABC transporter of the maltose system (38% identity, 60% similarity), and a third partial ORF for a putative aminoacylproline aminopeptidase that shared 42% identity and 53% similarity (out of 491 amino acids) with Streptomyces type II aminopeptidases. The downstream region included a putative DNA primase (46% identity, 58% similarity; based on comparisons to E. coli DNA primase) and a partial ORF for a putative aminoacyl-transferase (55% identity, 67% similarity; out of 289 amino acids, based on comparisons to a putative dihydrolipoamide acyltransferase component E2 of the pyruvate dehydrogenase complex in S. coelicolor).
Analysis of the nucleotide sequence of chiA revealed an ORF of 1,716 nucleotides that starts with an ATG and encodes a protein of 571 amino acids with a predicted molecular mass of 59.2 kDa and a theoretical pI of 8.3. The base composition of the ORF was 70 mol% G+C, which is slightly below the overall G+C content of C. uda DNA (76.3 mol%) (Stackebrandt and Kandler 1979). A putative cleavage site for a signal peptidase was identified between the two alanines at positions 53 and 54.
As shown in Fig. 4B, a potential ribosome-binding site, AGGA, was found upstream of the start codon, as well as two direct repeats flanked by two inverted repeats analogous to the repeats found in the promoter regions of many Streptomyces chitinases, which are thought to function as binding sites of regulatory proteins involved in the mechanisms of induction by chitin and catabolic repression by glucose (Saito et al. 1999, 2000). The similarity of this region with the promoter regions of Streptomyces chitinases was further supported by the identification of a −35 hexamer, TTGACC, located between the two direct repeats, which is conserved in the Streptomyces promoters (Saito et al. 2000).
Modular structure of ChiA
Computer analyses of the secondary structure of the protein coded by chiA identified a family 2 carbohydrate-binding module (CBM2), spanning amino acids 57–155, and a catalytic domain of family 18 of glycosyl hydrolases between amino acids 203 and 556 connected by a region rich in prolines and threonines (P-T domain) (Fig. 5). A typical family 18 active site (following a consensus sequence FDGIDVDWEY, in which the glutamic acid (E) presumably acts as the catalytic residue) was identified between amino acids 337 and 346 in the catalytic domain. The catalytic domain of the ChiA precursor showed greatest similarity to Chi63 of Cellulomonas sp. strain GM13 and Streptomyces chitinases (57–63% identity, 65–68% similarity, out of 349–354 amino acids) and thus seemed to be highly conserved among chitinases from actinobacteria. The CBM2 module was similar to other CBM2s present in chitinases from other high-G+C gram-positive bacteria, such as Chi63 of Cellulomonas sp. strain GM13 (81% identity, 89% similarity, out of 99 amino acids) and, to a lesser extent, Streptomyces chitinases (46–47% identity, 64–66% similarity, out of 98 amino acids), but it was also similar to CBM2s present in cellulases (e.g., CelE5 of Thermofibida fusca (44% identity, 55% similarity, out of 98 amino acids)), xylanases (e.g., Cellulomonas fimi xylanase (37% identity, 54% similarity, out of 91 amino acids)), and pectate lyases (e.g., PelA of Pseudomonas fluorescens (31% identity, 52% similarity, out of 96 amino acids)).
Modular structure of ChiA of C. uda and other bacterial chitinases. The modular organization of ChiA, including a signal peptide (white block), a carbohydrate-binding module (CBM2) (hatched block) and a family 18 catalytic domain (gray block), is similar to that of ChiC of S. lividans (Fujii and Miyashita 1993), except for the presence of a proline- and threonine-rich linker region (PT) in ChiA replacing the fibronectin type III domain (black arrow) of the Streptomyces chitinase. Family 18 chitinases, such as ChiA1 of Bacillus circulans (Watanabe et al. 1994) and ChiA and ChiB from Serratia marcescens (Brurger et al. 1996), all have a CBM5 that previously was identified as a chitin-binding domain (vertical lines). A polycystic kidney disease I (PKD) domain (white arrow) and unknown modules (black block) are also present in some chitinases. P-T linker modules also can be found in other glycosyl hydrolases such as in a chitinase from Aeromonas sp. strain 10S-24 (Ueda et al. 2003), where they link type-5 CBMs to each other and to a family 19 catalytic domain (gray dotted block)
The region between the CBM and the catalytic domain lacked type III repetitive sequences of fibronectin, such as those found as linker regions in many Streptomyces chitinases (Fig. 5). Instead, a low complexity P-T domain with a proline-rich extensine signature (constituted by four TNPTTGPT amino acid tandem repeats and a single SNPT sequence) was identified. P-T regions such as this, which are abundant in certain plant glycoproteins and are also found as linker regions in microbial modular enzymes (Lee et al. 2000), are thought to be highly glycosylated in the mature protein, thus contributing greatly to the molecular mass of the secreted protein.
