Abstract
In the present study, an aerobic bacterial strains OCS-A and OCS- B were isolated from an oil contaminated soil. The strains were identified to be Citrobacter freundi and Proteus mirabilis according to morphological, physiological and biochemical characteristics. The strains were able to degrade about 90% of 100 mg/L phenol within 80 h as sole carbon and energy source. The lag phase increased with increase in phenol concentration. Determination of metabolic intermediate 2-HMS, was done which indicate meta-cleavage pathway of phenol metabolism. Hence these isolates can be effectively used for bioremediation of phenol contaminated sites.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Now a day’s phenol is used by many industries in the manufacturing of products like insulation materials, adhesives, lacquers, paint, rubber, ink, dyes, illuminating gases, perfumes, soaps and toys. Phenol is one of the most widely used organic compounds in existence and is a basic structural unit for a variety of synthetic organic compounds including agricultural chemicals and pesticides, vegetables and coal. The German chemist, Runge isolated phenol from coal tar in 1834 and named karbolsaure (coal–oil acid or carbolic acid), though its composition was not known until 1841 (Nair et al. 2008).
Biodegradation of phenol has increased strong attention throughout the world due to its poisonous properties and unrelenting nature (Leung et al. 1997). Microbial uptake and mineralization of phenol has been studied extensively (Dua et al. 2002; Lovely 2003; Watanabe 2001). The microorganisms that are normally used in phenol degradation include Pseudomonas sp, Candida tropicalis, Azotobacter sp, Alcaligenes sp, Acinetobacter sp etc. (Basha et al. 2010). The biological degradation is accomplished through benzene ring cleavage mediated by intracellular enzymatic reaction (Kumar et al. 2004).
This study was aimed to evaluate the potential of phenol degraders from environment. Phenol degrading bacteria were isolated and their performance in treating the compound was investigated.
Materials and Methods
Two pure cultures used in this study were isolated from oil contaminated soil. The isolates were periodically subcultured on LB agar at 30°C and further maintained at 4°C. MP and MSM media were used in present study. MP medium composed of K2HPO4–2.75 gm, KH2PO4–2.25 gm (NH4)NO3–1.0 gm, MgCl2·6H2O–0.2 gm, NaCl–0.1 gm, FeCl3·6H2O–0.02 gm, CaCL2–0.01 gm (Watanabe et al. 1998). The minimal salt media (MSM) was used in this study. The medium was composed as: Na2HPO4 (6 g), KH2PO4 (3 g), NaCl (0.5 g), NH4Cl (1 g), CaCl2·2H2O (1 M) and MgSO47.H2O (1 M) in 1,000 mL double distilled water (Nagamani et al. 2009). The medium was autoclaved at 121°C for 15 min for sterilization. Phenol (10 mg/L) was separately sterilized and aseptically added into the sterile medium.
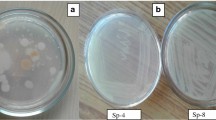
Sample was collected from contaminated soil from oil industry and added into MP and MSM media in which phenol (10 mg/L) was added as sole carbon source. It was incubated for 24 h at room temperature on 120 rpm shaker. The enriched sample was further transferred into freshly prepared enrichment media each week. Serially diluted final enriched media were spread-plated on LB agar supplemented with phenol (50 mg/L) and incubated at 30°C. The single colonies were streaked onto LB agar plates, incubated at 30°C overnight and the pure isolates were stored at 4°C until further use.
Colony morphology of the bacterium was determined by cultivating the isolates on MP and MSM medium supplemented with 50 mg/L phenol. Physiological and biochemical tests were performed including Gram’s staining, Acid fast staining, Endopore staining, Capsule staining, amylase, gelatinase, citrate utilization, indole test etc. (Aneja 2001). Bergey’s manual of determinative of bacteriology was used as reference to identify the isolates (Goodfeelow 1994).
The isolates OCS-A was inoculated into MP medium and OCS-C were inoculated into MSM containing 10 mg/L phenol as carbon source for 48 h shaken at 150 rpm. The acclimatized isolates were further exposed to higher concentration of phenol subsequently.
The OCS-A and OCS-C bacterial isolates were grown overnight in nutrient broth and the grown bacteria were centrifuged for the 15 min at 3000 rpm. The cells were resuspended in MP and MSM media, respectively having phenol (10 mg/L) as sole carbon source for degradation study. The reaction containing all components but devoid of bacterial inoculums were used as control. The concentration of phenol was increased from 10 to 100 mg/L.
The residual phenol concentration was determined by analyzing sample regularly after 4 h interval using UV visible spectrophotometer 1606 Systronics colorimetrically by 4-amino antipyrine method (Yang and Humphery 1975; APHA 1992). The cell growth (biomass) was monitored at 600 nm.
The bacterial strains OCS-A & OCS-C were spread on LB medium & incubated overnight. A 50 mg/L phenol solution was spread into the plate. The bacteria were incubated at 37°C for 25 min. The plates were eluted with a small amount of phosphate buffer (0.2 M, pH 7.0) & centrifuged at 10,000 rpm for 4°C. The supernatant was measured spectrophotometrically at 270 and 375 nm after removing the cells by centrifugation (Chen et al. 2003).
