Abstract
Objective
To define the potential of resistant gram-negative colonization surveillance to predict etiology of subsequent infection and improve adequacy of empiric antimicrobial treatment.
Design
Retrospective cohort study.
Setting
A mixed medical–surgical six-bed intensive care unit (ICU), from November 2003 to December 2006.
Patients
All patients having at least one episode of ventilator-associated pneumonia (VAP) or bloodstream infection (BSI) caused by a resistant gram-negative pathogen during the study period.
Interventions
Colonization surveillance of the respiratory tract and gastrointestinal tract was systematically performed in all ICU patients. Tracheal aspirates were obtained twice weekly and rectal swabs once weekly. Both tracheal and rectal samples were cultured in antibiotic-enriched media (containing ceftazidime, ciprofloxacin, imipenem or piperacillin/tazobactam), to focus on resistant gram-negative pathogen isolation.
Measurements and results
Colonization concordance between resistant, gram-negative pathogens of infectious episodes and previous, recent (≤7 days) colonization of the respiratory and gastrointestinal tract was determined, based on species identity and antimicrobial susceptibility. Concordance was 82% in VAP and 86% in BSI cases and was further confirmed by molecular testing of 15 randomly selected cases by REP-PCR. Previous colonization had high sensitivity and specificity in VAP, but was less specific in BSI cases. Knowledge of previous colonization improved the rate of adequate empiric antimicrobial treatment (91 vs. 40% in VAP and 86 vs. 50% in BSI cases, P < 0.05).
Conclusions
Colonization surveillance for resistant gram-negative microorganisms is predictive of subsequent infection etiology and can improve empiric antimicrobial treatment adequacy in a critical care setting.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Appropriate empiric antimicrobial treatment in sepsis directly affects mortality in the intensive care unit (ICU) [1] and consists of timely and adequate coverage of the implicated pathogen.
The choice of empiric antimicrobial treatment for critically ill patients is often challenging. These patients have usually long hospital stays, complex underlying pathology and previous antimicrobial exposure; they are commonly infected with drug-resistant pathogens, with often variable antimicrobial resistance patterns. Current guidelines suggest the using of local epidemiologic data and individual patient risk factors to prescribe empiric antibiotic treatment [2], in less than 1 h after onset of severe sepsis or septic shock [3].
Theoretically, colonization with a pathogen is a prerequisite for subsequent invasive disease. In critically ill patients, colonization of the respiratory and gastrointestinal tract with nosocomial flora occurs within 48–72 h after admission [4], to be variably followed by invasive disease. It has been hypothesized that colonization surveillance in such patients can provide early insight in the microbial etiology of subsequent infection, allowing thus the provision of adequate empiric treatment in a timely fashion. Several studies have addressed the issue [5–18]; there is however equivocal data on the clinical effectiveness of such an approach, in terms of adequacy of empiric antimicrobial therapy prescribed, of impact on survival or of its cost-effectiveness.
In our ICU, a systematic surveillance strategy has been adopted since 2003 to identify patients colonized with resistant gram-negative microorganisms. Resistant gram-negatives are implicated in the majority of infections in our setting, accounting for about 90% of ventilator-associated pneumonia (VAP) and 50% of bloodstream infection (BSI) etiology. In this context, colonization data has been used as a tool for empiric antibiotic therapy prescription in the setting of infection. We organized a retrospective study to investigate the hypothesis that systematic surveillance for gram-negative resistant pathogens in critically ill patients could predict microbial etiology of subsequent infection and improve the rate of appropriate empiric therapy. Part of this study has been presented before [19].
Materials and methods
Setting
The study was performed in the ICU of University General Hospital “ATTIKON,” a 700-bed hospital in Athens metropolitan area. During the study period, the ICU enlarged gradually, from 4 beds in 2004 to 12 beds by the end of 2006 (median 6 beds), serving a mixed medical and surgical population.
Study design
The study is a retrospective cohort of all episodes of ICU-acquired VAP and BSI caused by resistant gram-negative bacteria, from November 2003 to December 2006. VAP and BSI episodes were analyzed according to previous colonization and adequacy of empiric therapy received. The study design was approved by the Ethical Committee of the hospital.
