Abstract
Four bacterial strains designated as SNTP-1, NS-2 to NS-4 were isolated from selenium contaminated soils of Nawanshahr-Hoshiarpur region of Punjab, India, by enrichment technique and a consortium was developed using these isolates. The isolates were observed to be belonging to Bacillus sp. In soil microcosm, complete removal was observed by the consortium in selenite augmented soils while the rate of removal with consortia in selenate treatment was 72% after 120 days. Population survival of isolates showed stability at lower treatments and decline at higher levels of Se enrichment. The consortium can, thus, be used for removal of Se contaminated sites.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
The behavior of selenium compounds, ranging from being essential to highly toxic has enhanced the interest in understanding its fate in water, soil and biota. Selenate and selenite are the dominant bioavailable forms of selenium in soil and can be reduced and/or methylated through microbiological processes that drive reduction and methylation processes (Stolz et al. 2002) Over the last two decades there has been considerable interest in microbial removal of Se oxyanions [i.e. selenate (SeO4 2−) and selenite (SeO3 2−)] to volatile forms such as dimethyl selenide (DMSe) and dimethyl diselenide (DMDSe) and insoluble forms viz., elemental selenium (Se0) (Dungan and Frankenberger 2001). The understanding of mechanisms underlying bacterial assimilation and removal of selenium prerequisites the information on phenotypic and phylogenetic characterization of species involved (de Souza et al. 2001; Ghosh et al. 2008).
The present study attempts to characterize and examine the viability of four selenium tolerant rhizosphere aerobic bacterial isolates in soils supplemented with selenium oxyanions viz., selenate and selenite. The consortium of these isolates was examined for its potential to remove selenium supplemented soil at various concentrations in the form of selenate and selenite.
Materials and Methods
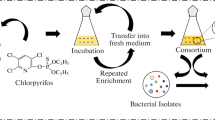
Selenium contaminated soil samples were collected from a seleniferous site bordering the Nawanshahr-Hoshiarpur region in North-west India (75°55 E; 31°56 N). Aerobic bacterial strains were isolated using repeated enrichment technique at 37°C in SeO3 2− and SeO4 2− supplemented tryptic soy agar (TSA-HiMedia). Red colonies, indicating reduction of Se oxyanions, were re-streaked on TSA without Se to confirm that the colour was not due to pigmentation. The pure cultures were isolated and maintained on Se supplemented plates. Four strains, observed to be tolerant up to 100 μg/g Se, were picked for constituting as a consortium for microcosm studies.
Morphological analysis, gram staining and biochemical characterization were carried out applying standard protocols (Krieg et al. 1984). Fatty acid methyl esters (FAMEs) were characterized in these strains at the Microbial Type Culture Collection (MTCC-IMTECH), Chandigarh, India using protocol as described by Pandey et al. (2002) and analyzed with Sherlock Microbial Identification Systems software (MIDI, USA).
Culture was grown in Luria broth overnight and DNA was isolated using standard protocols PCR amplication (Genamp PCR system, Applied Biosystem, USA) of 16S rRNA region was carried out using 16 S–F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 16 S-R (5′-ACGGGCGGTGTGTTC-3′) universal primers (Sambrook et al. 1989; Weisberg et al. 1991). The sequence for cloned genes was then generated using sequencing system (Applied Biosystems) at the DNA sequencing facility, University of Delhi, India. Sequence data from 16S rRNA genes was used to perform a preliminary phylogenetic analysis. The 16S rRNA sequence of SNTP-1, NS-2, NS-3 and NS-4 were compared with other sequences in GenBank using BLAST (www.ncbi.nlm.nih.gov/blast). The alignment was carried out with CLUSTALW and only un-ambiguous alignments were used in the phylogenetic analysis. Phylogenetic analysis was done using MEGA 4.0 following UPGMA method with bootstrap corrections (Tamura et al. 2007).
The conditions of the microcosm experiment such as soil moisture content and inoculum size were standardized, and the efficacy of the consortia for removal of Se oxyanions in soil was then tested. Soil samples (100 g of sandy loam with pH 7.3 ± 0.2; organic carbon, 0.273 ± 0.0008%; available phosphorus, 42.40 ± 2.43 mg/l; total nitrogen, 1568.0 ± 550.7 mg/l) were treated with selenate and selenite (2.5, 5.0 and 7.5 mg/kg soil). The consortium containing inoculum of the four Bacilli strains was grown in Luria broth (100 ml), harvested at log phase, and re-suspended in basal medium (10 ml).
