Abstract
In this study, arsenic resistant bacteria were isolated from sediments of an arsenic contaminated river. Arsenic tolerance of bacteria isolated was carried out by serial dilution on agar plate. Redox abilities were investigated using KMnO4. arsC and aox genes were detected by PCR and RT-PCR, respectively. Bacterial populations were identified by RapID system. Forty nine bacterial strains were isolated, of these, 55 % corresponded to the reducing bacteria, 4% to oxidizing bacteria, 8% presented both activities and in 33% of the bacteria none activity was detected. arsC gene was detected in 11 strains and aox genes were not detected. The activity of arsenic transforming microorganisms in river sediment has significant implications for the behavior of the metalloid.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Arsenic is a toxic metalloid naturally found as inorganic oxyanion arsenate As(V) and arsenite As(III) species. Arsenate is the predominant specie in oxygenated aqueous environment, whereas arsenite species predominate under anoxic or reduced conditions, being 100 times more toxic than As(V) (Al-Abed et al. 2007; Taerakul et al. 2007; Neff 1997). Arsenite has the ability to bind to sulfhydryl groups of proteins and dithiols such as glutaredoxin. On the other hand, arsenate is a chemical analog of phosphate and can inhibit oxidative phosphorylation (Ordoñez et al. 2005).
Microorganisms are known to play an important role in the biochemical cycle of arsenic, through its conversion to species with different solubility, mobility, bioavailability and toxicity (Silver and Phung 2005). Several bacteria involved in arsenic transformation processes-featuring reduction, oxidation and methylation mechanisms-have been reported (Ellis et al. 2001). Arsenate reduction could occur as a respiratory process or as part of arsenic resistance, through the gene arsC encoded arsenate reductase, arsenite oxidation or methylation as well as active extrusion of arsenite from the cell. Arsenite oxidation can be coupled to chemoautotrophic growth, or occurs as a detoxification mechanism (Silver and Phung 2005).
There are regions in the world where arsenic is present at very high levels due to natural geological conditions, as in the Bengal Basin and Northern Chile (Smedley and Kinniburgh 2002). Indeed, in the case of the Atacama Dessert in Northern Chile, arsenic is leached out from volcanic materials naturally present in watershed areas in the Andes, adding to antropogenic pollution due to mining activities. As a result, some rivers contain arsenic concentration levels in the range 0.1–1.5 mgL−1, becoming a serious health risks to the local population (Queirolo et al. 2000; Yañez et al. 2005).
Since the local microbial biota is expected to be adapted to those conditions, this work aims to characterize arsenic-resistant bacteria isolated from sediments obtained from a highly polluted river in Northern Chile.
Materials and Methods
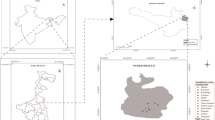
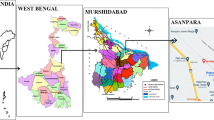
Sediment samples (upper 2 cm) were collected from Camarones River (18º57′S, 69º30′O) of Northern Chile, a narrow and shallow river featuring arsenic concentrations up to 1 mgL−1 (Yañez et al. 2005). Sediment samples were maintained at 4°C during transportation to laboratory for further processing.
Microcosm experiments were carried in two flask, each one containing 5 g of sediment and 400 mL of chemically definided medium (CDM) (Macur et al. 2004), enriched with As(III) (75 μM) and As(V) (250 μM) respectively. Then, samples were incubated at 20°C for 14 days.
Bacteria were isolated by adding 1 g of homogenized sediment to 10 mL of NaCl (0.85%), and shaking at 100 rpm for 5 min. The mixture was serially diluted and 0.1 mL aliquots were plated onto R2A agar and incubated at 25°C for 48 h (Battaglia-Brunet et al. 2002). After growth, a number of differents colonies were purified. Bacterial isolates were identified by RapID ONE System and RapID NF Plus System (Remel) according to the supplier’s instructions.
