Abstract
Osteoporosis is a common skeletal disease characterized by low bone mineral density (BMD), deterioration of bone microarchitecture and increased fracture risk. It is a complex disease that has high social and economic costs. Osteoporosis and its associated phenotypes are under the strong genetic control. Identification and characterization of specific loci or genes involved in determining osteoporosis and its associated phenotypes will contribute to a greater understanding of the pathogenesis of osteoporosis, and ultimately might lead to the development of better diagnosis, prevention and treatment strategies. Efforts to identify osteoporosis genes have focused on three approaches: animal models, candidate gene approach, and genome-wide scans. In this article, we review the current status for mapping and identification of genes for osteoporosis, with a focus on some promising regions and future prospects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Osteoporosis is a systemic skeletal disease characterized by low bone mineral density (BMD) and microarchitectural deterioration of bone tissue, with a consequent increase in bone fragility and susceptibility to fracture [1]. The World Health Organization (WHO) defines osteoporosis in post-menopausal Caucasian women as a value for bone mineral density (BMD) or bone mineral content that is more than 2.5 SD below the young gender and ethnicity-matched adult mean value [2]. According to the definition, osteoporosis affects 30% of postmenopausal white women in the USA, and the proportion rises to 70% in women over the age of 80 years [3]. The most common clinical outcomes of osteoporosis are fracture of the spine, hip and wrist. Of these, hip fractures are the most severe, leading to a 12–20% reduction of expected survival [4]. The direct cost for hip fractures was around $13.8 billion in the US in 1995 [5], and £942 million in the UK in 1998 [6]. With rapid economic development and aging of the population, the worldwide health and economic burden of osteoporosis will rise further in the future.
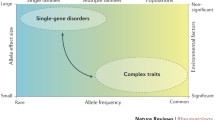
It is well established that BMD and other determinants of osteoporotic fracture are under strong genetic control. Identification and characterization of specific loci or genes involved in determining osteoporosis and associated phenotypes not only contribute to a greater understanding of the pathogenesis of osteoporosis, but also lead to the development of better diagnosis, prevention and treatment strategies of the disease. Genetic determination of osteoporosis may be monogenic or polygenic. In this review, we are mainly concerned with the polygenic form, although a limited few monogenic forms will also be briefly mentioned. Genetics research of osteoporosis represents one of the most active areas for bone biology research. Several reviews have nicely summarized the results of the candidate genes research [7,8,9,10]. As a complement to these, in this review, we first give an overview of the evidence that osteoporosis and associated traits have a genetic basis, then briefly summarize the main findings that come from linkage and association studies, with a focus on some promising chromosomal regions, and finally discuss the future challenges and directions.
Evidence for genetic determinants of osteoporosis and associated traits
Fracture is the ultimate consequence of osteoporosis. Ideally, scientists would perform genetic studies with fracture as an endpoint, and search for genes underlying the differential susceptibility to fracture. Genetic epidemiological studies have shown that a family history of fracture is a significant risk factor for fracture [11,12]. However, prospective 25-year follow-up of a nationwide cohort of elderly Finnish twins showed that susceptibility to osteoporotic fracture is not strongly influenced by genetic factors. In women, the pairwise concordance rate for fracture was 9.5% in monozygotic pairs and 7.9% in dizygotic pairs. In men, the figures were 9.9% and 2.3%, respectively [13]. Deng and colleagues [14] estimated that the narrow-sense heritability of Colles' fracture was approximately 0.25 in a cohort of white American women, thus accounting for approximately one-quarter of the variation in total Colles' fracture risk observed. Genetic factors, at most, account for about one third of the variance in the liability to fracture [15]. Fractures are relatively rare and tend to occur late in life. Although vertebral fractures are relatively common compared with hip and wrist fractures, they do not always come to clinical attention and their diagnosis may prove uncertain [16]. The relatively low heritability of osteoporotic fracture and difficulty in recruitment of an adequate sample in which to perform mapping studies of fracture in humans lead investigators to adopt alternative strategies using surrogate traits.
Bone strength is an ultimate measurement of resistance to fracture. It is mainly determined by BMD, bone size, and bone quality [7,17]. Bone strength cannot be directly measured in vivo in human. As BMD contributes substantially to bone strength and can be practically measured with marked sensitivity and precision, the evaluation of BMD is the most commonly used method for predicting fracture risk in humans. Consequently, the vast majority of genetic studies of osteoporosis to date have used BMD as a surrogate phenotype. BMD is a complex phenotype because it results from remodeling processes affecting bone compartments (endosteal and periosteal), and therefore bone size, and the process differ according to age and gender [18]. BMD in both sons and daughters correlates most closely with the average parental BMD [19]. Twin studies have shown that the heritability estimates of BMD ranged from 0.5–0.9 [20,21,22,23,24,25,26]. Since environmental influences can differ between generations considerably, heritability estimates of BMD in inter-generational studies have generally been lower than those reported in twin studies, ranging from 0.46 to 0.67 [19,27,28,29,30]. Most of the segregation analyses [28,31,32,33] suggest that there exits at least one major gene for population BMD variation. The genetic correlation between lumbar spine and femoral neck BMD is 0.64, and approximately one-third of the genetic influence on variance of femoral neck BMD is mediated through the same gene or genes that influence the lumbar spine [34]. Therefore, there are some common and specific genetic factors underlying the determination of BMD in various skeletal sites. However, a recent study indicated that genetic correlation between fracture and BMD is not significant despite a high phenotypic correlation between fracture and BMD [35]. Thus, all important risk factors for fracture need to be studied in order to find genes for osteoporotic fractures. In addition, there are obvious gender differences in the genetic components of BMD in mice [36]. Men generally have larger bone size and greater cortical mass than women [37], which is associated with considerable fracture risk reduction. Whether the genetic determinants of BMD in humans also would be influenced by gender remains to be elucidated.
Bone size is also an important determinant of osteoporotic fractures. A longer hip axis length is associated with increased hip fracture risk independent of BMD [38]. However, there have been few reports on the estimation of heritability of variation in bone size [39,40,41]. In 49 pedigrees with 703 subjects, after adjusting for sex, age, weight, height, lifestyle factors, and the significant interactions among these factors, heritability estimates were, respectively, 0.48, 0.64, and 0.6 for bone size at the hip, spine, and wrist [41]. In addition, forearm width and hip axis length are also highly heritable with heritability estimates of greater than 0.5 [20,25].
Quantitative ultrasound (QUS) measurements, including broadband ultrasound attenuation (BUA) and speed of sound (SOS), are measurements that reflect the quality aspects of bone. Subjects with lower BUA at baseline have a higher risk of hip and vertebral fractures, possibly independent of BMD [42]. Estimates of heritability based on twin studies for age- and weight-adjusted BUA and SOS are 0.74 and 0.82, respectively [25,26]. Bivariate genetic analysis indicated that the genetic correlation between BUA and BMD ranged between 0.43 and 0.51, whereas the environmental correlation ranged between 0.2 and 0.28 [26].
Bone formation and resorption markers may predict hip fracture in elderly white women [43]. Each standard deviation higher in bone specific alkaline phosphatase (BSAP) values was associated with a 4% lower level of BMD in both lumbar spine and femoral neck [44]. The genetic contribution to variation in bone turnover after attainment of peak bone mass is established [45,46,47], although the genetic effects on bone turnover are smaller than those on BMD and bone size. However, the genetic contribution to variation in rate of bone loss has not been shown consistently.
In order to understand the genetic basis for decreased bone strength, and ultimately osteoporotic fractures, one needs to assess the inheritance of, and identify the specific genes associated with, a multitude of skeletal traits, such as BMD, bone size, QUS, and bone turnover. If osteoporotic fractures are not studied as the phenotype, the genes identified need to be tested for relevance to osteoporotic fracture. The above results consistently support the hypothesis that genetic factors are a major determinant of BMD and, possibly, variance in bone size, bone turnover, and QUS measurements. This hypothesis has fueled most osteoporosis genetics investigations over the last decade.
The search for osteoporosis genes
Three major approaches to identifying genes for osteoporosis have been pursued: animal models, the candidate gene approach (association studies), and genome-wide scans (linkage studies) [42]. One major advantage of using an animal model is that it is possible to control for the heterogeneity of environmental factors in animals, which is otherwise impossible in human studies. The candidate gene approach tests for the association between a particular gene variant and osteoporosis (BMD variation), and depends on linkage disequilibrium of markers with functional mutations. It is generally prone to population admixture/stratification in yielding false positive or false negative results [48]. To overcome this problem, the transmission disequilibrium test (TDT) is employed to test specific candidate genes for both association and linkage [49]. Genome-wide scans test only linkage and are robust to population admixture/stratification. A disadvantage is that they have relatively low statistical power to detect genes with modest effects unless the sample size as reflected by the informative relative pairs is large. Genome scan not only guides candidate gene research by according greater priority to candidates that are located within regions of linkage, but also identifies novel chromosomal regions within which no known candidates have been recognized.