Results and Discussion
Two bacteria capable of degrading phenol were isolated from the soil sample collected near oil industry and coded as OCS-A and OCS-C. These isolates were enriched in MP and MSM media, respectively, supplemented with phenol. These strains were characterized using morphological and biochemical properties listed in Table 1. By comparing these characteristics with those mentioned in Bergey’s manual of systematic Bacteriology, the OCS-A was identified as Citrobacter freundii and OCS-C as Proteus mirabilis.
The degradation performance of selected strains was examined at different phenol concentrations. Serial exposure to increasing level of phenol concentration was used to determine the resistance of isolated strains.
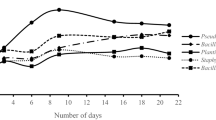
The time dependant changes of phenol concentrations in the media were evaluated. Strains OCS-A and OCS-C were able to degrade phenol almost completely (about 90%) within a relatively short time of 80 h. The phenol tolerance and growth of the isolates at elevated level of phenol concentration was investigated. Cells growing at high phenol concentration (100 mg/L) showed a longer lag time compared to those growing at low concentration (50 mg/L) (Figs. 1, 2, 3 and 4).
Phenol and 2-hydroxymuconic semialdehyde (2-HMS) are starting and ending compounds of the bioconversion which show absorbance maxima at 270 and 375 nm, respectively (Quilty and Farrell 1999). OCS-C showed absorbance (0.970) at 270 nm. It was less than control (3.325) but which was decreased gradually while absorbance at 375 nm was increased progressively (Fig. 5).
The respective increase and decrease in absorption as 270 and 375 nm indicated the formation of 2-hydroxymuconic semialdehyde as a result of extradiol ring cleavage of phenol (El-Sayed et al. 2003). These results indicate that OCS-C metabolized phenol by meta-pathway. The same pattern was followed for OCS-A.
In conclusion, we have isolated two bacteria with capability of phenol tolerance and degrading potential. Isolates were identified on the basis of Bergey’s manual of systematic Bacteriology which degraded phenol by meta cleavage pathway.
References
Aneja KR (2001) Experiments in microbiology plant pathology and biotechnology, 4th edn. 102, 106,112,245–275,278
APHA (American Public Health Association) (1992) Standard methods for the examination of water and wastewater, 18th edn. Washington, USA
Basha KM, Aravindan R, Thangavelu V (2010) Recent advances in the Biodegradation of Phenol: a review. Asian J Exp Biol Sci 1(2):219–234
Chen Y, Liu H, Chen H (2003) Characterization of phenol degradation by Comamonas testosteroni ZD4–1 and Pseudomonas aeuroginosa ZD4–3. Biomed Environ Sci 16:163–172
Dua M, Singh A, Sethunathan N, Johri AK (2002) Biotechnology and bioremediation: successes and limitations. Appl Microbiol Biot 59:143–152
El-Sayed W, Ibrahim MK, Abu-Shady M, El-Beaith F, Ohmura N, Saiki H, Ando A (2003) Isolation and characterization of phenol-catalyzing bacteria from a cooking plant. Biosci Biotechnol Biochem 67(9):2026–2029
Goodfeelow M (1994) Bergey’s manual of determinative bacteriology, 9th edn. Williams and Wilkins, London
Kumar A, Kumar S, Kumar S (2004) Biodegradation kinetics of phenol and cathecol using Pseudomonas putida 1194. Biochem Eng J 22:151–159
Leung KI, Tresse O, Errampalli D, Lee H, Trevors JT (1997) Mineralization of p-nitrophenol be pentachlorophenol—degrading Sphingomonas spp. FEMS Microbiol Lett 155:107–114
Lovely DR (2003) Cleaning up with genomics: applying molecular biology of self-bioremediation. Nat Rev Microbiol 1:35–44
Nagamani A, Soligala R, Lowry M (2009) Isolation and characterization of phenol degrading Xanthobacter flavus. Afr J Biotechnol 8(20):5449–5453
Nair IC, Jayachandran K, Shankar Shashidhar (2008) Biodegradation of phenol. Afr J Biotechnol 7(25):4951–4958
Quilty B, Farrell A (1999) Degradation of mono-chlorophenols by a mixed microbial community via a meta-cleavage pathway. Biodegradation 10:353–362
Watanabe ME (2001) Can bioremediation bounce back? Nat Biotechnol 19:1111–1115
Watanabe K, Teramoto M, Futamata H, Shigeaki H (1998) Molecular detection, isolation, and physiological characterization of functionally dominant phenol-degrading bacteria in activated sludge. Appl Environ Microbiol 1998:4396–4402
Yang RD, Humphery AE (1975) Dynamic and steady sate studies of phenol biodegradation in pure and mixed cultures. Biotechnol Bioeng 17:1211–1235
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mohite, B.V., Pawar, S.P. & Morankar, A. Isolation, Selection and Biodegradation Profile of Phenol Degrading Bacteria from Oil Contaminated Soil. Bull Environ Contam Toxicol 87, 143–146 (2011). https://doi.org/10.1007/s00128-011-0322-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00128-011-0322-z