The Minimum Inhibitory Concentration (MIC) of isolates was determined by the agar dilution technique,on LB agar plates amended with variable concentrations of sodium arsenite, between 0.5–100 mM and sodium arsenate between 0.5–1,000 mM. Each plate was inoculated with cell suspensions from fresh precultures to a final density of approximately 3 × 107 CFU mL−1 and incubated for 24 h at 25°C. An agar plate with bacteria and without the metalloid was used as a control. The MIC is defined as the lowest concentration that causes no visible growth (Valenzuela et al. 2008).
Redox transformation of arsenic was screened using the KMnO4 method described by Salmassi et al. (2002). Chemically defined medium (CDM) with As(III) (500 μM) or As(V) (1,000 μM) were inoculated with isolates, incubated at 25°C for 48 h and 60 μL of 0.01 M KMnO4 were added to 1 mL of the culture. A pink color indicated that arsenate was present, whereas a clear yellow or orange colored solution indicated that arsenite was present.
For arsC gene detection, genomic DNA was extracted from each isolate by freezing. The extracted DNA was used as template for the amplification by PCR. The primers sets utilized were arsC-1-F (5′ GTAATACGCTGGAGATGATCCG 3′) and arsC-1-R (5′ TTTTCCTGCTTCATCAACGAC 3′) corresponding to the ars operon from Escherichia coli according to the description of Saltikov y Olson (2002); and arsC-1-F (5′ AGTCCTGTTCATGTGYAC 3′) and arsC-1-R (5′ TGGCGTSGAAYGCCG 3′) corresponding to the ars operon of Pseudomonas aeruginosa and Pseudomonas putida according to the description of Saltikov and Olson (2002) and Macur et al. (2004). PCR products were separated by electrophoresis in 1.2% agarose gel and visualized by staining with ethidium bromide in UV transilluminator (Muller et al. 2003).
The aox gene detection was performed incubating the bacterial strains in R2A broth during 24 h; cultures were induced by addition of 250 mgL−1 of As(III) 12 h before extraction. The RT-PCR was carried out as described by Muller et al. (2003).
Results and Discussion
A total of 49 bacterial strains were isolated from sediments based on their abilities to grow in the presence of arsenic. Resistance to arsenic was defined as the ability to grow on agar containing either 7 mM As(III) or 20 mM As(V) at 25°C (Rokbani et al. 2007). Fourty five of 49 isolates were resistant to arsenite and all isolates were resistant to arsenate. The strains UC-9, UC-29, UC-90, UC-31, UC-54, UC-58, UC-101 and UC-72 were capable of tolerating arsenate above 1,000 mM. The strains UC-4, UC-34, UC-66, UC-5, UC-7, UC-39, UC-31, UC-45, UC-60 and UC-68 were capable of tolerating arsenite concentrations greater than 40 mM (Table 1a, b).
Arsenate and arsenite levels tested in this study are higher compared to those used in previous studies. Jackson et al. (2005) reported arsenic resistant levels of 400 and 10 mM of arsenate and arsenite, respectively. Studies conducted by Macur et al. (2004) reported arsenate resistance levels of 0.25 mM, from estuarine samples. Anderson and Cook (2004) obtained one isolate from arsenic-contaminated gold mine tailings that was able to grow at 100 mM arsenate (the maximum concentration tested), but the majority of their isolates showed growth inhibition at lower concentrations.
A qualitative KMnO4 screening technique was used to detect the oxidation of As(III) to As(V) or the reduction of As(V) to As(III). Twenty-seven out of the 49 selected isolates where able to reduce arsenic (55%); two strains was able to oxidize arsenic (4%); four show arsenic oxidizing as well as reducing capabilities (8%), and 16 bacteria had not detectable activity (33%) (data not shown). These results suggest that bacteria can influence the arsenic speciation in the environment.