Phylogenetic analysis of the CBM2 module of ChiA and homologous CBMs identified three distinct evolutionary groups (Fig. 6A). The substrate-binding domain of ChiA of C. uda was closely related to the CBMs of Chi63 of Cellulomonas sp. strain GM13 and of CelE5 of T. fusca, while the CBMs present in the Streptomyces chitinases formed an independent phylogenetic group. In contrast, the catalytic domain of the chitinases of the two cellulomonads branched in the same phylogenetic group as the catalytic domains of Streptomyces chitinases (Fig. 6B), suggesting that the catalytic domains of these actinobacteria derived from a sequence present in a common ancestor.
Phylogenetic relationships of the CBM2 module (a) and the catalytic domain (b) of ChiA and homologous protein modules from other hydrolases. Sequences were first aligned using the Clustal W program (Thompson et al. 1994). The phylogenetic tree was constructed and visualized using the Clustal W and the PHYLIP DRAWGRAM version 3.5c programs of the SDSC Biology Workbench. Phylogenetic groups are indicated by numbers at right. Abbreviations (protein designation and accession nos. are in parentheses): Cuda, C. uda (ChiA, AY008839); Csp., Cellulomonas sp. strain GM13 (Chi63, AF181718); Tfus, Thermofibida fusca (CelE5, L01577); Sliv, Streptomyces lividans (ChiC, D12647; CelB, U04629); Scoe, Streptomyces coelicolor (ChiC, AB017010; Chi, SC26G5; ChiD, AB017011); Spli, Streptomyces plicatus (Chi63, M82804); Speu, Streptomyces peucetius (ChiC, AF206633); Pflu, Pseudomonas fluorescens subsp. cellulosa (PelA, AF279264); Cfim, Cellulomonas fimi (CBP120, L38827; Xyl, L11080); Ccel, Clostridium cellulovorans (EngD, M37434); Mtub, Mycobacterium tuberculosis (RV1987, Z74025); Bcer, Bacillus cereus (ChiB, AB041932); Sther, Streptomyces thermoviolaceus (Chi40, D14536); Stmal, Stenotrophomonas maltophila (ChiA, AF014950); Dchi, Doohwaniella chitinasigens (Chi67, U81007); Jliv, Janthinobacterium lividum (Chi69, U07025); Asp., Arthrobacter sp. (ChiB, P250586); Bcir, Bacillus circulans WL-12 (ChiA1, A38368); Smar, Serratia marcescens (ChiA, AB015996)
Discussion
Although considerable information is available on the chitinase systems and chitinase genes from certain representative soil bacteria such as Streptomyces, little is known about the chitinolytic properties of other soil chitinolytic bacteria, which might contribute significantly to the diversity of chitinases present in agricultural soils (Metcalf et al. 2002). We recently found that the ability to degrade chitin might be widespread among species of Cellulomonas (Reguera and Leschine 2001), facultative aerobic soil actinobacteria traditionally characterized by their ability to degrade cellulose (Stackebrandt and Prauser 1992). Thus, cellulomonad chitinases are likely to play a pivotal role in the dynamics of chitin turnover in both oxic and anoxic terrestrial environments. In particular, the free-living bacterium C. uda efficiently degraded chitin, both aerobically and anaerobically, and utilized it as a source of carbon and nitrogen (Reguera and Leschine 2001). When grown in the presence of insoluble forms of chitin such as colloidal chitin, C. uda produced extracellular endo- and exochitinase activities. A major enzyme component, designated ChiA, was purified to homogeneity and confirmed to be an endochitinase that hydrolyzed insoluble substrates into long-chain chitooligosaccarides, while one (or more) as yet uncharacterized exochitinase(s) acted on the chitooligosaccharides to produce chitobiose. Presumably, C. uda cells, similar to other chitinolytic bacteria, have a transport mechanism to take up chitobiose and further metabolize it (Keyhani et al. 1996; Keyhani and Roseman 1997).