Surveillance methodology
Infection control policy in our ICU included twice weekly tracheal aspirates and once weekly rectal swabs. If patients were not intubated, then tracheal aspirates were substituted by pharyngeal swabs. Identification was performed using routine microbiological methodologies and an automated identification system (API ID32GN and ID32E system, bioMerieux, Marcy-L’Etoile, France). Besides routine culturing, all specimens were also streaked on antibiotic-enriched Mac-Conkey agar plates containing antibiotics, namely ceftazidime, ciprofloxacin, piperacillin/tazobactam and imipenem. Growth in any of the above media during the first 24 h of incubation suggested the presence of resistant pathogens and was reported to treating physicians the day after sampling, as a preliminary result [20]. Antibiotic susceptibility was determined by the disk diffusion method; in pathogenic strains isolated from infectious episodes, exact MICs were also determined by the agar dilution method (Broth Microdilution panels-Sensititre Oxoid). Culture results with formal microbial identification and resistance patterns were reported to the treating physicians, 2–3 days after sampling.
Definitions
Only VAP or BSI caused by gram-negative bacteria resistant to one or more of ceftazidime, ciprofloxacin, piperacillin/tazobactam or imipenem were studied.
VAP was suspected when a new or worsening pulmonary infiltrate associated with two of four clinical signs of infection (fever, leukocytosis, purulent sputum or worsening gas exchange) developed in a patient on mechanical ventilation for more than 48 h. VAP suspicion was followed by quantitative culture of a tracheal aspirate and/or bronchoalveolar lavage specimen, and pneumonia was diagnosed when quantitative cultures were ≥106 cfu/mL or 104 cfu/ mL, respectively.
BSI was defined as the isolation of a pathogen in at least one peripherally drawn blood culture. BSIs were considered ICU-acquired when diagnosed 48 h or more after ICU admission. Day of onset of infection was defined as the day when clinical deterioration of the patient developed, usually as a new septic episode, necessitating diagnostic specimens for culture and the initiation of empirical antimicrobial treatment.
Colonization concordance was defined as the isolation of the same pathogenic microorganism in prior respiratory or gastrointestinal tract colonization specimens (RT or GT, respectively) and the infectious focus of the patient (lung or bloodstream). Only the most recent colonization results (i.e. ≤7 preceding days), available at the time of the onset of the infection, were taken into account. Microbial concordance was based on species identification and identical antibiotic susceptibility testing. For randomly selected isolates, colonization concordance was confirmed using molecular typing by repetitive extragenic palindromic (REP)–PCR methodology.
Empiric antibiotic therapy was considered appropriate when the patient received at least one effective in vitro antibiotic against the isolated pathogen(s), during the first 24 h after the onset of the infectious episode. For extended-spectrum β-lactamase-producing enterobacteriaceae, either a carbapenem or colimycin was considered adequate therapy.
Antibiotic policy
In our ICU, there was a daily infectious disease consultation. The selection of empiric antibiotic regimens was based on the patient’s underlying pathology, length of previous hospitalization, colonization status, presumed infectious focus and hemodynamic status. Carbapenems, piperacillin/tazobactam, glycopeptides and colimycin were reserved for patients with nosocomial septic shock, or other risk factors for drug-resistant etiology of infection (hospital stay > 5 days, transfer from another hospital or ICU, prior receipt of extended-spectrum antibiotics or known drug-resistant pathogen colonization). Indications for antimicrobial treatment were discussed in daily rounds. Shorter duration treatments and de-escalation practices were encouraged, if indicated.
Statistical analysis
Continuous variables are presented as median (quartiles). Categorical variables are presented as number (percentage). Odds ratios (OR) and 95% confidence intervals (95% CI) are reported where appropriate. The Chi-square test or Fisher’s exact test were used for comparisons. Statistical analyses were executed with SPSS software (version 13.0; SPSS Inc., Chicago, IL). All tests are two-tailed and statistical significance is defined as P value less than 0.05.
Results
During the study period, 404 patients were admitted in the ICU. In total, 35 cases of VAP and 104 cases of BSI were diagnosed. Patient characteristics and microbiology of ICU-acquired infection are described in Table 1.
Analysis of ventilator-associated pneumonia episodes
During the study period, 31 VAP episodes caused by resistant gram-negative pathogens were diagnosed in 30 patients.