Animal models
Animal models, which offer controlled exposure, limited and consistent genetic variation, and unlimited size of sib-ships, hold considerable potential for understanding the genetics of osteoporosis and associated traits. A promising approach is to map quantitative traits in experimental animal models and then search syntenic regions of the human genome for genes defining these traits in humans. The genetics of osteoporosis and associated traits have been studied extensively in inbred strains of experimental animals [17,50,51,52,53,54,55,56,57,58]. Li et al. [17] identified six significant QTLs affecting bone breaking strength, of which three influence BMD, two influence bone quality, and one influences bone size. The QTL mapping results for BMD in experimental animals and the associated human homologous regions are summarized in Table 1. Notably, in several cases the same QTLs have been mapped in different crosses, using different but related phenotypes. Examples include cfh-Mit15 [17,50,54,55,58] and Mit291-Mit362 [17,51,58] on chromosome 1, Mit296-Il2ra [50,57] and Mit413-Ncvs42 [50,55,56] on chromosome 2, Mit124-Mit204 on chromosome 4 [17,55], Mit210-Mit80 on chromosome 7 [56,57], Mit242-Mit349 [51,55] and Mit36-Mit160 [50,52,57,58] on chromosome 11, Mit135-Mit16 on chromosome 13 [52,53,54], Mit132-Ptprg [17,50] and Mit160-Mit194 [55,58] on chromosome 14; Mit29-Atf4 on chromosome 15 [50,54], Rik29-Mit39 on chromosome 16 [50,55,57], and Mit185-Ncvs23 [17,50] on chromosome 18. The future challenge is to identify genes responsible for these effects and to determine the relevance of these regions to human osteoporosis and associated traits.
The candidate gene approach
There are three types of candidate genes: functional candidate genes, positional candidate genes, and expressional candidate genes. Functional candidate genes are based upon a priori knowledge of the phenotype and the potential function of the gene involved. Such knowledge may come from clinical observation or physiological studies of affected individuals, from studies of known disease-related process, from animal models of disease, and from pharmacogenetic studies. Positional candidate genes are genes targeted because of their location within regions identified through genetic linkage analyses. Expressional candidate genes are identified through differences in gene expression using genomic arrays. Candidate genes are commonly examined by association studies, using a case-control design. Since the first report of an association study between the α2-HS-glycoprotein (ASHG) gene and bone mass [59], there has been an extensive and growing list of candidate genes investigated for linkage and association with osteoporosis and associated traits. There are currently more than 200 genes that have been proposed as potential candidates for osteoporosis and associated traits [60,61].
Among the multiple candidate genes harboring polymorphic loci so far investigated in relation to BMD and fracture, the vitamin D receptor (VDR) gene has received the greatest attention. The relationship between the VDR genotype and BMD has been studied in Caucasians, East Asian, and Africans. A meta-analysis combining the results of 75 articles and abstracts published between 1994 and 1998 which examined the relationship between the VDR polymorphisms (BsmI, ApaI, TaqI, EcoRV, and FokI) and osteoporosis-related phenotypes (BMD, fracture, and QSU) have shown a highly significant association between VDR polymorphisms and BMD. Positive results were significantly more common in studies that included premenopausal rather than postmenopausal women, and the association may have been missed in some studies because of small sample size and other confounding factors [62].
Collagen type I (COL1A1) is the most abundant protein in bone, and mutations in the genes encoding collagen type Iα1 and collagen type Iα2 are estimated to be responsible for up to 90% of cases of the Mendelian disease osteogenesis imperfecta. Polymorphisms affecting the coding regions of the collagen type I genes are rare and do not appear to be associated with osteoporosis [63]. Grant et al. [64] described a G→T polymorphism in intron 1 of the COLIA1 gene at a binding site for the Sp1 transcription factor, and reported decreased BMD and increased fracture risk for carriers of the s allele in an analysis of 205 predominant postmenopausal British women. Since that time, numerous studies have been performed in both Caucasians and Asians. The unfavorable effect of the s allele has not been seen consistently across different studies, and a G→T polymorphism in intron 1 of the COLIA1 gene at a binding site for the Sp1 transcription factor does not appear to exist in Asians [65]. Recently, two meta-analyses about association of COL1A1 Sp1 polymorphism with BMD and/or the risk of prevalent fractures have been performed [66,67]. The main conclusions to emerge from these meta-analyses were that the COLIA1 Sp1 polymorphism showed a dose-response relationship to prevalence of fractures (increases stepwise from SS homozygotes to Ss heterozygotes and from Ss heterozygotes to ss homozygotes). Further, the association with fracture was stronger than expected on the basis of the observed differences in BMD. Because a large part of the inherited predisposition to fracture is due to inherited factors in bone density, and/or material quality of bone, authors concluded that the Sp1 effect on fractures may be mediated in part by its influence on bone quality other than BMD.
The relationship between the estrogen receptor α (ER-α) gene polymorphisms (TA, CA, PvuII, and XbaI) and BMD/fracture has been investigated extensively, and contrasting results were reported. A recent meta-analysis indicated that XX homozygotes (women carrying two copies of the gene variant without an XbaI restriction site) have higher BMD and also a decreased risk of fractures when compared with carriers of the χ allele, whereas the PvuII polymorphism is not associated with either BMD or fracture risk [68]. Notably, a significant gene-gene interaction between VDR and ER-α gene polymorphisms has been suggested by several authors [69,70,71]. In addition, several studies assessed whether genotypes of ER-α are associated with bone changes in women with and without hormone replacement therapy [72,73,74]. Although results are inconsistent, the information obtained should turn out to be helpful in choosing optimum therapy for osteoporosis for these different genotypes (genotype-specific treatment).
Other candidate genes investigated include, but are not limited to, the transformation growth factor β1 (TGFβ1) gene [75,76,77,78,79,80], the parathyroid hormone (PTH) receptor [81,82], the calcitonin receptor [83,84,85,86], the calcium-sensing receptor [87,88], the osteocalcin gene [82,89,90,91,92,93], the interleukin-6 (IL-6) genes [94,95,96,97,98,99], the insulin-like growth factor-I (IGF-I) genes [100,101,102,103,104], the apolipoprotein E gene [105,106,107], alpha 2 HS-glycoprotein (AHSG) gene [108], the interleukin-1 receptor antagonist gene [109], the androgen receptor (ADR) gene [90], the peroxisome proliferator-activated receptor gamma gene [110], tumor necrosis factor receptor 2 gene (TNFR2) [111], calcitonin genes [112], the P57 [113], methylenetetrahydrofolate reductase gene (MTHFR) [114], the aromatase (CYP19) gene [115], the Werner helicase (WRN) gene [116], the CC chemokine receptor-2 (CCR2) gene [117], the Klotho gene [118] and the runt-related gene 2 (RUNX2)/core binding factor A1 (CBFA1) gene [119].
Despite considerable efforts, it is still premature to draw conclusions about the potential influence of these genes on osteoporosis (BMD) and fracture. Results from association studies of candidate genes are often inconsistent. Reasons for this include false positive or negative results [48], small sample sizes and low statistical power, different sets of genes operating in different populations, variable linkage disequilibrium among populations [120], or low prior probability of the involvement of the gene in question in the overall risk of the disease [121]. To deliver robust results, some guidelines have been suggested. These include (1) significantly increased sample sizes; (2) incorporation of diverse study designs including case-control, family-based association studies and intermediate phenotype data sets [122]; (3) replication of findings in additional study groups of similar ethnic origin [123].
Whole-genome scans
Several whole genome-wide linkage studies have been conducted [124,125,126,127,128,129,130,131,132,133,134,135,136]. These results are summarized in Table 2. The following discussion will focus on some promising regions.
