Abstract
Aims/hypothesis
We assessed the predictive value of ectonucleotide pyrophosphatase/phosphodiesterase 1 gene (ENPP1) SNPs with regard to the risk of developing obesity and/or type 2 diabetes in a large French general population.
Methods
We genotyped the ENPP1 SNPs K121Q (rs1044498), IVS20delT-11 (rs1799774) and A/G+1044TGA (rs7754561) in 5,153 middle-aged participants of the Data from an Epidemiological Study on the Insulin Resistance Syndrome (DESIR) cohort.
Results
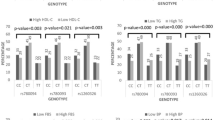
At baseline, the K121Q polymorphism was not associated either with BMI (p = 0.98) or with class I obesity (odds ratio [OR] 0.99, p = 0.81), but showed a borderline association with class II obesity (OR 1.65, p = 0.02). The K121Q variant was not associated with any trait during the 9-year follow-up. Pooled analyses both at baseline and at follow-up failed to show any association with hyperglycaemia (OR 1.08, p = 0.28) or type 2 diabetes (OR 1.15, p = 0.38). However, we did show an association of the Q121 allele with the risk of hyperglycaemia (OR 1.45, p = 0.001; n = 265) and type 2 diabetes (OR 1.65, p = 0.01; n = 103) in participants reporting a family history of type 2 diabetes. These results did not remain significant after a Bonferroni correction. The IVS20delT-11 and A/G+1044TGA polymorphisms and the three-allele risk haplotype (K121Q, IVS20delT-11 and A→G+1044TGA [QdelTG]) were not associated with any trait, either at baseline or at follow-up.
Conclusions/interpretation
In a general French population we did not find an association of the QdelTG risk haplotype with adult obesity and type 2 diabetes. We detected nominal evidence of association between the K121Q polymorphism and both severe adult obesity at baseline and the risk of hyperglycaemia or type 2 diabetes in participants with a family history of type 2 diabetes in pooled analyses both at baseline and follow-up.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Type 2 diabetes is closely linked to obesity. Indeed adults with a BMI ≥ 40 kg/m2 have a sevenfold increased risk of type 2 diabetes in comparison with lean individuals [1], but the molecular determinants of the obesity-related type 2 diabetes phenotype also named ‘diabesity’ are mostly unknown. Linkage analyses for traits associated with obesity [2–4] and type 2 diabetes [5–9] have reported a region potentially contributing to ‘diabesity’ on chromosome 6q16-q27. A plausible positional candidate gene in this region is the ecto-nucleotide pyrophosphatase phosphodiesterase 1 (ENPP1) gene. ENPP1 directly inhibits insulin-induced conformational changes of the insulin receptor, thereby affecting its activation and downstream signalling [10]. The functional K121Q missense polymorphism [11] has been associated with insulin resistance [12–16], type 2 diabetes [16–19], overweight [20], obesity [21] and ‘diabesity’ [22] in several studies with samples from diverse ethnic backgrounds. Recently, we studied more than 6,000 participants of European origin and showed an association between an ENPP1 three-allele risk haplotype (K121Q, IVS20delT-11 and A→G+1044TGA [QdelTG]) and childhood obesity, severe adult obesity and the associated type 2 diabetes [23]. The association between the ENPP1 three-allele risk haplotype and childhood obesity was recently confirmed in a German population [24], whereas the association with the at-risk haplotype for adult obesity and type 2 diabetes was not replicated in other well powered studies [25, 26]. Some other reports on the K121Q single nucleotide polymorphism (SNP) have shown inconsistent allelic associations with BMI [27] and obesity [28] or even no association with insulin resistance or type 2 diabetes [20, 29–34]. The effects of ENPP1 SNPs have not been investigated yet in the context of a prospective cohort study, said to be the ‘gold standard’ in genetic epidemiology [35]. Although small-sized genetic association studies for complex traits have often been challenged by irreproducibility [36], large case-control, familial and population-based prospective cohorts have provided reliable data on several common genetic variants in type 2 diabetes susceptibility [37–39]. Here we analysed the ENPP1 polymorphisms in a well-known general French population, the Data from an Epidemiological Study on the Insulin Resistance Syndrome (DESIR) study cohort [40, 41].