Colonization concordance for VAP pathogens
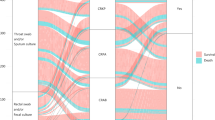
Recent colonization data were available in 28 out of 31 cases. In 23 out of 28 VAP episodes (82%), prior colonization of either the respiratory or gastrointestinal tract could accurately predict the implicated pathogens (Fig. 1). Both sites seemed to be equally important for pathogen prediction (69% for RT vs. 58% for GT) and data from both of them were additive. Median duration of prior colonization before VAP diagnosis was 7 days (5–17) for the RT and 9.5 days (6–23) for the GT. Concordance was high for most pathogens with no significant differences observed between them (Acinetobacter 84%, Pseudomonas 80%, Klebsiella 100%).
Operational characteristics of prior colonization as a diagnostic tool for VAP etiology diagnosis
The use of the presence of colonization of the major pathogens (Acinetobacter, Pseudomonas or Klebsiella) as a diagnostic test for the prediction of subsequent gram-negative resistant VAP etiology had high sensitivity and specificity, as shown in Table 2.
VAP pathogen prediction, adequacy of empiric therapy and ICU mortality
There were 27 VAP episodes with available data on empiric therapy. In 22 VAP cases out of 27 (81%), adequate empiric treatment was started in the first 24 h of infection. Of the five cases who did not receive adequate therapy, three were not predicted by previous colonization. Previous colonization permitted 91% empiric treatment adequacy, while failure of pathogen prediction by colonization resulted in only 40% treatment adequacy (P = 0.03, OR 15, 95% CI 0.9–254.0) (Table 3). No difference was observed in ICU mortality, independent of either pathogen prediction by previous colonization or receipt of adequate empiric therapy.
Analysis of bloodstream infection episodes
During the study period, 55 BSI episodes caused by resistant gram-negative pathogens were diagnosed in 33 patients. Two of them were considered secondary to an intraabdominal focus.
Colonization concordance for BSI pathogens
Available recent colonization data were available in 43 out of 55 bacteremias. In 37 out of the 43 BSI episodes (86%), prior colonization of either the respiratory or gastrointestinal tract could accurately predict the implicated pathogens (Fig. 1). Both sites seemed to be equally important for pathogen prediction (66% for RT vs. 70% for GT) and data from both of them were additive. Concordance was high for most pathogens with no significant differences observed between them (Acinetobacter 100%, Pseudomonas 93%, Klebsiella 86%).
Duration of colonization before bacteremia was variable for both the RT and GT. Median duration of colonization before BSI diagnosis was 11 days (6–27) for the RT and 15 days (6–29.5) for the GT.
Operational characteristics of prior colonization as a diagnostic tool for BSI etiology diagnosis
The use of the presence of colonization of the major pathogens (Acinetobacter, Pseudomonas or Klebsiella) as a diagnostic test for the prediction of subsequent gram-negative resistant VAP etiology had high sensitivity but lower specificity, as shown in Table 2.
BSI pathogen prediction, adequacy of empiric therapy and ICU mortality
There were 54 BSI episodes with available data on empiric therapy. In 42 BSI cases out of 54 (78%), adequate empiric treatment was started in the first 24 h of infection. Of the 12 cases who did not receive adequate therapy, six were not predicted by previous colonization. Previous colonization permitted 86% empiric treatment adequacy, while failure of pathogen prediction by colonization resulted in 50% treatment adequacy (P = 0.016, OR 6, 95% CI 1.5–24.9) (Table 3). No difference was observed in ICU mortality, independent of either pathogen prediction by previous colonization or receipt of adequate empiric therapy.
Molecular analysis of colonization concordance
In fifteen, randomly-selected cases, molecular typing by REP-PCR was performed to confirm concordance between pathogens and colonizers in bronchial secretions or stool. Among six VAP cases, four were caused by Acinetobacter baumannii and two were by Klebsiella pneumoniae; all pathogenic isolates were identical to previous colonizers in tracheal aspirates. Similarly, in nine BSI cases, five were caused by Klebsiella pneumoniae, two by Enterobacter cloacae, one by Acinetobacter baumannii and one more by Stenotrophomonas maltophilia; all isolates proved identical to previous colonizers in either bronchial secretions (in five cases) and/or rectal swabs (in five cases) (Fig. 2).