Chromosome 11q12–13
Significant linkage to chromosome 11q12–13 has been reported for three monogenic bone diseases. The first is osteoporosis-pseudoglioma syndrome (OPS), which is characterized by low bone mass, with childhood fractures and abnormal eye development. It was linked to chromosome 11q12–13 with a maximum LOD score of 5.99 achieved at marker D11S987 [137]. OPS has been shown to be caused by several different mutations in the gene for low-density lipoprotein receptor-related protein 5 (LRP5) [138]. The second monogenic bone disease is autosomal recessive osteopetrosis (arOP) that is characterized by osteosclerosis, deafness, blindness, and severe anaemia that are due to failure of osteoclast-mediated bone resorption. It is linked to chromosome 11q12–13 with a maximum LOD score of 5.9 achieved in two Bedouin pedigrees [139]. The T-cell immune regulator 1 (TCIRG1) gene was identified as one of the genes responsible for arOP [140,141]. Mutations in this gene may account for as many as 50% of the cases of recessive osteopetrosis [142]. Inactivation of TCIRG1 causes osteoclast-rich osteopetrosis in mice [143]. The third monogenic bone trait is an autosomal dominant trait characterized by high bone mass (HBM). Johnson et al. [144] reported it to be linked to a 30-cm region of 11q12–13 with a LOD score of 5.74 achieved at the marker D11S987. A G to T transversion in exon 3 of LRP5 results in the autosomal dominant high-bone-mass trait [145]. Remarkably, this mutation also causes an autosomal dominant syndrome characterized by high bone density, torus palatinus, and a wide, deep mandible [146]. Mice deficient in LRP5 have been reported to have low bone mass, low body weight, and abnormal eye vascularization [147]. This finding supports the critical role of this gene in skeletal integrity. It is of particular interest that variation in bone density in the general population was also linked to the chromosome region containing LRP5 [127,148]. However, Deng et al. [149] genotyped five markers in a genomic region of ~27 cM centering on D11S987 for 630 individuals from 53 human pedigrees, and did not find evidence of linkage of these five markers to BMD at the spine, hip and wrist and total body BMC. The maximum LOD score at these five markers was 0.25 and the maximum LOD score at D11S987 was 0.15. Karasik et al. [129] and others did not report linkage findings on chromosome 11q12–13 in the general population either. Whether common variants that alter the expression or function of LRP5 have a role in the risk of osteoporosis in the general population merits further studies.
Chromosome 1p36
Devoto et al. [124] reported a genome-wide scan in 149 members of seven large pedigrees. The strongest evidence of linkage was on chromosome 1p36, which was identified with two marker loci separated by 13.9 cM (D1S450 and D1S214) that gave LOD scores in the single-point non-parametric analysis of +3.51 and +2.62, respectively, for hip BMD. This finding was confirmed and extended in an expanded sample of 42 families by analyzing nine microsatellite markers spanning a 40 cM interval across the candidate region [150]. In addition, Albagha et al. [151] analyzed allele distribution of microsatellites in 54 women with high BMD and 54 women with low BMD, and found that markers on 1p36 were associated with differences in BMD. Recently, a genome-wide screen of 1097 unselected female UK twin pairs confirms the presence of QTLs for BMD at 1p36 [132]. It is also of interest to note that 1p36 showed suggestive evidence for linkage to BUA [133]. Plausible candidate genes include TNFR2, lysyl hydroxylase (PLOD), and MTHFR. Previous studies have indicated significant association of the polymorphism of the TNFR2 gene, and MTHFR gene with BMD [111,114,152].
Chromosome 1q21–32
An autosomal genome screen in 429 pre-menopausal Caucasian sister pairs found significant evidence of linkage to markers on chromosome 1q. The maximum LOD score (3.11) was attained at the 170 cM position of the Marshfield chromosome 1 map [127]. A genome wide screen of an additional 289 premenopausal Caucasian sister pairs yielded a LOD score of 3.2 at position 188 cM. Linkage analysis in an expanded sample of 570 white, which has partial overlap (281 white pairs) with the previously reported genome screen sample, yielded a maximum LOD score of 6.3 at position 188 cM [131]. In addition, 1q22 showed significant evidence of linkage with one-third distal area with a multi-point LOD score of 4.78 [135].
Chromosome 2p23-p24
Devoto et al. [124] reported a multi-point LOD score of 2.25 on 2p23-p24 for spinal BMD. Niu and colleagues [126] found linkage evidence of 2p23–24 with forearm BMD with LOD scores of 2.15 in a Chinese population. In addition, 2p25 also showed evidence of linkage with L1 area with a multi-point LOD score of 2.98 [135]. This region contains two genes of potential interest, pro-opiomelanocortin (POMC) and serine threonine kinase (STK).
Chromosome 4q25-q32
Devoto et al. [124] reported a LOD score of 2.28 near D4S1535 for hip BMD and a LOD score of 2.95 near D4S1539 for spine BMD. Deng et al. [130] reported a genomewide scan in 53 Caucasian pedigrees. They observed significant evidence of linkage to 4q25–32 for spine BMD with multi-point LOD scores of 3.08, and suggestive evidence of linkage for ultra-distal forearm BMD. Some additional support came from Duncan et al. [153], who found some evidence of linkage to 4q26 for femoral neck BMD.
Chromosome 6p11–21
Koller et al. [127] reported a LOD score of 1.94 near D6S462 for spine BMD. After a genome scan of 330 Caucasian families, Karasik et al. [129] have reported evidence of linkage to 6pter for both femoral neck and spine BMD at D6S2427.
Chromosome 12q12–24
Preliminary results of a genome scan in 286 members of ten large Mexican-American families identified low BMD loci on 12q24 [125]. Again in the Framingham osteoporosis study, Karasik et al. [129] have reported a LOD score of 2.08 (137 cM) for lumbar spine BMD at 12q23 in humans. Further support for linkage to 12q24 was reported by Deng et al. [130], with a multi-point LOD score of 2.96 for spine BMD. Drake et al. [56] detected a QTL for femur BMD with a LOD score greater than 2.3 in mice in regions homologous to human chromosome12q24. Plausible candidate genes include IGF1, T-box 3 (TBX3), TBX5 and nuclear transcription factor Y β (NFYB). IGF1 has previously been associated with BMD or osteoporosis in human populations [100,101]. Duncan et al. [153] reported a two-point LOD score of 1.7 at D12S83 for lumbar spine BMD. 12q13 achieved a LOD of 1.69 at D12S368 for hip BMD in two-point analysis [130]. This region contains a number of candidate genes, including VDR, integrin α7 (ITGA7), collagen type II α1 (COL2A1), and a cluster of homeobox (HOX). Interestingly, a QTL affecting osteochondrosis was located to a position between the interferon-γ (IFNG) and IGF-1 genes at pig chromosome 5 [154]. This region is homologous to human chromosome 12q14–24.
Chromosome 13q31–34
The genomic region 13q33–34 has previously been linked to forearm BMD with a LOD score of 1.67 [126]. 13q33 showed suggestive evidence of linkage with spine BMD [130]. Interestingly, in men, but not women, 13q34 (near marker D13S800) showed linkage with intertrochanter BMD with a LOD score of 3.1 [155]. Potential candidate genes in this region include collagen type IV α1 (COL4A1) and COL4A2.
Chromosome 17p12–13
Devoto et al. [124] reported a LOD score of 2.34 near D17S261 for hip BMD in humans. 17p12–13 showed suggestive evidence of linkage with ultra-distal forearm BMD [130]. In the region homologous to human chromosome 17p11–12 in mice, Beamer et al. [55] identified a QTL for femoral and vertebral BMD variation with LOD scores of 6.76 and 2.98, respectively, and Shimizu et al. [52] identified a QTL for femur peak bone mass with a LOD score of 10.8. Also, Benes et al. [57] reported a QTL for spine BMD (P=0.0001). A candidate gene, GLI3, was located in this region.
Chromosome 18q21–23
Devoto et al. [124] found linkage of hip BMD to D18S70 and D18S42 with a LOD score of 2.14 and 2.58, respectively. The gene responsible for familial expansile osteolysis (FEO), a rare, autosomal dominant bone disorder, has been linked to a region of chromosome 18q21.2–18q21.3 [156]. An 18-bp insertion in exon 1 of TNFRSF11A segregates with patients with FEO [157]. Cody et al. [158] documented a maximum two-point LOD score of 3.4, at marker D18S42, in a large pedigree with Paget disease of bone (PDB). Positive linkage of PDB to this region has also reported by Haslam et al. [159]. Recently, Good et al. [160] has identified a novel susceptibility locus of PDB at 18q23 (multipoint LOD score of 4.71 at marker D18S70) in a large subpedigree.
Prospects for gene discovery in osteoporosis
What next? The construction of a dense single nucleotide polymorphisms (SNPs) linkage map [161] and development of new technologies such as microarrays greatly facilitate identification of osteoporosis genes. The following aspects could be advanced.