Methods
Study population
Men and women aged 30 to 64 years participated in the DESIR cohort, a 9-year follow-up study that aims to clarify the development of the insulin resistance syndrome [42]. Participants were recruited from volunteers insured by the French social security system, which offers periodic health examinations free of charge. They came from ten health examination centres in the western central part of France. All participants signed an informed consent declaration. The ethics committee of Bicêtre Hospital (Le Kremlin-Bicêtre, France) approved the protocol. Ethnic origin cannot be legally documented in France. We estimated the proportion of participants with non-European ancestry from a subgroup of 654 DESIR participants, as previously described [43]. We genotyped 328 SNPs, which were spaced by at least 5 Mb and highly differentiated among individuals from different continents (Fst statistic of Wright > 0.2 based on the Perlegen dataset) [44]. Analysis using STRUCTURE software (http://pritch.bsd.uchicago.edu/structure.html) identified only two individuals of non-European ancestry in a total of 654 individuals. From this analysis, the proportion of participants having non-European ancestry was estimated to be 0.30% in the DESIR cohort. A total of 5,153 participants genotyped for ENPP1 K121Q, IVS20delT-11 and A→G+1044TGA polymorphisms had data available at baseline [men/women ratio 49.6:50.4; age (mean±SD): 47.2 ± 10.0 years, BMI: 24.7 ± 3.8 kg/m2]. Among them, 4,593 participants were normoglycaemic (fasting plasma glucose < 6.1 mmol/l), of whom 3,528 (77%) were followed for incident impaired fasting glucose and diabetes during a 9-year period. In addition, 3,807 men and women were non-obese (BMI < 27 kg2) at baseline, with 2,952 (78%) being followed for overweight and class I and class II obesity during the 9-year period. At the four 3-yearly examinations during the 9 years of follow-up, 2,005 participants were both non-obese and had normal glucose tolerance.
Glycaemic status was defined according to 1997 American Diabetes Association criteria [45] (normoglycaemia: glucose < 6.1 mmol/l; impaired fasting glucose, fasting plasma glucose 6.1–6.99 mmol/l; type 2 diabetes, fasting plasma glucose ≥ 7.0 mmol/l and/or treatment by hypoglycaemic agents). Participants with at least one first-degree relative with type 2 diabetes were declared as having a familial history of type 2 diabetes. This information was self-reported by the participant. Three classes of adiposity were used: overweight (BMI ≥ 27 kg/m2), class I obesity (BMI ≥ 30 kg/m2), class II obesity (BMI ≥ 35 kg/m2).
Measurements
Weight and height were measured by trained personnel and BMI (kg/m2) calculated. Venous blood samples were collected in the morning after a 12 h fasting period. Plasma glucose was assayed by the glucose oxidase method applied to fluoro-oxalated plasma, using a Technicon RA1000 (Bayer, Puteaux, France) or other automatic device (Kone Automate; Kone, Evry, France). HDL-cholesterol was measured by the phosphotungstic precipitation method and triacylglycerol using the enzymatic Trinder method. Participants were asked about previous myocardial infarctions and strokes.
Genotyping
K121Q and A→G+1044TGA genotyping was performed using SNPlex technology (Applied Biosystems: http://www.appliedbiosystems.com) based on the oligonucleotide ligation assay combined with multiplex PCR target amplification. The chemistry of the assay relies on a set of universal core reagent kits and a set of SNP-specific ligation probes allowing multiplex genotyping of 48 SNPs simultaneously in one unique sample. A quality control measure was included using specific internal controls for each step of the assay (according to the manufacturer’s instructions). Allelic discrimination was performed through capillary electrophoresis analysis using a DNA analyser (3730xl; Applied Biosystems) and GeneMapper 3.7 software (Applied Biosystems). This technology achieved 97% successful genotyping for the cohort. We genotyped IVS20delT-11 with the TaqMan (Applied Biosystems). As a quality control procedure we double-genotyped a subset of 742 individuals (14.4% of the whole sample) using Light Typer (Roche Diagnostics: http://www.rochediagnostics.fr) for K121Q and TaqMan for IVS20delT-11 and A→G+1044TGA SNPs. The concordance rates for K121Q, IVS20delT-11 and A→G+1044TGA were 99.7, 100 and 100% respectively. Probes for Light Typer were synthesised by TIB Molbiol Syntheselabor (http://www.tib-molbiol.de).