Some chitinolytic microorganisms, mainly gram-negative bacteria, have been found to possess chitinase genes organized in gene clusters (Shiro et al. 1996) while in others, mainly gram-positive bacteria such as Streptomyces, the chitinase genes, although co-regulated, appear to be scattered on the chromosome (Saito et al. 2000). The C. uda gene encoding ChiA was present in a chromosomal region flanked by a number of genes that are not directly involved in the degradation of chitin, a gene arrangement similar to that found in other gram-positive bacteria. The probe used to clone chiA failed to identify any additional chromosomal fragments that might contain sequences homologous to chiA, even under conditions of low stringency, suggesting that the exochitinase component(s) of the chitinase system of C. uda might be encoded by a gene (or genes) not homologous to chiA. One of the most striking features of the sequence of chiA was the presence of potential regulatory sequences upstream of the start ATG codon resembling the promoter regions of many Streptomyces chitinases (Saito et al. 2000). In S. coelicolor, five of its eight chitinase genes, which are found scattered on the chromosome, are induced in the presence of colloidal chitin and chitobiose and are subjected to catabolite repression by glucose. All five genes have conserved promoter regions containing conserved 12-bp direct repeats, TGGTC(C/T)(A/G)GACC(T/A), suggesting that these operator regions could be binding sites for the same regulatory proteins (Saito et al. 2000, 2001). However, the nucleotide sequence of the direct repeats identified upstream of chiA differed significantly from those in Streptomyces, suggesting possible differences in the regulation of chitinase gene expression in C. uda and Streptomyces species.
C. uda chiA codes for a modular enzyme consisting of a CBM2 and a family 18 catalytic domain linked by a P-T module, a modular organization also found in Chi63 of Cellulomonas sp. strain GM13 (GenBank accession no. AF181718). As illustrated in Fig. 5, the modular structure of ChiA differed from that of other actinobacterial chitinases, such as ChiC of S. lividans, in the lack of a fibronectin type III (Fn3) linker domain. Fn3 linker domains and structurally similar domains, such as the polycystic kidney disease I (PKD) domains, can also be found in chitinases from distantly related bacteria such as ChiA1 of Bacillus circulans and ChiA of Serratia marcescens, respectively (Fig. 5), and are thought to play a role in the optimal assembly of the substrate-binding and catalytic domains (Watanabe et al. 1994).
In ChiA, the linker region consisted of a P-T module similar to those found in other glycosyl hydrolases such as cellulase Cex of C. fimi (Gilkes et al. 1988) and a family-19 chitinase of Aeromonas sp. strain 10S-24 (Ueda et al. 2003). P-T modules are flexible protein regions thought to function as hinges to adapt the tertiary structure of chitinases to the substrate for a correct orientation of both the substrate-binding and catalytic modules (Cohen-Kupiec and Chet 1998). The P-T module found in ChiA was especially long (47 amino acids, 30 of which were prolines or threonines), which could translate into a more flexible protein, thus enabling it to recognize a broader range of substrates and, possibly, allowing for a more efficient binding and hydrolysis. In the mature protein, P-T modules also serve as glycosylation sites (Gilkes et al. 1988), which are thought to protect the enzymes from proteolytic attack and increase their affinity for the substrate (Ong et al. 1994). Glycosylation at P-T modules contributes to significant increases in mass (Lee et al. 2000), which might explain the discrepancy between the predicted molecular mass of the protein encoded by chiA (59.2 kDa, or 54 kDa upon processing of the putative signal peptide) and the molecular mass estimated for native ChiA (70 kDa).
In modular enzymes such as ChiA, each module or domain is characterized by a distinct three-dimensional protein structure or folding, suggesting that each may have evolved independently (Ponting et al. 1999). The catalytic domain of chiA was closely related to those found in chitinases of other high-G+C gram-positive bacteria such as chiC of S. lividans (Fujii and Miyashita 1993), chi63 of S. plicatus (Robbins et al. 1988) and chiC of S. coelicolor (Saito et al. 1999) (Fig. 6B). However, the CBM found in chiA of C. uda clustered in a phylogenetic group distinct from the CBMs of Streptomyces chitinases (Fig. 6A). The specific arrangement and nature of the modules identified in ChiA of C. uda and Chi63 of Cellulomonas sp. strain GM13 are likely to be a reflection of evolutionary adaptations to specific substrates and conditions present in the natural ecosystem inhabited by these actinobacteria.