REP-PCR molecular typing of 12 (A1–D12) Klebsiella pneumoniae isolates from four patients (A–D). Patient A bloodstream infection, A1 blood, A2 tracheal aspirate, A3 stool, A4 pus; patient B ventilator-associated pneumonia, B5 tracheal aspirate, B6 BAL; patient C ventilator-associated pneumonia, C7 tracheal aspirate, C8 stool, C9 BAL, C10 pus; patient D bloodstream infection, D11 stool, D12 blood
Discussion
This study was performed to evaluate the potential of serial surveillance of resistant, gram-negative microbial colonization to predict the etiologic pathogens of subsequent infection in critically ill patients. Colonization surveillance showed significant concordance with infectious pathogens and exhibited high sensitivity as a diagnostic test. More importantly, knowledge of previous colonization permitted higher rates of adequate empiric therapy, without however having a significant impact on ICU mortality.
Colonization surveillance has several potential advantages in an ICU. It can be used to detect newly admitted patients with resistant pathogen colonization, indicating the need for contact isolation measures. It can also aid in the early detection and follow-up of epidemics. Finally, it can provide insight in the prevailing mechanism of drug resistance in an ICU, whether due to horizontal spread between patients or due to increased antibiotic pressure and de novo resistance development in individual patients [21].
The use of colonization surveillance data for determination of empiric therapy has not been studied extensively in the literature and its practice is at best equivocal. Best sites for colonization surveillance are a matter of debate. Published studies have used cultures of the lower respiratory tract, oropharynx, nasopharynx, urine, intact skin, trauma, stool, removed catheter tips or drainage fluid in varying combinations. To focus on gram-negative pathogens and achieve a simple but sensitive enough surveillance protocol, we favored a combination of respiratory tract and gastrointestinal tract sampling. Tracheal aspirate cultures have a, theoretically, obvious relation to the development of respiratory infection, are easy to obtain and are commonly surveyed in ICU. Stool samples, on the other hand, mirror the content of the largest ecological niche for microbial flora in the human body. It seems that the gastrointestinal tract is among the first sites to get colonized in a critically ill patient. It serves as a deposit for subsequent urinary tract, skin or respiratory tract colonization and is also a major site of resistance development under antibiotic pressure [4]. Furthermore, intestinal flora has been linked to direct bloodstream invasion through the process of allothesis.
Interestingly enough, the value of intestinal colonization and subsequent bacteremia has not been adequately studied. In a study performed in very low birth weight infants, 18 out of 19 gram-negative BSI pathogens were genotypically identical to antecedent rectal colonizers [15]. In BSIs caused by extended-spectrum β-lactamase-producing Enterobacteriaceae, concordant rectal colonization was preceding in 76% of cases [14], similar to the 70% concordance for resistant gram-negatives reported in our study.
The frequency of colonization surveillance is also equivocal. It seems that the shorter the time interval between colonization sampling and infection, the better its predictive value [6]. Studies using bronchial surveillance less often than once weekly have a low predictive potential [6, 7]. According to our data, the time interval between colonization and subsequent infection is variable, but in a significant number of cases is less than a week, suggesting that surveillance more than once weekly of both RT and GT would add to the diagnostic accuracy.
In this study, we tried to corroborate the hypothesis that a colonization surveillance strategy can be used as a guide to empiric therapy in a clinically meaningful way. However, it should be noted that it has been performed in a special context of high incidence of antimicrobial resistance. Greek ICUs have a high rate of nosocomial infection [22], attributed to more severely admitted patients, longer length of stay, increased use of invasive devices and low nurse-to-patient ratios [23]. More than half of infections are caused by gram-negative bacteria, mainly Acinetobacter, Pseudomonas and Klebsiella species, most of them resistant to several groups of antibiotics [24]. The present study also does not provide any information on gram-positive colonization and its impact on subsequent infection prediction, considering their rarity in our setting. It is thus probable that our results might not be applicable in settings with different infection epidemiology or resistance patterns than ours.
The first step in adopting such a colonization surveillance strategy for empiric therapy prescription is to establish its effectiveness in predicting the responsible pathogens. Our data suggest that colonization surveillance can predict subsequent infection etiology in 82% of VAP cases and 86% of bacteremias. Several groups have reported similar colonization concordance rates, ranging at 61–88% [9, 11–13, 16–18] in VAP cases and 75–82% in BSI cases [8, 14, 15]. Lower concordance rates have been published from groups performing surveillance cultures once weekly or less [5, 6], as well as from studies with low incidence of drug-resistant colonization or infection [16].
Some reports also describe the operational characteristics of colonization data as a diagnostic tool to predict the pathogen of subsequent infection [5, 13, 17, 18]. In VAP cases, prior colonization seems to be both sensitive and specific, while in BSI cases, specificity is lower although sensitivity remains high. In a setting of high incidence of resistant gram-negative infection, the above diagnostic characteristics translate into adequate negative predictive value (87–96%); this could prove a useful tool in excluding resistant gram-negative infection, in the absence of preceding colonization.