Statistical analysis
Tests for deviation from Hardy-Weinberg equilibrium and for association were performed with the De Finetti program (http://ihg.gsf.de/cgi-bin/hw/hwa1.pl). Logistic regression was used for disease status, adjusting for the confounding variables baseline age, BMI and sex. To assess the effect of genotype on the incidence of new disease in participants who were healthy at baseline, we used non parametric Kaplan–Meier survival analysis models and the log-rank test to compare genotypes. SPSS 10.1 software (http://www.spss.com/fr) was used for general statistical analyses. For statistical power calculation we used the QUANTO program (http://hydra.usc.edu/GxE) [46].
Results
The genotypic distributions of all three SNPs were in Hardy–Weinberg equilibrium (p > 0.01). The Q121, IVS20delT-11 and A→G+1044TGA minor allele frequencies were 16.1, 23.4 and 28.4% in the whole cohort, in agreement with previously described data in European white populations. As ENPP1 SNPs have been suggested to be associated with obesity and type 2 diabetes, we selected as control persons the 2,005 DESIR participants who were non-obese (BMI < 27 kg/m2) and had normal fasting glucose values (<6.1 mmol/l) at baseline and at every subsequent 3-yearly examination during the 9-year period of follow-up.
We first analysed the K121Q polymorphism at baseline. The K121Q polymorphism was not associated with overweight [odds ratio (OR) 0.95, p = 0.67], class I obesity (OR 0.99, p = 0.81) or hyperglycaemia (OR 1.16, p = 0.11). In the whole population, the Q121 allele was not associated with the continuous trait BMI [p = 0.98; Electronic supplementary material (ESM) Table 1]. The proportion of Q121 allele carriers was higher in the class II obese group [OR 1.65 (95% CI 1.09–2.46) under an additive model, p = 0.02; Table 1]. The OR under a recessive model increased to 2.71 (95% CI 1.05–6.99; p = 0.03). Using the same model, we observed an association between K121Q and type 2 diabetes [OR 2.14 (95% CI 1.00–4.58), p = 0.04; Table 1]. In addition, we found a trend towards association of K121Q with a decreased HDL-cholesterol : triacylglycerol ratio (KK + KQ, 2.01 ± 0.02 vs QQ, 1.80 ± 0.09, arbitrary unit, p = 0.02) and with an increased frequency of stroke (KK + KQ, 0.4% vs QQ, 2.1%, p = 0.03) under a recessive model (ESM Table 1).
We then analysed the genotype distribution of K121Q in participants who were unaffected at baseline and developed obesity and/or hyperglycaemia during the 9-year follow-up period (Table 2). K121Q was not predictive of higher incidence of class I/class II obesity, hyperglycaemia or type 2 diabetes. The Q121 allele was marginally more prevalent in participants who developed overweight [OR 1.21 (95% CI 0.99–1.42), p = 0.06]. A non-parametric Kaplan–Meier survival analysis model over the 9-year follow-up showed no significant results with any trait (log-rank test, p > 0.05) and only a trend towards higher occurrence of overweight (log-rank test, p = 0.06) for Q121 carriers.