Despite the critical role that chitinolytic bacteria play in carbon and nitrogen cycling due to chitin turnover in soils, little is known about the diversity of chitinases within terrestrial systems. Advancing our understanding of the biochemistry and diversity of chitinolysis in soil ecosystems relies, in part, on more studies of the chitinolytic systems from relevant soil bacteria such as the cellulomonads, whose role in chitin decomposition in terrestrial environments only recently has been recognized.
Abbreviations
- CBM :
-
Carbohydrate-binding module
- P-T :
-
Proline- and threonine-rich domain
- Fn3 :
-
Type III repetitive sequences of fibronectin domain
- PKD :
-
Polycystic kidney disease I domain
References
Brurger MB, Nes IF, Eijsink VGH (1996) Comparative studies of chitinases A and B from Serratia marcescens. Microbiology 142:1581–1589
Cavedon K, Leschine SB, Canale-Parola E (1990) Cellulase system of a free-living, mesophilic clostridium (strain C7). J Bacteriol 172:4222–4230
Cohen-Kupiec R, Chet I (1998) The molecular biology of chitin digestion. Curr Op Biotechnol 9:270–277
Edman P, Begg G (1967) A protein sequenator. Eur J Biochem 244:80–91
Felse PA, Panda T (1999) Regulation and cloning of microbial chitinase genes. Appl Microbiol Biotechnol 51:141–151
Fujii T, Miyashita K (1993) Multiple domain structure in a chitinase gene (chiC) of Streptomyces lividans. J Gen Microbiol 4:677–686
Gilkes NR, Warren RAJ, Miller RC, Kilburn DG (1988) Precise excision of the cellulose binding domains from two Cellulomonas fimi cellulases by a homologous protease and the effect on catalysis. J Biol Chem 263:10401–10407
Gomori G (1955) Preparation of buffers for use in enzyme studies. In: Colowick SP, Kaplan NO (eds) Methods in enzymology. Academic Press, New York, pp 138–146
Henrissat B (1999) Classification of chitinase modules. In: Jolles P, Muzzarelli RAA (eds) Chitin and chitinases. Birkhauser Verlag, Basel, Switzerland, pp 137–156
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293:781–788
Heukeshoven J, Dernick R (1985) Simplified method for silver staining of proteins in polyacrylamide gels and the mechanism of silver staining. Electrophoresis 6:103–112
Hsu SC, Lockwood JL (1974) Powdered chitin as a selective medium for enumeration of actinomycetes in water and soil. Appl Microbiol 29:422–426
Jeuniaux C (1966) Chitinases. In: Neufeld EF, Ginsburg V (eds) Methods in enzymology. Academic, New York, pp 644–650
Keyhani NO, Roseman S (1997) Wild-type Escherichia coli grows on the chitin disaccharide, N,N’-diacetylchitobiose, by expressing the cel operon. Proc Natl Acad Sci USA 94:14367–14371
Keyhani ND, Wang L, Lee YC, Roseman S (1996) The chitin catabolic cascade in the marine bacterium Vibrio furnissii. Characterization of an N, N’-diacetyl-chitobiose transport system. J Biol Chem 271:33409–33439
Koga D, Mitsutomi M, Kono M, Matsumiya M (1999) Biochemistry of chitinases. In: Jolles P, Muzzarelli RAA (eds) Chitin and chitinases. Birkhauser Verlag, Basel, Switzerland, pp 111–123
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature (London) 227:680–685
Lee HS, Han DS, Choi SJ, Choi SW, Kim DS, Bai DH, Yu JH (2000) Purification, characterization, and primary structure of a chitinase from Pseudomonas sp. YHS-A2. Appl Microbiol Biotechnol 54:397–405
Leschine S, Canale-Parola E (1983) Mesophilic cellulolytic clostridia from freshwater environments. Appl Environ Microbiol 46:728–737
Metcalf AC, Krsek M, Gooday GW, Prosser JI, Wellington EMH (2002) Molecular analysis of a bacterial chitinolytic community in an upland pasture. Appl Environ Microbiol 68:5042–5050
Muzzarelli RAA (1977) Chitin. Pergamon Press, Oxford
O’Brien M, Colwell RR (1987) A rapid test for chitinase activity that uses 4-methylumbelliferyl-N-Acetyl-b-D-Glucosaminide. Appl Environ Microbiol 53:1718–1720