The next step in adopting such a colonization surveillance strategy is to demonstrate that it has meaningful clinical advantages, such as an improvement in the adequacy of empiric therapy prescription or in ICU outcomes. In our study, pathogen prediction resulted in higher rates of empiric therapy adequacy in the first 24 h of infection, for both VAP and BSI cases. Comparable results have been published in the literature [8, 9, 11, 12]. One study has also shown that pathogen prediction by colonization correlates with improved survival [11]. Our strategy could not demonstrate any significant effect on ICU mortality, as it was obviously not adequately powered for that.
A final, third step is to show that such a surveillance strategy is cost-effective in terms of laboratory workload or antibiotic use. Prescribing empiric therapy based on colonization results poses a definite antibiotic overuse danger, if antimicrobial treatment is targeted against colonizing microorganisms rather than infectious pathogens. Among few published data on the subject, three studies tried to clarify the issue by comparing antibiotic use based on a colonization surveillance strategy versus several hypothetical antimicrobial schemes, based on published guidelines [9, 12, 18]: in all three of them, empiric treatment based on surveillance results proved more adequate, while at the same time permitting significantly lower usage of broad spectrum antibiotics. However, no randomized, controlled trial has been performed to evaluate the impact of colonization surveillance on antibiotic use in clinical practice.
In conclusion, a systematic surveillance strategy for gram-negative resistant pathogen colonization can serve as a simple, rapid and effective aid to empiric antimicrobial treatment prescription in the ICU. However, more studies are needed to define the potential impact of the use of such an approach on mortality and its cost-effectiveness.
References
Kollef MH (2000) Inadequate antimicrobial treatment: an important determinant of outcome for hospitalized patients. Clin Infect Dis 31(Suppl 4):S131–S138
American Thoracic Society Documents (2005) Guidelines for the management of adults with hospital-acquired, ventilator-associated and healthcare-associated pneumonia. Am J Respir Crit Care Med 171:388–416
Dellinger RP, Carlet JM, Masur H, Gerlach H, Calandra T, Cohen J, Gea-Banacloche J, Keh D, Marshall JC, Parker MM, Ramsay G, Zimmerman JL, Vincent JL, Levy MM (2004) Surviving Sepsis Campaign guidelines for management of severe sepsis and septic shock. Intensive Care Med 30:536–555
Bonten MJ, Weinstein RA (1996) The role of colonization in the pathogenesis of nosocomial infections. Infect Control Hosp Epidemiol 17:193–200
Delclaux C, Roupie E, Blot F, Brochard L, Lemaire F, Brun-Buisson C (1997) Lower respiratory tract colonization and infection during acute respiratory distress syndrome. Am J Resp Crit Care Med 156:1092–1098
Hayon J, Figliolini C, Combes A, Trouillet JL, Kassis N, Dombret MC, Gibert C, Chastre J (2002) Role of serial routine microbiologic culture results in the initial management of ventilator-associated pneumonia. Am J Resp Crit Care Med 165:41–46
Bouza E, Perez A, Munoz P, Perez J, Rincon C, Sanchez C, Martin-Rabadan P, Riesgo M, the Cardiovascular Infection Study Group (2003) Ventilator-associated pneumonia after heart surgery: a prospective analysis and the value of surveillance. Crit Care Med 31:1964–1970
Blot S, Depuydt P, Vogelaers D, Decruyenaere J, De Waele J, Hoste E, Peleman R, Claeys G, Verschraegen G, Colardyn F, Vandewoude K (2005) Colonization status and appropriate antibiotic therapy for nosocomial bacteremia caused by antibiotic-resistant gram-negative bacteria in an intensive care unit. Infect Control Hosp Epidemiol 26:575–579
Michel F, Franceschini B, Berger P, Arnal JM, Gainnier M, Sainty JM, Papazian L (2005) Early antibiotic treatment for BAL-confirmed ventilator-associated pneumonia. Chest 127:589–597