To test the K121Q association with obesity and type 2 diabetes traits with more statistical power, we pooled participants affected at baseline (Table 1) with those developing the disease during the 9-year follow-up period (Table 2), for an overall case-control analysis (Table 3). We did not observe any consistent association of the K121Q polymorphism with overweight (OR 1.03, p = 0.58), class I obesity (OR 0.99, p = 0.92), class II obesity (OR 1.27, p = 0.21), hyperglycaemia (OR 1.08, p = 0.28) and type 2 diabetes (OR 1.15, p = 0.38). We then assessed the effect of K121Q polymorphism on hyperglycaemia and type 2 diabetes in a subgroup of participants with a family history of type 2 diabetes. Interestingly, and under an additive model, we confirmed an association between the K121Q polymorphism and both hyperglycaemia [OR 1.45 (95% CI 1.16–1.82), p = 0.001, n = 265] and type 2 diabetes [OR 1.61 (95% CI 1.11–2.19), p = 0.01, n = 103], this association being restricted to participants with a family history of type 2 diabetes. The OR for hyperglycaemia and type 2 diabetes increased markedly under a recessive model [1.94 (95% CI 1.06–3.53) and 2.93 (95% CI 1.36–6.33) respectively, p = 0.03 and p = 0.004]. In the logistic regression models, and taking into account baseline age, BMI and sex for hyperglycaemia and type 2 diabetes in participants with a family history of the disease, associations with K121Q remained significant (p < 0.05, data not shown).
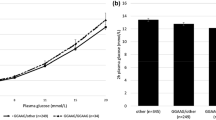
Due to the striking differences in ORs of K121Q for type 2 diabetes in general and type 2 diabetes in participants with a family history of the disease, we next classified the type 2 diabetic patients into two subgroups, namely with and without a family history of the disease (Fig. 1). Type 2 diabetic patients with a family history of the disease showed an increased Q121 allele frequency in comparison with lean normoglycaemic participants [22.3 vs 15.5%, OR 1.56 (95% CI 1.11–2.19), p = 0.009]. In contrast, type 2 diabetic individuals without a family history of the disease and control individuals had similar Q121 allele frequencies [14.3 vs 15.5%, OR 0.91 (95% CI 0.68–1.21), p = 0.51]. A case-only Q121 allele frequency comparison between familial and non-familial type 2 diabetes showed a significant difference [OR 1.72 (95% CI 1.12–2.63), p = 0.01].
Allele frequencies of the Q121 allele in lean normoglycaemic (open bars) and type 2 diabetic patients (black bar), stratified according to presence (dark grey bar) or not (light grey bar) of type 2 diabetes family history. OR for type 2 diabetic group vs lean normoglycaemic group was 1.11 (p = 0.37).Type 2 diabetic patients with a family history of the disease (n = 103) showed an increased Q121 allele frequency in comparison with lean normoglycaemic patients [22.3 vs 15.5%, OR 1.56 (95% CI 1.11–2.19), p = 0.009]. In contrast, type 2 diabetic individuals without a family history of the disease (n = 210) and control individuals had similar Q121 allele frequencies [14.3 vs 15.5%, OR 0.91 (95% CI 0.68–1.21), p = 0.51]. A case-only Q121 allele frequency comparison between familial and non-familial type 2 diabetes showed a significant difference [OR 1.72 (95% CI 1.12–2.63), p = 0.01]
IVS20delT-11 and A/G+1044TGA polymorphisms and the QdelTG haplotype were not associated with any trait, either at baseline or at follow-up (p > 0.05, data not shown).
Discussion
The ENPP1 gene analysis of this large prospectively followed French cohort does not support the role of K121Q in the development of obesity or type 2 diabetes. Our results suggest a putative association of the K121Q polymorphism with class II adult obesity at baseline and with glucose intolerance and type 2 diabetes restricted to participants with a self reported family history of type 2 diabetes. This is in agreement with data reported earlier in highly selected case-control and family samples [23]. We found higher Q121 allele prevalence among diabetic participants (diabetic at baseline plus those developing hyperglycaemia during the 9-year follow up) with a family history of type 2 diabetes than among other diabetic patients (22.3 vs 14.3%, respectively, p = 0.01). This observation confirms that familial forms of type 2 diabetes are a high-value resource to identify susceptibility genes, but may not be fully representative of common forms of the disease. A similar finding was concluded from the Botnia observational prospective study, which included 2,293 first-degree relatives of type 2 diabetic patients [47]. In this regard, the analysis of the peroxisome proliferator-activated receptor gamma gene (PPARG) Pro12Ala and the calpain 10 gene (CAPN10) SNP44 in our French familial type 2 diabetic cases that have been previously reported showed higher relative risks for the disease [48, 49] than previously found in non selected case-control studies [37, 39]. However, caution is needed, since this subgroup analysis was performed on a limited number of familial cases (hyperglycaemia, n = 265; type 2 diabetes, n = 103).