Microcosm was prepared with un-autoclaved soils supplemented with 2.5, 5.0 and 7.5 mg/kg of selenate and selenite. These were inoculated with standardized cell concentration of 3.9 × 109 cells/g soil. Moisture content of soils was maintained at 30% throughout the experiments. Antibiotic profiles of the isolates were studied to select the marker for screening of inoculated microorganisms from indigenous soil bacteria. Survival of populations of inoculated strains was also assessed by antibiotic plating and the rate of removal was estimated based on difference between the initial concentrations of augmented Se to that of residual concentration at different intervals in percentage.
Selenium estimation was carried out using instrumental neutron activation analysis at Bhabha Atomic Research Centre, Mumbai, India. The soil samples were dried at 40°C for 2–3 days for removal of moisture and were ground in mortar pestle. The samples were sieved using 0.2 mm mesh. Samples were air dried followed by oven at 40°C for 48 h before irradiation. IAEA SL-1 (sludge multi-element; Se concentration: 7.34 ± 0.18) and NRCC CRM DOLT-1 (Dogfish liver standard; Se concentration: 2.90 ± 0.08) were used as standards. Samples and reference materials weighing about 100 mg were packed in thin aluminum foils. The samples, the reference materials and blank silica were introduced into Harwell cans and irradiated in self-server position of CIRUS reactor (Bhabha Atomic Research Centre, Mumbai, India) for 7 h duration at a neutron flux of ~1013 cm−2s−1. The samples were allowed to cool for about ten days before radioactive assay and the samples were counted for 1–10 h depending on selenium concentration levels. The gamma-ray spectrometric measurements were carried out using a Compton suppressed spectrometer consisting of HPGe-BGO detector systems coupled to PC based 8 k MCA card (PHAST-BARC-India). The resolution of detector was 2.0 keV at 1332 keV of 60Co. The peak areas were determined using peak-fit software PHAST. Relative method of NAA was used to calculate Se concentrations in the sample (Lin et al. 1997).
Results and Discussion
The soil samples examined from the Se-impacted pockets of agricultural lands were found to be alkaline and Se levels averaging at 6.5 mg/kg which was above the stipulated level in agricultural soils i.e., 4.0 mg/kg (Sharma et al. 2009). Four aerobic isolates prominent in Se oxyanions reduction were selected and designated as SNTP-1, NS-2, NS-3 and NS-4. Morphological and biochemical characteristics were examined for isolates which indicated that three of the four strains were consistent with that of typical facultative anaerobic bacteria. The isolate, NS-3 showed negative results in catalase and nitrate test which was attributed to the absence of facultative anaerobic nature. Isolates were also found to be methyl red negative (acid production) and nitrate positive i.e. capable of anaerobic respiration. These observations corresponded with that of studies on characteristics of genus Bacillus (Foldes et al. 2001).
Homology analysis of 16S rRNA provides suitable genetic data that could be used to determine both close and very distant relationships (Bull et al. 1992). BLAST homology search for 16S rRNA gene sequences of isolates indicated 98–99% similarity with their closest matches. The sequences were compared with that of the published sequences of Bacilli and other distantly related strains available in the NCBI database. The 16S rRNA gene sequence of SNTP-1 showed 100% similarity with Bacillus pumilus, thus clustered along with other B. pumilus sequences. The 16S rRNA gene sequences determined for isolates under study were deposited in GenBank of NCBI data library under accession numbers SNTP1 (EU532490), NS-2 (EU622629), NS-3 (EU573774) and NS-4 (EU622630). The sequences of strains NS-2, NS-3 and NS-4 showed broad identity B. cereus and B. thuringiensis. A phylogenetic dendrogram was drawn following UPGMA method. The tree shows that the sequences of the test strains clearly belong to Bacilli clade consisting the closely similar sequences mentioned above (Fig. 1).