Fine mapping
The most crucial future aim is positional cloning of causal genes and identification of sequence variants within the coding or controlling regions of such genes. To achieve this, it will be essential to refine and to narrow the existing QTL to ~1 cM, a requisite size at which positional cloning becomes feasible. The chromosomal regions described so far are quite broad. It is recognized that the saturation of a candidate interval with ever more markers contribute very little to its narrowing by linkage [162,163]. Now, increased attention is turning to techniques of linkage disequilibrium (LD)-based association mapping with SNPs. SNPs allow the unification of the candidate gene approach and association-based fine mapping to identify gene(s) of interest. However, it is important to keep in mind that, even in the region narrowed, there is still the challenge of identifying the actual gene involved. There may be lack of LD even between polymorphic loci that mapped to the same gene [164]. On the contrary, even where association is demonstrated it might not indicate a contribution of that gene, but might rather reflect LD with polymorphisms in a neighboring gene [165]. Recently, haplotype blocks were found in the human genome [166,167,168]. The existence of haplotype blocks raises hope that whole genome association studies can be carried out with reasonable cost by genotyping only a small fraction of SNPs that represent most haplotype diversity within the blocks. However, haplotype blocks make identification of a true causal variant more difficult due to the underlying haplotype effect.
DNA microarray analysis and proteomics
Oligonucleotide and cDNA microarrays have revolutionized the study of differential gene expression in cells and tissues, enabling genome-wide screening of gene transcript variations. Failure to find mutations in candidate gene coding regions does not rule out a possible contribution of altered gene expression contributing to osteoporosis. The identification of differentially expressed transcripts in normal versus affected tissues may add to the process of gene discovery in osteoporosis. In the simplest case, the target gene of interest might be identified directly by characteristic changes in expression levels across a series of samples. Alternatively, statistical analysis of microarray data might aid gene discovery by detecting new metabolic disease pathways related to the target gene and facilitating identification of candidate genes [169]. Of course, some of the gene expression changes identified in this way may be a result of environmental factors, chance, or other confounding variables (false positive). Nevertheless, combining positional information and expression information will simplify the process of moving from putative linkage to gene identification. For example, microarray analysis led to the generation of a list of 175 cDNAs underexpressed by 2.5-fold or more in the fibroblasts of an affected individual (the Tangier disease, TD). By combining these data with linkage information that localized the disease gene to chromosome 9q between the markers WI-14706 and WI-4062, the candidate list was narrowed sufficiently to identify the gene ABC1, which did carry mutations [170]. DNA microarray technology has begun to be utilized to identify differentially expressed genes associated with osteoporosis [171]. It can be anticipated that more of these data will emerge in the near future.
Likewise, comparison of protein expression between normal and disease states would identify proteins relevant to the disease process, and provides obvious candidate genes as the source of inherited variation in susceptibility. DNA microarrays have limited utility for the analysis of biological fluids and for uncovering assayable biomarkers directly in the fluids. Numerous alterations may occur in proteins that are not reflected in changes at the RNA level. Since genes ultimately influence disease states through the protein products they encode, the field of proteomics could be a powerful means to help identify candidate genes that underlie genetic variation. The correlation among DNA sequence, mRNA and protein is low due to transcriptional control, translational control and post-translational modification. Strategies to incorporate DNA microarray and proteomics data into traditional linkage or candidate gene studies would improve the efficiency and capability of gene discovery in osteoporosis and in illuminating the functions of the genes and the pathways the genes and/or their products involved [172,173].
Investigation of gene-gene and gene-environment interactions
For complex human diseases such as osteoporosis, which are determined by the joint action of multiple genes and environmental factors, most current models treat separate disease loci as if they were independent of each other. Even though the individual effect of a gene may appear to be small, interactions with other genes and/or environments could make a substantial contribution to the final manifestation of the disease. Failure to recognize and accommodate such interactions may often mask the effects of individual gene. For example, Cox et al. [174] described an approach to assessing statistical interactions between different chromosomal regions where evidence for linkage at one region is taken into account in assessing the evidence for linkage elsewhere in the genome. Using this approach, they showed an interaction between loci on chromosomes 2 and 15 that increases susceptibility to non-insulin-dependent diabetes (NIDD1). Interestingly, conventional linkage analysis failed to detect linkage to chromosome 15 in the initial genome scan. In addition, Cordell et al. [175] described a multi-locus linkage method. They showed that multi-locus analysis not only increased power to detect linkage, but also assisted in determining the nature of the relation between disease loci (i.e. genetic heterogeneity versus epistasis). One of the most important goals of the next generation of genetic studies of osteoporosis is to determine which multi-locus genotypes create the highest risk for development of osteoporosis.
The examples of gene-gene and gene-environment interaction for the VDR gene have been described by others [69,70,71] and us [10]. Bone density at any age is the end result of peak bone density and subsequent loss, and thus reflects the sum of responses to various environmental exposures. If genetic factors modulate those responses to environments, these gene-enviroment interactions presumably accumulate over time with aging [7]. Strength and direction of the VDR allelic effects may relate to the genetic backgrounds in different studies and environmental factors such as calcium and vitamin D intakes. This at least partially explains inconsistent results of the relationship between VDR polymorphism and BMD across multiple studies.
International collaborations
Several whole genome-wide linkage studies have been conducted [124,125,126,127,128,129,130,131,132,133,134,135,136]. Sample sizes have generally been modest. The relatively small numbers studied would have tended both to limit the power of these genome screens to detect linkage and to increase the possibility of false positive errors. A common approach to enhance the power of any study is to utilize a larger sample size. Large samples may augment weak linkage signals found in small data sets and are less susceptible to random statistical fluctuations that may lead to false positive results in smaller samples. The most expedient approach to further progress in the identification of genes for osteoporosis and associated traits would be through the efforts of a consortium to merge and jointly analyze all extant data sets for linkage. The feasibility of this approach has been demonstrated in search of genes for type 1 diabetes [176]. However, since there may be considerable differences in sampling strategies, in phenotypic measurement, marker sets, and expected etiologic heterogeneity, sometimes it is not possible to directly pool the data from studies that are conducted independently without standardization. In this case, meta-analyses of multiple independent data sets have been proposed as an alternative [177]. We advocate that both significant and non-significant results of whole genome scans should be published to facilitate meta-analyses. On the other hand, multicenter genetic and family studies are rapidly evolving as a means of generating large samples of family data collected by using standardized protocols, for example, the Family Blood Pressure Program (FBPP) [178].
Conclusions
Considerable efforts have been made recently to investigate the genetic basis of osteoporosis. Numerous candidate genes have been tested for association and linkage with osteoporosis. However, these candidate genes are often neither essential nor sufficient to produce osteoporosis on their own. Whole genome studies have identified some regions that may harbor QTLs contributing to osteoporosis. However, there is currently little consensus about loci or identity of specific genes that confer genetic susceptibility to the development of osteoporosis in different populations. Furthermore, the transition from QTL detection to gene identification has proven difficult. Nevertheless, the successful identification of NOD2 [179,180] and calpain-10 susceptibility loci [181] for Crohn's disease and type II diabetes, as well as replication of some of linkage findings across multiple studies is encouraging. With the anatomy of the human genome at hand, sequence-based gene discovery is complementing, and will eventually replace, map-based gene discovery. Identifying sequence variations responsible for osteoporosis and understanding how these variations regulate the phenotypes will still be the major challenges in the future. We can be optimistic concerning the future of gene discovery for osteoporosis by the use of a combination of functional, positional, and expression information.