The difference in the genetic background of type 2 diabetic cases may in part explain discrepancies between ENPP1 K121Q SNP as highlighted in recent studies. In the present report, K121Q polymorphism was not associated with overweight, class I obesity, hyperglycaemia and type 2 diabetes, either at baseline or during the follow-up. Cohorts enriched in familial type 2 diabetes/obesity cases or obtained from ethnic groups with a high prevalence of type 2 diabetes/obesity are more likely to show positive association with ENPP1. Indeed, the most potent effect of ENPP1 on type 2 diabetes was found in French families with early-onset obesity and in southern Indians [17, 23], two populations showing severe insulin resistance. In addition, we selected a cohort of 752 French participants with type 2 diabetes and familial history of the disease in our previous report [23].
With regard to obesity, we only observed a borderline association with class II obesity at baseline and a trend towards higher occurrence of overweight in initially lean Q121 carriers over the 9-year follow-up. However, K121Q was not associated with overweight and class I obesity, either at baseline or during the follow-up. Power calculation with alpha level = 0.05 indicated a good power to detect association of class I obesity, hyperglycaemia and type 2 diabetes with K121Q in the whole sample (0.99, 0.99 and 0.86, respectively) and a reduced power to detect association of K121Q with class II obesity (0.59), considering our previously reported ORs in the French population [23]. Again, it is likely that cohorts enriched in familial obesity cases in our previous study probably led to overestimation of the relative risk of obesity [23]. Unfortunately, the absence of available data regarding the family history of obesity precluded a subgroup analysis of familial and non-familial obese participants in the DESIR cohort. The previously documented, highly significant association with class III obesity [23] was not tested, given the weak number of affected participants in this study (n = 28). These data indicate the limitations of the existing medium-sized general cohorts to explore the effect of SNP on the development of extreme phenotypes. We also observed a putative effect of the K121Q SNP on the atherogenic risk factor HDL:triacylglycerol ratio and on history of stroke. Although these data may be due to noise, they are in line with the previously reported association of K121Q with myocardial infarction [18, 50].
We did not replicate, in this cohort, our previous finding of an association between ENPP1 IVS20delT-11 and A/G+1044TGA SNPs and obesity [23], nor did we find a significant association between the QdelTG haplotype and both obesity and type 2 diabetes. Three recent reports have shown conflicting results about an effect of the QdelTG haplotype on glucose intolerance, type 2 diabetes and obesity [24–26]. It remains possible that our original results reported a false positive result, and further replication is needed to clarify the role of the QdelTG haplotype in comparison with K121Q analysed alone.
Importantly, our results would not be significant if subjected to stringent correction for multiple testing. Therefore, we cannot rule out a spurious positive association due to multiple testing.
In conclusion, our data do not confirm the association of the ENPP1 QdelTG haplotype with type 2 diabetes and obesity. We observed a possible association of the K121Q polymorphism both with severe adult obesity at baseline, and with hyperglycaemia and type 2 diabetes in individuals with a family history of the disease in pooled analyses at baseline and at follow-up.