Ohno T, Armand S, Hata T, Nikaidou N, Henrissat B, Mitsutomi M, Watanabe T (1996) A modular family 19 chitinase found in the prokaryotic organism Streptomyces griseus HUT 6037. J Bacteriol 178:5065–5070
Ong E, Kilburn DG, Miller Jr. RC, Warren AJ (1994) Streptomyces lividans glycosylates the linker region of a beta-1,4-glycanase from Cellulomonas fimi. J Bacteriol 176:999–1008
Ponting CP, Aravind L, Schultz JB, P., Koonin EV (1999) Eukaryotic signalling domain homologues in archaea and bacteria. Ancient ancestry and horizontal gene transfer. J Mol Biol 289:729–745
Reguera G, Leschine SB (2001) Chitin degradation by cellulolytic anaerobes and facultative aerobes from soils and sediments. FEMS Microbiol Lett 204:367–374
Robbins PW, Albright C, Benfield B (1988) Cloning and expression of a Streptomyces plicatus chitinase (chitinase-63) in Escherichia coli. J Biol Chem 263:443–447
Saito A, Fujii T, Yoneyama T, Redenbach M, Ohno T, Watanabe T, Miyashita K (1999) High-multiplicity of chitinase genes in Streptomyces coelicolor A3(2). Biosci Biotechnol Biochem 63:710–718
Saito A, Ishizaka M, Francisco Jr. PB, Fujii T, Miyashita K (2000) Transcriptional co-regulation of five chitinase genes scattered on the Streptomyces coelicolor A3(2) chromosome. Microbiology 146:2937–2946
Saito A, Miyashita K, Biukovic G, Schrepf H (2001) Characteristics of Streptomyces coelicolor A3(2) extracellular protein targeting chitin and chitosan. Appl Environ Microbiol 67:1268–1273
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Shiro M, Ueda M, Kawaguchi T, Arai M (1996) Cloning of a cluster of chitinase genes from Aeromonas sp. No. 10S-24. Biochim Biophys Acta 1305:44–48
Smith PK, Krohn RI, Hermanson GT, Mallia AK, Gartner FH, Provenzano MD, Fujimoto EK, Goeke NM, Olson BJ, Klenk DC (1985) Measurement of protein using bicinchoninic acid. Anal Biochem 150:76–85
Stackebrandt E, Kandler O (1979) Taxonomy of the genus Cellulomonas, based on phenotypic characters and deoxyribonucleic acid- deoxyribonucleic acid homology, and proposal of seven neotype strains. Int J Syst Bacteriol 29:273–282
Stackebrandt E, Prauser H (1992) The family Cellulomonadaceae. In: Balows A, Truper HG, Dworkin M, Harder W, Schleifer KH (eds) The prokaryotes, 2nd edn. Springer, Berlin Heidelberg New York
Svitil AL, Ni Chadhain SM, Moore JA, Kirchman DL (1997) Chitin degradation proteins produced by the marine bacterium Vibrio harveyi growing on different forms of chitin. Appl Environ Microbiol 63:408–413
Thompson JD, Higgins DG, Gilson TJ (1994) Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalies and weight matrix. Nucleic Acids Res 22:4674–4680
Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci USA 76:4350–4354
Trudel J, Asselin A (1989) Detection of chitinase activity after polyacrylamide gel electrophoresis. Anal Biochem 178:362–366
Ueda M, Kojima M, Yoshikawa T, Mitsuda N, Araki K, Kawaguchi T, Miyatake K, Arai M, Fukamizo T (2003) A novel type of family 19 chitinase from Aeromonas sp. No. 10S-24. Eur J Biochem 270:2513–2520
Watanabe T, Ito Y, Hashimoto M, Yamada T, Alam MM, Tanaka H (1994) The role of the C-terminal domain and type III domains of chitinase A1 from Bacillus circulans WL-12 in chitin degradation. J Bacteriol 176:4465–4472
Williams J, McGrath WJ, Mangel WF (2000) Sensitive method to identify and characterize proteinases in situ after SDS-PAGE. Biotechniques 29:1108–1113
Acknowledgements
We would like to express our appreciation to Dr. Elizabeth Stuart and Dr. Thomas Lessie for technical advice and helpful suggestions during different phases of this work. This work was supported by U.S. Department of Energy grants DE-FG02–88ER13898 and DE-FG02–02ER15330.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Reguera, G., Leschine, S.B. Biochemical and genetic characterization of ChiA, the major enzyme component for the solubilization of chitin by Cellulomonas uda . Arch Microbiol 180, 434–443 (2003). https://doi.org/10.1007/s00203-003-0611-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-003-0611-y