References
Consensus Development Conference (1993) Diagnosis, prophylaxis and treatment of osteoporosis. Am J Med 94:646–650
Kanis JA, Melton LJ, Christiansen C, Johnston, CC, Khaltaev N (1994) The diagnosis of osteoporosis. J Bone Miner Res 9:1137–1141
Melton LJ III (1995) How many women have osteoporosis now? J Bone Miner Res 10:175–177
Sexson SB, Lehner JT (1988) Factors affecting hip fracture mortality. J Orthopaed Trauma 1:298–305
Ray NF, Chan JK, Thamer M, Melton LJ III (1997) Medical expenditures for the treatment of osteoporotic fractures in the United States in 1995: report from the National Osteoporosis Foundation. J Bone Miner Res 12:24–35
Torgerson D, Cooper C (1998) Osteoporosis as a candidate for disease management: epidemiological and cost of illness considerations. Dis Manage Health Outcomes 3:207–214
Eisman JA (1999) Genetics of osteoporosis. Endocr Rev 788–804
Rizzoli R, Bonjour JP, Ferrari SL (2001) Osteoporosis, genetics and hormones. J Mol Endocrinol 26:79–94
Ralston SH (2002) Genetic control of susceptibility to osteoporosis. J Clin Endocrinol Metab 87:2460–2466
Liu YZ, Liu YJ, Recker RR, Deng HW (2003) Molecular genetic studies of gene identification for osteoporosis: the 2002 update. J Endocrinol (in press)
Cummings SR, Nevitt MC, Browner WS et al. (1995) Risk factors for hip fracture in white women. N Engl J Med 332:767–773
Keen RW, Hart DJ, Arden NK, Doyle DV, Spector TD (1999) Family history of appendicular fracture and risk of osteoporosis: a population-based study. Osteoporos Int 10:161–166
Kannus P, Palvanen M, Kaprio J, Parkkari J, Koskenvuo M (1999) Genetic factors and osteoporotic fractures in elderly people: prospective 25 year follow up of a nationwide cohort of elderly Finnish twins. BMJ 319:1334–1337
Deng HW, Chen WM, Recker S et al. (2000) Genetic determination of Colles' fractures and differential bone mass in women with and without Colles' fractures. J Bone Miner Res 15:1243–1252
MacGregor A, Snieder H, Spector TD (2000) Genetic factors and osteoporotic fractures in elderly people: twin data support genetic contribution to risk of fracture. BMJ 320:1669–1670
Blank RD (2001) Breaking down bone strength: a perspective on the future of skeletal genetics. J Bone Miner Res 16:1207–1211
Li X, Masinde G, Gu W, Wergedal J, Mohan S, Baylink DJ (2002) Genetic dissection of femur breaking strength in a large population (MRL/MpJ × SJL/J) of F2 mice: single QTL effects, epistasis, and pleiotropy. Genomics 79:734–740
Duan Y, Seeman E, Turner CH (2001) The biomechanical basis of vertebral body fragility in men and women. J Bone Miner Res 16:2276–2283
Krall EA, Dawson-Hughes B (1993) Heritable and life-style determinants of bone mineral density. J Bone Miner Res 8:1–9
Smith DM, Nance WE, Kang KW, Christian JC, Johnston CC Jr (1973) Genetic factors in determining bone mass. J Clin Invest 52:2800–2808
Dequeker J, Nijs J, Verstraeten A, Geusens P, Gevers G (1987) Genetic determinants of bone mineral content at the spine and radius: a twin study. Bone 8:207–209
Pocock NA, Eisman JA, Hopper JL, Yeates MG, Sambrook PN, Eberl S (1987) Genetic determinants of bone mass in adults. A twin study. J Clin Invest 80:706–710
Slemeda SW, Christian JCC, Williams CJ, Norton JA, Johnston Jr CC (1991) Genetic determinants of bone mass in adult woman: a reevaluation of the twin model and the potential importance of gene interaction on heritability estimates. J Bone Miner Res 6:561–567
Flicker L, Hopper JL, Rodgers L, Kaymakci B, Green RM, Wark JD (1995) Bone density determinants in elderly women: a twin study. J Bone Miner Res 10:1607–1613
Arden NK, Baker J, Hogg C, Baan K, Spector TD (1996) The heritability of bone mineral density, ultrasound of the calcaneus and hip axis length: a study of postmenopausal twins. J Bone Miner Res 11:530–534
Howard GM, Nguyen TV, Harris M, Kelly PJ, Eisman JA (1998) Genetic and environmental contributions to the association between quantitative ultrasound and bone mineral density measurements: a twin study. J Bone Miner Res 13:1318–1327
Sowers MR, Boehnke M, Jannausch ML, Crutchfield M, Corton G, Burns TL (1992) Familiality and partitioning the variability of femoral bone mineral density in woman of child-bearing age. Calcif Tissue Int 50:110–114
Gueguen R, Jouanny P, Guillemin F, Kuntz C, Pourel J, Siest G (1995) Segregation analysis and variance components analysis of bone mineral density in health families. J Bone Miner Res 10:2017–2022
Deng HW, Stegman MR, Davies M, Conway T, Recker RR (1999) Genetic determination of peak bone mass (PBM) at hip and spine and common familiar environmental effects on bone qualities. J Clin Densitom 2:251–263
Deng, HW, Chen WM, Conway T et al. (2000) Determination of BMD at hip and spine by genetic and life-style factors. Genet Epidemiol 19:160–177
Livshits G, Karasik D, Pavlovsky, O, Kobyliansky E (1999) Segregation analysis reveals a major gene effect in compact and cancellous bone mineral density in two populations. Hum Biol 71:155–172
Cardon LR, Garner C, Bennett ST et al. (2000) Evidence for a major gene for bone mineral density in idiopathic osteoporotic families. J Bone Miner Res 15:1132–1137
Deng HW, Livshits G, Yakovenko K et al. (2002) Evidence for a major gene for bone mineral density/content in human pedigrees identified via probands with extreme bone mineral density. Ann Hum Genet 66:61–74
Nguyen TV, Howard GM, Kelly PJ, Eisman JA (1998) Bone mass, lean mass and fat mass: same genes or same environments. Am J Epidemiol 147:3–16
Deng HW, Mahaney MC, Williams J (2002) Relevance of the genes for bone mass variation to susceptibility to osteoporotic fractures and its implications to gene search for complex human diseases. Genet Epidemiol 22:12–25
Orwoll ES, Belknap JK, Klein RF (2001) Gender specificity in the genetic determinants of peak bone mass. J Bone Miner Res 16:1962–1971
Lu PW, Cowell CT, Lloyd-Jones SA et al. (1996) Volumetric bone mineral density in normal subjects aged 5–27 years. J Clin Endocrinol Metab 81:1586–1590
Faulkner KG, Cummings SR, Black D, Palermo L, Gluer CC, Genant HK (1993) Simple measurement of femoral geometry predicts hip fracture: the study of osteoporotic fractures. J Bone Miner Res 8:1211–1217
Moller M, Horsman A, Harvald B, Hauge M, Henningsen K, Nordin BE (1978) Metacarpal morphometry in monozygotic dizygotic elderly twins. Calcif Tissue Res 25:197–201
Ferrari S, Rizzoli R, Slosman D, Bonjour JP (1998) Familial resemblance for bone mineral mass is expressed before puberty. J Clin Endocrinol Metab 83:358–361
Deng HW, Deng XT, Conway T, Xu FH, Heaney R, Recker RR (2002) Determination of bone size of hip, spine, and wrist in human pedigrees by genetic and lifestyle factors. J Clin Densitom 5:45–56
Nguyen TV, Blangero J, Eisman JA (2000) Genetic epidemiological approaches to the search for osteoporosis genes. J Bone Miner Res 15:392–401
Garnero P (2000) Markers of bone turnover for the prediction of fracture risk. Osteoporos Int 11:S55–65
Harris M, Nguyen TV, Howard GM, Kelly PJ, Eisman JA (1998) Genetic and environmental correlations between bone formation and bone mineral density: a twin study. Bone 22:141–145
Kelly PJ, Hopper JL, Macaskill GT, Pocock NA, Sambrook PN, Eisman JA (1991) Genetic factors in bone turnover. J Clin Endocrinol Metab 72:808–813
Tokita A, Kelly PJ, Nguyen TV et al. (1994) Genetic influences on type I collagen synthesis and degradation: further evidence for genetic regulation of bone turnover. J Clin Endocrinol Metab78:1461–1466
Garnero P, Arden NK, Griffiths G, Delmas PD, Spector TD (1996) Genetic influence on bone turnover in postmenopausal twins. J Clin Endocrinol Metab 81:140–146
Deng HW (2001) Population admixture may appear to mask, change or reverse genetic effects of genes underlying complex traits. Genetics 159:1319–1323
Allison DB (1997) Transmission-disequilibrium tests for quantitative traits. Am J Hum Genet 60:676–690
Klein RF, Mitchell SR, Phillips TJ, Belknap JK, Orwoll ES (1998) Quantitative trait loci affecting peak bone mineral density in mice. J Bone Miner Res 13:1648–1656
Klein RF, Carlos AS, Vartanian KA et al. (2001) Confirmation and fine mapping of chromosomal regions influencing peak bone mass in mice. J Bone Miner Res 16:1953–1961