Abbreviations
- DESIR:
-
Data from an Epidemiological Study on the Insulin Resistance Syndrome
- ENPP1:
-
ectonucleotide pyrophosphatase/phosphodiesterase 1
- OR:
-
odds ratio
- QdelTG:
-
three-allele risk haplotype (K121Q, IVS20delT-11 and A→G+1044TGA)
- SNP:
-
single nucleotide polymorphism
References
Mokdad AH, Ford ES, Bowman BA et al (2003) Prevalence of obesity, diabetes, and obesity-related health risk factors, 2001. JAMA 289:76–79
Atwood LD, Heard-Costa NL, Cupples LA, Jaquish CE, Wilson PW, D’Agostino RB (2002) Genomewide linkage analysis of body mass index across 28 years of the Framingham Heart Study. Am J Hum Genet 71:1044–1050
Meyre D, Lecoeur C, Delplanque J et al (2004) A genome-wide scan for childhood obesity-associated traits in French families shows significant linkage on chromosome 6q22.31–q23.2. Diabetes 53:803–811
Fox CS, Heard-Costa NL, Wilson PW, Levy D, D’Agostino RB Sr, Atwood LD (2004) Genome-wide linkage to chromosome 6 for waist circumference in the Framingham Heart Study. Diabetes 53:1399–1402
Ghosh S, Watanabe RM, Valle TT et al (2000) The Finland–United States investigation of non-insulin-dependent diabetes mellitus genetics (FUSION) study. I. An autosomal genome scan for genes that predispose to type 2 diabetes. Am J Hum Genet 67:1174–1185
Ehm MG, Karnoub MC, Sakul H et al (2000) Genomewide search for type 2 diabetes susceptibility genes in four American populations. Am J Hum Genet 66:1871–1881
Duggirala R, Blangero J, Almasy L et al (2001) A major locus for fasting insulin concentrations and insulin resistance on chromosome 6q with strong pleiotropic effects on obesity-related phenotypes in nondiabetic Mexican Americans. Am J Hum Genet 68:1149–1164
Abney M, Ober C, McPeek MS (2002) Quantitative-trait homozygosity and association mapping and empirical genomewide significance in large, complex pedigrees: fasting serum-insulin level in the Hutterites. Am J Hum Genet 70:920–934
Xiang K, Wang Y, Zheng T et al (2004) Genome-wide search for type 2 diabetes/impaired glucose homeostasis susceptibility genes in the Chinese: significant linkage to chromosome 6q21–q23 and chromosome 1q21–q24. Diabetes 53:228–234
Maddux BA, Goldfine ID (2000) Membrane glycoprotein PC-1 inhibition of insulin receptor function occurs via direct interaction with the receptor alpha-subunit. Diabetes 49:13–19
Costanzo BV, Trischitta V, Di Paola R et al (2001) The Q allele variant (GLN121) of membrane glycoprotein PC-1 interacts with the insulin receptor and inhibits insulin signaling more effectively than the common K allele variant (LYS121). Diabetes 50:831–836
Pizzuti A, Frittitta L, Argiolas A et al (1999) A polymorphism (K121Q) of the human glycoprotein PC-1 gene coding region is strongly associated with insulin resistance. Diabetes 48:1881–1884
Gu HF, Almgren P, Lindholm E et al (2000) Association between the human glycoprotein PC-1 gene and elevated glucose and insulin levels in a paired-sibling analysis. Diabetes 49:1601–1603
Frittitta L, Baratta R, Spampinato D et al (2001) The Q121 PC-1 variant and obesity have additive and independent effects in causing insulin resistance. J Clin Endocrinol Metab 86:5888–5891
Kubaszek A, Pihlajamaki J, Karhapaa P, Vauhkonen I, Laakso M (2003) The K121Q polymorphism of the PC-1 gene is associated with insulin resistance but not with dyslipidemia. Diabetes Care 26:464–467
Kubaszek A, Markkanen A, Eriksson JG et al (2004) The association of the K121Q polymorphism of the plasma cell glycoprotein-1 gene with type 2 diabetes and hypertension depends on size at birth. J Clin Endocrinol Metab 89:2044–2047
Abate N, Chandalia M, Satija P et al (2005) ENPP1/PC-1 K121Q polymorphism and genetic susceptibility to type 2 diabetes. Diabetes 54:1207–1213