Eleven out of the 49 (22,4%) arsenic-resistant isolates presented a PCR product of approximately 370 bp, expected size related to arsenate reductase (Saltikov and Olson 2002) (Fig. 1). The ars system is a widely studied arsenic resistance mechanism, whose arsC gene codifies a soluble enzyme, arsenate reductase, that catalyzes arsenate reduction to arsenite (Jackson and Dugas 2003).
Electrophoresis in gel of agarose from PCR products using primers arsC-1-F y arsC-1-R. Line 1 S. paucimobilis; 2 P. agglomerans; 3 E. americana; 4 E. hermanii;5 C. testosteroni; 6 H. alvei; 7 M. osloensis; 8 S. odorifera; 9 P. agglomerans; 10 E. aerogenes; 11 S. plymuthica; 12 S. odorifera; 13 P. stutzeri; 14 B. cepacia;15 E. cloacae; 16 C. Meningosepticum; MP, DNA ladder
Various researchers have already reported that amplification of the ars genes and unknown chromosomal ars areas during a PCR is influenced by many factors, such as the type of primer used, conformational variation in the extracted DNA, thermal cyclic conditions and the composition of buffers or agents (Chang et al. 2007). Ford et al. (2005) failed to correlate the level of arsenate resistance with the prevalence of ars genes in a set of aerobic and anaerobic bacteria.
Although the oxidation of As(III) to As(V) is known to be regulated by the presence of arsenite oxidase enzyme, the arsenite-oxidase gen was not detected in this study (see Table 1a, b). There was not amplification of any genes for arsenite oxidase in the isolates, despite the fact that arsenite oxidizing activity was experimentally shown. Such activity could be explained by other alternative mechanisms of arsenic resistance. Nevertheless, it is possible that the resistance arsenic genes in the investigated bacteria could be different to those described in the literature.
A total of 49 isolates were identified by RapID ONE System and RapID NF Plus System (Remel) and distributed between three mayor bacterial lineages (Table 1a, b). γ-Proteobacteria (26 isolates), β-Proteobacteria (13 isolates) and α-Proteobacteria (5 isolates). Most genera recovered from the sediments were: Moraxella, Alcaligenes, Comamonas, Burkordelia, Sphingomonas, Pantoea, Erwinia, Serratia, Enterobacter and Pseudomonas. The genera identified were not previously reported in literature as arsenic resistant.
This is the first work that describe arsenic resistant bacteria, isolated from sediments from Camarones river, involved on the arsenic biogeochemical cycles. These arsenic-resistant bacteria could grow in the presence of arsenite and arsenate, and could play an important role in natural arsenic speciation. Futher studies are required to understand the physiological role of biological transformation of arsenic.
References
Al-Abed SR, Jegadeesan G, Purandare J, Allen D (2007) Arsenic release from iron rich mineral processing waste: influence of pH and redox potential. Chemosphere 66:775–782
Anderson C, Cook G (2004) Isolation and characterization of arsenate-reducing bacteria from arsenic-contaminated sites in New Zealand. Curr Microbiol 48:341–347
Battaglia-Brunet F, Dictor MC, Garrido F, Crouzet C, Morin D, Dekeyser K, Clarens M, Baranger P (2002) An arsenic(III)-oxidizing bacterial population: selection, characterization and performance in reactors. J Appl Microbiol 93:656–667
Chang JS, Yoon IH, Kim KW (2007) Isolation and ars detoxification of arsenite-oxidizing bacteria from abandoned arsenic-contaminated mines. J Microbiol Biotech 17:812–821
Ellis PJ, Conrads T, Hille R, Kuhn P (2001) Crystal structure of the 100 kDa arsenite oxidase from Alcaligenes faecalis in two crystal forms at 1.64 A and 2.03 A. Structure 9:125–132