Galoisy-Guibal L, Soubirou JL, Desjeux G, Dusseau JY, Eve O, Escarment J, Ecochard R (2006) Screening for multidrug-resistant bacteria as a predictive test for subsequent onset of nosocomial infection. Infect Control Hosp Epidemiol 27:1233–1241
Depuydt P, Benoit D, Vogelaers D, Claeys G, Verschraegen G, Vandewoude K, Decruyenaere J, Blot S (2006) Outcome in bacteremia associated with nosocomial pneumonia and the impact of pathogen prediction by tracheal surveillance cultures. Intensive Care Med 32:1773–1781
Depuydt PO, Blot SI, Benoit DD, Claeys GW, Verschraegen GL, Vandewoude KH, Vogelaers DP, Decruyenaere JM, Colardyn FA (2006) Antimicrobial resistance in nosocomial bloodstream infection associated with pneumonia and the value of systematic surveillance cultures in an adult intensive care unit. Crit Care Med 34:653–659
Malacarne P, Corini M, Maremmani P, Viaggi B, Verdigi S (2007) Diagnostic characteristics of routine surveillance cultures of endotracheal aspirate samples in cases of late-onset ventilator-associated pneumonia due to Acinetobacter baumannii. Infect Control Hosp Epidemiol 28:867–869
Reddy P, Malczynski M, Obias A, Reiner S, Jin N, Huang J, Noskin GA, Zembower T (2007) Screening for extended spectrum β-lactamase producing Enterobacteriaceae among high-risk patients and rates of subsequent bacteremia. Clin Infect Dis 45:846–852
Graham P, Della-Latta P, Wu F, Zhou J, Saiman L (2007) The gastrointestinal tract serves as the reservoir for gram-negative pathogens in very low birth weight infants. Pediatr Infect Dis J 26:1153–1156
Sanders K, Adhikari N, Friedrich J, Day A, Jiang X, Heyland D, for the Canadian Critical Care Trials Group (2008) Previous cultures are not clinically useful for guiding empiric antibiotics in suspected ventilator-associated pneumonia: secondary analysis from a randomized trial. J Crit Care 23:58–63
Boots RJ, Philips GE, George N, Faoagali JL (2008) Surveillance culture utility and safety using low-volume blind bronchoalveolar lavage in the diagnosis of ventilator-associated pneumonia. Respirology 13:87–96
Depuydt P, Benoit D, Vogelaers D, Decruyenaere J, Vandijck D, Claeys G, Verschraegen G, Blot S (2008) Systematic surveillance cultures as a tool to predict involvement of multidrug antibiotic resistant bacteria in ventilator-associated pneumonia. Intensive Care Med 34:675–682
Papadomichelakis E, Kontopidou F, Kopterides P, Mavrou I, Paramythiotou E, Poulakou G, Antoniadou A, Giamarellou H, Armaganidis A (2007) A three-year retrospective study of systematic colonization surveillance in the ICU. Poster presented in the 20th ESICM Annual Congress, Berlin. Intensive Care Med 33(Suppl 2):S131
Poulakou G, Plachouras D, Kontopidou F, Antoniadou A, Papadomichelakis E, Armaganidis A, Giamarellou H (2008) Rapid surveillance of multidrug resistance in ICU patients, by use of antibiotic-containing agar plates. Poster presented in the 18th ECCMID, Barcelona. Clin Microbiol Infect 14(Suppl 4):S609–S610
Nijssen S, Boetsma M, Bonten M (2006) Potential confounding in evaluating infection-control interventions in hospital settings: changing antibiotic prescription. Clin Infect Dis 43:616–623
Vincent JL (2000) Microbial resistance: lessons from the EPIC study. Intensive Care Med 26:S3–S8
Vincent JL, Suter P, Bihari D, Bruining H (1997) Organization of intensive care units in Europe: lessons from the EPIC study. Intensive Care Med 23:1181–1184
Dima S, Kritsotakis EI, Roumbelaki M, Metalidis S, Karabinis A, Maguina N, Klouva F, Levidiotou S, Zakynthinos E, Kioumis J, Gikas A (2007) Device-associated nosocomial infection rates in intensive care units in Greece. Infect Control Hosp Epidemiol 28:602–605
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is discussed in the editorial available at: doi:10.1007/s00134-008-1249-7.
Rights and permissions
About this article
Cite this article
Papadomichelakis, E., Kontopidou, F., Antoniadou, A. et al. Screening for resistant gram-negative microorganisms to guide empiric therapy of subsequent infection. Intensive Care Med 34, 2169–2175 (2008). https://doi.org/10.1007/s00134-008-1247-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00134-008-1247-9