Species of Bacilli are divided into six groups (A–F) based on the FAME studies (Kaneda 1977). The dominant cellular fatty acids present in four isolates were 13-methyltetradacanoin (iso-C15:O)—44.32% (SNTP-1), 26.38% (NS-2), 20.12% (NS-3), 30.52% (NS-4) followed by 12-Methyltetradecanoic (anteiso-C15) -35.07% (SNTP-1), 4.57% (NS-2), 5.91% (NS-3). Strains NS-2, NS-3 and NS-4 belong to Kaneda group E (B. anthracis, B. cereus, B. thuringiensis) in which iso-C15:O (19–31%) occur as the most abundant fatty acid with small proportion of unsaturated fatty acids (7–12%). Identification of all isolates based on MIDI similarity indices obtained from TSBA50 5.00 version library match showed all isolates belong to Bacilli group and SNTP-1 showed close match with strain Bacillus pumillus strain and NS-3 with Bacillus cereus while NS-2 and NS-4 belongs to Bacillus sp.
In the present study, 2 antibiotics viz., nalidixic acid and carbenicillin did not inhibit the growth of bacteria at concentration of up to 90 mcg and 300 mcg respectively, indicating resistance of the isolates to these antibiotics. Resistance to a group of selective antibiotics can effectively be used as a screening factor to track the survival rate of isolate present amongst the native bacteria (Lawrence 2000). Keeping this in view, resistance of the isolates to carbenicillin, was used as an antibiotic marker for microcosm studies.
In soil microcosm studies, observations on population survival based on carbenicillin resistance, showed an increase in colony forming units (CFU) in all treatments from 40 to 120 days studies. The microbial population in microcosm exposed to 7.5 mg/kg selenite did not vary (5.7; 6.1 and 5.6 (×105) cfu/gm) significantly across the duration of 40, 80 and 120 days (Fig. 2a, b).
Rate of selenium removal [a selenite and b selenate] and corresponding viability of the consortia in soils supplemented with 7.5 mg/kg of Se. The gray tone and white tone in graphs represent CFU count and rate of removal (%), respectively. NI represents soils without inoculum of consortia and I represents soils augumented with inoculum
Across all treatments, maximum survival rate was observed in soil supplemented with 2.5 mg/kg during 120 day period. The corresponding trends in selenium removal over time, on exposure to 7.5 mg/kg of selenium oxyanions, are also represented in Fig. 2a, b. Determination of Se concentration in soils supplemented with 2.5 mg/kg selenite revealed 32%, 52% and almost complete removal of Se across 40, 80 after 120 days as compared to control (without inoculum) in which it was 40% after 120 days. Similarly, the extent of removal of Se from soils was observed to be near complete in inoculated soils supplemented with 5 mg/kg of selenite where as in control soils with same level of Se supplementation it was 26% after 120 days. At 7.5 mg/kg of selenite, the level of removal was relatively less with only 80% when compared to initial concentration. In inoculated soils treated with 2.5 mg/kg of selenate, the removal was 24% at 40 days increasing to 60% at 120 days as compare to control which was 28% after 120 days. The rate of removal followed a similar trend even in soils supplemented with 5.0 mg/kg and 7.5 mg/kg.
The observations, thus, present an insight on the characteristics of four selenium tolerant isolates obtained from the seleniferous soils. Phylogenetic studies revealed that isolates were belonging to Bacilli group. During soil removal studies, the consortium of these isolates was able to effectively remove Se from soils contaminated with selenium oxyanions along with increase in survival rate of inoculated consortia with time. High rate of removal was achieved in soils augmented with selenite as compare to selenate, indicating better bioaccessibility of selenite over selenate to soil microbiota.
Microorganisms can mobilize and remove metals/metalloids through diverse variety of mechanisms (Gadd 2004). They can take up the elements and accumulate them in their biomass via intracellular sequestration or precipitation, or adsorb them onto cell walls and exopolymers released into their surroundings (Zaidi and Musarrat 2004). Similarly, the fate of selenium in contaminated soils is linked to the activity of soil bacteria, which is the primary method by which soluble and biologically active forms of selenium can be effectively assimilated and removed from ambient environment (Stolz et al. 2002), as observed in the present study. The role of rhizosphere processes in removal of inorganic contaminants, with the exception of some studies regarding selenium, is largely unexplored and needs to be addressed before the potential of this approach can be properly evaluated. Soil microorganisms are known to convert some metals and metalloids (i.e. arsenic, boron, antimony, selenium, tin, tellurium, lead, mercury) to their volatile species. In the case of selenium, the volatile methylated species are less toxic than inorganic forms (Wilber 1980). This microbial conversion is usually considered as a detoxification mechanism by which the microorganisms decrease the toxicity of the surrounding microenvironment. However, the use of micro-organisms for bioremediation requires an understanding of all physiological, microbiological, ecological, biochemical and molecular aspects involved in pollutant transformation (Iranzo et al. 2001). This study augments to limited reports available on selenium reduction potential of microbial strains and consortia from seleniferous tropical soils.