Shimizu M, Higuchi K, Bennett B et al. (1999) Identification of peak bone mass QTL in spontaneously osteoporotic mouse strain. Mamm Genome 10:81–87
Shimizu M, Higuchi K, Kasai S et al. (2001) Chromosome 13 locus, Pbd2, regulates bone density in mice. J Bone Miner Res 12:1972–1982
Beamer WG, Shultz KL, Churchill GA et al. (1999) Quantitative trait loci for bone density in C57BL/6J and CAST/EiJ inbred mice. Mamm Genome 10:1043–1049
Beamer WG, Shultz KL, Donahue LA et al. (2001) Quantitative trait loci for femoral and lumbar vertebral bone mineral density in C57BL/6J and C3H/HeJ inbred strains of mice. J Bone Miner Res 16:1195–1206
Drake TA, Schadt E, Hannani K et al. (2001) Genetic loci determining bone density in mice with diet-induced atherosclerosis. Physiol Genom 5:205–215
Benes H, Weinstein RS, Zheng W et al. (2000) Chromosomal mapping of osteopenia-associated quantitative trait loci using closely related mouse strains. J Bone Miner Res 15:626–633
Masinde GL, Li X, Gu W, Wergedal J, Mohan S, Baylink DJ (2002) Quantitative trait loci for bone density in mice: the genes determining total skeletal density and femur density show little overlap in F2 mice. Calcif Tissue Int 71:421–428
Eichner JE, Friedrich CA, Cauley JA et al. (2000) Alpha 2-HS glycoprotein phenotypes and quantitative hormone and bone measures in postmenopausal women. Calcif Tissue Int 47:345–349
Ho NC, Jia L, Driscoll CC, Gutter EM, Francomano CA (1999) A skeletal gene database. J Bone Miner Res 15:2095–2122
Uitterlinden AG, van Leeuwen JPTM, Pols HAP (2001) Genetics and genomics of osteoporosis. In: Marcus R, Feldman D, Kelsey J (eds) Osteoporosis, vol 1. Academic Press, New York, pp 639–667
Gong G, Stern HS, Cheng SC et al. (1999) The association of bone mineral density with vitamin D receptor gene polymorphisms. Osteoporos Int 9:55–64
Spotila LD, Colige A, Sereda L et al. (1994) Mutation analysis of coding sequences for type I procollagen in individuals with low bone density. J Bone Miner Res 9:923–932
Grant SFA, Reid DM, Blake G, Herd R, Fogelman I, Ralston SH (1996) Reduced bone density and osteoporosis associated with a polymorphic Sp1 binding site in the cillagen type Iα1 gene. Nat Genet 14:203–205
Lei SF, Deng FY, Liu XH et al. (2003) Polymorphisms of four bone mineral density candidate genes in Chinese populations and comparison with other populations of different ethnicity. J Bone Miner Metab21:34–42
Mann V, Hobson EE, Li B et al. (2001) A COL1A1 Sp1 binding site polymorphism predisposes to osteoporotic fracture by affecting bone density and quality. J Clin Invest 107:899–907
Efstathiadou Z, Tsataoulis A, Ioannidis JPA (2001) Association of collagen Iα1 Sp1 polymorphism with the risk of prevalent fractures: a meta-analysis. J Bone Miner Res 16:1586–1592
Ioannidis JP, Stavrou I, Trikalinos TA et al. (2002) Association of polymorphisms of the estrogen receptor α gene with bone mineral density and fracture risk in women: a meta-analysis. J Bone Miner Res 17:2048–2060
Willing M, Sowers M, Aron D et al. (1998) Bone mineral density and its change in white women: estrogen and vitamin D receptor genotypes and their interaction. J Bone Miner Res13:695–705
Gennari L, Becherini L, Masi L et al. (1998) Vitamin D and estrogen receptor allelic variants in Italian postmenopausal women: evidence of multiple gene contribution to bone mineral density. J Clin Endocrinol Metab 83:939–944
Deng HW, Li J, Li JL, Johnson M, Gong G, Recker RR (1999) Association of vdr and estrogen receptor genotypes with bone mass in postmenopausal Caucasian women: different conclusions with different analyses and the implications. Osteoporos Int 9:499–507
Mizunuma H, Hosoi T, Okano H et al. (1997) Estrogen receptor gene polymorphism and bone mineral density at the lumber spine of pre- and postmenopausal women. Bone 21:379–383
Han KO, Moon IG, Kang YS, Chung HY, Min HK, Han IK (1997) Nonassociation of estrogen receptor genotypes with bone mineral density and estrogen responsiveness to hormone replacement therapy in Korean postmenopausal women. J Clin Endocrinol Metab 82:991–995
Deng HW, Li J, Li JL, Johnson M, Davies M, Recker RR (1998) Change of bone mass in postmenopausal Caucasian women with and without hormone replacement therapy is associated with vitamin D receptor and estrogen receptor genotypes. Hum Genet 103:576–585
Langdahl BL, Knudsen JY, Jensen HK, Gregersen N, Eriksen EF (1997) A sequence variation: 713–8delC in the transforming growth factor-ß1 gene has higher prevalence in osteoporotic women than in normal women and is associated with very low bone mass in osteoporotic women and increased bone turnover in both osteoporotic and normal women. Bone 20:289–294
Yamada Y, Miyauchi A, Goto J et al. (1998) Association of a polymorphism of the transforming growth factor-β1 gene with genetic susceptibility to osteoporosis in postmenopausal Japanese women. J Bone Miner Res 13:1569–1576
Yamada Y (2000) Association of a Leu (10)→Pro polymorphism of the transforming growth factor-beta1 with genetic susceptibility to osteoporosis and spinal osteoarthritis. Mech Ageing Dev 116:113–123
Yamada Y, Miyauchi A, Takagi Y, Tanaka M, Mizuno M, Harada A (2001) Association of the C509→T polymorphism, alone or in combination with the T869→C polymorphism of the transforming growth factor-beta1 gene with bone mineral density and genetic susceptibility to osteoporosis in Japanese women. J Mol Med 79:149–156
Bertoldo F, D'Agruma L, Furlan F et al. (2000) Transforming growth factor-beta1 gene polymorphism, bone turnover, and bone mass in Italian postmenopausal women. J Bone Miner Res 15:634–639
Keen RW, Snieder H, Molloy H et al. (2001) Evidence of association and linkage disequilibrium between a novel polymorphism in the transforming growth factor beta1 gene and hip bone mineral density: a study of female twins. Rheumatology 40:48–54
Hosoi T, Miyao M, Inoue S et al. (1999) Association study of parathyroid hormone gene polymorphism and bone mineral density in Japanese postmenopausal women. Calcif Tissue Int 64:205–208
Deng HW, Shen H, Xu FH et al. (2002) Tests of linkage and/or association of genes for vitamin D receptor, steocalcin, and parathyroid hormone with bone mineral density. J Bone Miner Res 17:678–686
Masi L, Becherini L, Colli E et al. (1998) Polymorphisms of the calcitonin receptor gene are associated with bone mineral density in postmenopausal Italian women. Biochem Biophys Res Commun 248:190–195
Taboulet J, Frenkian M, Frendo JL, Feingold N, Jullienne A, De Vernejoul MC (1998) Calcitonin receptor polymorphism is associated with a decreased fracture risk in post-menopausal women. Hum Mol Genet 7:2129–2133
Braga V, Mottes M, Mirandola S et al. (2000) Association of CTR and COLIA1 alleles with BMD values in peri- and postmenopausal women. Calcif Tissue Int 67:361–366
Nakamura M, Morimoto S, Zhang Z et al. (2001) Calcitonin receptor gene polymorphism in Japanese women: correlation with body mass and bone mineral density. Calcif Tissue Int 68:211–215
Tsukamoto K, Orimo H, Hosoi T et al. (2000) Association of bone mineral density with polymorphism of the human calcium-sensing receptor locus. Calcif Tissue Int 66:181–183
Takacs I, Speer G, Bajnok E et al. (2002) Lack of association between calcium-sensing receptor gene "A986S" polymorphism and bone mineral density in Hungarian postmenopausal women. Bone 30:849–852
Dohi Y, Iki M, Ohgushi H et al. (1998) A novel polymorphism in the promoter region for the human osteocalcin gene: the possibility of a correlation with bone mineral density in postmenopausal Japanese women. J Bone Miner Res 13:1633–1639
Sowers M, Willing M, Burns T et al. (1999) Genetic markers, bone mineral density, and serum osteocalcin levels. J Bone Miner Res 14:1411–1419
Raymond MH, Schutte BC, Torner JC, Burns TL, Willing MC (1999) Osteocalcin: genetic and physical mapping of the human gene BGLAP and its potential role in postmenopausal osteoporosis. Genomics 60:210–217
Tsukamoto K, Orimo H, Hosoi T et al. (2000) Association of bone mineral density with polymorphism of the human matrix Gla protein locus in elderly women. J Bone Miner Metab 18:27–30