Ford T, Jay J, Patel A, Kile M, Prommasith P, Galloway T, Sanger R, Smith K, Depledge M (2005) Use of ecotoxicological tools to evaluate the health of new Bedford harbor sediments: a microbial biomarker approach. Environ Health Perspect 113:186–191
Jackson CR, Dugas SL (2003) Phylogenetic analysis of bacterial and archael arsC gene sequences suggest an ancient, common origin for arsenate reductase. BMC Evol Biol 3:18–23
Jackson CR, Harrison KG, Dugas SL (2005) Enumeration and characterization of culturable arsenate resistant bacteria in a large estuary. Syst Appl Microbiol 28:727–734
Macur RE, Colin J, Botero I, Mc Dermott TR, Inskeep W (2004) Bacterial populations associated with the oxidation and reduction of arsenic in an unsaturated soil. Environ Sci Technol 38:101–111
Muller D, Lièvremont D, Simeonova D, Hubert JC, Lett MC (2003) Arsenite oxidase aox genes from a metal-resistant B-proteobacterium. J Bacteriol 185:135–141
Neff JM (1997) Ecotoxicology of arsenic in marine environment. Environ Toxicol Chem 16:917–927
Ordoñez E, Letek M, Valbuena N, Gil JA, Mateos LM (2005) Analysis of genes involved in arsenic resistance in Corynebacterium glutamicum ATCC 13032. Appl Environ Microbiol 71:6206–6215
Queirolo F, Stegen S, Mondaca J, Cortes R, Rojas R, Contreras C, Muñoz L, Schwuger MJ, Ostapczuk P (2000) Total arsenic, lead, cadmium, copper and zinc in some salt rivers in the northern Andes of Antofagasta, Chile. Sci Total Environ 255:75–84
Rokbani A, Bauda P, Billard P (2007) Diversity of arsenite transporter genes from arsenic-resistant soil bacteria. Res Microbiol 158:128–137
Salmassi TM, Venkateswaren K, Satomi M, Nealson K, Newman D, Hering J (2002) Oxidation of arsenite by Agrobacterium albertimagni, AOL15, sp. Nov, isolated from Hotcreek, California. Geomicrobiol J 19:53–66
Saltikov C, Olson B (2002) Homology of Escherichia coli R773 arsA, arsB, and arsC genes in arsenic-resistant bacteria isolated from raw sewage and arsenic-enriched creek waters. Appl Environ Microbiol 68:280–288
Silver S, Phung T (2005) Genes and enzymes involved in bacterial oxidation and reduction of inorganic arsenic. Appl Environ Microbiol 71:599–608
Smedley PL, Kinniburgh DG (2002) A review of the source, behaviour and distribution of arsenic in natural waters. Appl Geochem 17:517–568
Taerakul P, Sun P, Golightly DW, Walker HW, Weavers LK, Zand B, Butalia T, Thomas TJ, Gupta H, Fan LS (2007) Characterization and re-use potential of by-products generated from the Ohio state carbonation and ash reactivation (OSCAR). Process Fuel 86:541–553
Valenzuela C, Campos VL, Yañez J, Zaror C, Mondaca MA (2008) Isolation of arsenite-oxidizing bacteria from arsenic-enriched sediments from Camarones river, northern Chile. Bull Environ Contam Toxicol 82:593–596
Yañez J, Fierro V, Mansilla H, Figueroa L, Cornejo L, Barnes R (2005) Arsenic speciation in human hair: a new perspective for epidemiological assessment in chronic arsenicism. J Environ Monit 7:1335–1341
Acknowledgments
This work has been supported by Dirección de Investigación, Universidad de Concepción (Proyect N° 204.036.027-1.0) and FONDECYT (Proyect N° 1050088).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Escalante, G., Campos, V.L., Valenzuela, C. et al. Arsenic Resistant Bacteria Isolated from Arsenic Contaminated River in the Atacama Desert (Chile). Bull Environ Contam Toxicol 83, 657–661 (2009). https://doi.org/10.1007/s00128-009-9868-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00128-009-9868-4