References
Bull AT, Goodfellow M, Slater JH (1992) Biodiversity as a source of innovation in biotechnology. Annu Rev Microbiol 46:219–252
De Souza MP, Amini A, Dojka MA, Pickering LJ, Dawson SC, Pace NR, Terry N (2001) Identification and characterization of bacteria in a selenium-contaminated hypersaline evaporation pond. Appl Environ Microbiol 67:3785–3794
Dungan RS, Frankenberger WT (2001) Bioremoval of selenium by Enterobacter cloacae SLD1a–1: formation of dimethlselenide. Biogeochemistry 55:73–86
Foldes T, Banhegyi I, Herpai Z, Varga L, Szigeti J (2001) Isolation of Bacillus strains from the rhizosphere of cereals and in vitro screening for antagonism against phytopathogenic, food-borne pathogenic and spoilage microorganisms. J Appl Microbiol 89:840–846
Gadd GM (2004) Microbial influence on metal mobility and application to bioremediation. Geoderma 122:109–119
Ghosh A, Mohod AM, Paknikar KM, Jain RK (2008) Isolation and characterization of selenite and selenate tolerant microorganisms from selenium contaminated sites. World J Microbiol Biotechnol 24:1607–1611
Iranzo M, Sainz-Pardo I, Boluda R, Sanchez J, Mormeneo S (2001) The use of microorganisms in environmental remediation. Ann Microbiol 51:135–143
Kaneda T (1977) Fatty acids of the genus Bacillus: an example of branched-chain preference. Bacteriol Rev 41:391–418
Krieg NR, Holt JG, Murray RGE (1984) Bergey’s manual of systematic bacteriology, vol 1-2. Williams and Wilkins, Baltimore
Lawrence JG (2000) Clustering of antibiotic resistance genes: Beyond the selfish operon. ASM News 66:281–286
Lin X, Baumgartner F, Li X (1997) The program “MULTINAA” for various standardization methods in neutron activation analysis. J Radioanal Nucl Chem 215:179–191
Pandey KK, Mayilraj S, Chakrabarti T (2002) Pseudomonas indica sp. Nov., a novel butane-utilizing species. Int J Syst Evol Microbiol 52:1559–1567
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor, New York
Sharma N, Prakash R, Srivastava A, Sadana US, Acharya R, Prakash NT, Reddy AVR (2009) Profile of selenium in soil and crops in seleniferous area of Punjab, India by neutron activation analysis. J Radioanal Nucl Chem 281:59–62
Stolz JF, Basu P, Oremland RS (2002) Microbial removal of elements in the case of arsenic and selenium. Int Microbiol 5:201–207
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA 4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Weisberg WA, Barns SM, Pelletier DA, Lane DJ (1991) 16S rRNA amplification for phylogenetic study. J Bacteriol 173:697–703
Wilber CG (1980) Toxicology of selenium: a review. Clin Toxicol 17:171–230
Zaidi S, Musarrat J (2004) Characterisation and nickel sorption kinetics of a new metal hyper-accumulator, Bacillus sp. J Environ Sci Health 39:681–691
Acknowledgments
The authors acknowledge the funding provided by Board of Research in Nuclear Sciences (BRNS), Govt. of India reactor facilities provided by Scientific Officers, Research Reactors, BARC, Mumbai.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Prakash, N.T., Sharma, N., Prakash, R. et al. Removal of Selenium from Se Enriched Natural Soils by a Consortium of Bacillus Isolates. Bull Environ Contam Toxicol 85, 214–218 (2010). https://doi.org/10.1007/s00128-010-0061-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00128-010-0061-6