Chen HY, Tsai HD, Chen WC, Wu JY, Tsai FJ, Tsai CH (2001) Relation of polymorphism in the promotor region for the human osteocalcin gene to bone mineral density and occurrence of osteoporosis in postmenopausal Chinese women in Taiwan. J Clin Lab Anal 15:251–255
Murray RE, McGuigan F, Grant SFA, Reid DM, Ralson SH (1997) Polymorphisms of the interleukin-6 gene are associated with bone mineral density. Bone 21:89–92
Tsukamoto K, Yoshida H, Watanabe S et al. (1999) Association of radial bone mineral density with CA repeat polymorphism at the interleukin 6 locus in postmenopausal Japanese women. J Hum Genet 44:148–151
Ota N, Hunt SC, Nakajima T et al. (1999) Linkage of interleukin 6 locus to human osteopenia by sibling pair analysis. Hum Genet 105:253–257
Ota N, Nakajima T, Nakazawa I et al. (2001) A nucleotide variant in the promoter region of the interleukin-6 gene associated with decreased bone mineral density. J Hum Genet 46:267–272
Ferrari SL, Garnero P, Emond S, Montgomery H, Humphries SE, Greenspan SL (2001) A functional polymorphic variant in the interleukin-6 gene promoter associated with low bone resorption in postmenopausal women. Arthr Rheum 44:196–201
Garnero P, Borel O, Sornay-Rendu E et al. (2002) Association between a functional interleukin-6 gene polymorphism and peak bone mineral density and postmenopausal bone loss in women: the OFELY study. Bone 31:43–50
Rosen CJ, Kurkland ES, Vereault D et al. (1998) Association between serum insulin-like growth factor-1 (IGF-1) and a simple sequence repeat in IGF-1 gene: Implications for genetic studies of bone mineral density. J Clin Endocrinol Metab 83:2286–2290
Meulenbelt I, Bijkerk C, Micdema HS et al. (1998) A genetic association study of the IGF-1 gene and radiological osteoarthritis in a population-based cohort study (the Rotterdam study). Ann Rheum Dis 57:371–374
Miyao M, Hosoi T, Tnoue S et al. (1998) Polymorphism of insulin-like growth factor I gene and bone mineral density. Calcif Tissue Int 63:306–311
Takacs I, Koller DL, Peacock M et al. (1999) Sibling pair linkage and association studies between bone mineral density and the insulin-like growth factor I gene locus. J Clin Endocrinol Metab 84:4467–4471
Kim JG, Roh KR, Lee JY (2002) The relationship among serum insulin-like growth factor-I, insulin-like growth factor-I gene polymorphism, and bone mineral density in postmenopausal women in Korea. Am J Obstet Gynecol 186:345–350
Shiraki M, Shiraki Y, Aoki C et al. (1997) Association of bone mineral density with apoplipoprotein E phenotype. J Bone Miner Res 12:1438–1445
Cauley JA, Zmuda JM, Kuller LH, Ferrell RE, Wisniewski SR, Cummings SR (1999) Apolipoprotein E polymorphism: a new genetic marker of hip fracture risk—the study of osteoporotic fractures. J Bone Miner Res 14:1175–1181
Heikkinen AM, Kroger H, Niskanen L et al. (2000) Does apolipoprotein E genotype relate to BMD and bone markers in postmenopausal women? Maturitas 34:33–41
Zmuda JM, Eichner JE, Ferrell RE, Bauer DC, Kuller H, Cauley JA (1998) Genetic variation in alpha 2 HS-glycoprotein is related to calcaneal broadband ultrasound attenuation in older women. Calcif Tissue Int 63:5–8
Keen RW, Woodford-Richens KL, Lanchbury JS, Spector TD (1998) Allelic variation at the interleukin-1 receptor antagonist gene is associated with early postmenopausal bone loss at the spine. Bone 23:367–371
Ogawa S, Urano T, Hosoi T et al. (1999) Association of bone mineral density with a polymorphism of the peroxisome proliferator-activated receptor gamma gene: PPARgamma expression in osteoblasts. Biochem Biophys Res Commun 260:122–126
Spotila LD, Rodriguez H, Koch M et al. (2000) Association of a polymorphism in the TNFR2 gene with low bone mineral density. J Bone Miner Res 15:1376–1383
Miyao M, Hosoi T, Emi M et al. (2000) Association of bone mineral density with a dinucleotide repeat polymorphism at the calcitonin (CT) locus. J Hum Genet 45:346–350
Urano T, Hosoi T, Shiraki M, Toyoshima H, Ouchi Y, Inoue S (2000) Possible involvement of the p57kip2 gene in bone metabolism. Biochem Biophys Res Commun 269: 422–426
Miyao M, Morita H, Hosoi T et al. (2000) Association of Methylenetetrahydrofolate reductase (MTHFR) polymorphism with bone mineral density in postmenopausal Japanese women. Calcif Tissue Int 66:190–194
Masi L, Becherini L, Gennari L et al. (2001) Polymorphism of the aromatase gene in postmenopausal Italian women: distribution and correlation with bone mass and fracture risk. J Clin Endocrinol Metab 86:2263–2269
Ogata N, Shiraki M, Hosoi T, Koshizuka Y, Nakamura K, Kawaguchi H (2001) A polymorphic variant at the Werner helicase (WRN) gene is associated with bone density, but not spondylosis, in postmenopausal women. J Bone Miner Metab 19:296–301
Yamada Y, Ando F, Niino N, Shimokata H (2002) Association of a polymorphism of the CC chemokine receptor-2 gene with bone mineral density. Genomics 80:8–12
Ogata N, Matsumura Y, Shiraki M et al. (2002) Association of Klotho gene polymorphism with bone density and spondylosis of the lumbar spine in postmenopausal women. Bone 31:37–42
Vaughan T, Pasco JA, Kotowicz MA, Nicholson GC, Morrison NA (2002) Alleles of RUNX2/CBFA1 gene are associated with differences in bone mineral density and risk of fracture. J Bone Miner Res 17:1527–1534
Hirschhorn JN, Lohmueller K, Byrne E, Hirschhorn K (2002) A comprehensive review of genetic association studies. Genet Med 4:45–61
Risch NJ (2000) Searching for genetic determinants in the new millennium. Nature 405:847–856
Cardon LR, Bell JI (2001) Association study designs for complex diseases. Nat Rev Genet 2:91–99
Cooper DN, Nussbaum RL, Krawezak M (2002) Proposed guidelines for papers describing DNA polymorphism-disease associations. Hum Genet 110:207–208
Devoto M, Shimoya K, Caminis J et al. (1998) First-stage autosomal genome screen in extended pedigrees suggests genes predisposing to low bone mineral density on chromosomes 1p, 2p and 4q. Eur J Hum Genet 6:151–157
Mitchell BD, Bauer RL, Perez R et al. (1998) Genome-wide scan for loci influencing bone density in Mexican Americans. Am J Hum Genet 63:A301 (Abstract 1741)
Niu T, Chen C, Cordel H et al. (1999) A genome-wide scan for loci linked to forearm bone mineral density. Hum Genet 104:226–233
Koller DL, Econs MJ, Rodriguez LA et al. (2000) Genome scan for QTLs contributing to normal variation in bone mineral density and osteoporosis J Clin Endocrinol Metab 85:3116–3120
Styrkarsdottir U, Jonasson K, Johannsdottir KH et al. (2001) Evidence for a locus of a major osteoporosis gene in an icelandic linkage study. Bone 28:S72
Karasik D, Myers RH, Cupples LA et al. (2002) Genome screen for quantitative trait loci contributing to normal variation in bone mineral density: the Framingham study. J Bone Miner Res 17:1718–1727
Deng HW, Xu FH, Huang QY et al. (2002) A whole-genome linkage scan suggests several genomic regions potentially containing quantitative trait loci for Osteoporosis. J Clin Endocrinol Metab 87:5151–5159
Econs MJ, Koller DL, Hui SL et al. (2002) Lumbar Spine BMD is Linked to Genetic Markers on Chromosome 1q. J Bone Miner Res 17:S189
Wilson SG, Reed PW, Bansal A et al. (2003) Comparison of genome screens for two independent cohorts provides replication of suggestive linkage of bone mineral density to 3p21 and 1p36. Am J Hum Genet 72:144–155
Karasik D, Myers RH, Hannan MT et al. (2001) Mapping of quantitative ultrasound of the calcaneus to chromosomes 1 and 5 by genome-wide linkage analysis. J Bone Miner Res 16:S167
Koller DL, Liu G, Econs MJ et al. (2001) Genome screen for QTLs contributing to normal variation in femoral structure. J Bone Miner Res 16:985–991
Huang QY, Xu FH, Shen H et al. (2002) Genome scan for QTLs underlying bone size variation at ten refined skeletal sites: genetic heterogeneity and the value of the subdivision of traits. Am J Hum Genet 71:431
Deng HW, Shen H, Xu FH et al. (2003) Several genomic regions potentially containing QTLs for bone size variation were identified in a whole-genome linkage scan. Am J Med Genet 119A:121–131
Gong Y, Vikkula M, Boon L et al. (1996) Osteoporosis-pseudoglioma syndrome, a disorder affecting skeletal strength and vision, is assigned to chromosome region 11q12–13. Am J Hum Genet 59:146–151