Bacci S, Ludovico O, Prudente S et al (2005) The K121Q polymorphism of the ENPP1/PC-1 gene is associated with insulin resistance/atherogenic phenotypes, including earlier onset of type 2 diabetes and myocardial infarction. Diabetes 54:3021–3025
Willer CJ, Bonnycastle LL, Conneely KN et al (2007) Screening of 134 single nucleotide polymorphisms (SNPs) previously associated with type 2 diabetes replicates association with 12 SNPs in nine genes. Diabetes 56:256–264
Grarup N, Urhammer SA, Ek J et al (2006) Studies of the relationship between the ENPP1 K121Q polymorphism and type 2 diabetes, insulin resistance and obesity in 7,333 Danish white subjects. Diabetologia 49:2097–2104
Wan C, Zhang T, Wang B et al (2006) Obesity risk associated with the K121Q polymorphism of the glycoprotein PC-1 gene. Diabetes Obes Metab 8:703–708
Bochenski J, Placha G, Wanic K et al (2006) New polymorphism of ENPP1 (PC-1) is associated with increased risk of type 2 diabetes among obese individuals. Diabetes 55:2626–2630
Meyre D, Bouatia-Naji N, Tounian A et al (2005) Variants of ENPP1 are associated with childhood and adult obesity and increase the risk of glucose intolerance and type 2 diabetes. Nat Genet 37:863–867
Bottcher Y, Korner A, Reinehr T et al (2006) ENPP1 variants and haplotypes predispose to early onset obesity and impaired glucose and insulin metabolism in German obese children. J Clin Endocrinol Metab 91:4948–4952
Weedon MN, Shields B, Hitman G et al (2006) No evidence of association of ENPP1 variants with type 2 diabetes or obesity in a study of 8,089 U.K. Caucasians. Diabetes 55:3175–3179
Lyon HN, Florez JC, Bersaglieri T et al (2006) Common variants in the ENPP1 gene are not reproducibly associated with diabetes or obesity. Diabetes 55:3180–3184
Prudente S, Chandalia M, Morini E et al (2007) The Q121/Q121 genotype of ENPP1/PC-1 is associated with lower BMI in non-diabetic whites. Obesity (Silver Spring) 15:1–4
Matsuoka N, Patki A, Tiwari HK et al (2006) Association of K121Q polymorphism in ENPP1 (PC-1) with BMI in Caucasian and African-American adults. Int J Obes (Lond) 30:233–237
Rasmussen SK, Urhammer SA, Pizzuti A et al (2000) The K121Q variant of the human PC-1 gene is not associated with insulin resistance or type 2 diabetes among Danish Caucasians. Diabetes 49:1608–1611
Gonzalez-Sanchez JL, Martinez-Larrad MT, Fernandez-Perez C, Kubaszek A, Laakso M, Serrano-Rios M (2003) K121Q PC-1 gene polymorphism is not associated with insulin resistance in a Spanish population. Obes Res 11:603–605
Laukkanen O, Pihlajamaki J, Lindstrom J et al (2004) Common polymorphisms in the genes regulating the early insulin signalling pathway: effects on weight change and the conversion from impaired glucose tolerance to Type 2 diabetes. The Finnish Diabetes Prevention Study. Diabetologia 47:871–877
Morrison JA, Gruppo R, Glueck CJ et al (2004) Population-specific alleles: the polymorphism (K121Q) of the human glycoprotein PC-1 gene is strongly associated with race but not with insulin resistance in black and white children. Metabolism 53:465–468
Keshavarz P, Inoue H, Sakamoto Y et al (2006) No evidence for association of the ENPP1 (PC-1) K121Q variant with risk of type 2 diabetes in a Japanese population. J Hum Genet 51:559–566
Gouni-Berthold I, Giannakidou E, Faust M, Berthold HK, Krone W (2006) The K121Q polymorphism of the plasma cell glycoprotein-1 gene is not associated with diabetes mellitus type 2 in German Caucasians. Horm Metab Res 38:524–529
Hattersley AT, McCarthy MI (2005) What makes a good genetic association study? Lancet 366:1315–1323
Hirschhorn JN, Lohmueller K, Byrne E, Hirschhorn K (2002) A comprehensive review of genetic association studies. Genet Med 4:45–61
Altshuler D, Hirschhorn JN, Klannemark M et al (2000) The common PPARgamma Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat Genet 1:76–80
Gloyn AL, Weedon MN, Owen KR et al (2003) Large-scale association studies of variants in genes encoding the pancreatic beta-cell KATP channel subunits Kir6.2 (KCNJ11) and SUR1 (ABCC8) confirm that the KCNJ11 E23K variant is associated with type 2 diabetes. Diabetes 52:568–572
Weedon MN, Schwarz PE, Horikawa Y et al (2003) Meta-analysis and a large association study confirm a role for calpain-10 variation in type 2 diabetes susceptibility. Am J Hum Genet 73:1208–1212
Jaziri R, Lobbens S, Aubert R et al (2006) The PPARG Pro12Ala polymorphism is associated with a decreased risk of developing hyperglycemia over 6 years and combines with the effect of the APM1 G-11391A single nucleotide polymorphism: The Data From an Epidemiological Study on the Insulin Resistance Syndrome (D.E.S.I.R.) Study. Diabetes 55:1157–1162
Cauchi S, Meyre D, Choquet H et al (2006) TCF7L2 variation predicts hyperglycemia incidence in a French general population: The Data From an Epidemiological Study on the Insulin Resistance Syndrome (D.E.S.I.R.) Study. Diabetes 55:3189–3192
Balkau B (1996) An epidemiologic survey from a network of French Health Examination Centres, (D.E.S.I.R.): epidemiologic data on the insulin resistance syndrome. Rev Epidemiol Sante Publique 44:373–375 [article in French]
Sladek R, Rocheleau G, Rung J et al (2007) A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445:881–885
Hinds DA, Stuve LL, Nilsen GB et al (2005) Whole-genome patterns of common DNA variation in three human populations. Science 307:1072–1079
The Expert Committee on the Diagnosis and Classification of Diabetes Mellitus (1997) Report of the Expert Committee on the diagnosis and classification of diabetes mellitus. Diabetes Care 20:1183–1197
Gauderman WJ (2002) Sample size requirements for matched case-control studies of gene–environment interaction. Stat Med 21:35–50
Lyssenko V, Almgren P, Anevski D et al (2005) Genetic prediction of future type 2 diabetes. PLoS Med 2:1299–1308
Ghoussaini M, Meyre D, Lobbens S et al (2005) Implication of the Pro12Ala polymorphism of the PPAR-gamma 2 gene in type 2 diabetes and obesity in the French population. BMC Med Genet 6:11
Tsuchiya T, Schwarz PE, Bosque-Plata LD et al (2006) Association of the calpain-10 gene with type 2 diabetes in Europeans: results of pooled and meta-analyses. Mol Genet Metab 89:174–184
Endler G, Mannhalter C, Sunder-Plassmann H et al (2002) The K121Q polymorphism in the plasma cell membrane glycoprotein 1 gene predisposes to early myocardial infarction. J Mol Med 80:791–795
Acknowledgements
The cohort was supported by: (1) co-operative contracts between INSERM, CNAMTS, Novartis Pharma, Lilly and Sanofi-Aventis; (2) INSERM Réseaux en Santé Publique and INSERM Interactions entre les determinants de la santé; (3) Association Diabète Risque Vasculaire, the Fédèration Française de Cardiologie, La Fondation de France, ALFEDIAM and ONIVINS; and (4) Ardix Medical, Bayer Diagnostics, Becton Dickinson, Cardionics, Lilly, Merck Santé, Novo Nordisk, Pierre Fabre, Roche, Sanofi and Topcon.
Duality of interest
The authors declare that there is no duality of interest associated with this manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Meyre, D., Bouatia-Naji, N., Vatin, V. et al. ENPP1 K121Q polymorphism and obesity, hyperglycaemia and type 2 diabetes in the prospective DESIR Study. Diabetologia 50, 2090–2096 (2007). https://doi.org/10.1007/s00125-007-0787-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-007-0787-9