Gong Y, Slee RB, Fukai N et al. (2001) LDL receptor-related protein 5 (LRP5) affects bone accrual and eye development. Cell 107:513–523
Heaney C, Carmi R, Dushkin H, Sheffield V, Beier DR (1997) Genetic mapping of recessive osteopetropsis to 11q12–13. Am J Hum Genet 61:A12
Frattini A, Orchard PJ, Sobacchi C et al. (2000) Defects in TCIRGI subunit of the vacuolar proton pump are responsible for a subset of human autosomal recessive osteopetrosis. Nat Genet 25:343–346
Kornak U, Schulz A, Friedrich W et al. (2000) Mutations in the a3 subunit of the vacuolar H(+)-ATPase cause infantile malignant osteopetrosis. Hum Mol Genet 9:2059–2063
Sobacchi C, Frattini A, Orchard P, et al. (2001) The mutational spectrum of human malignant autosomal recessive osteopetrosis. Hum Mol Genet 10:1767–1773
Li YP, Chen W, Liang Y, Li E, Stashenko P (1999) Apt6I-deficient mice exhibit severe osteopetrosis due to loss of osteoclast-mediated extracellular acidification. Nat Genet 23:447–451
Johnson ML, Gong GD, Kimberling W, Recker SM, Kimmel DB, Recker RR (1997) Linkage of a gene causing high bone mass to human chromosome 11 (11q12–13). Am J Hum Genet 60:1326–1332
Little RD, Carulli JP, Del Mastro RJ, et al. (2002) A mutation in the LDL receptor-related protein 5 gene results in the autosomal dominant high-bone-mass trait. Am J Hum Genet 70:11–19
Boyden LM, Mao J, Belsky J et al. (2002) High bone density due to a mutation in LDL-receptor-related protein 5. N Engl J Med 346:1513–1521
Levasseur R, Kato M, Patel MS, Chan L, Karsenty G (2001) Low bone mass, low body weight and abnormal eye vascularization in mice deficient in Lrp5, the gene mutated in human osteoporosis pseudoglioma syndrome (OPS). J Bone Miner Res 16:S152
Koller DL, Rodriguez LA, Christian JC et al. (1999) Linkage of a QTL contributing to normal variation in bone mineral density to chromosome 11q12–13. J Bone Miner Res 13:1903–1908
Deng HW, Xu FH, Conway T (2001) Is population BMD variation linked to the marker D11S987 on chromosome 11q12–13? J Clin Endocrinol Metab 86:3735–3741
Devoto M, Specchia C, Li HH et al. (2001) Variance component linkage analysis indicates a QTL for femoral neck bone mineral density on chromosomes 1p36. Hum Mol Genet 10:2447–2452
Albagha OM, McGuigan FEA, Reid D, Ralston SH (1999) Association mapping of a locus for regulation of bone mass in the normal population using DNA pooling. J Bone Miner Res 14:S142
Albagha OM, Tasker PN, McGuigan FEA, Reid D, Ralston SH (2002) Linkage disequilibrium between polymorphisms in the human TNFRSF1B gene and their association with bone mass in perimenopausal women. Hum Mol Genet 11:2289–2295
Duncan EL, Brown MA, Sinsheimer J et al. (1999) Suggestive linkage of the parathyroid receptor type 1 to osteoporosis. J Bone Miner Res14:1993–1999
Andersson-Eklund L, Uhlhorn H, Lundeheim N, Dalin G, Andersson L (2000) Mapping quantitative trait loci for principal components of bone measurements and osteochondrosis scores in a wild boar × large white intercross. Genet Res 75:223–230
Mitchell BD, Kammerer CM, Schneider JL et al. (2001) A quantitative trait locus on chromosome 4p influences variation in bone mineral density at the wrist and hip. J Bone Miner Res 16:S167
Hughes AE, Shearman AM, Weber JL et al. (1994) Genetic linkage of familial expansile osteolysis to chromosome 18q. Hum Mol Genet 3:359–361
Hughes AE, Ralston SH, Marken J et al. (2000) Mutations in TNFRSF11A, affecting the signal peptide of RANK, cause familial expansile osteolysis. Nat Genet 24:45–48
Cody JD, Singer FR, Roodman GD et al. (1997) Genetic linkage of Paget disease of the bone to chromosome 18q. Am J Hum Genet 61:1117–1122
Haslam SI, Van Hul W, Morales-Piga A et al. (1998) Paget's disease of bone: evidence for a susceptibility locus on chromosome 18q and for genetic heterogeneity. J Bone Miner Res 13:911–917
Good DA, Busfield F, Fletcher BH et al. (2002) Linkage of Paget disease of bone to a novel region on human chromosome 18q23. Am J Hum Genet 70:517–525
The International SNP Map Working Group (2001) A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409:928–933
Feakes R, Sawcer S, Chataway J et al. (1999) Exploring the dense mapping of a region of potential linkage in complex disease: an example in multiple sclerosis. Genet Epidemiol 17:51–63
van Heel DA, McGovern DPB, Cardon LR et al. (2002) Fine mapping of the IBD1 locus did not identify Crohn disease-associated NOD2 variants: implications for complex disease genetics. Am J Med Genet 111:253–259
Olavesen MG, Hampe J, Mirza MM et al. (2000) Analysis of single-nucleotide polymorphism in the interleukin-4 receptor gene for association with inflammatory bowel disease. Immunogenetics 51:1–7
Martin ER, Lai EH, Gilbert JR et al. (2000) SNPing away at complex diseases: analysis of single-nucleotide polymorphisms around APOE in Alzheimer disease. Am J Hum Genet 67:383–394
Daly MJ, Rioux JD, Schaffner SF, Hudson TJ, Lander ES (2001) High-resolution haplotype structure in the human genome. Nat Genet 29:229–232
Patil N, Berno AJ, Hinds DA et al. (2001) Blocks of limited haplotype diversity revealed by high-resolution scanning of human chromosome 21. Science 294:1719–1723
Stephens JC, Schneider JA, Tanguay DA et al. (2001) Haplotype variation and linkage disequilibrium in 313 human genes. Science 293:489–493
Meltzer PS (2001) Spotting the target: microarrays for disease gene discovery. Curr Opin Genet Dev 11:258–263
Lawn RM, Wade DP, Garvin MR et al. (1999) The Tangier disease gene product ABC1 controls the cellular apolipoprotein-mediated lipid removal pathway. J Clin Invest 104:R25–R31
Kim S, Kim M, Kim J, Choi J, Shin H, Park E (2002) Expression profiling of genes involved in osteoporosis using DNA microarray. J Bone Miner Res 17:S320
Sellers TA, Yates JR (2003) Review of proteomics with application to genetic epidemiology. Genet Epidemiol 24:83–98
Hanash S (2003) Disease proteomics. Nature 422:226–232
Cox NJ, Frigge M, Nicolae DL et al. (1999) Loci on chromosomes 2 (NIDD1) and 15 interact to increase susceptibility to diabetes in Mexican Americans. Nature Genet 21:213–215
Cordell HJ, Wedig GC, Jacobs KB, Elston RC (2000) Multilocus linkage tests based on affected relative pairs. Am J Hum Genet 66:1273–1286
Cox NJ, Wapelhorst B, Morrison VA et al. (2001) Seven regions of the genome show evidence of linkage to type 1 diabetes in a consensus analysis of 767 multiplex families. Am J Hum Genet 69:820–830
Allison DB, Heo M (1998) Meta-analysis of linkage data under worst-case conditions: a demonstration using the human OB region. Genetics 148:859–865
The FBPP Investigators (2002) Multi-center genetic study of hypertension: the family blood pressure program (FBPP). Hypertension 39:3–9
Hugot JP, Chamaillard M, Zouali H et al. (2001) Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn's disease. Nature 411:599–603
Ogura Y, Bonen DK, Inohara N et al. (2001) A frameshift mutation in NOD2 associated with susceptibility to Crohn's disease. Nature 411:603–606
Horikawa Y, Oda N, Cox NJ et al. (2000) Genetic variation in the gene encoding calpain-10 is associated with type 2 diabetes mellitus. Nat Genet 26:163–175
Acknowledgements
Investigators of this work are partially supported by grants from Health Future Foundation, NIH grants (K01 AR02170-01, R01 AR45349-01, R01 GM60402-01A1, P01 DC01813-07), grants from State of Nebraska Cancer and Smoking Related Disease Research Program (LB595) and the Nebraska Tobacco Settlement Fund (LB692), US Department of Energy grant DE-FG03–00ER63000/A00, Creighton University, grants (30025025, 30170504, 30230210) from National Science Foundation of China, a Seed Fund (25000106) and a key grant from the Ministry of Education of People's Republic of China, a grant (25000612) from HuNan Normal University, a grant (81017) from Huo Ying Dong Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huang, QY., Recker, R.R. & Deng, HW. Searching for osteoporosis genes in the post-genome era: progress and challenges. Osteoporos Int 14, 701–715 (2003). https://doi.org/10.1007/s00198-003-1445-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00198-003-1445-9